Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
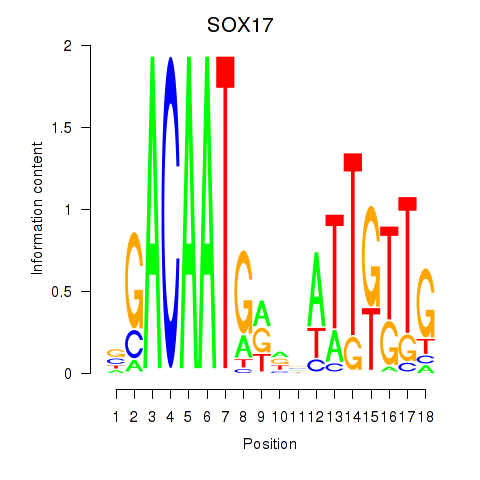
Results for SOX17
Z-value: 1.43
Transcription factors associated with SOX17
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SOX17
|
ENSG00000164736.5 | SOX17 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SOX17 | hg19_v2_chr8_+_55370487_55370503 | -0.34 | 1.9e-01 | Click! |
Activity profile of SOX17 motif
Sorted Z-values of SOX17 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SOX17
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_116708302 | 3.11 |
ENST00000375320.1 ENST00000359492.2 ENST00000375329.2 ENST00000375323.1 |
APOA1 |
apolipoprotein A-I |
| chr17_-_64225508 | 2.75 |
ENST00000205948.6 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chr7_-_22234381 | 2.56 |
ENST00000458533.1 |
RAPGEF5 |
Rap guanine nucleotide exchange factor (GEF) 5 |
| chr7_-_29186008 | 2.33 |
ENST00000396276.3 ENST00000265394.5 |
CPVL |
carboxypeptidase, vitellogenic-like |
| chr21_-_33975547 | 2.33 |
ENST00000431599.1 |
C21orf59 |
chromosome 21 open reading frame 59 |
| chr20_+_61448376 | 2.25 |
ENST00000343916.3 |
COL9A3 |
collagen, type IX, alpha 3 |
| chr12_+_69742121 | 2.20 |
ENST00000261267.2 ENST00000549690.1 ENST00000548839.1 |
LYZ |
lysozyme |
| chr10_-_69597915 | 2.03 |
ENST00000225171.2 |
DNAJC12 |
DnaJ (Hsp40) homolog, subfamily C, member 12 |
| chr4_+_69681710 | 2.00 |
ENST00000265403.7 ENST00000458688.2 |
UGT2B10 |
UDP glucuronosyltransferase 2 family, polypeptide B10 |
| chr9_+_117092149 | 1.97 |
ENST00000431067.2 ENST00000412657.1 |
ORM2 |
orosomucoid 2 |
| chr14_-_55369525 | 1.91 |
ENST00000543643.2 ENST00000536224.2 ENST00000395514.1 ENST00000491895.2 |
GCH1 |
GTP cyclohydrolase 1 |
| chr16_+_2867228 | 1.85 |
ENST00000005995.3 ENST00000574813.1 |
PRSS21 |
protease, serine, 21 (testisin) |
| chrX_+_49178536 | 1.82 |
ENST00000442437.2 |
GAGE12J |
G antigen 12J |
| chr13_-_46756351 | 1.81 |
ENST00000323076.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chrX_+_49216659 | 1.70 |
ENST00000415752.1 |
GAGE12I |
G antigen 12I |
| chr8_-_17752912 | 1.55 |
ENST00000398054.1 ENST00000381840.2 |
FGL1 |
fibrinogen-like 1 |
| chrX_-_6453159 | 1.48 |
ENST00000381089.3 ENST00000398729.1 |
VCX3A |
variable charge, X-linked 3A |
| chr4_+_100495864 | 1.46 |
ENST00000265517.5 ENST00000422897.2 |
MTTP |
microsomal triglyceride transfer protein |
| chr8_-_17752996 | 1.45 |
ENST00000381841.2 ENST00000427924.1 |
FGL1 |
fibrinogen-like 1 |
| chr4_+_128554081 | 1.43 |
ENST00000335251.6 ENST00000296461.5 |
INTU |
inturned planar cell polarity protein |
| chr2_+_219283815 | 1.42 |
ENST00000248444.5 ENST00000454069.1 ENST00000392114.2 |
VIL1 |
villin 1 |
| chrX_+_49235708 | 1.40 |
ENST00000381725.1 |
GAGE2B |
G antigen 2B |
| chr15_-_20193370 | 1.39 |
ENST00000558565.2 |
IGHV3OR15-7 |
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr7_+_29234028 | 1.38 |
ENST00000222792.6 |
CHN2 |
chimerin 2 |
| chr4_+_71063641 | 1.34 |
ENST00000514097.1 |
ODAM |
odontogenic, ameloblast asssociated |
| chr18_-_24283586 | 1.31 |
ENST00000579458.1 |
U3 |
Small nucleolar RNA U3 |
| chr1_-_109935819 | 1.31 |
ENST00000538502.1 |
SORT1 |
sortilin 1 |
| chr6_-_32920794 | 1.26 |
ENST00000395305.3 ENST00000395303.3 ENST00000374843.4 ENST00000429234.1 |
HLA-DMA XXbac-BPG181M17.5 |
major histocompatibility complex, class II, DM alpha Uncharacterized protein |
| chrX_-_40036520 | 1.25 |
ENST00000406200.2 ENST00000378455.4 ENST00000342274.4 |
BCOR |
BCL6 corepressor |
| chr11_+_125365110 | 1.21 |
ENST00000527818.1 |
AP000708.1 |
AP000708.1 |
| chr12_-_71533055 | 1.21 |
ENST00000552128.1 |
TSPAN8 |
tetraspanin 8 |
| chr22_-_29107919 | 1.20 |
ENST00000434810.1 ENST00000456369.1 |
CHEK2 |
checkpoint kinase 2 |
| chr1_+_100315613 | 1.18 |
ENST00000361915.3 |
AGL |
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase |
| chr2_+_86668464 | 1.18 |
ENST00000409064.1 |
KDM3A |
lysine (K)-specific demethylase 3A |
| chr7_+_29234101 | 1.16 |
ENST00000435288.2 |
CHN2 |
chimerin 2 |
| chr9_-_100935043 | 1.16 |
ENST00000343933.5 |
CORO2A |
coronin, actin binding protein, 2A |
| chr16_+_33629600 | 1.15 |
ENST00000562905.2 |
IGHV3OR16-13 |
immunoglobulin heavy variable 3/OR16-13 (non-functional) |
| chr10_-_115614127 | 1.14 |
ENST00000369305.1 |
DCLRE1A |
DNA cross-link repair 1A |
| chr18_+_20513782 | 1.13 |
ENST00000399722.2 ENST00000399725.2 ENST00000399721.2 ENST00000583594.1 |
RBBP8 |
retinoblastoma binding protein 8 |
| chr11_-_35440579 | 1.12 |
ENST00000606205.1 |
SLC1A2 |
solute carrier family 1 (glial high affinity glutamate transporter), member 2 |
| chr14_-_107219365 | 1.12 |
ENST00000424969.2 |
IGHV3-74 |
immunoglobulin heavy variable 3-74 |
| chrX_+_49197607 | 1.10 |
ENST00000402590.3 |
GAGE2E |
G antigen 2E |
| chr1_-_205091115 | 1.09 |
ENST00000264515.6 ENST00000367164.1 |
RBBP5 |
retinoblastoma binding protein 5 |
| chr12_+_25205446 | 1.07 |
ENST00000557489.1 ENST00000354454.3 ENST00000536173.1 |
LRMP |
lymphoid-restricted membrane protein |
| chrX_+_49363665 | 1.04 |
ENST00000381700.6 |
GAGE1 |
G antigen 1 |
| chr7_+_135242652 | 1.03 |
ENST00000285968.6 ENST00000440390.2 |
NUP205 |
nucleoporin 205kDa |
| chr16_+_2867164 | 1.02 |
ENST00000455114.1 ENST00000450020.3 |
PRSS21 |
protease, serine, 21 (testisin) |
| chr12_+_25205568 | 1.00 |
ENST00000548766.1 ENST00000556887.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr2_-_89399845 | 1.00 |
ENST00000479981.1 |
IGKV1-16 |
immunoglobulin kappa variable 1-16 |
| chrX_+_8432871 | 0.98 |
ENST00000381032.1 ENST00000453306.1 ENST00000444481.1 |
VCX3B |
variable charge, X-linked 3B |
| chr20_-_62199427 | 0.95 |
ENST00000427522.2 |
HELZ2 |
helicase with zinc finger 2, transcriptional coactivator |
| chr6_+_32605195 | 0.94 |
ENST00000374949.2 |
HLA-DQA1 |
major histocompatibility complex, class II, DQ alpha 1 |
| chr6_+_32605134 | 0.92 |
ENST00000343139.5 ENST00000395363.1 ENST00000496318.1 |
HLA-DQA1 |
major histocompatibility complex, class II, DQ alpha 1 |
| chr2_-_170430366 | 0.92 |
ENST00000453153.2 ENST00000445210.1 |
FASTKD1 |
FAST kinase domains 1 |
| chr1_+_146714291 | 0.91 |
ENST00000431239.1 ENST00000369259.3 ENST00000369258.4 ENST00000361293.5 |
CHD1L |
chromodomain helicase DNA binding protein 1-like |
| chr2_-_158300556 | 0.90 |
ENST00000264192.3 |
CYTIP |
cytohesin 1 interacting protein |
| chrX_-_8139308 | 0.87 |
ENST00000317103.4 |
VCX2 |
variable charge, X-linked 2 |
| chr19_-_55720824 | 0.86 |
ENST00000376350.3 ENST00000263434.5 |
PTPRH |
protein tyrosine phosphatase, receptor type, H |
| chr2_-_216003127 | 0.84 |
ENST00000412081.1 ENST00000272895.7 |
ABCA12 |
ATP-binding cassette, sub-family A (ABC1), member 12 |
| chr12_+_25205666 | 0.82 |
ENST00000547044.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr1_-_160492994 | 0.81 |
ENST00000368055.1 ENST00000368057.3 ENST00000368059.3 |
SLAMF6 |
SLAM family member 6 |
| chr4_+_170581213 | 0.80 |
ENST00000507875.1 |
CLCN3 |
chloride channel, voltage-sensitive 3 |
| chr12_+_9822331 | 0.80 |
ENST00000545918.1 ENST00000543300.1 ENST00000261339.6 ENST00000466035.2 |
CLEC2D |
C-type lectin domain family 2, member D |
| chr12_-_498620 | 0.79 |
ENST00000399788.2 ENST00000382815.4 |
KDM5A |
lysine (K)-specific demethylase 5A |
| chr20_+_36405665 | 0.79 |
ENST00000373469.1 |
CTNNBL1 |
catenin, beta like 1 |
| chr15_+_93749295 | 0.75 |
ENST00000599897.1 |
AC112693.2 |
AC112693.2 |
| chr6_+_31916733 | 0.74 |
ENST00000483004.1 |
CFB |
complement factor B |
| chr2_+_187371440 | 0.73 |
ENST00000445547.1 |
ZC3H15 |
zinc finger CCCH-type containing 15 |
| chr6_-_26216872 | 0.73 |
ENST00000244601.3 |
HIST1H2BG |
histone cluster 1, H2bg |
| chr15_-_33360085 | 0.72 |
ENST00000334528.9 |
FMN1 |
formin 1 |
| chr15_-_47426320 | 0.72 |
ENST00000557832.1 |
FKSG62 |
FKSG62 |
| chr11_-_85565906 | 0.70 |
ENST00000544076.1 |
AP000974.1 |
CDNA FLJ26432 fis, clone KDN01418; Uncharacterized protein |
| chr11_-_3818688 | 0.70 |
ENST00000355260.3 ENST00000397004.4 ENST00000397007.4 ENST00000532475.1 |
NUP98 |
nucleoporin 98kDa |
| chr2_+_71357434 | 0.70 |
ENST00000244230.2 |
MPHOSPH10 |
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr16_-_71523236 | 0.68 |
ENST00000288177.5 ENST00000569072.1 |
ZNF19 |
zinc finger protein 19 |
| chr2_-_157198860 | 0.68 |
ENST00000409572.1 |
NR4A2 |
nuclear receptor subfamily 4, group A, member 2 |
| chr16_+_29467780 | 0.68 |
ENST00000395400.3 |
SULT1A4 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 4 |
| chr6_+_167704838 | 0.67 |
ENST00000366829.2 |
UNC93A |
unc-93 homolog A (C. elegans) |
| chr17_+_41132564 | 0.67 |
ENST00000361677.1 ENST00000589705.1 |
RUNDC1 |
RUN domain containing 1 |
| chr2_-_89417335 | 0.67 |
ENST00000490686.1 |
IGKV1-17 |
immunoglobulin kappa variable 1-17 |
| chr9_-_35650900 | 0.67 |
ENST00000259608.3 |
SIT1 |
signaling threshold regulating transmembrane adaptor 1 |
| chr2_+_90139056 | 0.66 |
ENST00000492446.1 |
IGKV1D-16 |
immunoglobulin kappa variable 1D-16 |
| chr6_+_10556215 | 0.66 |
ENST00000316170.3 |
GCNT2 |
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme (I blood group) |
| chr6_+_167704798 | 0.66 |
ENST00000230256.3 |
UNC93A |
unc-93 homolog A (C. elegans) |
| chr1_+_158801095 | 0.65 |
ENST00000368141.4 |
MNDA |
myeloid cell nuclear differentiation antigen |
| chr2_+_33661382 | 0.64 |
ENST00000402538.3 |
RASGRP3 |
RAS guanyl releasing protein 3 (calcium and DAG-regulated) |
| chr2_+_231090433 | 0.64 |
ENST00000486687.2 ENST00000350136.5 ENST00000392045.3 ENST00000417495.3 ENST00000343805.6 ENST00000420434.3 |
SP140 |
SP140 nuclear body protein |
| chr14_+_77882741 | 0.63 |
ENST00000595520.1 |
FKSG61 |
FKSG61 |
| chrX_+_48380205 | 0.63 |
ENST00000446158.1 ENST00000414061.1 |
EBP |
emopamil binding protein (sterol isomerase) |
| chr6_-_135271260 | 0.62 |
ENST00000265605.2 |
ALDH8A1 |
aldehyde dehydrogenase 8 family, member A1 |
| chr1_+_43124087 | 0.60 |
ENST00000304979.3 ENST00000372550.1 ENST00000440068.1 |
PPIH |
peptidylprolyl isomerase H (cyclophilin H) |
| chr5_+_44809027 | 0.60 |
ENST00000507110.1 |
MRPS30 |
mitochondrial ribosomal protein S30 |
| chr21_+_44589118 | 0.58 |
ENST00000291554.2 |
CRYAA |
crystallin, alpha A |
| chr13_+_37581115 | 0.58 |
ENST00000481013.1 |
EXOSC8 |
exosome component 8 |
| chr6_-_45345597 | 0.57 |
ENST00000371460.1 ENST00000371459.1 |
SUPT3H |
suppressor of Ty 3 homolog (S. cerevisiae) |
| chr1_-_6662919 | 0.54 |
ENST00000377658.4 ENST00000377663.3 |
KLHL21 |
kelch-like family member 21 |
| chr14_-_106518922 | 0.54 |
ENST00000390598.2 |
IGHV3-7 |
immunoglobulin heavy variable 3-7 |
| chr3_-_58200398 | 0.54 |
ENST00000318316.3 ENST00000460422.1 ENST00000483681.1 |
DNASE1L3 |
deoxyribonuclease I-like 3 |
| chr7_+_95115210 | 0.54 |
ENST00000428113.1 ENST00000325885.5 |
ASB4 |
ankyrin repeat and SOCS box containing 4 |
| chr6_-_135271219 | 0.53 |
ENST00000367847.2 ENST00000367845.2 |
ALDH8A1 |
aldehyde dehydrogenase 8 family, member A1 |
| chr2_+_89184868 | 0.53 |
ENST00000390243.2 |
IGKV4-1 |
immunoglobulin kappa variable 4-1 |
| chr3_+_98451093 | 0.52 |
ENST00000483910.1 ENST00000460774.1 |
ST3GAL6 |
ST3 beta-galactoside alpha-2,3-sialyltransferase 6 |
| chr17_-_55038375 | 0.52 |
ENST00000240316.4 |
COIL |
coilin |
| chr8_+_21916710 | 0.51 |
ENST00000523266.1 ENST00000519907.1 |
DMTN |
dematin actin binding protein |
| chr2_+_103035102 | 0.50 |
ENST00000264260.2 |
IL18RAP |
interleukin 18 receptor accessory protein |
| chr2_-_58468437 | 0.50 |
ENST00000403676.1 ENST00000427708.2 ENST00000403295.3 ENST00000446381.1 ENST00000417361.1 ENST00000233741.4 ENST00000402135.3 ENST00000540646.1 ENST00000449070.1 |
FANCL |
Fanconi anemia, complementation group L |
| chr2_+_120687335 | 0.49 |
ENST00000544261.1 |
PTPN4 |
protein tyrosine phosphatase, non-receptor type 4 (megakaryocyte) |
| chr19_+_41882598 | 0.48 |
ENST00000447302.2 ENST00000544232.1 ENST00000542945.1 ENST00000540732.1 |
TMEM91 CTC-435M10.3 |
transmembrane protein 91 2-oxoisovalerate dehydrogenase subunit alpha, mitochondrial; Uncharacterized protein |
| chr15_-_28778117 | 0.47 |
ENST00000525590.2 ENST00000329523.6 |
GOLGA8G |
golgin A8 family, member G |
| chr19_-_49956728 | 0.46 |
ENST00000601825.1 ENST00000596049.1 ENST00000599366.1 ENST00000597415.1 |
PIH1D1 |
PIH1 domain containing 1 |
| chr2_+_234580499 | 0.46 |
ENST00000354728.4 |
UGT1A9 |
UDP glucuronosyltransferase 1 family, polypeptide A9 |
| chr17_+_41158742 | 0.46 |
ENST00000415816.2 ENST00000438323.2 |
IFI35 |
interferon-induced protein 35 |
| chr17_-_41132410 | 0.46 |
ENST00000409446.3 ENST00000453594.1 ENST00000409399.1 ENST00000421990.2 |
PTGES3L PTGES3L-AARSD1 |
prostaglandin E synthase 3 (cytosolic)-like PTGES3L-AARSD1 readthrough |
| chr19_-_19843900 | 0.46 |
ENST00000344099.3 |
ZNF14 |
zinc finger protein 14 |
| chr8_-_17942432 | 0.46 |
ENST00000381733.4 ENST00000314146.10 |
ASAH1 |
N-acylsphingosine amidohydrolase (acid ceramidase) 1 |
| chr2_+_234580525 | 0.46 |
ENST00000609637.1 |
UGT1A1 |
UDP glucuronosyltransferase 1 family, polypeptide A8 |
| chr6_+_26217159 | 0.45 |
ENST00000303910.2 |
HIST1H2AE |
histone cluster 1, H2ae |
| chr11_+_3819049 | 0.45 |
ENST00000396986.2 ENST00000300730.6 ENST00000532535.1 ENST00000396993.4 ENST00000396991.2 ENST00000532523.1 ENST00000459679.1 ENST00000464261.1 ENST00000464906.2 ENST00000464441.1 |
PGAP2 |
post-GPI attachment to proteins 2 |
| chr2_+_71357744 | 0.42 |
ENST00000498451.2 |
MPHOSPH10 |
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr14_+_39703084 | 0.41 |
ENST00000553728.1 |
RP11-407N17.3 |
cTAGE family member 5 isoform 4 |
| chr6_-_119256311 | 0.41 |
ENST00000316316.6 |
MCM9 |
minichromosome maintenance complex component 9 |
| chrX_+_70503037 | 0.41 |
ENST00000535149.1 |
NONO |
non-POU domain containing, octamer-binding |
| chr12_-_57443886 | 0.41 |
ENST00000300119.3 |
MYO1A |
myosin IA |
| chr4_-_87028478 | 0.40 |
ENST00000515400.1 ENST00000395157.3 |
MAPK10 |
mitogen-activated protein kinase 10 |
| chr3_+_98451275 | 0.40 |
ENST00000265261.6 ENST00000497008.1 |
ST3GAL6 |
ST3 beta-galactoside alpha-2,3-sialyltransferase 6 |
| chr1_+_13641973 | 0.40 |
ENST00000330087.5 |
PRAMEF15 |
PRAME family member 15 |
| chr19_+_10765003 | 0.40 |
ENST00000407004.3 ENST00000589998.1 ENST00000589600.1 |
ILF3 |
interleukin enhancer binding factor 3, 90kDa |
| chr6_+_126240442 | 0.39 |
ENST00000448104.1 ENST00000438495.1 ENST00000444128.1 |
NCOA7 |
nuclear receptor coactivator 7 |
| chr16_-_70239683 | 0.39 |
ENST00000601706.1 |
AC009060.1 |
Uncharacterized protein |
| chr1_+_13359819 | 0.38 |
ENST00000376168.1 |
PRAMEF5 |
PRAME family member 5 |
| chr22_-_32058166 | 0.38 |
ENST00000435900.1 ENST00000336566.4 |
PISD |
phosphatidylserine decarboxylase |
| chr7_+_16828866 | 0.38 |
ENST00000597084.1 |
AC073333.1 |
Uncharacterized protein |
| chr11_-_7695437 | 0.38 |
ENST00000533558.1 ENST00000527542.1 ENST00000531096.1 |
CYB5R2 |
cytochrome b5 reductase 2 |
| chr6_+_27107053 | 0.38 |
ENST00000354348.2 |
HIST1H4I |
histone cluster 1, H4i |
| chr14_-_68066975 | 0.37 |
ENST00000559581.1 ENST00000560722.1 ENST00000559415.1 ENST00000216452.4 |
PIGH |
phosphatidylinositol glycan anchor biosynthesis, class H |
| chr1_-_13007420 | 0.35 |
ENST00000376189.1 |
PRAMEF6 |
PRAME family member 6 |
| chr3_-_11610255 | 0.35 |
ENST00000424529.2 |
VGLL4 |
vestigial like 4 (Drosophila) |
| chr4_-_186877806 | 0.35 |
ENST00000355634.5 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr2_+_232573222 | 0.35 |
ENST00000341369.7 ENST00000409683.1 |
PTMA |
prothymosin, alpha |
| chr4_-_170897045 | 0.35 |
ENST00000508313.1 |
RP11-205M3.3 |
RP11-205M3.3 |
| chr2_-_74735707 | 0.34 |
ENST00000233630.6 |
PCGF1 |
polycomb group ring finger 1 |
| chr3_+_185304059 | 0.34 |
ENST00000427465.2 |
SENP2 |
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr7_-_91808845 | 0.34 |
ENST00000343318.5 |
LRRD1 |
leucine-rich repeats and death domain containing 1 |
| chr10_+_76871353 | 0.32 |
ENST00000542569.1 |
SAMD8 |
sterile alpha motif domain containing 8 |
| chr5_-_16509101 | 0.32 |
ENST00000399793.2 |
FAM134B |
family with sequence similarity 134, member B |
| chr2_+_10262857 | 0.31 |
ENST00000304567.5 |
RRM2 |
ribonucleotide reductase M2 |
| chr4_-_70361579 | 0.31 |
ENST00000512583.1 |
UGT2B4 |
UDP glucuronosyltransferase 2 family, polypeptide B4 |
| chr19_+_10764937 | 0.31 |
ENST00000449870.1 ENST00000318511.3 ENST00000420083.1 |
ILF3 |
interleukin enhancer binding factor 3, 90kDa |
| chr1_-_13005246 | 0.31 |
ENST00000415464.2 |
PRAMEF6 |
PRAME family member 6 |
| chr5_+_135468516 | 0.31 |
ENST00000507118.1 ENST00000511116.1 ENST00000545279.1 ENST00000545620.1 |
SMAD5 |
SMAD family member 5 |
| chr10_-_72362515 | 0.30 |
ENST00000373209.2 ENST00000441259.1 |
PRF1 |
perforin 1 (pore forming protein) |
| chr20_-_33539655 | 0.30 |
ENST00000451957.2 |
GSS |
glutathione synthetase |
| chr6_-_26659913 | 0.30 |
ENST00000480036.1 ENST00000415922.2 |
ZNF322 |
zinc finger protein 322 |
| chr1_-_45805667 | 0.29 |
ENST00000488731.2 ENST00000435155.1 |
MUTYH |
mutY homolog |
| chr1_+_10509971 | 0.29 |
ENST00000320498.4 |
CORT |
cortistatin |
| chrX_+_100645812 | 0.29 |
ENST00000427805.2 ENST00000553110.3 ENST00000392994.3 ENST00000409338.1 ENST00000409170.3 |
RPL36A RPL36A-HNRNPH2 |
ribosomal protein L36a RPL36A-HNRNPH2 readthrough |
| chr1_+_171217622 | 0.29 |
ENST00000433267.1 ENST00000367750.3 |
FMO1 |
flavin containing monooxygenase 1 |
| chr3_+_111260954 | 0.28 |
ENST00000283285.5 |
CD96 |
CD96 molecule |
| chr1_+_84873913 | 0.28 |
ENST00000370662.3 |
DNASE2B |
deoxyribonuclease II beta |
| chr17_-_61959202 | 0.28 |
ENST00000449787.2 ENST00000456543.2 ENST00000423893.2 ENST00000332800.7 |
GH2 |
growth hormone 2 |
| chr2_-_231090344 | 0.27 |
ENST00000540870.1 ENST00000416610.1 |
SP110 |
SP110 nuclear body protein |
| chr6_-_25042231 | 0.27 |
ENST00000510784.2 |
FAM65B |
family with sequence similarity 65, member B |
| chr19_-_22379753 | 0.27 |
ENST00000397121.2 |
ZNF676 |
zinc finger protein 676 |
| chr1_+_45805728 | 0.27 |
ENST00000539779.1 |
TOE1 |
target of EGR1, member 1 (nuclear) |
| chr6_-_49755019 | 0.27 |
ENST00000304801.3 |
PGK2 |
phosphoglycerate kinase 2 |
| chr1_-_45805607 | 0.26 |
ENST00000372104.1 ENST00000448481.1 ENST00000483127.1 ENST00000528013.2 ENST00000456914.2 |
MUTYH |
mutY homolog |
| chr11_-_102496063 | 0.26 |
ENST00000260228.2 |
MMP20 |
matrix metallopeptidase 20 |
| chr11_-_3818932 | 0.26 |
ENST00000324932.7 ENST00000359171.4 |
NUP98 |
nucleoporin 98kDa |
| chrX_-_109590174 | 0.26 |
ENST00000372054.1 |
GNG5P2 |
guanine nucleotide binding protein (G protein), gamma 5 pseudogene 2 |
| chr1_-_145076186 | 0.26 |
ENST00000369348.3 |
PDE4DIP |
phosphodiesterase 4D interacting protein |
| chr4_-_70361615 | 0.25 |
ENST00000305107.6 |
UGT2B4 |
UDP glucuronosyltransferase 2 family, polypeptide B4 |
| chr17_+_68100989 | 0.25 |
ENST00000585558.1 ENST00000392670.1 |
KCNJ16 |
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr4_-_168155577 | 0.24 |
ENST00000541354.1 ENST00000509854.1 ENST00000512681.1 ENST00000421836.2 ENST00000510741.1 ENST00000510403.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr8_-_95220775 | 0.23 |
ENST00000441892.2 ENST00000521491.1 ENST00000027335.3 |
CDH17 |
cadherin 17, LI cadherin (liver-intestine) |
| chr7_-_102257139 | 0.23 |
ENST00000521076.1 ENST00000462172.1 ENST00000522801.1 ENST00000449970.2 ENST00000262940.7 |
RASA4 |
RAS p21 protein activator 4 |
| chr1_+_174844645 | 0.23 |
ENST00000486220.1 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr17_+_27181956 | 0.22 |
ENST00000254928.5 ENST00000580917.2 |
ERAL1 |
Era-like 12S mitochondrial rRNA chaperone 1 |
| chr17_-_39646116 | 0.22 |
ENST00000328119.6 |
KRT36 |
keratin 36 |
| chr1_+_45805342 | 0.22 |
ENST00000372090.5 |
TOE1 |
target of EGR1, member 1 (nuclear) |
| chr6_+_136172820 | 0.22 |
ENST00000308191.6 |
PDE7B |
phosphodiesterase 7B |
| chrX_-_38186811 | 0.22 |
ENST00000318842.7 |
RPGR |
retinitis pigmentosa GTPase regulator |
| chr18_+_22040593 | 0.22 |
ENST00000256906.4 |
HRH4 |
histamine receptor H4 |
| chr12_-_323689 | 0.21 |
ENST00000428720.1 |
SLC6A12 |
solute carrier family 6 (neurotransmitter transporter), member 12 |
| chr9_+_100174232 | 0.21 |
ENST00000355295.4 |
TDRD7 |
tudor domain containing 7 |
| chr20_+_47662805 | 0.21 |
ENST00000262982.2 ENST00000542325.1 |
CSE1L |
CSE1 chromosome segregation 1-like (yeast) |
| chr6_-_133079022 | 0.21 |
ENST00000525289.1 ENST00000326499.6 |
VNN2 |
vanin 2 |
| chr16_+_16434185 | 0.20 |
ENST00000524823.2 |
AC138969.4 |
Protein PKD1P1 |
| chr17_+_57297807 | 0.20 |
ENST00000284116.4 ENST00000581140.1 ENST00000581276.1 |
GDPD1 |
glycerophosphodiester phosphodiesterase domain containing 1 |
| chr19_+_7011509 | 0.20 |
ENST00000377296.3 |
AC025278.1 |
Uncharacterized protein |
| chr4_-_122085469 | 0.19 |
ENST00000057513.3 |
TNIP3 |
TNFAIP3 interacting protein 3 |
| chr1_+_24969755 | 0.19 |
ENST00000447431.2 ENST00000374389.4 |
SRRM1 |
serine/arginine repetitive matrix 1 |
| chr1_+_171217677 | 0.19 |
ENST00000402921.2 |
FMO1 |
flavin containing monooxygenase 1 |
| chr10_-_91295304 | 0.19 |
ENST00000341233.4 ENST00000371790.4 |
SLC16A12 |
solute carrier family 16, member 12 |
| chr2_-_220174166 | 0.19 |
ENST00000409251.3 ENST00000451506.1 ENST00000295718.2 ENST00000446182.1 |
PTPRN |
protein tyrosine phosphatase, receptor type, N |
| chr5_-_10308125 | 0.18 |
ENST00000296658.3 |
CMBL |
carboxymethylenebutenolidase homolog (Pseudomonas) |
| chr19_+_15852203 | 0.18 |
ENST00000305892.1 |
OR10H3 |
olfactory receptor, family 10, subfamily H, member 3 |
| chr19_-_19887185 | 0.18 |
ENST00000590092.1 ENST00000589910.1 ENST00000589508.1 ENST00000586645.1 ENST00000586816.1 ENST00000588223.1 ENST00000586885.1 |
LINC00663 |
long intergenic non-protein coding RNA 663 |
| chr3_+_39093481 | 0.18 |
ENST00000302313.5 ENST00000544962.1 ENST00000396258.3 ENST00000418020.1 |
WDR48 |
WD repeat domain 48 |
| chr22_+_42017987 | 0.17 |
ENST00000405506.1 |
XRCC6 |
X-ray repair complementing defective repair in Chinese hamster cells 6 |
| chr9_-_21239978 | 0.17 |
ENST00000380222.2 |
IFNA14 |
interferon, alpha 14 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.0 | 2.3 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 3.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 2.5 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 0.7 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.0 | 1.3 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.0 | 0.7 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.0 | 0.5 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.9 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.7 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.0 | 1.7 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 0.4 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 0.3 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 0.3 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.7 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.6 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.3 | PID CMYB PATHWAY | C-MYB transcription factor network |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 3.1 | GO:0034365 | discoidal high-density lipoprotein particle(GO:0034365) |
| 0.4 | 2.2 | GO:0005594 | collagen type IX trimer(GO:0005594) |
| 0.3 | 1.0 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.2 | 3.0 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.2 | 1.1 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.2 | 1.0 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.2 | 2.7 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.2 | 0.6 | GO:0071001 | U4/U6 snRNP(GO:0071001) |
| 0.1 | 3.1 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.1 | 0.8 | GO:0097209 | epidermal lamellar body(GO:0097209) |
| 0.1 | 0.8 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 0.6 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.1 | 1.1 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.1 | 1.8 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 0.8 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.1 | 0.3 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 0.1 | 1.4 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.1 | 0.5 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 0.1 | 2.9 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 0.5 | GO:0005827 | polar microtubule(GO:0005827) |
| 0.1 | 0.6 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.1 | 0.6 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.1 | 0.6 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.1 | 0.4 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.0 | 0.0 | GO:0097059 | CNTFR-CLCF1 complex(GO:0097059) |
| 0.0 | 4.8 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.0 | 0.3 | GO:0044194 | cytolytic granule(GO:0044194) |
| 0.0 | 2.9 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.0 | 0.8 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.0 | 1.3 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 0.4 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.0 | 0.5 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.0 | 1.1 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 0.2 | GO:0043564 | Ku70:Ku80 complex(GO:0043564) |
| 0.0 | 0.9 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.0 | 0.3 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.0 | 2.3 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 1.6 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 0.1 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.0 | 3.1 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.1 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 0.2 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 2.3 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 1.0 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.2 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 1.2 | GO:0016529 | sarcoplasmic reticulum(GO:0016529) |
| 0.0 | 0.2 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.3 | GO:0071141 | SMAD protein complex(GO:0071141) |
| 0.0 | 2.1 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 0.1 | GO:0005683 | U7 snRNP(GO:0005683) pICln-Sm protein complex(GO:0034715) |
| 0.0 | 0.4 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 1.1 | GO:0000784 | nuclear chromosome, telomeric region(GO:0000784) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 3.1 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 0.5 | 2.7 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.4 | 2.2 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.3 | 2.2 | GO:0030020 | extracellular matrix structural constituent conferring tensile strength(GO:0030020) |
| 0.3 | 1.9 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.3 | 1.3 | GO:0010465 | nerve growth factor receptor activity(GO:0010465) |
| 0.2 | 0.9 | GO:0052798 | beta-galactoside alpha-2,3-sialyltransferase activity(GO:0052798) |
| 0.2 | 0.8 | GO:0034040 | lipid-transporting ATPase activity(GO:0034040) |
| 0.2 | 0.6 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.2 | 0.8 | GO:0034648 | histone demethylase activity (H3-dimethyl-K4 specific)(GO:0034648) |
| 0.2 | 2.6 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.2 | 1.1 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.1 | 1.2 | GO:0090599 | alpha-glucosidase activity(GO:0090599) |
| 0.1 | 2.3 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.1 | 1.1 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.1 | 1.4 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.1 | 0.6 | GO:0032406 | MutLbeta complex binding(GO:0032406) MutSbeta complex binding(GO:0032408) |
| 0.1 | 0.8 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.1 | 1.9 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.1 | 0.4 | GO:0004609 | phosphatidylserine decarboxylase activity(GO:0004609) |
| 0.1 | 0.5 | GO:0070139 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.1 | 0.5 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.1 | 0.3 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.1 | 1.2 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.1 | 1.1 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.1 | 0.7 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 0.1 | 0.3 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.1 | 0.3 | GO:0004618 | phosphoglycerate kinase activity(GO:0004618) |
| 0.1 | 0.3 | GO:0061731 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.1 | 1.2 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.1 | 1.1 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.1 | 2.0 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 0.4 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.1 | 0.5 | GO:0001165 | RNA polymerase I upstream control element sequence-specific DNA binding(GO:0001165) |
| 0.1 | 2.5 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 0.2 | GO:0035673 | oligopeptide transmembrane transporter activity(GO:0035673) |
| 0.1 | 0.2 | GO:0032089 | NACHT domain binding(GO:0032089) |
| 0.1 | 2.9 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 1.3 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.0 | 0.6 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.2 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.0 | 0.2 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.0 | 0.6 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 0.1 | GO:0004914 | interleukin-5 receptor activity(GO:0004914) |
| 0.0 | 0.4 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 0.2 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) gamma-aminobutyric acid transmembrane transporter activity(GO:0015185) |
| 0.0 | 0.5 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.0 | 0.3 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.0 | 0.9 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 0.1 | GO:0052591 | sn-glycerol-3-phosphate:ubiquinone oxidoreductase activity(GO:0052590) sn-glycerol-3-phosphate:ubiquinone-8 oxidoreductase activity(GO:0052591) |
| 0.0 | 0.7 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.0 | 0.1 | GO:0071209 | U7 snRNA binding(GO:0071209) |
| 0.0 | 0.3 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.0 | 0.7 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 1.5 | GO:0005548 | phospholipid transporter activity(GO:0005548) |
| 0.0 | 0.2 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 1.0 | GO:0004532 | exoribonuclease activity(GO:0004532) |
| 0.0 | 0.3 | GO:0016594 | glycine binding(GO:0016594) |
| 0.0 | 2.5 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 1.4 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.1 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.0 | 0.9 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.0 | 0.5 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.6 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.2 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.0 | 0.2 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.0 | 0.1 | GO:0030249 | guanylate cyclase regulator activity(GO:0030249) |
| 0.0 | 0.2 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 4.2 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.2 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.0 | 0.7 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 0.3 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.0 | 0.2 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.0 | 0.3 | GO:0022829 | wide pore channel activity(GO:0022829) |
| 0.0 | 0.2 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.1 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.0 | 0.8 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.0 | 0.1 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.0 | 1.3 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 0.2 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.0 | 0.2 | GO:0005328 | neurotransmitter:sodium symporter activity(GO:0005328) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 3.1 | GO:0010903 | negative regulation of very-low-density lipoprotein particle remodeling(GO:0010903) |
| 0.6 | 1.9 | GO:0051066 | dihydrobiopterin metabolic process(GO:0051066) |
| 0.4 | 2.2 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.4 | 1.3 | GO:0070171 | negative regulation of tooth mineralization(GO:0070171) |
| 0.4 | 1.2 | GO:0072428 | signal transduction involved in intra-S DNA damage checkpoint(GO:0072428) response to bisphenol A(GO:1903925) cellular response to bisphenol A(GO:1903926) |
| 0.4 | 1.2 | GO:0007290 | spermatid nucleus elongation(GO:0007290) |
| 0.4 | 4.2 | GO:0034196 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 0.3 | 1.3 | GO:0060054 | positive regulation of epithelial cell proliferation involved in wound healing(GO:0060054) |
| 0.3 | 1.3 | GO:0002503 | peptide antigen assembly with MHC class II protein complex(GO:0002503) |
| 0.3 | 1.4 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.2 | 2.0 | GO:1904469 | positive regulation of tumor necrosis factor secretion(GO:1904469) |
| 0.2 | 0.7 | GO:0072092 | ureteric bud invasion(GO:0072092) |
| 0.2 | 0.7 | GO:0021986 | epithalamus development(GO:0021538) habenula development(GO:0021986) |
| 0.2 | 0.8 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.2 | 0.8 | GO:0031064 | negative regulation of histone deacetylation(GO:0031064) |
| 0.2 | 1.0 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.2 | 0.6 | GO:0006711 | estrogen catabolic process(GO:0006711) |
| 0.2 | 1.1 | GO:0010792 | DNA double-strand break processing involved in repair via single-strand annealing(GO:0010792) |
| 0.2 | 0.8 | GO:0048388 | endosomal lumen acidification(GO:0048388) |
| 0.2 | 0.5 | GO:1903939 | negative regulation of histone H3-K9 dimethylation(GO:1900110) regulation of TORC2 signaling(GO:1903939) |
| 0.2 | 1.2 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.1 | 0.6 | GO:0034473 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 0.1 | 1.1 | GO:0070777 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.1 | 0.4 | GO:0030033 | microvillus assembly(GO:0030033) |
| 0.1 | 1.8 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 0.4 | GO:0002370 | natural killer cell cytokine production(GO:0002370) regulation of natural killer cell cytokine production(GO:0002727) |
| 0.1 | 0.5 | GO:0070301 | cellular response to hydrogen peroxide(GO:0070301) |
| 0.1 | 0.6 | GO:0033490 | cholesterol biosynthetic process via desmosterol(GO:0033489) cholesterol biosynthetic process via lathosterol(GO:0033490) |
| 0.1 | 1.2 | GO:0042904 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.1 | 1.1 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.1 | 1.4 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.1 | 0.7 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.1 | 0.5 | GO:0070560 | protein secretion by platelet(GO:0070560) |
| 0.1 | 0.6 | GO:0045007 | depurination(GO:0045007) |
| 0.1 | 0.2 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 0.1 | 0.2 | GO:1990502 | dense core granule maturation(GO:1990502) |
| 0.1 | 0.5 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.1 | 0.7 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.1 | 0.5 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.1 | 1.0 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.1 | 0.5 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.1 | 0.2 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 0.1 | 0.3 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.1 | 0.4 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.0 | 0.8 | GO:0001787 | natural killer cell proliferation(GO:0001787) |
| 0.0 | 2.9 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 3.1 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.0 | 0.5 | GO:2001214 | positive regulation of vasculogenesis(GO:2001214) |
| 0.0 | 0.3 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.0 | 0.1 | GO:0038043 | interleukin-5-mediated signaling pathway(GO:0038043) |
| 0.0 | 0.3 | GO:0002418 | immune response to tumor cell(GO:0002418) |
| 0.0 | 0.2 | GO:0008065 | establishment of blood-nerve barrier(GO:0008065) |
| 0.0 | 0.3 | GO:0070170 | regulation of tooth mineralization(GO:0070170) |
| 0.0 | 0.3 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.0 | 0.3 | GO:0035518 | histone H2A monoubiquitination(GO:0035518) |
| 0.0 | 0.9 | GO:0006295 | nucleotide-excision repair, preincision complex stabilization(GO:0006293) nucleotide-excision repair, DNA incision, 3'-to lesion(GO:0006295) |
| 0.0 | 1.2 | GO:0005980 | glycogen catabolic process(GO:0005980) |
| 0.0 | 0.8 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) |
| 0.0 | 0.2 | GO:0071475 | cellular hyperosmotic salinity response(GO:0071475) |
| 0.0 | 0.1 | GO:0021589 | hindbrain structural organization(GO:0021577) cerebellum structural organization(GO:0021589) |
| 0.0 | 0.1 | GO:0072709 | cellular response to sorbitol(GO:0072709) |
| 0.0 | 0.7 | GO:0030889 | negative regulation of B cell proliferation(GO:0030889) |
| 0.0 | 1.0 | GO:0018146 | keratan sulfate biosynthetic process(GO:0018146) |
| 0.0 | 2.6 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.0 | 0.4 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.0 | 0.1 | GO:0006127 | glycerophosphate shuttle(GO:0006127) |
| 0.0 | 0.3 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 0.5 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 0.1 | GO:0071676 | negative regulation of mononuclear cell migration(GO:0071676) |
| 0.0 | 1.2 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.0 | 0.8 | GO:0006505 | GPI anchor metabolic process(GO:0006505) GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 0.2 | GO:1902525 | regulation of protein monoubiquitination(GO:1902525) |
| 0.0 | 0.2 | GO:0015824 | proline transport(GO:0015824) |
| 0.0 | 2.6 | GO:0006903 | vesicle targeting(GO:0006903) |
| 0.0 | 0.6 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.0 | 0.4 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.0 | 0.1 | GO:1902310 | positive regulation of peptidyl-serine dephosphorylation(GO:1902310) |
| 0.0 | 1.1 | GO:1904837 | beta-catenin-TCF complex assembly(GO:1904837) |
| 0.0 | 0.2 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 0.2 | GO:0050713 | negative regulation of interleukin-1 beta secretion(GO:0050713) |
| 0.0 | 0.3 | GO:0035090 | maintenance of apical/basal cell polarity(GO:0035090) maintenance of epithelial cell apical/basal polarity(GO:0045199) |
| 0.0 | 2.1 | GO:0008585 | female gonad development(GO:0008585) |
| 0.0 | 0.7 | GO:0043029 | T cell homeostasis(GO:0043029) |
| 0.0 | 1.9 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.0 | 0.0 | GO:0032696 | negative regulation of interleukin-13 production(GO:0032696) |
| 0.0 | 0.6 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.0 | GO:0019853 | L-ascorbic acid biosynthetic process(GO:0019853) |
| 0.0 | 0.3 | GO:0051290 | deoxyribonucleotide biosynthetic process(GO:0009263) protein heterotetramerization(GO:0051290) |
| 0.0 | 0.6 | GO:0042026 | protein refolding(GO:0042026) |
| 0.0 | 0.8 | GO:0016445 | somatic diversification of immunoglobulins(GO:0016445) |
| 0.0 | 0.1 | GO:0060041 | retina development in camera-type eye(GO:0060041) |
| 0.0 | 0.0 | GO:0035482 | gastric motility(GO:0035482) gastric emptying(GO:0035483) |
| 0.0 | 0.1 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.0 | 0.4 | GO:0061049 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 0.3 | GO:0060397 | JAK-STAT cascade involved in growth hormone signaling pathway(GO:0060397) |
| 0.0 | 0.2 | GO:0015812 | gamma-aminobutyric acid transport(GO:0015812) |
| 0.0 | 0.1 | GO:0015705 | iodide transport(GO:0015705) |
| 0.0 | 0.2 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.0 | 0.1 | GO:0072584 | caveolin-mediated endocytosis(GO:0072584) |
| 0.0 | 0.9 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.0 | 0.1 | GO:2000825 | positive regulation of androgen receptor activity(GO:2000825) |
| 0.0 | 1.4 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.0 | 0.1 | GO:0070315 | G1 to G0 transition involved in cell differentiation(GO:0070315) |
| 0.0 | 0.2 | GO:0034308 | primary alcohol metabolic process(GO:0034308) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.6 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 2.2 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.1 | 1.9 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.1 | 1.9 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.1 | 0.6 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.1 | 1.2 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.1 | 1.2 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.1 | 5.1 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.8 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.9 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.7 | REACTOME INITIAL TRIGGERING OF COMPLEMENT | Genes involved in Initial triggering of complement |
| 0.0 | 1.1 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 0.6 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.0 | 1.0 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.0 | 0.5 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.7 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.0 | 1.5 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 0.5 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.0 | 0.6 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 1.3 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 2.6 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.1 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 0.0 | 0.3 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.0 | 0.1 | REACTOME VIF MEDIATED DEGRADATION OF APOBEC3G | Genes involved in Vif-mediated degradation of APOBEC3G |
| 0.0 | 0.3 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |