Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
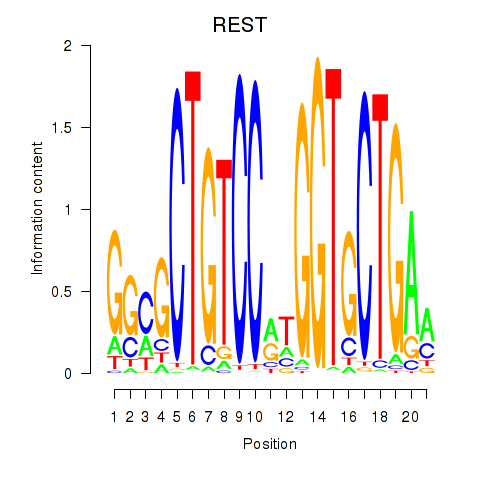
Results for REST
Z-value: 0.61
Transcription factors associated with REST
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
REST
|
ENSG00000084093.11 | REST |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| REST | hg19_v2_chr4_+_57774042_57774114 | 0.19 | 4.7e-01 | Click! |
Activity profile of REST motif
Sorted Z-values of REST motif
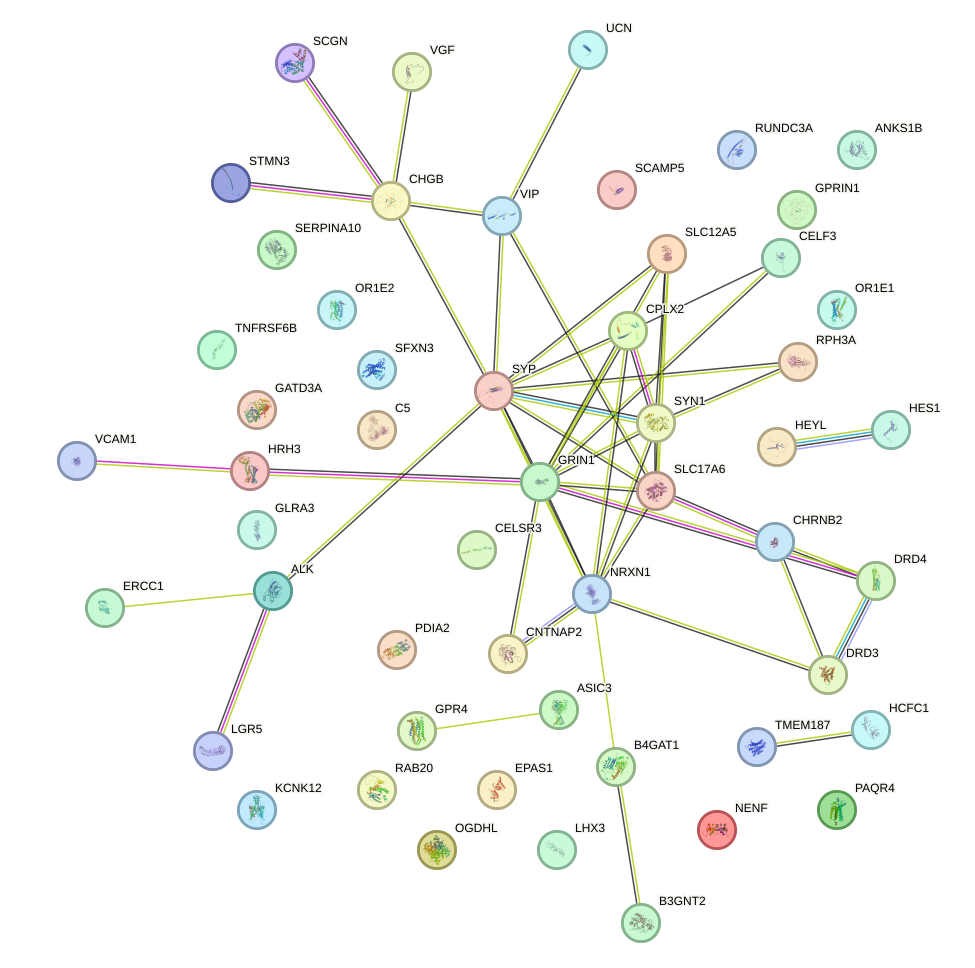
Network of associatons between targets according to the STRING database.
First level regulatory network of REST
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chrX_-_153141302 | 1.24 |
ENST00000361699.4 ENST00000543994.1 ENST00000370057.3 ENST00000538883.1 ENST00000361981.3 |
L1CAM |
L1 cell adhesion molecule |
| chr9_-_123812542 | 1.20 |
ENST00000223642.1 |
C5 |
complement component 5 |
| chr12_+_71833550 | 1.08 |
ENST00000266674.5 |
LGR5 |
leucine-rich repeat containing G protein-coupled receptor 5 |
| chr3_+_193853927 | 0.84 |
ENST00000232424.3 |
HES1 |
hes family bHLH transcription factor 1 |
| chr3_-_8686479 | 0.84 |
ENST00000544814.1 ENST00000427408.1 |
SSUH2 |
ssu-2 homolog (C. elegans) |
| chr13_-_111214015 | 0.82 |
ENST00000267328.3 |
RAB20 |
RAB20, member RAS oncogene family |
| chr14_-_94759595 | 0.65 |
ENST00000261994.4 |
SERPINA10 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 |
| chr14_-_94759361 | 0.65 |
ENST00000393096.1 |
SERPINA10 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 |
| chr14_-_94759408 | 0.65 |
ENST00000554723.1 |
SERPINA10 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 |
| chr3_-_48700310 | 0.60 |
ENST00000164024.4 ENST00000544264.1 |
CELSR3 |
cadherin, EGF LAG seven-pass G-type receptor 3 |
| chr11_-_66115032 | 0.49 |
ENST00000311181.4 |
B3GNT1 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 1 |
| chr6_+_25652432 | 0.47 |
ENST00000377961.2 |
SCGN |
secretagogin, EF-hand calcium binding protein |
| chrX_-_47479246 | 0.39 |
ENST00000295987.7 ENST00000340666.4 |
SYN1 |
synapsin I |
| chr12_+_71833756 | 0.37 |
ENST00000536515.1 ENST00000540815.2 |
LGR5 |
leucine-rich repeat containing G protein-coupled receptor 5 |
| chr20_+_62327996 | 0.37 |
ENST00000369996.1 |
TNFRSF6B |
tumor necrosis factor receptor superfamily, member 6b, decoy |
| chr10_-_50970382 | 0.36 |
ENST00000419399.1 ENST00000432695.1 |
OGDHL |
oxoglutarate dehydrogenase-like |
| chr10_-_50970322 | 0.36 |
ENST00000374103.4 |
OGDHL |
oxoglutarate dehydrogenase-like |
| chr20_+_36012051 | 0.36 |
ENST00000373567.2 |
SRC |
v-src avian sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog |
| chr12_+_175930 | 0.32 |
ENST00000538872.1 ENST00000326261.4 |
IQSEC3 |
IQ motif and Sec7 domain 3 |
| chr4_+_158141899 | 0.30 |
ENST00000264426.9 ENST00000506284.1 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr7_+_145813453 | 0.29 |
ENST00000361727.3 |
CNTNAP2 |
contactin associated protein-like 2 |
| chr2_+_46524537 | 0.28 |
ENST00000263734.3 |
EPAS1 |
endothelial PAS domain protein 1 |
| chr15_+_75287861 | 0.27 |
ENST00000425597.3 ENST00000562327.1 ENST00000568018.1 ENST00000562212.1 ENST00000567920.1 ENST00000566872.1 ENST00000361900.6 ENST00000545456.1 |
SCAMP5 |
secretory carrier membrane protein 5 |
| chr2_-_51259292 | 0.27 |
ENST00000401669.2 |
NRXN1 |
neurexin 1 |
| chr5_+_175298573 | 0.27 |
ENST00000512824.1 |
CPLX2 |
complexin 2 |
| chr2_-_51259229 | 0.25 |
ENST00000405472.3 |
NRXN1 |
neurexin 1 |
| chr12_-_12837423 | 0.25 |
ENST00000540510.1 |
GPR19 |
G protein-coupled receptor 19 |
| chr15_-_83378611 | 0.24 |
ENST00000542200.1 |
AP3B2 |
adaptor-related protein complex 3, beta 2 subunit |
| chr2_-_47798044 | 0.23 |
ENST00000327876.4 |
KCNK12 |
potassium channel, subfamily K, member 12 |
| chr2_-_30143525 | 0.22 |
ENST00000431873.1 |
ALK |
anaplastic lymphoma receptor tyrosine kinase |
| chr6_+_25652501 | 0.22 |
ENST00000334979.6 |
SCGN |
secretagogin, EF-hand calcium binding protein |
| chr4_+_158141806 | 0.18 |
ENST00000393815.2 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr6_+_110299501 | 0.18 |
ENST00000414000.2 |
GPR6 |
G protein-coupled receptor 6 |
| chr4_+_158141843 | 0.17 |
ENST00000509417.1 ENST00000296526.7 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr11_+_64808675 | 0.17 |
ENST00000529996.1 |
SAC3D1 |
SAC3 domain containing 1 |
| chr11_+_22359562 | 0.17 |
ENST00000263160.3 |
SLC17A6 |
solute carrier family 17 (vesicular glutamate transporter), member 6 |
| chr12_+_113229543 | 0.16 |
ENST00000447659.2 |
RPH3A |
rabphilin 3A homolog (mouse) |
| chr5_+_175298674 | 0.16 |
ENST00000514150.1 |
CPLX2 |
complexin 2 |
| chr17_-_3301704 | 0.16 |
ENST00000322608.2 |
OR1E1 |
olfactory receptor, family 1, subfamily E, member 1 |
| chr5_+_175298487 | 0.15 |
ENST00000393745.3 |
CPLX2 |
complexin 2 |
| chr10_+_26505179 | 0.15 |
ENST00000376261.3 |
GAD2 |
glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) |
| chrX_+_153237740 | 0.15 |
ENST00000369982.4 |
TMEM187 |
transmembrane protein 187 |
| chrX_-_153237258 | 0.14 |
ENST00000310441.7 |
HCFC1 |
host cell factor C1 (VP16-accessory protein) |
| chr2_+_226265364 | 0.14 |
ENST00000272907.6 |
NYAP2 |
neuronal tyrosine-phosphorylated phosphoinositide-3-kinase adaptor 2 |
| chr3_-_113897545 | 0.13 |
ENST00000467632.1 |
DRD3 |
dopamine receptor D3 |
| chr7_-_100808394 | 0.12 |
ENST00000445482.2 |
VGF |
VGF nerve growth factor inducible |
| chr12_+_113229737 | 0.12 |
ENST00000551052.1 ENST00000415485.3 |
RPH3A |
rabphilin 3A homolog (mouse) |
| chr12_+_113229452 | 0.11 |
ENST00000389385.4 |
RPH3A |
rabphilin 3A homolog (mouse) |
| chr3_-_113897899 | 0.10 |
ENST00000383673.2 ENST00000295881.7 |
DRD3 |
dopamine receptor D3 |
| chr1_-_175712665 | 0.10 |
ENST00000263525.2 |
TNR |
tenascin R |
| chr10_-_102790852 | 0.09 |
ENST00000470414.1 ENST00000370215.3 |
PDZD7 |
PDZ domain containing 7 |
| chr12_+_52695617 | 0.09 |
ENST00000293525.5 |
KRT86 |
keratin 86 |
| chr17_+_42385927 | 0.09 |
ENST00000426726.3 ENST00000590941.1 ENST00000225441.7 |
RUNDC3A |
RUN domain containing 3A |
| chr4_+_96012614 | 0.09 |
ENST00000264568.4 |
BMPR1B |
bone morphogenetic protein receptor, type IB |
| chr9_+_140033862 | 0.09 |
ENST00000350902.5 ENST00000371550.4 ENST00000371546.4 ENST00000371555.4 ENST00000371553.3 ENST00000371559.4 ENST00000371560.3 |
GRIN1 |
glutamate receptor, ionotropic, N-methyl D-aspartate 1 |
| chr6_+_153071925 | 0.08 |
ENST00000367244.3 ENST00000367243.3 |
VIP |
vasoactive intestinal peptide |
| chr12_-_57443886 | 0.08 |
ENST00000300119.3 |
MYO1A |
myosin IA |
| chr10_+_102790980 | 0.07 |
ENST00000393459.1 ENST00000224807.5 |
SFXN3 |
sideroflexin 3 |
| chr16_+_335680 | 0.07 |
ENST00000435833.1 |
PDIA2 |
protein disulfide isomerase family A, member 2 |
| chr15_-_83378526 | 0.07 |
ENST00000535348.1 ENST00000535359.1 |
AP3B2 |
adaptor-related protein complex 3, beta 2 subunit |
| chr2_-_51259528 | 0.07 |
ENST00000404971.1 |
NRXN1 |
neurexin 1 |
| chr20_-_62284766 | 0.06 |
ENST00000370053.1 |
STMN3 |
stathmin-like 3 |
| chr1_-_151689259 | 0.06 |
ENST00000420342.1 ENST00000290583.4 |
CELF3 |
CUGBP, Elav-like family member 3 |
| chr2_-_27531313 | 0.06 |
ENST00000296099.2 |
UCN |
urocortin |
| chr1_+_154540246 | 0.06 |
ENST00000368476.3 |
CHRNB2 |
cholinergic receptor, nicotinic, beta 2 (neuronal) |
| chr20_+_44657845 | 0.05 |
ENST00000243964.3 |
SLC12A5 |
solute carrier family 12 (potassium/chloride transporter), member 5 |
| chr1_+_212606219 | 0.05 |
ENST00000366988.3 |
NENF |
neudesin neurotrophic factor |
| chr11_+_637246 | 0.05 |
ENST00000176183.5 |
DRD4 |
dopamine receptor D4 |
| chr20_+_5892037 | 0.04 |
ENST00000378961.4 |
CHGB |
chromogranin B (secretogranin 1) |
| chr16_+_3019246 | 0.04 |
ENST00000318782.8 ENST00000293978.8 |
PAQR4 |
progestin and adipoQ receptor family member IV |
| chr17_-_3337135 | 0.04 |
ENST00000248384.1 |
OR1E2 |
olfactory receptor, family 1, subfamily E, member 2 |
| chr15_-_83378638 | 0.04 |
ENST00000261722.3 |
AP3B2 |
adaptor-related protein complex 3, beta 2 subunit |
| chr7_-_100808843 | 0.04 |
ENST00000249330.2 |
VGF |
VGF nerve growth factor inducible |
| chrX_-_49056635 | 0.04 |
ENST00000472598.1 ENST00000538567.1 ENST00000479808.1 ENST00000263233.4 |
SYP |
synaptophysin |
| chr2_-_51259641 | 0.04 |
ENST00000406316.2 ENST00000405581.1 |
NRXN1 |
neurexin 1 |
| chr7_+_150748288 | 0.03 |
ENST00000490540.1 |
ASIC3 |
acid-sensing (proton-gated) ion channel 3 |
| chr1_-_40105617 | 0.03 |
ENST00000372852.3 |
HEYL |
hes-related family bHLH transcription factor with YRPW motif-like |
| chr19_-_45927622 | 0.03 |
ENST00000300853.3 ENST00000589165.1 |
ERCC1 |
excision repair cross-complementing rodent repair deficiency, complementation group 1 (includes overlapping antisense sequence) |
| chr10_+_26505594 | 0.02 |
ENST00000259271.3 |
GAD2 |
glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) |
| chr4_-_175750364 | 0.02 |
ENST00000340217.5 ENST00000274093.3 |
GLRA3 |
glycine receptor, alpha 3 |
| chr20_-_60795316 | 0.01 |
ENST00000317393.6 |
HRH3 |
histamine receptor H3 |
| chr9_-_139096955 | 0.01 |
ENST00000371748.5 |
LHX3 |
LIM homeobox 3 |
| chr12_-_99288536 | 0.01 |
ENST00000549797.1 ENST00000333732.7 ENST00000341752.7 |
ANKS1B |
ankyrin repeat and sterile alpha motif domain containing 1B |
| chr15_-_22473353 | 0.01 |
ENST00000557788.2 |
IGHV4OR15-8 |
immunoglobulin heavy variable 4/OR15-8 (non-functional) |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.2 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.7 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.0 | 1.2 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.0 | 0.4 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.0 | 0.8 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.0 | 0.4 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.0 | 0.5 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.0 | 1.2 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 0.8 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.7 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 0.3 | 0.8 | GO:0021558 | midbrain-hindbrain boundary morphogenesis(GO:0021555) trochlear nerve development(GO:0021558) regulation of timing of neuron differentiation(GO:0060164) |
| 0.3 | 2.0 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.2 | 0.8 | GO:0090385 | phagosome-lysosome fusion(GO:0090385) |
| 0.1 | 0.6 | GO:0036515 | serotonergic neuron axon guidance(GO:0036515) |
| 0.1 | 1.5 | GO:2001013 | epithelial cell proliferation involved in renal tubule morphogenesis(GO:2001013) |
| 0.1 | 0.3 | GO:0071109 | superior temporal gyrus development(GO:0071109) |
| 0.1 | 0.4 | GO:0071393 | cellular response to progesterone stimulus(GO:0071393) |
| 0.1 | 0.2 | GO:0050883 | musculoskeletal movement, spinal reflex action(GO:0050883) |
| 0.1 | 0.6 | GO:0097116 | gephyrin clustering involved in postsynaptic density assembly(GO:0097116) |
| 0.1 | 0.1 | GO:0035483 | gastric motility(GO:0035482) gastric emptying(GO:0035483) |
| 0.1 | 0.2 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.4 | GO:0097091 | synaptic vesicle clustering(GO:0097091) |
| 0.0 | 0.1 | GO:0019046 | release from viral latency(GO:0019046) |
| 0.0 | 0.3 | GO:0043129 | surfactant homeostasis(GO:0043129) |
| 0.0 | 0.1 | GO:0043449 | olfactory learning(GO:0008355) cellular alkene metabolic process(GO:0043449) |
| 0.0 | 0.2 | GO:0006540 | glutamate decarboxylation to succinate(GO:0006540) |
| 0.0 | 0.6 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.0 | 0.4 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.0 | 0.2 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.0 | 0.1 | GO:0048686 | regulation of sprouting of injured axon(GO:0048686) regulation of axon extension involved in regeneration(GO:0048690) |
| 0.0 | 0.7 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 0.2 | GO:0098700 | neurotransmitter loading into synaptic vesicle(GO:0098700) |
| 0.0 | 0.1 | GO:0001550 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) |
| 0.0 | 0.0 | GO:0050968 | detection of chemical stimulus involved in sensory perception of pain(GO:0050968) |
| 0.0 | 0.3 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 0.1 | GO:0042320 | regulation of circadian sleep/wake cycle, REM sleep(GO:0042320) synaptic transmission involved in micturition(GO:0060084) |
| 0.0 | 0.9 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.0 | 0.6 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.0 | 0.1 | GO:0050910 | detection of mechanical stimulus involved in sensory perception of sound(GO:0050910) |
| 0.0 | 0.1 | GO:0030644 | cellular chloride ion homeostasis(GO:0030644) |
| 0.0 | 0.2 | GO:0090520 | sphingosine-1-phosphate signaling pathway(GO:0003376) sphingolipid mediated signaling pathway(GO:0090520) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.1 | 0.7 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.1 | 0.4 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.3 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.6 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.0 | 0.4 | GO:0008430 | selenium binding(GO:0008430) |
| 0.0 | 0.2 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.0 | 0.2 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 0.0 | 0.2 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.0 | 0.1 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.0 | 0.6 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.8 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.0 | 1.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.1 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.0 | 0.2 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.0 | 0.1 | GO:0010853 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
| 0.0 | 0.3 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 0.4 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.1 | GO:0043995 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 0.7 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) |
| 0.1 | 0.7 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.1 | 0.6 | GO:0070554 | synaptobrevin 2-SNAP-25-syntaxin-3-complexin complex(GO:0070554) |
| 0.1 | 1.1 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.4 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.0 | 0.1 | GO:0072534 | perineuronal net(GO:0072534) |
| 0.0 | 0.4 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.0 | 0.1 | GO:0002139 | stereocilia coupling link(GO:0002139) |
| 0.0 | 0.3 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.0 | 1.7 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.0 | 0.4 | GO:0000795 | synaptonemal complex(GO:0000795) |
| 0.0 | 0.1 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |