Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
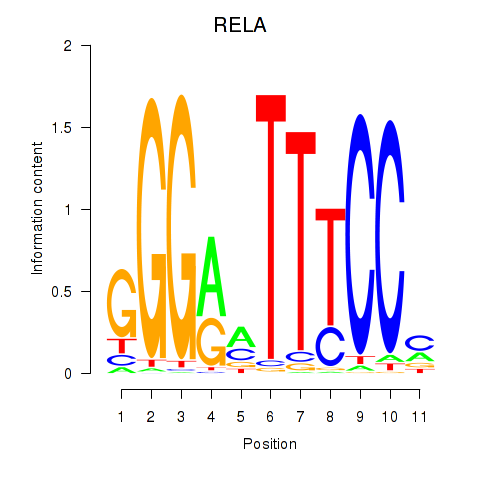
Results for RELA
Z-value: 1.33
Transcription factors associated with RELA
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RELA
|
ENSG00000173039.14 | RELA |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RELA | hg19_v2_chr11_-_65430251_65430399 | 0.19 | 4.8e-01 | Click! |
Activity profile of RELA motif
Sorted Z-values of RELA motif
Network of associatons between targets according to the STRING database.
First level regulatory network of RELA
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_149792295 | 5.93 |
ENST00000518797.1 ENST00000524315.1 ENST00000009530.7 ENST00000377795.3 |
CD74 |
CD74 molecule, major histocompatibility complex, class II invariant chain |
| chr11_+_102188272 | 5.47 |
ENST00000532808.1 |
BIRC3 |
baculoviral IAP repeat containing 3 |
| chr11_+_102188224 | 5.24 |
ENST00000263464.3 |
BIRC3 |
baculoviral IAP repeat containing 3 |
| chr12_-_9913489 | 5.10 |
ENST00000228434.3 ENST00000536709.1 |
CD69 |
CD69 molecule |
| chr14_+_75988851 | 5.09 |
ENST00000555504.1 |
BATF |
basic leucine zipper transcription factor, ATF-like |
| chr19_-_6591113 | 4.89 |
ENST00000423145.3 ENST00000245903.3 |
CD70 |
CD70 molecule |
| chr6_+_138188551 | 4.68 |
ENST00000237289.4 ENST00000433680.1 |
TNFAIP3 |
tumor necrosis factor, alpha-induced protein 3 |
| chr22_-_37545972 | 4.40 |
ENST00000216223.5 |
IL2RB |
interleukin 2 receptor, beta |
| chr6_+_32605134 | 4.37 |
ENST00000343139.5 ENST00000395363.1 ENST00000496318.1 |
HLA-DQA1 |
major histocompatibility complex, class II, DQ alpha 1 |
| chr6_+_32605195 | 4.34 |
ENST00000374949.2 |
HLA-DQA1 |
major histocompatibility complex, class II, DQ alpha 1 |
| chr11_-_58345569 | 3.97 |
ENST00000528954.1 ENST00000528489.1 |
LPXN |
leupaxin |
| chr14_-_35873856 | 3.76 |
ENST00000553342.1 ENST00000216797.5 ENST00000557140.1 |
NFKBIA |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha |
| chr20_+_44746885 | 3.35 |
ENST00000372285.3 |
CD40 |
CD40 molecule, TNF receptor superfamily member 5 |
| chr7_+_69064300 | 3.08 |
ENST00000342771.4 |
AUTS2 |
autism susceptibility candidate 2 |
| chr4_-_76944621 | 2.85 |
ENST00000306602.1 |
CXCL10 |
chemokine (C-X-C motif) ligand 10 |
| chr6_-_32636145 | 2.80 |
ENST00000399084.1 |
HLA-DQB1 |
major histocompatibility complex, class II, DQ beta 1 |
| chr3_-_158390282 | 2.70 |
ENST00000264265.3 |
LXN |
latexin |
| chr19_+_4229495 | 2.65 |
ENST00000221847.5 |
EBI3 |
Epstein-Barr virus induced 3 |
| chr4_+_103422471 | 2.61 |
ENST00000226574.4 ENST00000394820.4 |
NFKB1 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 |
| chr22_-_37640277 | 2.60 |
ENST00000401529.3 ENST00000249071.6 |
RAC2 |
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr6_-_31550192 | 2.57 |
ENST00000429299.2 ENST00000446745.2 |
LTB |
lymphotoxin beta (TNF superfamily, member 3) |
| chr4_-_40631859 | 2.56 |
ENST00000295971.7 ENST00000319592.4 |
RBM47 |
RNA binding motif protein 47 |
| chr10_+_30722866 | 2.44 |
ENST00000263056.1 |
MAP3K8 |
mitogen-activated protein kinase kinase kinase 8 |
| chr20_-_4795747 | 2.41 |
ENST00000379376.2 |
RASSF2 |
Ras association (RalGDS/AF-6) domain family member 2 |
| chr14_+_75988768 | 2.40 |
ENST00000286639.6 |
BATF |
basic leucine zipper transcription factor, ATF-like |
| chr2_-_191885686 | 2.39 |
ENST00000432058.1 |
STAT1 |
signal transducer and activator of transcription 1, 91kDa |
| chr22_-_37640456 | 2.25 |
ENST00000405484.1 ENST00000441619.1 ENST00000406508.1 |
RAC2 |
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr16_+_50776021 | 2.20 |
ENST00000566679.2 ENST00000564634.1 ENST00000398568.2 |
CYLD |
cylindromatosis (turban tumor syndrome) |
| chr14_+_103243813 | 2.10 |
ENST00000560371.1 ENST00000347662.4 ENST00000392745.2 ENST00000539721.1 ENST00000560463.1 |
TRAF3 |
TNF receptor-associated factor 3 |
| chr6_+_29691198 | 2.09 |
ENST00000440587.2 ENST00000434407.2 |
HLA-F |
major histocompatibility complex, class I, F |
| chr6_+_31540056 | 2.07 |
ENST00000418386.2 |
LTA |
lymphotoxin alpha |
| chr12_+_7055631 | 2.04 |
ENST00000543115.1 ENST00000399448.1 |
PTPN6 |
protein tyrosine phosphatase, non-receptor type 6 |
| chr7_-_24797032 | 2.03 |
ENST00000409970.1 ENST00000409775.3 |
DFNA5 |
deafness, autosomal dominant 5 |
| chrX_-_73072534 | 2.00 |
ENST00000429829.1 |
XIST |
X inactive specific transcript (non-protein coding) |
| chr14_+_61789382 | 1.97 |
ENST00000555082.1 |
PRKCH |
protein kinase C, eta |
| chr6_-_29527702 | 1.95 |
ENST00000377050.4 |
UBD |
ubiquitin D |
| chr6_+_29691056 | 1.87 |
ENST00000414333.1 ENST00000334668.4 ENST00000259951.7 |
HLA-F |
major histocompatibility complex, class I, F |
| chr10_+_12391481 | 1.87 |
ENST00000378847.3 |
CAMK1D |
calcium/calmodulin-dependent protein kinase ID |
| chr19_+_42381173 | 1.85 |
ENST00000221972.3 |
CD79A |
CD79a molecule, immunoglobulin-associated alpha |
| chr7_-_24797546 | 1.85 |
ENST00000414428.1 ENST00000419307.1 ENST00000342947.3 |
DFNA5 |
deafness, autosomal dominant 5 |
| chr1_+_111770278 | 1.80 |
ENST00000369748.4 |
CHI3L2 |
chitinase 3-like 2 |
| chr1_+_111770232 | 1.78 |
ENST00000369744.2 |
CHI3L2 |
chitinase 3-like 2 |
| chr19_+_42381337 | 1.77 |
ENST00000597454.1 ENST00000444740.2 |
CD79A |
CD79a molecule, immunoglobulin-associated alpha |
| chr6_+_29910301 | 1.77 |
ENST00000376809.5 ENST00000376802.2 |
HLA-A |
major histocompatibility complex, class I, A |
| chr19_+_2476116 | 1.74 |
ENST00000215631.4 ENST00000587345.1 |
GADD45B |
growth arrest and DNA-damage-inducible, beta |
| chr19_-_7766991 | 1.73 |
ENST00000597921.1 ENST00000346664.5 |
FCER2 |
Fc fragment of IgE, low affinity II, receptor for (CD23) |
| chr19_-_11688447 | 1.72 |
ENST00000590420.1 |
ACP5 |
acid phosphatase 5, tartrate resistant |
| chr2_+_61108650 | 1.67 |
ENST00000295025.8 |
REL |
v-rel avian reticuloendotheliosis viral oncogene homolog |
| chr2_-_163175133 | 1.64 |
ENST00000421365.2 ENST00000263642.2 |
IFIH1 |
interferon induced with helicase C domain 1 |
| chr20_+_9494987 | 1.55 |
ENST00000427562.2 ENST00000246070.2 |
LAMP5 |
lysosomal-associated membrane protein family, member 5 |
| chr10_+_104155450 | 1.54 |
ENST00000471698.1 ENST00000189444.6 |
NFKB2 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) |
| chr4_-_76928641 | 1.53 |
ENST00000264888.5 |
CXCL9 |
chemokine (C-X-C motif) ligand 9 |
| chr16_+_50775971 | 1.51 |
ENST00000311559.9 ENST00000564326.1 ENST00000566206.1 |
CYLD |
cylindromatosis (turban tumor syndrome) |
| chr5_-_150460539 | 1.48 |
ENST00000520931.1 ENST00000520695.1 ENST00000521591.1 ENST00000518977.1 |
TNIP1 |
TNFAIP3 interacting protein 1 |
| chr19_-_11688500 | 1.47 |
ENST00000433365.2 |
ACP5 |
acid phosphatase 5, tartrate resistant |
| chr17_-_34207295 | 1.45 |
ENST00000463941.1 ENST00000293272.3 |
CCL5 |
chemokine (C-C motif) ligand 5 |
| chr16_+_50730910 | 1.40 |
ENST00000300589.2 |
NOD2 |
nucleotide-binding oligomerization domain containing 2 |
| chr5_-_150460914 | 1.35 |
ENST00000389378.2 |
TNIP1 |
TNFAIP3 interacting protein 1 |
| chr15_+_85923797 | 1.33 |
ENST00000559362.1 |
AKAP13 |
A kinase (PRKA) anchor protein 13 |
| chr1_+_156123359 | 1.30 |
ENST00000368284.1 ENST00000368286.2 ENST00000438830.1 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr3_-_4793274 | 1.30 |
ENST00000414938.1 |
EGOT |
eosinophil granule ontogeny transcript (non-protein coding) |
| chr22_-_39268308 | 1.28 |
ENST00000407418.3 |
CBX6 |
chromobox homolog 6 |
| chr19_+_45504688 | 1.28 |
ENST00000221452.8 ENST00000540120.1 ENST00000505236.1 |
RELB |
v-rel avian reticuloendotheliosis viral oncogene homolog B |
| chr3_+_53195136 | 1.28 |
ENST00000394729.2 ENST00000330452.3 |
PRKCD |
protein kinase C, delta |
| chr1_+_156123318 | 1.28 |
ENST00000368285.3 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr1_-_209825674 | 1.26 |
ENST00000367030.3 ENST00000356082.4 |
LAMB3 |
laminin, beta 3 |
| chr2_+_61108771 | 1.21 |
ENST00000394479.3 |
REL |
v-rel avian reticuloendotheliosis viral oncogene homolog |
| chr17_-_7590745 | 1.21 |
ENST00000514944.1 ENST00000503591.1 ENST00000455263.2 ENST00000420246.2 ENST00000445888.2 ENST00000509690.1 ENST00000604348.1 ENST00000269305.4 |
TP53 |
tumor protein p53 |
| chr6_+_106534192 | 1.20 |
ENST00000369091.2 ENST00000369096.4 |
PRDM1 |
PR domain containing 1, with ZNF domain |
| chr6_+_135502466 | 1.14 |
ENST00000367814.4 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr22_-_39268192 | 1.07 |
ENST00000216083.6 |
CBX6 |
chromobox homolog 6 |
| chr8_+_38758737 | 1.06 |
ENST00000521746.1 ENST00000420274.1 |
PLEKHA2 |
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 2 |
| chr5_-_150466692 | 1.05 |
ENST00000315050.7 ENST00000523338.1 ENST00000522100.1 |
TNIP1 |
TNFAIP3 interacting protein 1 |
| chr16_+_50775948 | 1.02 |
ENST00000569681.1 ENST00000569418.1 ENST00000540145.1 |
CYLD |
cylindromatosis (turban tumor syndrome) |
| chr3_+_57261743 | 0.99 |
ENST00000288266.3 |
APPL1 |
adaptor protein, phosphotyrosine interaction, PH domain and leucine zipper containing 1 |
| chr15_+_85923856 | 0.94 |
ENST00000560302.1 ENST00000394518.2 ENST00000361243.2 ENST00000560256.1 |
AKAP13 |
A kinase (PRKA) anchor protein 13 |
| chr12_+_11802753 | 0.93 |
ENST00000396373.4 |
ETV6 |
ets variant 6 |
| chr10_+_104154229 | 0.91 |
ENST00000428099.1 ENST00000369966.3 |
NFKB2 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) |
| chr9_-_136344197 | 0.91 |
ENST00000414172.1 ENST00000371897.4 |
SLC2A6 |
solute carrier family 2 (facilitated glucose transporter), member 6 |
| chr6_+_292051 | 0.88 |
ENST00000344450.5 |
DUSP22 |
dual specificity phosphatase 22 |
| chrX_-_30595959 | 0.86 |
ENST00000378962.3 |
CXorf21 |
chromosome X open reading frame 21 |
| chr1_-_205744574 | 0.81 |
ENST00000367139.3 ENST00000235932.4 ENST00000437324.2 ENST00000414729.1 |
RAB7L1 |
RAB7, member RAS oncogene family-like 1 |
| chr1_+_26869597 | 0.79 |
ENST00000530003.1 |
RPS6KA1 |
ribosomal protein S6 kinase, 90kDa, polypeptide 1 |
| chr2_-_89442621 | 0.75 |
ENST00000492167.1 |
IGKV3-20 |
immunoglobulin kappa variable 3-20 |
| chr14_+_103589789 | 0.72 |
ENST00000558056.1 ENST00000560869.1 |
TNFAIP2 |
tumor necrosis factor, alpha-induced protein 2 |
| chr1_-_54304212 | 0.72 |
ENST00000540001.1 |
NDC1 |
NDC1 transmembrane nucleoporin |
| chr11_+_10476851 | 0.68 |
ENST00000396553.2 |
AMPD3 |
adenosine monophosphate deaminase 3 |
| chr1_-_54303949 | 0.66 |
ENST00000234725.8 |
NDC1 |
NDC1 transmembrane nucleoporin |
| chr4_-_185395672 | 0.64 |
ENST00000393593.3 |
IRF2 |
interferon regulatory factor 2 |
| chr19_+_10381769 | 0.63 |
ENST00000423829.2 ENST00000588645.1 |
ICAM1 |
intercellular adhesion molecule 1 |
| chr6_+_87865262 | 0.62 |
ENST00000369577.3 ENST00000518845.1 ENST00000339907.4 ENST00000496806.2 |
ZNF292 |
zinc finger protein 292 |
| chr11_-_72853091 | 0.62 |
ENST00000311172.7 ENST00000409314.1 |
FCHSD2 |
FCH and double SH3 domains 2 |
| chr2_+_97203082 | 0.60 |
ENST00000454558.2 |
ARID5A |
AT rich interactive domain 5A (MRF1-like) |
| chr1_-_202130702 | 0.60 |
ENST00000309017.3 ENST00000477554.1 ENST00000492451.1 |
PTPN7 |
protein tyrosine phosphatase, non-receptor type 7 |
| chr1_-_54303934 | 0.58 |
ENST00000537333.1 |
NDC1 |
NDC1 transmembrane nucleoporin |
| chr1_-_236030216 | 0.58 |
ENST00000389794.3 ENST00000389793.2 |
LYST |
lysosomal trafficking regulator |
| chr6_+_144471643 | 0.57 |
ENST00000367568.4 |
STX11 |
syntaxin 11 |
| chr9_+_19230433 | 0.55 |
ENST00000434457.2 ENST00000602925.1 |
DENND4C |
DENN/MADD domain containing 4C |
| chr1_+_46640750 | 0.54 |
ENST00000372003.1 |
TSPAN1 |
tetraspanin 1 |
| chr1_+_63833261 | 0.53 |
ENST00000371108.4 |
ALG6 |
ALG6, alpha-1,3-glucosyltransferase |
| chr2_+_233925064 | 0.53 |
ENST00000359570.5 ENST00000538935.1 |
INPP5D |
inositol polyphosphate-5-phosphatase, 145kDa |
| chr1_-_205744205 | 0.52 |
ENST00000446390.2 |
RAB7L1 |
RAB7, member RAS oncogene family-like 1 |
| chr19_+_41281060 | 0.52 |
ENST00000594436.1 ENST00000597784.1 |
MIA |
melanoma inhibitory activity |
| chr2_+_97202480 | 0.52 |
ENST00000357485.3 |
ARID5A |
AT rich interactive domain 5A (MRF1-like) |
| chr17_-_4852332 | 0.52 |
ENST00000572383.1 |
PFN1 |
profilin 1 |
| chr12_-_54653313 | 0.51 |
ENST00000550411.1 ENST00000439541.2 |
CBX5 |
chromobox homolog 5 |
| chr17_+_16318909 | 0.51 |
ENST00000577397.1 |
TRPV2 |
transient receptor potential cation channel, subfamily V, member 2 |
| chr2_+_208394616 | 0.50 |
ENST00000432329.2 ENST00000353267.3 ENST00000445803.1 |
CREB1 |
cAMP responsive element binding protein 1 |
| chr11_-_3862206 | 0.50 |
ENST00000351018.4 |
RHOG |
ras homolog family member G |
| chr1_+_101185290 | 0.49 |
ENST00000370119.4 ENST00000347652.2 ENST00000294728.2 ENST00000370115.1 |
VCAM1 |
vascular cell adhesion molecule 1 |
| chr17_+_16318850 | 0.48 |
ENST00000338560.7 |
TRPV2 |
transient receptor potential cation channel, subfamily V, member 2 |
| chr19_-_4831701 | 0.47 |
ENST00000248244.5 |
TICAM1 |
toll-like receptor adaptor molecule 1 |
| chr5_+_150591678 | 0.47 |
ENST00000523466.1 |
GM2A |
GM2 ganglioside activator |
| chr2_+_163175394 | 0.47 |
ENST00000446271.1 ENST00000429691.2 |
GCA |
grancalcin, EF-hand calcium binding protein |
| chr1_-_65432171 | 0.46 |
ENST00000342505.4 |
JAK1 |
Janus kinase 1 |
| chr12_+_49761273 | 0.46 |
ENST00000551540.1 ENST00000552918.1 ENST00000548777.1 ENST00000547865.1 ENST00000552171.1 |
SPATS2 |
spermatogenesis associated, serine-rich 2 |
| chr19_-_51472031 | 0.45 |
ENST00000391808.1 |
KLK6 |
kallikrein-related peptidase 6 |
| chr4_+_114214125 | 0.44 |
ENST00000509550.1 |
ANK2 |
ankyrin 2, neuronal |
| chr1_-_151319710 | 0.43 |
ENST00000290524.4 ENST00000437327.1 ENST00000452513.2 ENST00000368870.2 ENST00000452671.2 |
RFX5 |
regulatory factor X, 5 (influences HLA class II expression) |
| chr7_-_93520259 | 0.42 |
ENST00000222543.5 |
TFPI2 |
tissue factor pathway inhibitor 2 |
| chr12_+_49761224 | 0.42 |
ENST00000553127.1 ENST00000321898.6 |
SPATS2 |
spermatogenesis associated, serine-rich 2 |
| chr1_-_209824643 | 0.40 |
ENST00000391911.1 ENST00000415782.1 |
LAMB3 |
laminin, beta 3 |
| chr7_-_93520191 | 0.40 |
ENST00000545378.1 |
TFPI2 |
tissue factor pathway inhibitor 2 |
| chr15_+_57884086 | 0.39 |
ENST00000380569.2 ENST00000380561.2 ENST00000574161.1 ENST00000572390.1 ENST00000396180.1 ENST00000380560.2 |
GCOM1 |
GRINL1A complex locus 1 |
| chr19_+_45251804 | 0.38 |
ENST00000164227.5 |
BCL3 |
B-cell CLL/lymphoma 3 |
| chr4_-_174256276 | 0.37 |
ENST00000296503.5 |
HMGB2 |
high mobility group box 2 |
| chr12_+_5019061 | 0.37 |
ENST00000382545.3 |
KCNA1 |
potassium voltage-gated channel, shaker-related subfamily, member 1 (episodic ataxia with myokymia) |
| chr22_-_28315115 | 0.36 |
ENST00000455418.3 ENST00000436663.1 ENST00000320996.10 ENST00000335272.5 |
PITPNB |
phosphatidylinositol transfer protein, beta |
| chr20_+_18488528 | 0.36 |
ENST00000377465.1 |
SEC23B |
Sec23 homolog B (S. cerevisiae) |
| chr8_-_70747205 | 0.36 |
ENST00000260126.4 |
SLCO5A1 |
solute carrier organic anion transporter family, member 5A1 |
| chr15_+_57884199 | 0.36 |
ENST00000587652.1 ENST00000380568.3 ENST00000380565.4 ENST00000380563.2 |
GCOM1 MYZAP POLR2M |
GRINL1A complex locus 1 myocardial zonula adherens protein polymerase (RNA) II (DNA directed) polypeptide M |
| chr18_-_12884259 | 0.36 |
ENST00000353319.4 ENST00000327283.3 |
PTPN2 |
protein tyrosine phosphatase, non-receptor type 2 |
| chr12_-_56727487 | 0.35 |
ENST00000548043.1 ENST00000425394.2 |
PAN2 |
PAN2 poly(A) specific ribonuclease subunit homolog (S. cerevisiae) |
| chr12_-_28124903 | 0.35 |
ENST00000395872.1 ENST00000354417.3 ENST00000201015.4 |
PTHLH |
parathyroid hormone-like hormone |
| chr10_+_12391685 | 0.34 |
ENST00000378845.1 |
CAMK1D |
calcium/calmodulin-dependent protein kinase ID |
| chr3_-_49131473 | 0.34 |
ENST00000430979.1 ENST00000357496.2 ENST00000437939.1 |
QRICH1 |
glutamine-rich 1 |
| chr2_+_208394658 | 0.33 |
ENST00000421139.1 |
CREB1 |
cAMP responsive element binding protein 1 |
| chr5_+_133984462 | 0.33 |
ENST00000398844.2 ENST00000322887.4 |
SEC24A |
SEC24 family member A |
| chr6_-_30712313 | 0.32 |
ENST00000376377.2 ENST00000259874.5 |
IER3 |
immediate early response 3 |
| chr11_-_790060 | 0.32 |
ENST00000330106.4 |
CEND1 |
cell cycle exit and neuronal differentiation 1 |
| chr4_-_151936416 | 0.31 |
ENST00000510413.1 ENST00000507224.1 |
LRBA |
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr1_-_209979375 | 0.31 |
ENST00000367021.3 |
IRF6 |
interferon regulatory factor 6 |
| chr2_+_208394455 | 0.30 |
ENST00000430624.1 |
CREB1 |
cAMP responsive element binding protein 1 |
| chr2_+_44396000 | 0.29 |
ENST00000409895.4 ENST00000409432.3 ENST00000282412.4 ENST00000378551.2 ENST00000345249.4 |
PPM1B |
protein phosphatase, Mg2+/Mn2+ dependent, 1B |
| chr13_-_52027134 | 0.29 |
ENST00000311234.4 ENST00000425000.1 ENST00000463928.1 ENST00000442263.3 ENST00000398119.2 |
INTS6 |
integrator complex subunit 6 |
| chr1_-_41950342 | 0.29 |
ENST00000372587.4 |
EDN2 |
endothelin 2 |
| chr21_+_34775181 | 0.27 |
ENST00000290219.6 |
IFNGR2 |
interferon gamma receptor 2 (interferon gamma transducer 1) |
| chr5_+_140787600 | 0.27 |
ENST00000520790.1 |
PCDHGB6 |
protocadherin gamma subfamily B, 6 |
| chr12_+_53662073 | 0.27 |
ENST00000553219.1 ENST00000257934.4 |
ESPL1 |
extra spindle pole bodies homolog 1 (S. cerevisiae) |
| chr12_-_49259643 | 0.27 |
ENST00000309739.5 |
RND1 |
Rho family GTPase 1 |
| chr11_-_128392085 | 0.25 |
ENST00000526145.2 ENST00000531611.1 ENST00000319397.6 ENST00000345075.4 ENST00000535549.1 |
ETS1 |
v-ets avian erythroblastosis virus E26 oncogene homolog 1 |
| chr1_-_202129105 | 0.25 |
ENST00000367279.4 |
PTPN7 |
protein tyrosine phosphatase, non-receptor type 7 |
| chr10_+_13141585 | 0.25 |
ENST00000378764.2 |
OPTN |
optineurin |
| chr21_+_34775698 | 0.24 |
ENST00000381995.1 |
IFNGR2 |
interferon gamma receptor 2 (interferon gamma transducer 1) |
| chr9_+_82187630 | 0.24 |
ENST00000265284.6 |
TLE4 |
transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) |
| chr12_-_56727676 | 0.24 |
ENST00000547572.1 ENST00000257931.5 ENST00000440411.3 |
PAN2 |
PAN2 poly(A) specific ribonuclease subunit homolog (S. cerevisiae) |
| chr2_+_208394794 | 0.24 |
ENST00000536726.1 ENST00000374397.4 ENST00000452474.1 |
CREB1 |
cAMP responsive element binding protein 1 |
| chr18_+_3252265 | 0.24 |
ENST00000580887.1 ENST00000536605.1 |
MYL12A |
myosin, light chain 12A, regulatory, non-sarcomeric |
| chr4_+_140222609 | 0.24 |
ENST00000296543.5 ENST00000398947.1 |
NAA15 |
N(alpha)-acetyltransferase 15, NatA auxiliary subunit |
| chr20_+_18488137 | 0.23 |
ENST00000450074.1 ENST00000262544.2 ENST00000336714.3 ENST00000377475.3 |
SEC23B |
Sec23 homolog B (S. cerevisiae) |
| chr9_+_82187487 | 0.22 |
ENST00000435650.1 ENST00000414465.1 ENST00000376537.4 ENST00000376534.4 |
TLE4 |
transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) |
| chr5_-_141257954 | 0.22 |
ENST00000456271.1 ENST00000394536.3 ENST00000503492.1 ENST00000287008.3 |
PCDH1 |
protocadherin 1 |
| chr13_+_47127322 | 0.21 |
ENST00000389798.3 |
LRCH1 |
leucine-rich repeats and calponin homology (CH) domain containing 1 |
| chr7_+_143013198 | 0.21 |
ENST00000343257.2 |
CLCN1 |
chloride channel, voltage-sensitive 1 |
| chr11_-_6633799 | 0.20 |
ENST00000299424.4 |
TAF10 |
TAF10 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 30kDa |
| chr21_+_34775772 | 0.20 |
ENST00000405436.1 |
IFNGR2 |
interferon gamma receptor 2 (interferon gamma transducer 1) |
| chr9_+_36572851 | 0.20 |
ENST00000298048.2 ENST00000538311.1 ENST00000536987.1 ENST00000545008.1 ENST00000536860.1 ENST00000536329.1 ENST00000541717.1 ENST00000543751.1 |
MELK |
maternal embryonic leucine zipper kinase |
| chr10_+_22605304 | 0.20 |
ENST00000475460.2 ENST00000602390.1 ENST00000489125.2 ENST00000456711.1 ENST00000444869.1 |
COMMD3-BMI1 COMMD3 |
COMMD3-BMI1 readthrough COMM domain containing 3 |
| chr11_-_66313699 | 0.19 |
ENST00000526986.1 ENST00000310442.3 |
ZDHHC24 |
zinc finger, DHHC-type containing 24 |
| chr2_+_203776937 | 0.19 |
ENST00000402905.3 ENST00000414490.1 ENST00000431787.1 ENST00000444724.1 ENST00000414857.1 ENST00000430899.1 ENST00000445120.1 ENST00000441569.1 ENST00000432024.1 ENST00000443740.1 ENST00000414439.1 ENST00000428585.1 ENST00000545253.1 ENST00000545262.1 ENST00000447539.1 ENST00000456821.2 ENST00000434998.1 ENST00000320443.8 |
CARF |
calcium responsive transcription factor |
| chr19_-_51471362 | 0.18 |
ENST00000376853.4 ENST00000424910.2 |
KLK6 |
kallikrein-related peptidase 6 |
| chr10_+_22605374 | 0.17 |
ENST00000448361.1 |
COMMD3 |
COMM domain containing 3 |
| chr4_+_74735102 | 0.17 |
ENST00000395761.3 |
CXCL1 |
chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) |
| chr7_-_16844611 | 0.17 |
ENST00000401412.1 ENST00000419304.2 |
AGR2 |
anterior gradient 2 |
| chr12_+_53662110 | 0.16 |
ENST00000552462.1 |
ESPL1 |
extra spindle pole bodies homolog 1 (S. cerevisiae) |
| chr14_+_22984601 | 0.15 |
ENST00000390509.1 |
TRAJ28 |
T cell receptor alpha joining 28 |
| chr6_-_136847099 | 0.15 |
ENST00000438100.2 |
MAP7 |
microtubule-associated protein 7 |
| chr1_-_202129704 | 0.15 |
ENST00000476061.1 ENST00000544762.1 ENST00000467283.1 ENST00000464870.1 ENST00000435759.2 ENST00000486116.1 ENST00000543735.1 ENST00000308986.5 ENST00000477625.1 |
PTPN7 |
protein tyrosine phosphatase, non-receptor type 7 |
| chr2_-_136743039 | 0.15 |
ENST00000537273.1 |
DARS |
aspartyl-tRNA synthetase |
| chr3_+_5163905 | 0.14 |
ENST00000256496.3 ENST00000419534.2 |
ARL8B |
ADP-ribosylation factor-like 8B |
| chr10_+_74870253 | 0.14 |
ENST00000544879.1 ENST00000537969.1 ENST00000372997.3 |
NUDT13 |
nudix (nucleoside diphosphate linked moiety X)-type motif 13 |
| chr15_-_71055878 | 0.14 |
ENST00000322954.6 |
UACA |
uveal autoantigen with coiled-coil domains and ankyrin repeats |
| chr6_-_30710510 | 0.13 |
ENST00000376389.3 |
FLOT1 |
flotillin 1 |
| chr22_+_41347363 | 0.13 |
ENST00000216225.8 |
RBX1 |
ring-box 1, E3 ubiquitin protein ligase |
| chr14_+_24439148 | 0.12 |
ENST00000543805.1 ENST00000534993.1 |
DHRS4L2 |
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr10_+_74870206 | 0.12 |
ENST00000357321.4 ENST00000349051.5 |
NUDT13 |
nudix (nucleoside diphosphate linked moiety X)-type motif 13 |
| chr11_-_44331679 | 0.12 |
ENST00000329255.3 |
ALX4 |
ALX homeobox 4 |
| chr1_-_1822495 | 0.12 |
ENST00000378609.4 |
GNB1 |
guanine nucleotide binding protein (G protein), beta polypeptide 1 |
| chr1_-_155658085 | 0.12 |
ENST00000311573.5 ENST00000438245.2 |
YY1AP1 |
YY1 associated protein 1 |
| chr19_-_51471381 | 0.12 |
ENST00000594641.1 |
KLK6 |
kallikrein-related peptidase 6 |
| chr9_+_82188077 | 0.11 |
ENST00000425506.1 |
TLE4 |
transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) |
| chr16_+_67282853 | 0.11 |
ENST00000299798.11 |
SLC9A5 |
solute carrier family 9, subfamily A (NHE5, cation proton antiporter 5), member 5 |
| chr12_-_57504069 | 0.10 |
ENST00000543873.2 ENST00000554663.1 ENST00000557635.1 |
STAT6 |
signal transducer and activator of transcription 6, interleukin-4 induced |
| chr5_+_112312416 | 0.09 |
ENST00000389063.2 |
DCP2 |
decapping mRNA 2 |
| chr3_+_141205852 | 0.09 |
ENST00000286364.3 ENST00000452898.1 |
RASA2 |
RAS p21 protein activator 2 |
| chr1_-_203274418 | 0.09 |
ENST00000457348.1 |
RP11-134P9.1 |
long intergenic non-protein coding RNA 1136 |
| chr11_+_18287801 | 0.09 |
ENST00000532858.1 ENST00000405158.2 |
SAA1 |
serum amyloid A1 |
| chr18_-_31802282 | 0.09 |
ENST00000535475.1 |
NOL4 |
nucleolar protein 4 |
| chr2_+_162016916 | 0.09 |
ENST00000405852.1 |
TANK |
TRAF family member-associated NFKB activator |
| chr11_+_65190245 | 0.08 |
ENST00000499732.1 ENST00000501122.2 ENST00000601801.1 |
NEAT1 |
nuclear paraspeckle assembly transcript 1 (non-protein coding) |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 5.7 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.4 | 21.3 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.4 | 8.7 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.3 | 3.8 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.2 | 6.0 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.2 | 2.6 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.1 | 5.6 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 2.6 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 1.9 | REACTOME TAK1 ACTIVATES NFKB BY PHOSPHORYLATION AND ACTIVATION OF IKKS COMPLEX | Genes involved in TAK1 activates NFkB by phosphorylation and activation of IKKs complex |
| 0.1 | 3.6 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.1 | 2.4 | REACTOME CD28 DEPENDENT PI3K AKT SIGNALING | Genes involved in CD28 dependent PI3K/Akt signaling |
| 0.1 | 1.0 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 3.2 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.1 | 3.4 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.1 | 1.4 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.1 | 4.5 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.1 | 1.2 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.0 | 0.9 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.0 | 5.9 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 3.2 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 2.4 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 1.6 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 2.7 | REACTOME ACTIVATION OF NF KAPPAB IN B CELLS | Genes involved in Activation of NF-kappaB in B Cells |
| 0.0 | 0.8 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.0 | 0.7 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 0.2 | REACTOME TRAF6 MEDIATED NFKB ACTIVATION | Genes involved in TRAF6 mediated NF-kB activation |
| 0.0 | 0.4 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.9 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 0.6 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 0.5 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.1 | REACTOME VIF MEDIATED DEGRADATION OF APOBEC3G | Genes involved in Vif-mediated degradation of APOBEC3G |
| 0.0 | 0.5 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 0.6 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 0.5 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 5.9 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 1.0 | 4.1 | GO:0004911 | interleukin-2 receptor activity(GO:0004911) interleukin-2 binding(GO:0019976) |
| 0.9 | 9.4 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.9 | 11.5 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.8 | 4.2 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.7 | 4.0 | GO:0046979 | TAP2 binding(GO:0046979) |
| 0.6 | 3.2 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.3 | 3.6 | GO:0008061 | chitinase activity(GO:0004568) chitin binding(GO:0008061) |
| 0.3 | 13.7 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.2 | 3.2 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.2 | 10.2 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.2 | 1.7 | GO:0019863 | IgE binding(GO:0019863) |
| 0.2 | 1.5 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.2 | 0.5 | GO:0016314 | phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 0.2 | 1.4 | GO:1990763 | arrestin family protein binding(GO:1990763) |
| 0.2 | 1.8 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.2 | 2.6 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.1 | 0.7 | GO:0004906 | interferon-gamma receptor activity(GO:0004906) |
| 0.1 | 1.2 | GO:0000990 | transcription factor activity, core RNA polymerase binding(GO:0000990) |
| 0.1 | 0.4 | GO:0097677 | STAT family protein binding(GO:0097677) |
| 0.1 | 1.1 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.1 | 5.5 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.1 | 2.7 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 1.8 | GO:0030881 | beta-2-microglobulin binding(GO:0030881) |
| 0.1 | 1.1 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.1 | 3.8 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.1 | 2.9 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 0.4 | GO:0044378 | non-sequence-specific DNA binding, bending(GO:0044378) |
| 0.1 | 3.9 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.1 | 1.8 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.1 | 0.5 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.1 | 2.0 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 0.7 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.1 | 2.4 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.1 | 0.9 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.1 | 0.5 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.1 | 0.5 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.1 | 0.3 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.1 | 2.2 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 0.9 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.1 | 1.5 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.5 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.0 | 1.1 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.0 | 1.1 | GO:0046965 | retinoid X receptor binding(GO:0046965) |
| 0.0 | 0.6 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.1 | GO:0047391 | alkylglycerophosphoethanolamine phosphodiesterase activity(GO:0047391) |
| 0.0 | 0.5 | GO:0046527 | glucosyltransferase activity(GO:0046527) |
| 0.0 | 2.5 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 0.2 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.0 | 0.7 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.1 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.3 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 6.5 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.3 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 0.1 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.0 | 0.4 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.0 | 0.6 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.2 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.1 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.0 | 0.3 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 0.5 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.1 | GO:0015386 | sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.1 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 13.4 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.8 | 5.9 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 0.7 | 2.0 | GO:0070762 | nuclear pore transmembrane ring(GO:0070762) |
| 0.6 | 11.5 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.5 | 5.7 | GO:0042612 | MHC protein complex(GO:0042611) MHC class I protein complex(GO:0042612) |
| 0.5 | 3.6 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.5 | 3.3 | GO:0043196 | varicosity(GO:0043196) |
| 0.4 | 2.5 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.2 | 0.7 | GO:0071746 | IgA immunoglobulin complex(GO:0071745) IgA immunoglobulin complex, circulating(GO:0071746) monomeric IgA immunoglobulin complex(GO:0071748) polymeric IgA immunoglobulin complex(GO:0071749) secretory IgA immunoglobulin complex(GO:0071751) |
| 0.2 | 1.7 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.2 | 0.6 | GO:0031251 | PAN complex(GO:0031251) |
| 0.2 | 1.4 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.2 | 2.0 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.2 | 0.5 | GO:0071065 | alpha9-beta1 integrin-vascular cell adhesion molecule-1 complex(GO:0071065) |
| 0.1 | 0.5 | GO:0005889 | hydrogen:potassium-exchanging ATPase complex(GO:0005889) |
| 0.1 | 1.3 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.1 | 4.7 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.1 | 0.5 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.1 | 4.9 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.0 | 0.1 | GO:0043224 | nuclear SCF ubiquitin ligase complex(GO:0043224) |
| 0.0 | 0.9 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 13.1 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 1.9 | GO:0002102 | podosome(GO:0002102) |
| 0.0 | 0.5 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.7 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 1.9 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.2 | GO:0031415 | NatA complex(GO:0031415) |
| 0.0 | 0.3 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 1.4 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 1.3 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 9.5 | GO:0098857 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.0 | 2.4 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.0 | 1.3 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 1.7 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.0 | 0.6 | GO:0031594 | neuromuscular junction(GO:0031594) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 4.7 | GO:1990108 | protein linear deubiquitination(GO:1990108) |
| 1.6 | 4.7 | GO:0034146 | B-1 B cell homeostasis(GO:0001922) toll-like receptor 5 signaling pathway(GO:0034146) regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070428) |
| 1.4 | 1.4 | GO:1902523 | positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 1.3 | 3.9 | GO:0052250 | modulation of signal transduction in other organism(GO:0044501) modulation by symbiont of host signal transduction pathway(GO:0052027) modulation of signal transduction in other organism involved in symbiotic interaction(GO:0052250) modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade(GO:0085032) |
| 1.3 | 3.8 | GO:0070427 | nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070427) |
| 1.2 | 10.7 | GO:0070424 | regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070424) |
| 1.2 | 5.9 | GO:0002905 | mature B cell apoptotic process(GO:0002901) regulation of mature B cell apoptotic process(GO:0002905) negative regulation of mature B cell apoptotic process(GO:0002906) |
| 0.8 | 3.1 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.8 | 3.8 | GO:0002925 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) |
| 0.8 | 2.3 | GO:1900169 | regulation of glucocorticoid mediated signaling pathway(GO:1900169) |
| 0.8 | 6.0 | GO:0050859 | negative regulation of B cell receptor signaling pathway(GO:0050859) |
| 0.7 | 2.8 | GO:0002268 | follicular dendritic cell differentiation(GO:0002268) |
| 0.7 | 4.9 | GO:0038110 | interleukin-2-mediated signaling pathway(GO:0038110) |
| 0.7 | 3.3 | GO:0033590 | response to cobalamin(GO:0033590) |
| 0.7 | 2.0 | GO:0051300 | spindle pole body duplication(GO:0030474) spindle pole body organization(GO:0051300) spindle pole body localization(GO:0070631) establishment of spindle pole body localization(GO:0070632) spindle pole body localization to nuclear envelope(GO:0071789) establishment of spindle pole body localization to nuclear envelope(GO:0071790) |
| 0.6 | 3.2 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.6 | 2.4 | GO:0072183 | negative regulation by virus of viral protein levels in host cell(GO:0046725) negative regulation of nephron tubule epithelial cell differentiation(GO:0072183) negative regulation of metanephric nephron tubule epithelial cell differentiation(GO:0072308) negative regulation of epithelial cell differentiation involved in kidney development(GO:2000697) |
| 0.6 | 5.2 | GO:0045075 | interleukin-12 biosynthetic process(GO:0042090) regulation of interleukin-12 biosynthetic process(GO:0045075) |
| 0.6 | 5.7 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 0.6 | 2.8 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 0.5 | 2.1 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.4 | 6.7 | GO:0060263 | regulation of respiratory burst(GO:0060263) |
| 0.4 | 2.6 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.4 | 3.2 | GO:0034351 | regulation of glial cell apoptotic process(GO:0034350) negative regulation of glial cell apoptotic process(GO:0034351) |
| 0.4 | 1.2 | GO:0051097 | negative regulation of helicase activity(GO:0051097) oligodendrocyte apoptotic process(GO:0097252) |
| 0.4 | 1.9 | GO:0070842 | aggresome assembly(GO:0070842) |
| 0.4 | 1.1 | GO:1904897 | regulation of hepatic stellate cell proliferation(GO:1904897) positive regulation of hepatic stellate cell proliferation(GO:1904899) hepatic stellate cell proliferation(GO:1990922) |
| 0.4 | 7.0 | GO:0072540 | T-helper 17 cell lineage commitment(GO:0072540) |
| 0.4 | 1.5 | GO:0010536 | positive regulation of activation of Janus kinase activity(GO:0010536) positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) negative regulation of macrophage apoptotic process(GO:2000110) |
| 0.3 | 2.6 | GO:0045078 | positive regulation of interferon-gamma biosynthetic process(GO:0045078) |
| 0.3 | 3.6 | GO:0006032 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.3 | 2.4 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.3 | 4.5 | GO:0032688 | negative regulation of interferon-beta production(GO:0032688) |
| 0.2 | 1.2 | GO:0033078 | extrathymic T cell differentiation(GO:0033078) |
| 0.2 | 0.5 | GO:0045359 | interferon-beta biosynthetic process(GO:0045350) regulation of interferon-beta biosynthetic process(GO:0045357) positive regulation of interferon-beta biosynthetic process(GO:0045359) |
| 0.2 | 1.4 | GO:0034670 | chemotaxis to arachidonic acid(GO:0034670) response to arachidonic acid(GO:1904550) |
| 0.2 | 0.8 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 0.2 | 0.6 | GO:0033364 | mast cell secretory granule organization(GO:0033364) |
| 0.2 | 1.1 | GO:0022614 | membrane to membrane docking(GO:0022614) |
| 0.2 | 0.5 | GO:0045409 | negative regulation of interleukin-6 biosynthetic process(GO:0045409) regulation of neutrophil differentiation(GO:0045658) negative regulation of neutrophil differentiation(GO:0045659) |
| 0.2 | 0.9 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.2 | 1.4 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.1 | 1.6 | GO:0039530 | MDA-5 signaling pathway(GO:0039530) |
| 0.1 | 0.4 | GO:0036309 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.1 | 0.3 | GO:0060584 | positive regulation of the force of heart contraction by chemical signal(GO:0003099) regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) |
| 0.1 | 0.4 | GO:0051685 | maintenance of ER location(GO:0051685) |
| 0.1 | 14.0 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.1 | 0.7 | GO:0006196 | AMP catabolic process(GO:0006196) |
| 0.1 | 2.7 | GO:0050965 | detection of temperature stimulus involved in sensory perception(GO:0050961) detection of temperature stimulus involved in sensory perception of pain(GO:0050965) |
| 0.1 | 0.4 | GO:0050966 | detection of mechanical stimulus involved in sensory perception of pain(GO:0050966) |
| 0.1 | 0.5 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.1 | 1.7 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.1 | 2.6 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.1 | 0.7 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.1 | 0.3 | GO:0021590 | cerebellum maturation(GO:0021590) cerebellar cortex maturation(GO:0021699) |
| 0.1 | 0.1 | GO:0048619 | embryonic hindgut morphogenesis(GO:0048619) |
| 0.1 | 0.5 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.1 | 1.7 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.1 | 0.5 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 0.0 | 0.2 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.0 | 0.3 | GO:1904417 | regulation of xenophagy(GO:1904415) positive regulation of xenophagy(GO:1904417) |
| 0.0 | 0.9 | GO:0050858 | negative regulation of antigen receptor-mediated signaling pathway(GO:0050858) negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.0 | 0.2 | GO:1903899 | positive regulation of PERK-mediated unfolded protein response(GO:1903899) |
| 0.0 | 0.3 | GO:2000189 | positive regulation of cholesterol homeostasis(GO:2000189) |
| 0.0 | 2.7 | GO:0060113 | inner ear receptor cell differentiation(GO:0060113) |
| 0.0 | 3.7 | GO:0035690 | cellular response to drug(GO:0035690) |
| 0.0 | 0.4 | GO:0045842 | positive regulation of mitotic metaphase/anaphase transition(GO:0045842) positive regulation of mitotic sister chromatid separation(GO:1901970) positive regulation of metaphase/anaphase transition of cell cycle(GO:1902101) |
| 0.0 | 0.2 | GO:0060355 | positive regulation of cell adhesion molecule production(GO:0060355) |
| 0.0 | 0.4 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 0.0 | 0.8 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.0 | 0.2 | GO:0035865 | cellular response to potassium ion(GO:0035865) |
| 0.0 | 2.9 | GO:0042100 | B cell proliferation(GO:0042100) |
| 0.0 | 0.7 | GO:0060334 | regulation of interferon-gamma-mediated signaling pathway(GO:0060334) |
| 0.0 | 0.5 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 0.1 | GO:0002295 | T-helper cell lineage commitment(GO:0002295) |
| 0.0 | 0.9 | GO:0035428 | hexose transmembrane transport(GO:0035428) glucose transmembrane transport(GO:1904659) |
| 0.0 | 0.3 | GO:1904996 | PML body organization(GO:0030578) positive regulation of leukocyte adhesion to vascular endothelial cell(GO:1904996) |
| 0.0 | 0.8 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.0 | 0.7 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 0.6 | GO:0031629 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.0 | 1.5 | GO:0090003 | regulation of establishment of protein localization to plasma membrane(GO:0090003) |
| 0.0 | 0.1 | GO:0006422 | aspartyl-tRNA aminoacylation(GO:0006422) |
| 0.0 | 0.7 | GO:0090503 | RNA phosphodiester bond hydrolysis, exonucleolytic(GO:0090503) |
| 0.0 | 0.2 | GO:0019227 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.0 | 0.3 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.0 | 0.4 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.0 | 0.2 | GO:0070365 | hepatocyte differentiation(GO:0070365) |
| 0.0 | 1.0 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.0 | 3.1 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.0 | 1.0 | GO:0071391 | cellular response to estrogen stimulus(GO:0071391) |
| 0.0 | 0.9 | GO:0071357 | type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 0.0 | 0.5 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 0.1 | GO:0010265 | SCF complex assembly(GO:0010265) |
| 0.0 | 0.6 | GO:1904837 | beta-catenin-TCF complex assembly(GO:1904837) |
| 0.0 | 0.3 | GO:0002076 | osteoblast development(GO:0002076) |
| 0.0 | 0.1 | GO:0097190 | apoptotic signaling pathway(GO:0097190) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 26.2 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.2 | 2.8 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.2 | 1.6 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.2 | 3.4 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.1 | 3.0 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.1 | 4.3 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.1 | 6.2 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.1 | 6.8 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.1 | 2.2 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.1 | 3.3 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.1 | 2.6 | PID EPO PATHWAY | EPO signaling pathway |
| 0.1 | 3.4 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.1 | 3.7 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 1.7 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.1 | 3.2 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.1 | 1.1 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.1 | 2.7 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
| 0.0 | 1.1 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.0 | 1.2 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.0 | 0.6 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 1.4 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.3 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 0.4 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.0 | 0.3 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 0.4 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |