Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
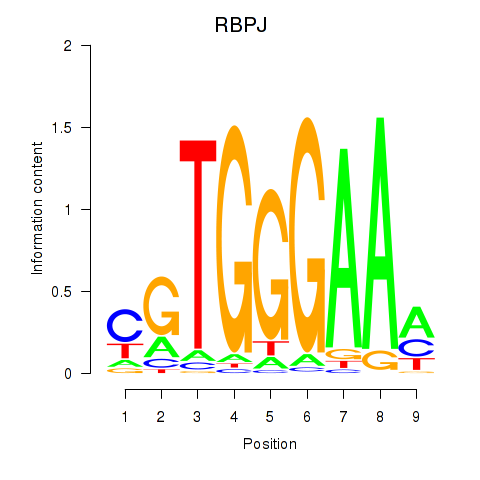
Results for RBPJ
Z-value: 0.59
Transcription factors associated with RBPJ
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RBPJ
|
ENSG00000168214.16 | RBPJ |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RBPJ | hg19_v2_chr4_+_26321284_26321377, hg19_v2_chr4_+_26322409_26322479 | 0.14 | 6.2e-01 | Click! |
Activity profile of RBPJ motif
Sorted Z-values of RBPJ motif
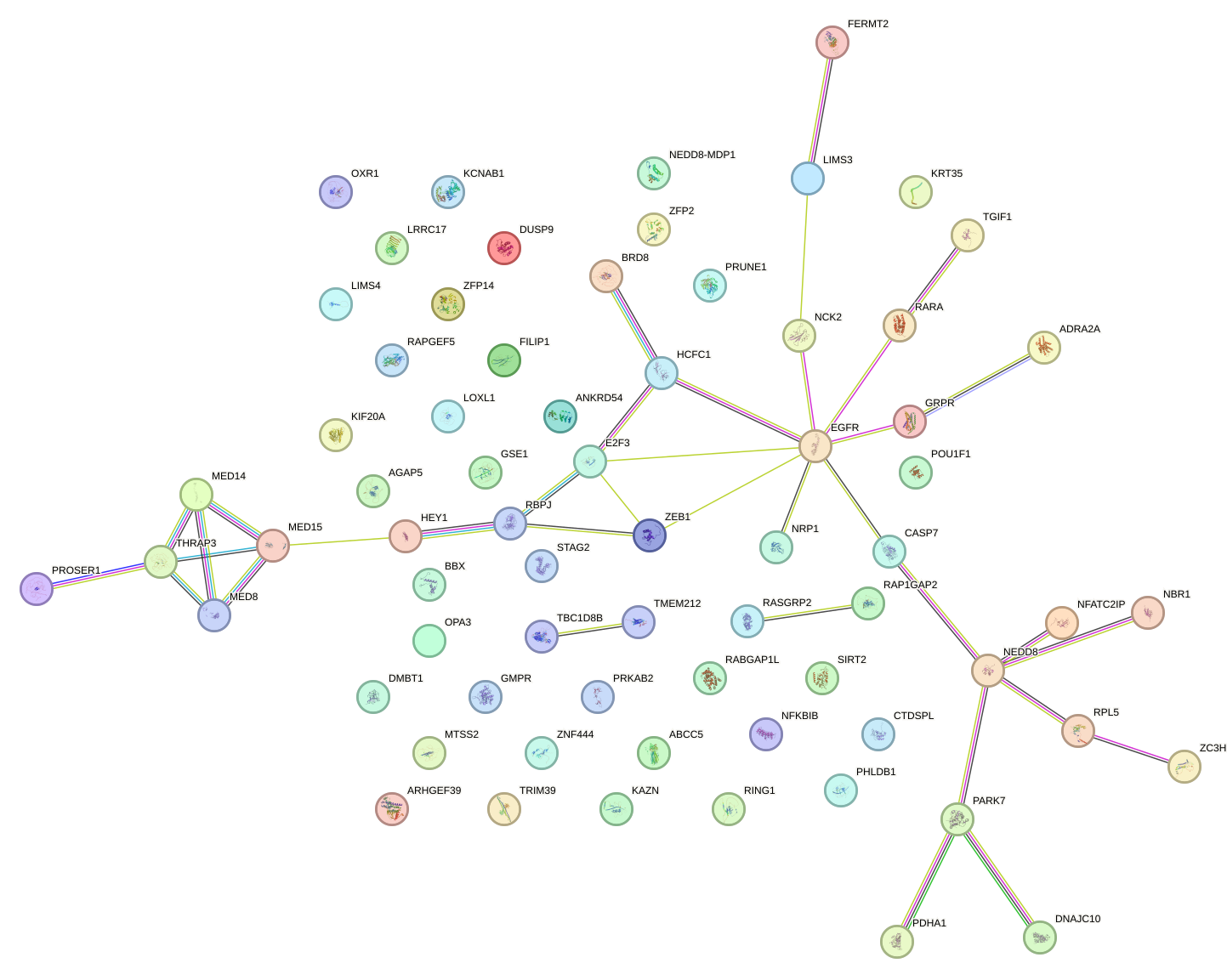
Network of associatons between targets according to the STRING database.
First level regulatory network of RBPJ
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_-_70719925 | 0.86 |
ENST00000338779.6 |
MTSS1L |
metastasis suppressor 1-like |
| chr9_+_706842 | 0.74 |
ENST00000382293.3 |
KANK1 |
KN motif and ankyrin repeat domains 1 |
| chr10_+_124320156 | 0.74 |
ENST00000338354.3 ENST00000344338.3 ENST00000330163.4 ENST00000368909.3 ENST00000368955.3 ENST00000368956.2 |
DMBT1 |
deleted in malignant brain tumors 1 |
| chr1_-_146644036 | 0.72 |
ENST00000425272.2 |
PRKAB2 |
protein kinase, AMP-activated, beta 2 non-catalytic subunit |
| chr10_+_124320195 | 0.72 |
ENST00000359586.6 |
DMBT1 |
deleted in malignant brain tumors 1 |
| chr8_-_80680078 | 0.71 |
ENST00000337919.5 ENST00000354724.3 |
HEY1 |
hes-related family bHLH transcription factor with YRPW motif 1 |
| chr7_+_102553430 | 0.68 |
ENST00000339431.4 ENST00000249377.4 |
LRRC17 |
leucine rich repeat containing 17 |
| chrX_+_152907913 | 0.67 |
ENST00000370167.4 |
DUSP9 |
dual specificity phosphatase 9 |
| chr1_-_146644122 | 0.64 |
ENST00000254101.3 |
PRKAB2 |
protein kinase, AMP-activated, beta 2 non-catalytic subunit |
| chr14_-_53417732 | 0.54 |
ENST00000399304.3 ENST00000395631.2 ENST00000341590.3 ENST00000343279.4 |
FERMT2 |
fermitin family member 2 |
| chr12_-_90049878 | 0.52 |
ENST00000359142.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr16_+_85645007 | 0.51 |
ENST00000405402.2 |
GSE1 |
Gse1 coiled-coil protein |
| chr15_+_74218787 | 0.48 |
ENST00000261921.7 |
LOXL1 |
lysyl oxidase-like 1 |
| chr12_-_90049828 | 0.47 |
ENST00000261173.2 ENST00000348959.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr7_+_55177416 | 0.47 |
ENST00000450046.1 ENST00000454757.2 |
EGFR |
epidermal growth factor receptor |
| chr3_+_38017264 | 0.44 |
ENST00000436654.1 |
CTDSPL |
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase-like |
| chr11_+_112832202 | 0.44 |
ENST00000534015.1 |
NCAM1 |
neural cell adhesion molecule 1 |
| chr10_+_31608054 | 0.38 |
ENST00000320985.10 ENST00000361642.5 ENST00000560721.2 ENST00000558440.1 ENST00000424869.1 ENST00000542815.3 |
ZEB1 |
zinc finger E-box binding homeobox 1 |
| chr17_+_2699697 | 0.38 |
ENST00000254695.8 ENST00000366401.4 ENST00000542807.1 |
RAP1GAP2 |
RAP1 GTPase activating protein 2 |
| chr10_-_51371321 | 0.35 |
ENST00000602930.1 |
AGAP8 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 8 |
| chr11_+_112832133 | 0.35 |
ENST00000524665.1 |
NCAM1 |
neural cell adhesion molecule 1 |
| chr17_-_36891830 | 0.33 |
ENST00000578487.1 |
PCGF2 |
polycomb group ring finger 2 |
| chr2_+_3642545 | 0.32 |
ENST00000382062.2 ENST00000236693.7 ENST00000349077.4 |
COLEC11 |
collectin sub-family member 11 |
| chr17_+_73452545 | 0.31 |
ENST00000314256.7 |
KIAA0195 |
KIAA0195 |
| chr18_+_3451584 | 0.29 |
ENST00000551541.1 |
TGIF1 |
TGFB-induced factor homeobox 1 |
| chrX_+_123094369 | 0.28 |
ENST00000455404.1 ENST00000218089.9 |
STAG2 |
stromal antigen 2 |
| chr3_+_142442841 | 0.28 |
ENST00000476941.1 ENST00000273482.6 |
TRPC1 |
transient receptor potential cation channel, subfamily C, member 1 |
| chr7_-_22259845 | 0.28 |
ENST00000420196.1 |
RAPGEF5 |
Rap guanine nucleotide exchange factor (GEF) 5 |
| chr22_+_20861858 | 0.27 |
ENST00000414658.1 ENST00000432052.1 ENST00000425759.2 ENST00000292733.7 ENST00000542773.1 ENST00000263205.7 ENST00000406969.1 ENST00000382974.2 |
MED15 |
mediator complex subunit 15 |
| chr16_-_53737795 | 0.26 |
ENST00000262135.4 ENST00000564374.1 ENST00000566096.1 |
RPGRIP1L |
RPGRIP1-like |
| chr16_-_53737722 | 0.26 |
ENST00000569716.1 ENST00000562588.1 ENST00000562230.1 ENST00000379925.3 ENST00000563746.1 ENST00000568653.3 |
RPGRIP1L |
RPGRIP1-like |
| chr11_+_112832090 | 0.25 |
ENST00000533760.1 |
NCAM1 |
neural cell adhesion molecule 1 |
| chr15_+_40697988 | 0.23 |
ENST00000487418.2 ENST00000479013.2 |
IVD |
isovaleryl-CoA dehydrogenase |
| chr17_+_38474489 | 0.23 |
ENST00000394089.2 ENST00000425707.3 |
RARA |
retinoic acid receptor, alpha |
| chr1_+_15272271 | 0.22 |
ENST00000400797.3 |
KAZN |
kazrin, periplakin interacting protein |
| chr4_-_164534657 | 0.22 |
ENST00000339875.5 |
MARCH1 |
membrane-associated ring finger (C3HC4) 1, E3 ubiquitin protein ligase |
| chr7_-_27169801 | 0.21 |
ENST00000511914.1 |
HOXA4 |
homeobox A4 |
| chrX_+_106045891 | 0.21 |
ENST00000357242.5 ENST00000310452.2 ENST00000481617.2 ENST00000276175.3 |
TBC1D8B |
TBC1 domain family, member 8B (with GRAM domain) |
| chr17_+_41322483 | 0.20 |
ENST00000341165.6 ENST00000586650.1 ENST00000422280.1 |
NBR1 |
neighbor of BRCA1 gene 1 |
| chr16_-_57831676 | 0.20 |
ENST00000465878.2 ENST00000539578.1 ENST00000561524.1 |
KIFC3 |
kinesin family member C3 |
| chr19_+_56652556 | 0.20 |
ENST00000337080.3 |
ZNF444 |
zinc finger protein 444 |
| chr19_+_56652686 | 0.19 |
ENST00000592949.1 |
ZNF444 |
zinc finger protein 444 |
| chr12_-_16759711 | 0.19 |
ENST00000447609.1 |
LMO3 |
LIM domain only 3 (rhombotin-like 2) |
| chr16_-_57836321 | 0.19 |
ENST00000569112.1 ENST00000562311.1 ENST00000445690.2 ENST00000379655.4 |
KIFC3 |
kinesin family member C3 |
| chr19_-_39390440 | 0.17 |
ENST00000249396.7 ENST00000414941.1 ENST00000392081.2 |
SIRT2 |
sirtuin 2 |
| chr19_-_39390350 | 0.16 |
ENST00000447739.1 ENST00000358931.5 ENST00000407552.1 |
SIRT2 |
sirtuin 2 |
| chrX_+_19362011 | 0.16 |
ENST00000379806.5 ENST00000545074.1 ENST00000540249.1 ENST00000423505.1 ENST00000417819.1 ENST00000422285.2 ENST00000355808.5 ENST00000379805.3 |
PDHA1 |
pyruvate dehydrogenase (lipoamide) alpha 1 |
| chr9_-_35665165 | 0.15 |
ENST00000343259.3 ENST00000378387.3 |
ARHGEF39 |
Rho guanine nucleotide exchange factor (GEF) 39 |
| chr6_+_30297306 | 0.15 |
ENST00000420746.1 ENST00000513556.1 |
TRIM39 TRIM39-RPP21 |
tripartite motif containing 39 TRIM39-RPP21 readthrough |
| chr3_-_183735731 | 0.15 |
ENST00000334444.6 |
ABCC5 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr14_+_24701628 | 0.14 |
ENST00000355299.4 ENST00000559836.1 |
GMPR2 |
guanosine monophosphate reductase 2 |
| chr2_-_219134822 | 0.14 |
ENST00000444053.1 ENST00000248450.4 |
AAMP |
angio-associated, migratory cell protein |
| chr22_-_38245304 | 0.13 |
ENST00000609454.1 |
ANKRD54 |
ankyrin repeat domain 54 |
| chr2_+_109271481 | 0.13 |
ENST00000542845.1 ENST00000393314.2 |
LIMS1 |
LIM and senescent cell antigen-like domains 1 |
| chr10_+_112836779 | 0.13 |
ENST00000280155.2 |
ADRA2A |
adrenoceptor alpha 2A |
| chrX_+_16141667 | 0.13 |
ENST00000380289.2 |
GRPR |
gastrin-releasing peptide receptor |
| chr1_-_8000872 | 0.13 |
ENST00000377507.3 |
TNFRSF9 |
tumor necrosis factor receptor superfamily, member 9 |
| chr19_+_50706866 | 0.12 |
ENST00000440075.2 ENST00000376970.2 ENST00000425460.1 ENST00000599920.1 ENST00000601313.1 |
MYH14 |
myosin, heavy chain 14, non-muscle |
| chr1_+_36690011 | 0.12 |
ENST00000354618.5 ENST00000469141.2 ENST00000478853.1 |
THRAP3 |
thyroid hormone receptor associated protein 3 |
| chr8_-_101348408 | 0.12 |
ENST00000519527.1 ENST00000522369.1 |
RNF19A |
ring finger protein 19A, RBR E3 ubiquitin protein ligase |
| chr11_-_64512803 | 0.12 |
ENST00000377489.1 ENST00000354024.3 |
RASGRP2 |
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr5_+_137514687 | 0.11 |
ENST00000394894.3 |
KIF20A |
kinesin family member 20A |
| chr14_-_24701539 | 0.11 |
ENST00000534348.1 ENST00000524927.1 ENST00000250495.5 |
NEDD8-MDP1 NEDD8 |
NEDD8-MDP1 readthrough neural precursor cell expressed, developmentally down-regulated 8 |
| chr3_+_171561127 | 0.11 |
ENST00000334567.5 ENST00000450693.1 |
TMEM212 |
transmembrane protein 212 |
| chr11_+_118478313 | 0.11 |
ENST00000356063.5 |
PHLDB1 |
pleckstrin homology-like domain, family B, member 1 |
| chr10_+_115439630 | 0.11 |
ENST00000369318.3 |
CASP7 |
caspase 7, apoptosis-related cysteine peptidase |
| chr19_+_42746927 | 0.11 |
ENST00000378108.1 |
AC006486.1 |
AC006486.1 |
| chrX_-_40594755 | 0.11 |
ENST00000324817.1 |
MED14 |
mediator complex subunit 14 |
| chr6_+_20403997 | 0.10 |
ENST00000535432.1 |
E2F3 |
E2F transcription factor 3 |
| chr5_+_137514834 | 0.10 |
ENST00000508792.1 ENST00000504621.1 |
KIF20A |
kinesin family member 20A |
| chr16_+_28962128 | 0.10 |
ENST00000564978.1 ENST00000320805.4 |
NFATC2IP |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 2 interacting protein |
| chr10_-_38146510 | 0.10 |
ENST00000395867.3 |
ZNF248 |
zinc finger protein 248 |
| chr2_+_187350973 | 0.10 |
ENST00000544130.1 |
ZC3H15 |
zinc finger CCCH-type containing 15 |
| chrX_-_153236819 | 0.09 |
ENST00000354233.3 |
HCFC1 |
host cell factor C1 (VP16-accessory protein) |
| chr1_+_174769006 | 0.09 |
ENST00000489615.1 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr19_-_46088068 | 0.09 |
ENST00000263275.4 ENST00000323060.3 |
OPA3 |
optic atrophy 3 (autosomal recessive, with chorea and spastic paraplegia) |
| chr9_-_35691017 | 0.09 |
ENST00000378292.3 |
TPM2 |
tropomyosin 2 (beta) |
| chr3_-_87325728 | 0.09 |
ENST00000350375.2 |
POU1F1 |
POU class 1 homeobox 1 |
| chr1_+_150980989 | 0.09 |
ENST00000368935.1 |
PRUNE |
prune exopolyphosphatase |
| chr6_+_33176265 | 0.09 |
ENST00000374656.4 |
RING1 |
ring finger protein 1 |
| chr3_+_107241783 | 0.09 |
ENST00000415149.2 ENST00000402543.1 ENST00000325805.8 ENST00000427402.1 |
BBX |
bobby sox homolog (Drosophila) |
| chr5_-_137514288 | 0.09 |
ENST00000454473.1 ENST00000418329.1 ENST00000455658.2 ENST00000230901.5 ENST00000402931.1 |
BRD8 |
bromodomain containing 8 |
| chr2_+_204193101 | 0.08 |
ENST00000430418.1 ENST00000424558.1 ENST00000261016.6 |
ABI2 |
abl-interactor 2 |
| chr17_-_39637392 | 0.08 |
ENST00000246639.2 ENST00000393989.1 |
KRT35 |
keratin 35 |
| chr16_-_57831914 | 0.08 |
ENST00000421376.2 |
KIFC3 |
kinesin family member C3 |
| chr11_-_64512469 | 0.08 |
ENST00000377485.1 |
RASGRP2 |
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr8_+_107738240 | 0.07 |
ENST00000449762.2 ENST00000297447.6 |
OXR1 |
oxidation resistance 1 |
| chr3_+_155860751 | 0.07 |
ENST00000471742.1 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr10_+_115439699 | 0.07 |
ENST00000369315.1 |
CASP7 |
caspase 7, apoptosis-related cysteine peptidase |
| chr2_+_106361333 | 0.07 |
ENST00000233154.4 ENST00000451463.2 |
NCK2 |
NCK adaptor protein 2 |
| chr1_+_150980889 | 0.07 |
ENST00000450884.1 ENST00000271620.3 ENST00000271619.8 ENST00000368937.1 ENST00000431193.1 ENST00000368936.1 |
PRUNE |
prune exopolyphosphatase |
| chr6_-_31510181 | 0.07 |
ENST00000458640.1 ENST00000396172.1 ENST00000417556.2 |
DDX39B |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39B |
| chr19_+_39390320 | 0.07 |
ENST00000576510.1 |
NFKBIB |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr19_-_36870087 | 0.06 |
ENST00000270001.7 |
ZFP14 |
ZFP14 zinc finger protein |
| chr11_-_64512273 | 0.06 |
ENST00000377497.3 ENST00000377487.1 ENST00000430645.1 |
RASGRP2 |
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr16_+_30075595 | 0.06 |
ENST00000563060.2 |
ALDOA |
aldolase A, fructose-bisphosphate |
| chr4_-_73935409 | 0.06 |
ENST00000507544.2 ENST00000295890.4 |
COX18 |
COX18 cytochrome C oxidase assembly factor |
| chr5_+_178322893 | 0.06 |
ENST00000361362.2 ENST00000520660.1 ENST00000520805.1 |
ZFP2 |
ZFP2 zinc finger protein |
| chr10_-_33623310 | 0.06 |
ENST00000395995.1 ENST00000374823.5 ENST00000374821.5 ENST00000374816.3 |
NRP1 |
neuropilin 1 |
| chr16_+_30075463 | 0.06 |
ENST00000562168.1 ENST00000569545.1 |
ALDOA |
aldolase A, fructose-bisphosphate |
| chr19_+_39390587 | 0.06 |
ENST00000572515.1 ENST00000392079.3 ENST00000575359.1 ENST00000313582.5 |
NFKBIB |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr17_-_34257731 | 0.05 |
ENST00000431884.2 ENST00000425909.3 ENST00000394528.3 ENST00000430160.2 |
RDM1 |
RAD52 motif 1 |
| chr1_+_8021954 | 0.05 |
ENST00000377491.1 ENST00000377488.1 |
PARK7 |
parkinson protein 7 |
| chr4_+_26321284 | 0.05 |
ENST00000506956.1 ENST00000512671.1 ENST00000345843.3 ENST00000342295.1 |
RBPJ |
recombination signal binding protein for immunoglobulin kappa J region |
| chr2_+_183580954 | 0.05 |
ENST00000264065.7 |
DNAJC10 |
DnaJ (Hsp40) homolog, subfamily C, member 10 |
| chr5_-_137514333 | 0.05 |
ENST00000411594.2 ENST00000430331.1 |
BRD8 |
bromodomain containing 8 |
| chr1_-_43855444 | 0.05 |
ENST00000372455.4 |
MED8 |
mediator complex subunit 8 |
| chrX_-_153236620 | 0.05 |
ENST00000369984.4 |
HCFC1 |
host cell factor C1 (VP16-accessory protein) |
| chrX_+_123095546 | 0.04 |
ENST00000371157.3 ENST00000371145.3 ENST00000371144.3 |
STAG2 |
stromal antigen 2 |
| chr1_+_93297582 | 0.04 |
ENST00000370321.3 |
RPL5 |
ribosomal protein L5 |
| chr6_-_76203345 | 0.04 |
ENST00000393004.2 |
FILIP1 |
filamin A interacting protein 1 |
| chr13_-_39612176 | 0.04 |
ENST00000352251.3 ENST00000350125.3 |
PROSER1 |
proline and serine rich 1 |
| chr3_+_32726774 | 0.03 |
ENST00000538368.1 |
CNOT10 |
CCR4-NOT transcription complex, subunit 10 |
| chr14_-_53258180 | 0.03 |
ENST00000554230.1 |
GNPNAT1 |
glucosamine-phosphate N-acetyltransferase 1 |
| chr2_-_79315112 | 0.03 |
ENST00000305089.3 |
REG1B |
regenerating islet-derived 1 beta |
| chr14_-_53258314 | 0.03 |
ENST00000216410.3 ENST00000557604.1 |
GNPNAT1 |
glucosamine-phosphate N-acetyltransferase 1 |
| chr5_+_178368186 | 0.03 |
ENST00000320129.3 ENST00000519564.1 |
ZNF454 |
zinc finger protein 454 |
| chr19_+_50528971 | 0.03 |
ENST00000598809.1 ENST00000595661.1 ENST00000391821.2 |
ZNF473 |
zinc finger protein 473 |
| chr16_+_21964662 | 0.03 |
ENST00000561553.1 ENST00000565331.1 |
UQCRC2 |
ubiquinol-cytochrome c reductase core protein II |
| chr15_-_70390191 | 0.03 |
ENST00000559191.1 |
TLE3 |
transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) |
| chr3_-_87325612 | 0.03 |
ENST00000561167.1 ENST00000560656.1 ENST00000344265.3 |
POU1F1 |
POU class 1 homeobox 1 |
| chr8_-_67974552 | 0.02 |
ENST00000357849.4 |
COPS5 |
COP9 signalosome subunit 5 |
| chr11_-_8959758 | 0.02 |
ENST00000531618.1 |
ASCL3 |
achaete-scute family bHLH transcription factor 3 |
| chr2_+_187350883 | 0.01 |
ENST00000337859.6 |
ZC3H15 |
zinc finger CCCH-type containing 15 |
| chr19_-_17356697 | 0.01 |
ENST00000291442.3 |
NR2F6 |
nuclear receptor subfamily 2, group F, member 6 |
| chr10_+_118305435 | 0.01 |
ENST00000369221.2 |
PNLIP |
pancreatic lipase |
| chr5_+_132009675 | 0.01 |
ENST00000231449.2 ENST00000350025.2 |
IL4 |
interleukin 4 |
| chr3_+_193853927 | 0.01 |
ENST00000232424.3 |
HES1 |
hes family bHLH transcription factor 1 |
| chr1_+_93297622 | 0.01 |
ENST00000315741.5 |
RPL5 |
ribosomal protein L5 |
| chr11_-_101454658 | 0.01 |
ENST00000344327.3 |
TRPC6 |
transient receptor potential cation channel, subfamily C, member 6 |
| chr17_-_16118835 | 0.01 |
ENST00000582357.1 ENST00000436828.1 ENST00000411510.1 ENST00000268712.3 |
NCOR1 |
nuclear receptor corepressor 1 |
| chr6_+_42531798 | 0.01 |
ENST00000372903.2 ENST00000372899.1 ENST00000372901.1 |
UBR2 |
ubiquitin protein ligase E3 component n-recognin 2 |
| chr2_+_219824357 | 0.01 |
ENST00000302625.4 |
CDK5R2 |
cyclin-dependent kinase 5, regulatory subunit 2 (p39) |
| chr1_+_43855560 | 0.01 |
ENST00000562955.1 |
SZT2 |
seizure threshold 2 homolog (mouse) |
| chr21_-_31538971 | 0.01 |
ENST00000286808.3 |
CLDN17 |
claudin 17 |
| chr12_+_175930 | 0.01 |
ENST00000538872.1 ENST00000326261.4 |
IQSEC3 |
IQ motif and Sec7 domain 3 |
| chr7_+_76090993 | 0.01 |
ENST00000425780.1 ENST00000456590.1 ENST00000451769.1 ENST00000324432.5 ENST00000307569.8 ENST00000457529.1 ENST00000446600.1 ENST00000413936.2 ENST00000423646.1 ENST00000438930.1 ENST00000430490.2 |
DTX2 |
deltex homolog 2 (Drosophila) |
| chr15_-_70390213 | 0.01 |
ENST00000557997.1 ENST00000317509.8 ENST00000442299.2 |
TLE3 |
transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) |
| chr12_-_4758159 | 0.00 |
ENST00000545990.2 |
AKAP3 |
A kinase (PRKA) anchor protein 3 |
| chr5_-_88119580 | 0.00 |
ENST00000539796.1 |
MEF2C |
myocyte enhancer factor 2C |
| chr14_+_22337014 | 0.00 |
ENST00000390436.2 |
TRAV13-1 |
T cell receptor alpha variable 13-1 |
Gene Ontology Analysis
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.2 | 0.5 | GO:0097489 | multivesicular body, internal vesicle lumen(GO:0097489) |
| 0.1 | 1.5 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.1 | 1.4 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 1.0 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 0.4 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.0 | 0.3 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.4 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.0 | 0.1 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.0 | 0.1 | GO:1902560 | GMP reductase complex(GO:1902560) |
| 0.0 | 0.1 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.0 | 0.2 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 0.0 | 0.5 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.0 | 0.4 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.1 | GO:0045180 | basal cortex(GO:0045180) |
| 0.0 | 0.1 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.1 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.0 | 0.1 | GO:1990635 | proximal dendrite(GO:1990635) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 0.7 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:2000393 | negative regulation of lamellipodium morphogenesis(GO:2000393) |
| 0.2 | 1.0 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.2 | 0.5 | GO:0043006 | activation of phospholipase A2 activity by calcium-mediated signaling(GO:0043006) |
| 0.1 | 1.5 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.1 | 0.7 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.1 | 0.5 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.1 | 0.3 | GO:2000777 | positive regulation of oocyte maturation(GO:1900195) negative regulation of NLRP3 inflammasome complex assembly(GO:1900226) positive regulation of proteasomal ubiquitin-dependent protein catabolic process involved in cellular response to hypoxia(GO:2000777) |
| 0.1 | 0.4 | GO:2000820 | negative regulation of transcription from RNA polymerase II promoter involved in smooth muscle cell differentiation(GO:2000820) |
| 0.1 | 1.4 | GO:0006853 | carnitine shuttle(GO:0006853) |
| 0.1 | 0.2 | GO:0060010 | Sertoli cell fate commitment(GO:0060010) |
| 0.0 | 0.4 | GO:0045602 | negative regulation of endothelial cell differentiation(GO:0045602) |
| 0.0 | 0.1 | GO:0019046 | release from viral latency(GO:0019046) |
| 0.0 | 0.1 | GO:0035625 | negative regulation of epinephrine secretion(GO:0032811) epidermal growth factor-activated receptor transactivation by G-protein coupled receptor signaling pathway(GO:0035625) |
| 0.0 | 0.1 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 0.0 | 0.5 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.0 | 0.2 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.0 | 0.3 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 0.0 | 0.1 | GO:0060133 | somatotropin secreting cell development(GO:0060133) |
| 0.0 | 0.2 | GO:0006552 | leucine catabolic process(GO:0006552) |
| 0.0 | 0.3 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.0 | 0.2 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 0.0 | 0.1 | GO:2001183 | negative regulation of interleukin-10 secretion(GO:2001180) negative regulation of interleukin-12 secretion(GO:2001183) |
| 0.0 | 0.9 | GO:0097178 | ruffle assembly(GO:0097178) |
| 0.0 | 0.5 | GO:0033622 | integrin activation(GO:0033622) |
| 0.0 | 0.1 | GO:1902958 | peptidyl-cysteine oxidation(GO:0018171) enzyme active site formation(GO:0018307) neuron intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:0036482) detoxification of mercury ion(GO:0050787) positive regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902958) regulation of protein K48-linked deubiquitination(GO:1903093) negative regulation of protein K48-linked deubiquitination(GO:1903094) regulation of dopamine biosynthetic process(GO:1903179) positive regulation of dopamine biosynthetic process(GO:1903181) regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903383) negative regulation of hydrogen peroxide-induced neuron intrinsic apoptotic signaling pathway(GO:1903384) negative regulation of ubiquitin-specific protease activity(GO:2000157) |
| 0.0 | 0.2 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 0.0 | 0.1 | GO:0072554 | arterial endothelial cell fate commitment(GO:0060844) blood vessel lumenization(GO:0072554) positive regulation of ephrin receptor signaling pathway(GO:1901189) positive regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901297) positive regulation of canonical Wnt signaling pathway involved in heart development(GO:1905068) |
| 0.0 | 0.7 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.1 | GO:2000435 | regulation of protein neddylation(GO:2000434) negative regulation of protein neddylation(GO:2000435) |
| 0.0 | 0.3 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.0 | 0.1 | GO:0006041 | glucosamine metabolic process(GO:0006041) |
| 0.0 | 0.1 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.0 | 0.2 | GO:0000920 | cell separation after cytokinesis(GO:0000920) |
| 0.0 | 0.1 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.0 | 0.1 | GO:0036493 | positive regulation of translation in response to endoplasmic reticulum stress(GO:0036493) |
| 0.0 | 0.1 | GO:1904261 | regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904259) positive regulation of basement membrane assembly involved in embryonic body morphogenesis(GO:1904261) basement membrane assembly involved in embryonic body morphogenesis(GO:2001197) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0034739 | histone deacetylase activity (H4-K16 specific)(GO:0034739) |
| 0.1 | 1.4 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.1 | 0.5 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.1 | 0.3 | GO:0035939 | microsatellite binding(GO:0035939) |
| 0.1 | 1.5 | GO:0038187 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.0 | 0.7 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 0.1 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) |
| 0.0 | 0.2 | GO:0004739 | pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.0 | 1.0 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.5 | GO:0016641 | oxidoreductase activity, acting on the CH-NH2 group of donors, oxygen as acceptor(GO:0016641) |
| 0.0 | 0.2 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.0 | 0.1 | GO:0016657 | GMP reductase activity(GO:0003920) oxidoreductase activity, acting on NAD(P)H, nitrogenous group as acceptor(GO:0016657) |
| 0.0 | 0.3 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 0.3 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.0 | 0.4 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.0 | 0.9 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 0.2 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.0 | 0.3 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 0.3 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 0.2 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.0 | 0.1 | GO:0016532 | superoxide dismutase copper chaperone activity(GO:0016532) cupric ion binding(GO:1903135) |
| 0.0 | 0.3 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.0 | 0.1 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.0 | 0.1 | GO:0046972 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
| 0.0 | 0.5 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.0 | 0.1 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.0 | 1.0 | GO:0001618 | virus receptor activity(GO:0001618) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 0.7 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.5 | REACTOME SHC1 EVENTS IN EGFR SIGNALING | Genes involved in SHC1 events in EGFR signaling |
| 0.0 | 0.6 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.0 | 0.5 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.9 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |