Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
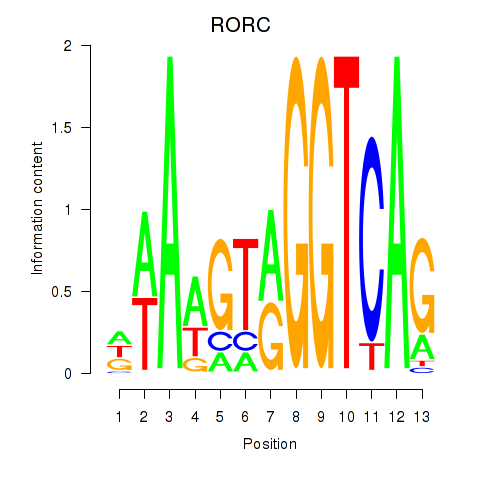
Results for RORC
Z-value: 0.68
Transcription factors associated with RORC
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RORC
|
ENSG00000143365.12 | RORC |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RORC | hg19_v2_chr1_-_151804222_151804228, hg19_v2_chr1_-_151804314_151804348, hg19_v2_chr1_-_151798546_151798590 | -0.16 | 5.4e-01 | Click! |
Activity profile of RORC motif
Sorted Z-values of RORC motif
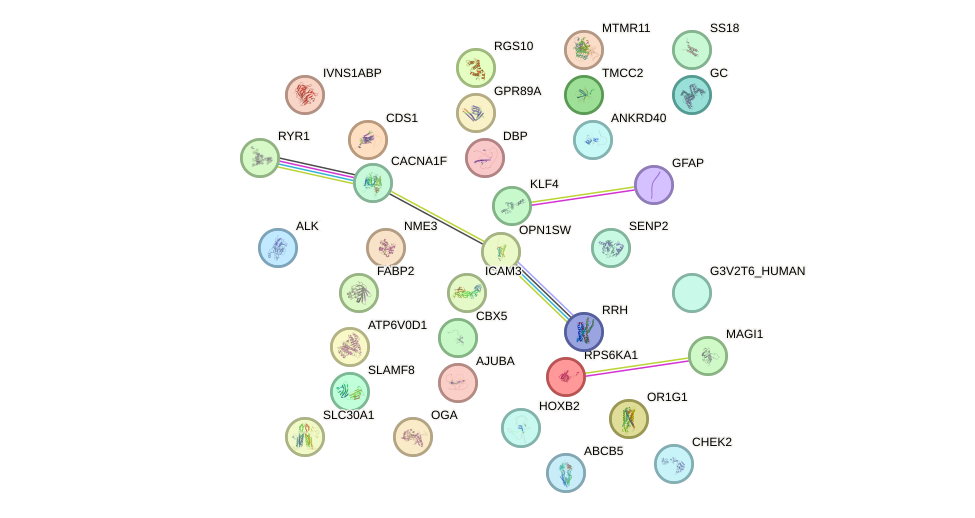
Network of associatons between targets according to the STRING database.
First level regulatory network of RORC
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_+_93964158 | 2.71 |
ENST00000549206.1 |
SOCS2 |
suppressor of cytokine signaling 2 |
| chr12_+_93963590 | 2.67 |
ENST00000340600.2 |
SOCS2 |
suppressor of cytokine signaling 2 |
| chr10_-_121296045 | 1.41 |
ENST00000392865.1 |
RGS10 |
regulator of G-protein signaling 10 |
| chr17_-_46623441 | 1.40 |
ENST00000330070.4 |
HOXB2 |
homeobox B2 |
| chr3_+_185303962 | 0.82 |
ENST00000296257.5 |
SENP2 |
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr3_+_185304059 | 0.74 |
ENST00000427465.2 |
SENP2 |
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr9_-_110251836 | 0.69 |
ENST00000374672.4 |
KLF4 |
Kruppel-like factor 4 (gut) |
| chr18_-_23671139 | 0.66 |
ENST00000579061.1 ENST00000542420.2 |
SS18 |
synovial sarcoma translocation, chromosome 18 |
| chr1_-_153522562 | 0.65 |
ENST00000368714.1 |
S100A4 |
S100 calcium binding protein A4 |
| chr16_-_1821496 | 0.57 |
ENST00000564628.1 ENST00000563498.1 |
NME3 |
NME/NM23 nucleoside diphosphate kinase 3 |
| chr2_+_160590469 | 0.56 |
ENST00000409591.1 |
MARCH7 |
membrane-associated ring finger (C3HC4) 7, E3 ubiquitin protein ligase |
| chr16_-_67514982 | 0.51 |
ENST00000565835.1 ENST00000540149.1 ENST00000290949.3 |
ATP6V0D1 |
ATPase, H+ transporting, lysosomal 38kDa, V0 subunit d1 |
| chr6_-_111927062 | 0.44 |
ENST00000359831.4 |
TRAF3IP2 |
TRAF3 interacting protein 2 |
| chr1_+_26869597 | 0.43 |
ENST00000530003.1 |
RPS6KA1 |
ribosomal protein S6 kinase, 90kDa, polypeptide 1 |
| chr10_-_105845674 | 0.42 |
ENST00000353479.5 ENST00000369733.3 |
COL17A1 |
collagen, type XVII, alpha 1 |
| chr7_+_112063192 | 0.40 |
ENST00000005558.4 |
IFRD1 |
interferon-related developmental regulator 1 |
| chr11_-_19223523 | 0.39 |
ENST00000265968.3 |
CSRP3 |
cysteine and glycine-rich protein 3 (cardiac LIM protein) |
| chr4_-_120243545 | 0.39 |
ENST00000274024.3 |
FABP2 |
fatty acid binding protein 2, intestinal |
| chr1_+_114447763 | 0.38 |
ENST00000369563.3 |
DCLRE1B |
DNA cross-link repair 1B |
| chr19_-_10450287 | 0.36 |
ENST00000589261.1 ENST00000590569.1 ENST00000589580.1 ENST00000589249.1 |
ICAM3 |
intercellular adhesion molecule 3 |
| chr8_-_95449155 | 0.30 |
ENST00000481490.2 |
FSBP |
fibrinogen silencer binding protein |
| chr1_-_150207017 | 0.27 |
ENST00000369119.3 |
ANP32E |
acidic (leucine-rich) nuclear phosphoprotein 32 family, member E |
| chr1_+_205197304 | 0.24 |
ENST00000358024.3 |
TMCC2 |
transmembrane and coiled-coil domain family 2 |
| chr1_-_185286461 | 0.22 |
ENST00000367498.3 |
IVNS1ABP |
influenza virus NS1A binding protein |
| chr11_+_13299186 | 0.20 |
ENST00000527998.1 ENST00000396441.3 ENST00000533520.1 ENST00000529825.1 ENST00000389707.4 ENST00000401424.1 ENST00000529388.1 ENST00000530357.1 ENST00000403290.1 ENST00000361003.4 ENST00000389708.3 ENST00000403510.3 ENST00000482049.1 |
ARNTL |
aryl hydrocarbon receptor nuclear translocator-like |
| chr7_+_20686946 | 0.20 |
ENST00000443026.2 ENST00000406935.1 |
ABCB5 |
ATP-binding cassette, sub-family B (MDR/TAP), member 5 |
| chr14_-_23451467 | 0.20 |
ENST00000555074.1 ENST00000361265.4 |
RP11-298I3.5 AJUBA |
RP11-298I3.5 ajuba LIM protein |
| chr12_+_7167980 | 0.19 |
ENST00000360817.5 ENST00000402681.3 |
C1S |
complement component 1, s subcomponent |
| chr10_-_103578182 | 0.19 |
ENST00000439817.1 |
MGEA5 |
meningioma expressed antigen 5 (hyaluronidase) |
| chr1_-_145826450 | 0.19 |
ENST00000462900.2 |
GPR89A |
G protein-coupled receptor 89A |
| chr12_-_54652060 | 0.19 |
ENST00000552562.1 |
CBX5 |
chromobox homolog 5 |
| chr1_-_211752073 | 0.17 |
ENST00000367001.4 |
SLC30A1 |
solute carrier family 30 (zinc transporter), member 1 |
| chr10_-_103578162 | 0.17 |
ENST00000361464.3 ENST00000357797.5 ENST00000370094.3 |
MGEA5 |
meningioma expressed antigen 5 (hyaluronidase) |
| chr2_-_148778323 | 0.16 |
ENST00000440042.1 ENST00000535373.1 ENST00000540442.1 ENST00000536575.1 |
ORC4 |
origin recognition complex, subunit 4 |
| chr3_-_164914640 | 0.16 |
ENST00000241274.3 |
SLITRK3 |
SLIT and NTRK-like family, member 3 |
| chr4_+_85504075 | 0.14 |
ENST00000295887.5 |
CDS1 |
CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 1 |
| chr1_+_171810606 | 0.14 |
ENST00000358155.4 ENST00000367733.2 ENST00000355305.5 ENST00000367731.1 |
DNM3 |
dynamin 3 |
| chr17_-_3030875 | 0.14 |
ENST00000328890.2 |
OR1G1 |
olfactory receptor, family 1, subfamily G, member 1 |
| chr7_-_128415844 | 0.13 |
ENST00000249389.2 |
OPN1SW |
opsin 1 (cone pigments), short-wave-sensitive |
| chr3_-_65583561 | 0.13 |
ENST00000460329.2 |
MAGI1 |
membrane associated guanylate kinase, WW and PDZ domain containing 1 |
| chr13_+_109248500 | 0.13 |
ENST00000356711.2 |
MYO16 |
myosin XVI |
| chr17_-_42992856 | 0.12 |
ENST00000588316.1 ENST00000435360.2 ENST00000586793.1 ENST00000588735.1 ENST00000588037.1 ENST00000592320.1 ENST00000253408.5 |
GFAP |
glial fibrillary acidic protein |
| chrX_-_110655391 | 0.11 |
ENST00000356915.2 ENST00000356220.3 |
DCX |
doublecortin |
| chrX_-_43832711 | 0.10 |
ENST00000378062.5 |
NDP |
Norrie disease (pseudoglioma) |
| chr1_+_159796534 | 0.10 |
ENST00000289707.5 |
SLAMF8 |
SLAM family member 8 |
| chr12_+_4671352 | 0.09 |
ENST00000542744.1 |
DYRK4 |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 4 |
| chrX_-_49089771 | 0.08 |
ENST00000376251.1 ENST00000323022.5 ENST00000376265.2 |
CACNA1F |
calcium channel, voltage-dependent, L type, alpha 1F subunit |
| chr19_+_1269324 | 0.07 |
ENST00000589710.1 ENST00000588230.1 ENST00000413636.2 ENST00000586472.1 ENST00000589686.1 ENST00000444172.2 ENST00000587323.1 ENST00000320936.5 ENST00000587896.1 ENST00000589235.1 ENST00000591659.1 |
CIRBP |
cold inducible RNA binding protein |
| chr19_-_49140609 | 0.07 |
ENST00000601104.1 |
DBP |
D site of albumin promoter (albumin D-box) binding protein |
| chr1_-_149908710 | 0.06 |
ENST00000439741.2 ENST00000361405.6 ENST00000406732.3 |
MTMR11 |
myotubularin related protein 11 |
| chr1_-_62190793 | 0.05 |
ENST00000371177.2 ENST00000606498.1 |
TM2D1 |
TM2 domain containing 1 |
| chr17_-_48785216 | 0.03 |
ENST00000285243.6 |
ANKRD40 |
ankyrin repeat domain 40 |
| chr19_+_38924316 | 0.02 |
ENST00000355481.4 ENST00000360985.3 ENST00000359596.3 |
RYR1 |
ryanodine receptor 1 (skeletal) |
| chr2_-_30144432 | 0.02 |
ENST00000389048.3 |
ALK |
anaplastic lymphoma receptor tyrosine kinase |
| chr4_+_110749143 | 0.01 |
ENST00000317735.4 |
RRH |
retinal pigment epithelium-derived rhodopsin homolog |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:0021569 | rhombomere 3 development(GO:0021569) |
| 0.3 | 5.4 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.2 | 0.7 | GO:0071409 | negative regulation of muscle hyperplasia(GO:0014740) cellular response to cycloheximide(GO:0071409) |
| 0.2 | 1.6 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.1 | 0.4 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 0.1 | 0.4 | GO:0031627 | telomeric loop formation(GO:0031627) |
| 0.1 | 0.4 | GO:0035995 | detection of muscle stretch(GO:0035995) |
| 0.1 | 1.4 | GO:0007213 | G-protein coupled acetylcholine receptor signaling pathway(GO:0007213) |
| 0.1 | 0.4 | GO:2000491 | positive regulation of hepatic stellate cell activation(GO:2000491) |
| 0.1 | 0.2 | GO:0090403 | oxidative stress-induced premature senescence(GO:0090403) |
| 0.1 | 0.2 | GO:0048749 | compound eye development(GO:0048749) |
| 0.0 | 0.6 | GO:0006228 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 0.0 | 0.1 | GO:0016185 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) |
| 0.0 | 0.2 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.7 | GO:0097150 | neuronal stem cell population maintenance(GO:0097150) |
| 0.0 | 0.2 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.0 | 0.1 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) |
| 0.0 | 0.5 | GO:0036295 | cellular response to increased oxygen levels(GO:0036295) |
| 0.0 | 0.6 | GO:0002643 | regulation of tolerance induction(GO:0002643) |
| 0.0 | 0.4 | GO:0006044 | N-acetylglucosamine metabolic process(GO:0006044) |
| 0.0 | 0.1 | GO:0006655 | phosphatidylglycerol biosynthetic process(GO:0006655) |
| 0.0 | 0.4 | GO:0001783 | B cell apoptotic process(GO:0001783) |
| 0.0 | 0.2 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.0 | 0.4 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | GO:0033179 | proton-transporting V-type ATPase, V0 domain(GO:0033179) |
| 0.0 | 0.1 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) |
| 0.0 | 0.7 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 0.2 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 0.4 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.2 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.7 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.0 | 0.3 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.0 | 1.6 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 0.2 | GO:0005664 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 5.4 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.0 | 0.7 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.5 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.0 | 0.2 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.0 | 0.2 | REACTOME CDC6 ASSOCIATION WITH THE ORC ORIGIN COMPLEX | Genes involved in CDC6 association with the ORC:origin complex |
| 0.0 | 0.1 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 0.4 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.0 | 0.2 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 5.4 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.0 | 1.6 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 0.4 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 5.4 | GO:0008269 | JAK pathway signal transduction adaptor activity(GO:0008269) |
| 0.4 | 1.6 | GO:0070139 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.1 | 0.7 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 0.1 | 0.5 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.4 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.0 | 0.1 | GO:0004605 | phosphatidate cytidylyltransferase activity(GO:0004605) |
| 0.0 | 1.4 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.4 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 0.4 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.0 | 0.6 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.0 | 0.2 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.0 | 0.6 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 0.1 | GO:0031802 | type 5 metabotropic glutamate receptor binding(GO:0031802) |
| 0.0 | 0.2 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.0 | 0.2 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.0 | 0.1 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.2 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |