Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
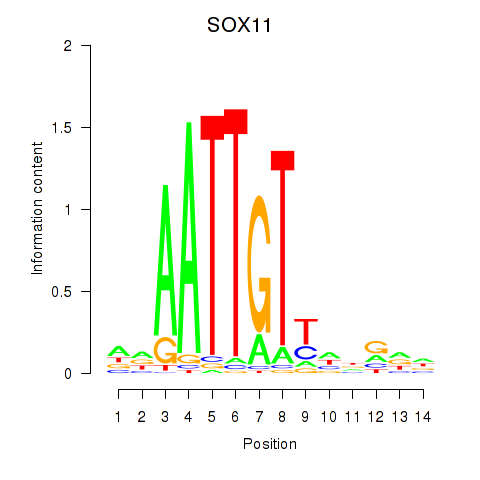
Results for SOX11
Z-value: 0.82
Transcription factors associated with SOX11
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SOX11
|
ENSG00000176887.5 | SOX11 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SOX11 | hg19_v2_chr2_+_5832799_5832799 | -0.05 | 8.5e-01 | Click! |
Activity profile of SOX11 motif
Sorted Z-values of SOX11 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SOX11
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_225266743 | 3.25 |
ENST00000409685.3 |
FAM124B |
family with sequence similarity 124B |
| chr2_-_225266711 | 2.57 |
ENST00000389874.3 |
FAM124B |
family with sequence similarity 124B |
| chr4_-_90759440 | 2.46 |
ENST00000336904.3 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr6_-_49604545 | 1.53 |
ENST00000371175.4 ENST00000229810.7 |
RHAG |
Rh-associated glycoprotein |
| chr10_-_95241951 | 1.17 |
ENST00000358334.5 ENST00000359263.4 ENST00000371488.3 |
MYOF |
myoferlin |
| chr10_-_95242044 | 1.17 |
ENST00000371501.4 ENST00000371502.4 ENST00000371489.1 |
MYOF |
myoferlin |
| chr7_-_56118981 | 1.16 |
ENST00000419984.2 ENST00000413218.1 ENST00000424596.1 |
PSPH |
phosphoserine phosphatase |
| chr3_+_69812701 | 1.13 |
ENST00000472437.1 |
MITF |
microphthalmia-associated transcription factor |
| chr1_+_196621002 | 1.03 |
ENST00000367429.4 ENST00000439155.2 |
CFH |
complement factor H |
| chr2_-_190044480 | 0.99 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr20_-_7921090 | 0.97 |
ENST00000378789.3 |
HAO1 |
hydroxyacid oxidase (glycolate oxidase) 1 |
| chr1_+_196621156 | 0.93 |
ENST00000359637.2 |
CFH |
complement factor H |
| chr6_+_26183958 | 0.89 |
ENST00000356530.3 |
HIST1H2BE |
histone cluster 1, H2be |
| chr4_-_73434498 | 0.89 |
ENST00000286657.4 |
ADAMTS3 |
ADAM metallopeptidase with thrombospondin type 1 motif, 3 |
| chrX_-_15619076 | 0.86 |
ENST00000252519.3 |
ACE2 |
angiotensin I converting enzyme 2 |
| chr10_-_5446786 | 0.75 |
ENST00000479328.1 ENST00000380419.3 |
TUBAL3 |
tubulin, alpha-like 3 |
| chr6_-_116381918 | 0.73 |
ENST00000606080.1 |
FRK |
fyn-related kinase |
| chr7_+_98923505 | 0.62 |
ENST00000432884.2 ENST00000262942.5 |
ARPC1A |
actin related protein 2/3 complex, subunit 1A, 41kDa |
| chr8_-_13134045 | 0.62 |
ENST00000512044.2 |
DLC1 |
deleted in liver cancer 1 |
| chr2_+_102928009 | 0.61 |
ENST00000404917.2 ENST00000447231.1 |
IL1RL1 |
interleukin 1 receptor-like 1 |
| chr8_+_19796381 | 0.60 |
ENST00000524029.1 ENST00000522701.1 ENST00000311322.8 |
LPL |
lipoprotein lipase |
| chr11_+_12399071 | 0.57 |
ENST00000539723.1 ENST00000550549.1 |
PARVA |
parvin, alpha |
| chrX_-_13835147 | 0.56 |
ENST00000493677.1 ENST00000355135.2 |
GPM6B |
glycoprotein M6B |
| chr12_+_32655048 | 0.55 |
ENST00000427716.2 ENST00000266482.3 |
FGD4 |
FYVE, RhoGEF and PH domain containing 4 |
| chr3_-_107777208 | 0.53 |
ENST00000398258.3 |
CD47 |
CD47 molecule |
| chr15_+_49447947 | 0.53 |
ENST00000327171.3 ENST00000560654.1 |
GALK2 |
galactokinase 2 |
| chr12_+_28410128 | 0.52 |
ENST00000381259.1 ENST00000381256.1 |
CCDC91 |
coiled-coil domain containing 91 |
| chr1_+_65730385 | 0.49 |
ENST00000263441.7 ENST00000395325.3 |
DNAJC6 |
DnaJ (Hsp40) homolog, subfamily C, member 6 |
| chr4_-_110624564 | 0.47 |
ENST00000352981.3 ENST00000265164.2 ENST00000505486.1 |
CASP6 |
caspase 6, apoptosis-related cysteine peptidase |
| chr17_-_66951474 | 0.45 |
ENST00000269080.2 |
ABCA8 |
ATP-binding cassette, sub-family A (ABC1), member 8 |
| chr19_+_39989535 | 0.44 |
ENST00000356433.5 |
DLL3 |
delta-like 3 (Drosophila) |
| chr4_-_170679024 | 0.43 |
ENST00000393381.2 |
C4orf27 |
chromosome 4 open reading frame 27 |
| chr20_-_32308028 | 0.43 |
ENST00000409299.3 ENST00000217398.3 ENST00000344022.3 |
PXMP4 |
peroxisomal membrane protein 4, 24kDa |
| chr9_+_27109133 | 0.40 |
ENST00000519097.1 ENST00000380036.4 |
TEK |
TEK tyrosine kinase, endothelial |
| chr12_+_60058458 | 0.39 |
ENST00000548610.1 |
SLC16A7 |
solute carrier family 16 (monocarboxylate transporter), member 7 |
| chr2_+_47596287 | 0.39 |
ENST00000263735.4 |
EPCAM |
epithelial cell adhesion molecule |
| chr3_-_18466026 | 0.39 |
ENST00000417717.2 |
SATB1 |
SATB homeobox 1 |
| chr1_-_241799232 | 0.38 |
ENST00000366553.1 |
CHML |
choroideremia-like (Rab escort protein 2) |
| chr15_-_37392086 | 0.37 |
ENST00000561208.1 |
MEIS2 |
Meis homeobox 2 |
| chr3_+_113616317 | 0.37 |
ENST00000440446.2 ENST00000488680.1 |
GRAMD1C |
GRAM domain containing 1C |
| chr13_-_99630233 | 0.36 |
ENST00000376460.1 ENST00000442173.1 |
DOCK9 |
dedicator of cytokinesis 9 |
| chr5_+_135496675 | 0.35 |
ENST00000507637.1 |
SMAD5 |
SMAD family member 5 |
| chr1_+_145524891 | 0.33 |
ENST00000369304.3 |
ITGA10 |
integrin, alpha 10 |
| chr6_-_36515177 | 0.33 |
ENST00000229812.7 |
STK38 |
serine/threonine kinase 38 |
| chr17_-_34344991 | 0.32 |
ENST00000591423.1 |
CCL23 |
chemokine (C-C motif) ligand 23 |
| chr4_-_159080806 | 0.31 |
ENST00000590648.1 |
FAM198B |
family with sequence similarity 198, member B |
| chr8_-_37707356 | 0.29 |
ENST00000520601.1 ENST00000521170.1 ENST00000220659.6 |
BRF2 |
BRF2, RNA polymerase III transcription initiation factor 50 kDa subunit |
| chr2_+_169757750 | 0.29 |
ENST00000375363.3 ENST00000429379.2 ENST00000421979.1 |
G6PC2 |
glucose-6-phosphatase, catalytic, 2 |
| chr4_+_183164574 | 0.28 |
ENST00000511685.1 |
TENM3 |
teneurin transmembrane protein 3 |
| chr5_+_140213815 | 0.28 |
ENST00000525929.1 ENST00000378125.3 |
PCDHA7 |
protocadherin alpha 7 |
| chr6_-_133055815 | 0.27 |
ENST00000509351.1 ENST00000417437.2 ENST00000414302.2 ENST00000423615.2 ENST00000427187.2 ENST00000275223.3 ENST00000519686.2 |
VNN3 |
vanin 3 |
| chr5_+_169780485 | 0.27 |
ENST00000377360.4 |
KCNIP1 |
Kv channel interacting protein 1 |
| chr2_-_219157250 | 0.27 |
ENST00000434015.2 ENST00000444183.1 ENST00000420341.1 ENST00000453281.1 ENST00000258412.3 ENST00000440422.1 |
TMBIM1 |
transmembrane BAX inhibitor motif containing 1 |
| chr17_+_6347729 | 0.26 |
ENST00000572447.1 |
FAM64A |
family with sequence similarity 64, member A |
| chr11_+_57425209 | 0.25 |
ENST00000533905.1 ENST00000525602.1 ENST00000302731.4 |
CLP1 |
cleavage and polyadenylation factor I subunit 1 |
| chr6_-_133055896 | 0.25 |
ENST00000367927.5 ENST00000425515.2 ENST00000207771.3 ENST00000392393.3 ENST00000450865.2 ENST00000392394.2 |
VNN3 |
vanin 3 |
| chr17_+_6347761 | 0.24 |
ENST00000250056.8 ENST00000571373.1 ENST00000570337.2 ENST00000572595.2 ENST00000576056.1 |
FAM64A |
family with sequence similarity 64, member A |
| chr4_+_169013666 | 0.24 |
ENST00000359299.3 |
ANXA10 |
annexin A10 |
| chr1_-_205744574 | 0.24 |
ENST00000367139.3 ENST00000235932.4 ENST00000437324.2 ENST00000414729.1 |
RAB7L1 |
RAB7, member RAS oncogene family-like 1 |
| chr1_-_205744205 | 0.23 |
ENST00000446390.2 |
RAB7L1 |
RAB7, member RAS oncogene family-like 1 |
| chr11_-_95657231 | 0.23 |
ENST00000409459.1 ENST00000352297.7 ENST00000393223.3 ENST00000346299.5 |
MTMR2 |
myotubularin related protein 2 |
| chr9_-_21142144 | 0.23 |
ENST00000380229.2 |
IFNW1 |
interferon, omega 1 |
| chr2_-_37068530 | 0.20 |
ENST00000593798.1 |
AC007382.1 |
Uncharacterized protein |
| chr2_+_166326157 | 0.20 |
ENST00000421875.1 ENST00000314499.7 ENST00000409664.1 |
CSRNP3 |
cysteine-serine-rich nuclear protein 3 |
| chr6_-_46048116 | 0.20 |
ENST00000185206.6 |
CLIC5 |
chloride intracellular channel 5 |
| chr8_-_54752406 | 0.19 |
ENST00000520188.1 |
ATP6V1H |
ATPase, H+ transporting, lysosomal 50/57kDa, V1 subunit H |
| chr5_+_140248518 | 0.19 |
ENST00000398640.2 |
PCDHA11 |
protocadherin alpha 11 |
| chr17_-_34345002 | 0.19 |
ENST00000293280.2 |
CCL23 |
chemokine (C-C motif) ligand 23 |
| chr6_-_11779174 | 0.18 |
ENST00000379413.2 |
ADTRP |
androgen-dependent TFPI-regulating protein |
| chr19_-_14911023 | 0.18 |
ENST00000248073.2 |
OR7C1 |
olfactory receptor, family 7, subfamily C, member 1 |
| chr6_-_49712123 | 0.17 |
ENST00000263045.4 |
CRISP3 |
cysteine-rich secretory protein 3 |
| chr6_-_11779014 | 0.17 |
ENST00000229583.5 |
ADTRP |
androgen-dependent TFPI-regulating protein |
| chr8_+_24241789 | 0.17 |
ENST00000256412.4 ENST00000538205.1 |
ADAMDEC1 |
ADAM-like, decysin 1 |
| chr3_-_87325728 | 0.16 |
ENST00000350375.2 |
POU1F1 |
POU class 1 homeobox 1 |
| chr16_+_67261008 | 0.15 |
ENST00000304800.9 ENST00000563953.1 ENST00000565201.1 |
TMEM208 |
transmembrane protein 208 |
| chr11_-_82611448 | 0.15 |
ENST00000393399.2 ENST00000313010.3 |
PRCP |
prolylcarboxypeptidase (angiotensinase C) |
| chr18_+_22040593 | 0.15 |
ENST00000256906.4 |
HRH4 |
histamine receptor H4 |
| chr5_+_108083517 | 0.15 |
ENST00000281092.4 ENST00000536402.1 |
FER |
fer (fps/fes related) tyrosine kinase |
| chr5_+_148521381 | 0.15 |
ENST00000504238.1 |
ABLIM3 |
actin binding LIM protein family, member 3 |
| chr10_-_127505167 | 0.15 |
ENST00000368786.1 |
UROS |
uroporphyrinogen III synthase |
| chr7_-_122635754 | 0.15 |
ENST00000249284.2 |
TAS2R16 |
taste receptor, type 2, member 16 |
| chr21_+_43619796 | 0.15 |
ENST00000398457.2 |
ABCG1 |
ATP-binding cassette, sub-family G (WHITE), member 1 |
| chr7_-_81399329 | 0.14 |
ENST00000453411.1 ENST00000444829.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr1_+_149754227 | 0.14 |
ENST00000444948.1 ENST00000369168.4 |
FCGR1A |
Fc fragment of IgG, high affinity Ia, receptor (CD64) |
| chr17_-_79895097 | 0.14 |
ENST00000402252.2 ENST00000583564.1 ENST00000585244.1 ENST00000337943.5 ENST00000579698.1 |
PYCR1 |
pyrroline-5-carboxylate reductase 1 |
| chr18_+_22040620 | 0.14 |
ENST00000426880.2 |
HRH4 |
histamine receptor H4 |
| chr16_-_21663919 | 0.14 |
ENST00000569602.1 |
IGSF6 |
immunoglobulin superfamily, member 6 |
| chr5_+_140753444 | 0.14 |
ENST00000517434.1 |
PCDHGA6 |
protocadherin gamma subfamily A, 6 |
| chr2_-_176046391 | 0.14 |
ENST00000392541.3 ENST00000409194.1 |
ATP5G3 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C3 (subunit 9) |
| chr9_-_95166841 | 0.13 |
ENST00000262551.4 |
OGN |
osteoglycin |
| chr12_-_10959892 | 0.13 |
ENST00000240615.2 |
TAS2R8 |
taste receptor, type 2, member 8 |
| chr3_-_47950745 | 0.13 |
ENST00000429422.1 |
MAP4 |
microtubule-associated protein 4 |
| chr4_+_88754069 | 0.12 |
ENST00000395102.4 ENST00000497649.2 |
MEPE |
matrix extracellular phosphoglycoprotein |
| chr2_+_109237717 | 0.12 |
ENST00000409441.1 |
LIMS1 |
LIM and senescent cell antigen-like domains 1 |
| chr9_+_71986182 | 0.12 |
ENST00000303068.7 |
FAM189A2 |
family with sequence similarity 189, member A2 |
| chr8_+_24241969 | 0.12 |
ENST00000522298.1 |
ADAMDEC1 |
ADAM-like, decysin 1 |
| chr7_-_81399411 | 0.11 |
ENST00000423064.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr4_+_72204755 | 0.11 |
ENST00000512686.1 ENST00000340595.3 |
SLC4A4 |
solute carrier family 4 (sodium bicarbonate cotransporter), member 4 |
| chr6_-_49712147 | 0.11 |
ENST00000433368.2 ENST00000354620.4 |
CRISP3 |
cysteine-rich secretory protein 3 |
| chr2_-_178128250 | 0.11 |
ENST00000448782.1 ENST00000446151.2 |
NFE2L2 |
nuclear factor, erythroid 2-like 2 |
| chr3_+_101504200 | 0.11 |
ENST00000422132.1 |
NXPE3 |
neurexophilin and PC-esterase domain family, member 3 |
| chr1_-_120935894 | 0.11 |
ENST00000369383.4 ENST00000369384.4 |
FCGR1B |
Fc fragment of IgG, high affinity Ib, receptor (CD64) |
| chr8_+_9413410 | 0.10 |
ENST00000520408.1 ENST00000310430.6 ENST00000522110.1 |
TNKS |
tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase |
| chr8_+_104831472 | 0.10 |
ENST00000262231.10 ENST00000507740.1 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr11_+_7110165 | 0.09 |
ENST00000306904.5 |
RBMXL2 |
RNA binding motif protein, X-linked-like 2 |
| chr1_-_115292591 | 0.09 |
ENST00000438362.2 |
CSDE1 |
cold shock domain containing E1, RNA-binding |
| chr17_-_79895154 | 0.09 |
ENST00000405481.4 ENST00000585215.1 ENST00000577624.1 ENST00000403172.4 |
PYCR1 |
pyrroline-5-carboxylate reductase 1 |
| chr14_+_39703112 | 0.09 |
ENST00000555143.1 ENST00000280082.3 |
MIA2 |
melanoma inhibitory activity 2 |
| chr1_-_150669500 | 0.08 |
ENST00000271732.3 |
GOLPH3L |
golgi phosphoprotein 3-like |
| chr8_+_104831554 | 0.08 |
ENST00000408894.2 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr11_-_118047376 | 0.07 |
ENST00000278947.5 |
SCN2B |
sodium channel, voltage-gated, type II, beta subunit |
| chr17_-_10276319 | 0.07 |
ENST00000252172.4 ENST00000418404.3 |
MYH13 |
myosin, heavy chain 13, skeletal muscle |
| chr5_+_172571445 | 0.07 |
ENST00000231668.9 ENST00000351486.5 ENST00000352523.6 ENST00000393770.4 |
BNIP1 |
BCL2/adenovirus E1B 19kDa interacting protein 1 |
| chr14_-_92198403 | 0.07 |
ENST00000553329.1 ENST00000256343.3 |
CATSPERB |
catsper channel auxiliary subunit beta |
| chr8_+_105352050 | 0.07 |
ENST00000297581.2 |
DCSTAMP |
dendrocyte expressed seven transmembrane protein |
| chr4_+_88754113 | 0.06 |
ENST00000560249.1 ENST00000540395.1 ENST00000511670.1 ENST00000361056.3 |
MEPE |
matrix extracellular phosphoglycoprotein |
| chr2_-_20251744 | 0.06 |
ENST00000175091.4 |
LAPTM4A |
lysosomal protein transmembrane 4 alpha |
| chr7_-_81399355 | 0.05 |
ENST00000457544.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr3_-_133380731 | 0.05 |
ENST00000260810.5 |
TOPBP1 |
topoisomerase (DNA) II binding protein 1 |
| chr17_+_62461569 | 0.04 |
ENST00000603557.1 ENST00000605096.1 |
MILR1 |
mast cell immunoglobulin-like receptor 1 |
| chr15_-_64673630 | 0.04 |
ENST00000558008.1 ENST00000559519.1 ENST00000380258.2 |
KIAA0101 |
KIAA0101 |
| chr10_+_32873190 | 0.04 |
ENST00000375025.4 |
C10orf68 |
Homo sapiens coiled-coil domain containing 7 (CCDC7), transcript variant 5, mRNA. |
| chrX_+_100645812 | 0.04 |
ENST00000427805.2 ENST00000553110.3 ENST00000392994.3 ENST00000409338.1 ENST00000409170.3 |
RPL36A RPL36A-HNRNPH2 |
ribosomal protein L36a RPL36A-HNRNPH2 readthrough |
| chr10_+_5238793 | 0.03 |
ENST00000263126.1 |
AKR1C4 |
aldo-keto reductase family 1, member C4 |
| chrX_-_7895755 | 0.03 |
ENST00000444736.1 ENST00000537427.1 ENST00000442940.1 |
PNPLA4 |
patatin-like phospholipase domain containing 4 |
| chr12_+_50794592 | 0.03 |
ENST00000293618.8 ENST00000429001.3 ENST00000548174.1 ENST00000548697.1 ENST00000548993.1 ENST00000398473.2 ENST00000522085.1 ENST00000518444.1 ENST00000551886.1 |
LARP4 |
La ribonucleoprotein domain family, member 4 |
| chr3_-_87325612 | 0.03 |
ENST00000561167.1 ENST00000560656.1 ENST00000344265.3 |
POU1F1 |
POU class 1 homeobox 1 |
| chr6_+_29068386 | 0.03 |
ENST00000377171.3 |
OR2J1 |
olfactory receptor, family 2, subfamily J, member 1 (gene/pseudogene) |
| chr11_-_64684672 | 0.02 |
ENST00000377264.3 ENST00000421419.2 |
ATG2A |
autophagy related 2A |
| chr15_-_78913628 | 0.02 |
ENST00000348639.3 |
CHRNA3 |
cholinergic receptor, nicotinic, alpha 3 (neuronal) |
| chr7_-_82792215 | 0.01 |
ENST00000333891.9 ENST00000423517.2 |
PCLO |
piccolo presynaptic cytomatrix protein |
| chr2_+_187371440 | 0.01 |
ENST00000445547.1 |
ZC3H15 |
zinc finger CCCH-type containing 15 |
| chr12_-_10955226 | 0.01 |
ENST00000240687.2 |
TAS2R7 |
taste receptor, type 2, member 7 |
| chrX_+_102192200 | 0.01 |
ENST00000218249.5 |
RAB40AL |
RAB40A, member RAS oncogene family-like |
| chr8_+_41386761 | 0.01 |
ENST00000523277.2 |
GINS4 |
GINS complex subunit 4 (Sld5 homolog) |
| chr1_-_111061797 | 0.01 |
ENST00000369771.2 |
KCNA10 |
potassium voltage-gated channel, shaker-related subfamily, member 10 |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0035379 | carbon dioxide transmembrane transporter activity(GO:0035379) |
| 0.5 | 2.5 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.3 | 1.0 | GO:0016899 | oxidoreductase activity, acting on the CH-OH group of donors, oxygen as acceptor(GO:0016899) |
| 0.2 | 1.2 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.2 | 0.6 | GO:0017129 | triglyceride binding(GO:0017129) |
| 0.2 | 0.5 | GO:0002113 | interleukin-33 binding(GO:0002113) |
| 0.2 | 0.5 | GO:0033858 | N-acetylgalactosamine kinase activity(GO:0033858) |
| 0.1 | 0.9 | GO:0008241 | peptidyl-dipeptidase activity(GO:0008241) |
| 0.1 | 0.5 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.1 | 0.4 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.1 | 2.0 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.1 | 0.5 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.1 | 0.3 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.1 | 0.3 | GO:0098639 | collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.1 | 0.3 | GO:0051733 | ATP-dependent polydeoxyribonucleotide 5'-hydroxyl-kinase activity(GO:0046404) polydeoxyribonucleotide kinase activity(GO:0051733) ATP-dependent polynucleotide kinase activity(GO:0051734) |
| 0.1 | 0.3 | GO:0001016 | transcription factor activity, RNA polymerase III transcription factor binding(GO:0001007) RNA polymerase III regulatory region DNA binding(GO:0001016) |
| 0.1 | 0.2 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.1 | 0.5 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.1 | 0.5 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.0 | 0.1 | GO:0034041 | sterol-transporting ATPase activity(GO:0034041) |
| 0.0 | 0.2 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.0 | 0.3 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 0.3 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.0 | 0.4 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 0.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.0 | 1.1 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.2 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.0 | 0.3 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
| 0.0 | 0.2 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.0 | 0.4 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.0 | 0.5 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.3 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.0 | 0.1 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.0 | 0.4 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.2 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 0.9 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.6 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 1.2 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.0 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.0 | 1.4 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 1.5 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.0 | 0.2 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 3.4 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.6 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 1.9 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.4 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.7 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.2 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 0.4 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 0.5 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.0 | 0.3 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.3 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.5 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0035378 | carbon dioxide transmembrane transport(GO:0035378) |
| 0.5 | 2.5 | GO:0051585 | negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) regulation of peroxidase activity(GO:2000468) positive regulation of peroxidase activity(GO:2000470) |
| 0.2 | 1.0 | GO:1903224 | regulation of endodermal cell differentiation(GO:1903224) |
| 0.2 | 2.3 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.2 | 0.9 | GO:0003051 | angiotensin-mediated drinking behavior(GO:0003051) |
| 0.2 | 0.3 | GO:0043405 | regulation of MAP kinase activity(GO:0043405) negative regulation of MAP kinase activity(GO:0043407) |
| 0.1 | 0.9 | GO:1900748 | positive regulation of vascular endothelial growth factor signaling pathway(GO:1900748) |
| 0.1 | 0.4 | GO:2000048 | negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 0.1 | 1.0 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 0.1 | 0.6 | GO:0051581 | negative regulation of neurotransmitter uptake(GO:0051581) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 0.1 | 0.5 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 0.1 | 1.2 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.1 | 0.6 | GO:0010890 | positive regulation of sequestering of triglyceride(GO:0010890) lipoprotein particle mediated signaling(GO:0055095) low-density lipoprotein particle mediated signaling(GO:0055096) |
| 0.1 | 2.0 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.1 | 0.4 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.1 | 0.3 | GO:1902044 | regulation of Fas signaling pathway(GO:1902044) |
| 0.1 | 0.5 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 0.1 | 0.3 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.1 | 0.2 | GO:2000645 | negative regulation of receptor catabolic process(GO:2000645) |
| 0.1 | 0.5 | GO:0090160 | Golgi to lysosome transport(GO:0090160) |
| 0.1 | 0.4 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.1 | 0.5 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.1 | 0.5 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.1 | 0.2 | GO:0006624 | vacuolar protein processing(GO:0006624) |
| 0.1 | 0.2 | GO:0036215 | response to stem cell factor(GO:0036215) cellular response to stem cell factor stimulus(GO:0036216) Kit signaling pathway(GO:0038109) |
| 0.0 | 0.4 | GO:0006848 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.0 | 0.1 | GO:0009726 | detection of endogenous stimulus(GO:0009726) |
| 0.0 | 0.2 | GO:0060133 | somatotropin secreting cell development(GO:0060133) |
| 0.0 | 0.3 | GO:0030423 | targeting of mRNA for destruction involved in RNA interference(GO:0030423) siRNA loading onto RISC involved in RNA interference(GO:0035087) |
| 0.0 | 0.4 | GO:0018344 | protein geranylgeranylation(GO:0018344) |
| 0.0 | 0.5 | GO:0042908 | xenobiotic transport(GO:0042908) |
| 0.0 | 0.6 | GO:1900119 | positive regulation of execution phase of apoptosis(GO:1900119) |
| 0.0 | 0.3 | GO:0060665 | regulation of branching involved in salivary gland morphogenesis by mesenchymal-epithelial signaling(GO:0060665) |
| 0.0 | 0.2 | GO:0002353 | regulation of thyroid hormone mediated signaling pathway(GO:0002155) kinin cascade(GO:0002254) plasma kallikrein-kinin cascade(GO:0002353) |
| 0.0 | 0.6 | GO:0071670 | smooth muscle cell chemotaxis(GO:0071670) |
| 0.0 | 0.2 | GO:0055129 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.0 | 0.1 | GO:0006780 | uroporphyrinogen III biosynthetic process(GO:0006780) uroporphyrinogen III metabolic process(GO:0046502) |
| 0.0 | 0.1 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.0 | 0.5 | GO:0008228 | opsonization(GO:0008228) |
| 0.0 | 0.1 | GO:1901377 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) regulation of glutathione biosynthetic process(GO:1903786) positive regulation of glutathione biosynthetic process(GO:1903788) |
| 0.0 | 0.5 | GO:0006012 | galactose metabolic process(GO:0006012) |
| 0.0 | 1.1 | GO:0030318 | melanocyte differentiation(GO:0030318) |
| 0.0 | 0.4 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.0 | 0.1 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.0 | 0.2 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 0.2 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 0.3 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.1 | GO:0072675 | multinuclear osteoclast differentiation(GO:0072674) osteoclast fusion(GO:0072675) |
| 0.0 | 0.1 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.0 | 0.6 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.1 | GO:0046684 | response to pyrethroid(GO:0046684) |
| 0.0 | 0.2 | GO:0033141 | positive regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033141) |
| 0.0 | 0.3 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.5 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.3 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 1.0 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 1.3 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 2.3 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.0 | 0.3 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.5 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 0.3 | GO:0034680 | integrin alpha10-beta1 complex(GO:0034680) |
| 0.1 | 0.4 | GO:0005968 | Rab-protein geranylgeranyltransferase complex(GO:0005968) |
| 0.1 | 2.5 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.1 | 0.3 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.0 | 0.2 | GO:0000221 | vacuolar proton-transporting V-type ATPase, V1 domain(GO:0000221) |
| 0.0 | 0.6 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.0 | 3.0 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 0.6 | GO:0042627 | chylomicron(GO:0042627) |
| 0.0 | 1.0 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 0.3 | GO:0071141 | SMAD protein complex(GO:0071141) |
| 0.0 | 0.1 | GO:0036128 | CatSper complex(GO:0036128) |
| 0.0 | 0.4 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.0 | 1.0 | GO:0035580 | specific granule lumen(GO:0035580) |