Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
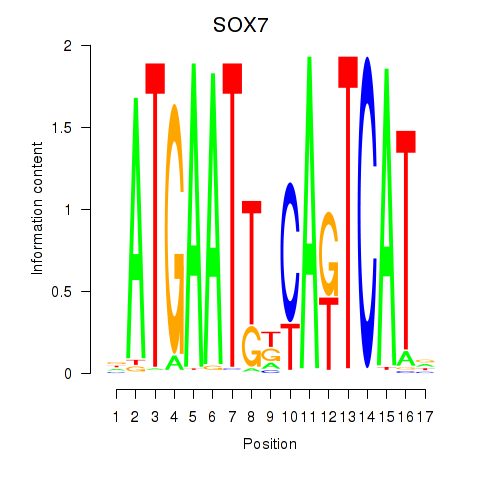
Results for SOX7
Z-value: 0.65
Transcription factors associated with SOX7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SOX7
|
ENSG00000171056.6 | SOX7 |
|
SOX7
|
ENSG00000258724.1 | SOX7 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SOX7 | hg19_v2_chr8_-_10588010_10588030 | 0.32 | 2.3e-01 | Click! |
Activity profile of SOX7 motif
Sorted Z-values of SOX7 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SOX7
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_95158644 | 1.50 |
ENST00000237858.6 |
GLRX |
glutaredoxin (thioltransferase) |
| chr22_+_23101182 | 1.32 |
ENST00000390312.2 |
IGLV2-14 |
immunoglobulin lambda variable 2-14 |
| chr14_-_106573756 | 1.17 |
ENST00000390601.2 |
IGHV3-11 |
immunoglobulin heavy variable 3-11 (gene/pseudogene) |
| chr5_-_95158375 | 0.96 |
ENST00000512469.2 ENST00000379979.4 ENST00000505427.1 ENST00000508780.1 |
GLRX |
glutaredoxin (thioltransferase) |
| chr11_+_7618413 | 0.87 |
ENST00000528883.1 |
PPFIBP2 |
PTPRF interacting protein, binding protein 2 (liprin beta 2) |
| chr21_-_15918618 | 0.69 |
ENST00000400564.1 ENST00000400566.1 |
SAMSN1 |
SAM domain, SH3 domain and nuclear localization signals 1 |
| chr21_+_34144411 | 0.56 |
ENST00000382375.4 ENST00000453404.1 ENST00000382378.1 ENST00000477513.1 |
C21orf49 |
chromosome 21 open reading frame 49 |
| chr6_-_111804905 | 0.52 |
ENST00000358835.3 ENST00000435970.1 |
REV3L |
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr8_-_102803163 | 0.51 |
ENST00000523645.1 ENST00000520346.1 ENST00000220931.6 ENST00000522448.1 ENST00000522951.1 ENST00000522252.1 ENST00000519098.1 |
NCALD |
neurocalcin delta |
| chr1_+_100598691 | 0.50 |
ENST00000370143.1 ENST00000370141.2 |
TRMT13 |
tRNA methyltransferase 13 homolog (S. cerevisiae) |
| chr12_-_90049878 | 0.46 |
ENST00000359142.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr12_-_90049828 | 0.41 |
ENST00000261173.2 ENST00000348959.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr16_+_33020496 | 0.40 |
ENST00000565407.2 |
IGHV3OR16-8 |
immunoglobulin heavy variable 3/OR16-8 (non-functional) |
| chr19_-_22379753 | 0.34 |
ENST00000397121.2 |
ZNF676 |
zinc finger protein 676 |
| chr16_-_33647696 | 0.32 |
ENST00000558425.1 ENST00000569103.2 |
RP11-812E19.9 |
Uncharacterized protein |
| chr8_+_77593448 | 0.31 |
ENST00000521891.2 |
ZFHX4 |
zinc finger homeobox 4 |
| chr10_+_5005445 | 0.29 |
ENST00000380872.4 |
AKR1C1 |
aldo-keto reductase family 1, member C1 |
| chr12_-_57443886 | 0.27 |
ENST00000300119.3 |
MYO1A |
myosin IA |
| chr6_+_131894284 | 0.25 |
ENST00000368087.3 ENST00000356962.2 |
ARG1 |
arginase 1 |
| chr10_-_5046042 | 0.25 |
ENST00000421196.3 ENST00000455190.1 |
AKR1C2 |
aldo-keto reductase family 1, member C2 |
| chr2_-_166930131 | 0.18 |
ENST00000303395.4 ENST00000409050.1 ENST00000423058.2 ENST00000375405.3 |
SCN1A |
sodium channel, voltage-gated, type I, alpha subunit |
| chrX_+_69353284 | 0.17 |
ENST00000342206.6 ENST00000356413.4 |
IGBP1 |
immunoglobulin (CD79A) binding protein 1 |
| chr6_+_42531798 | 0.15 |
ENST00000372903.2 ENST00000372899.1 ENST00000372901.1 |
UBR2 |
ubiquitin protein ligase E3 component n-recognin 2 |
| chr18_+_32820990 | 0.15 |
ENST00000601719.1 ENST00000591206.1 ENST00000330501.7 ENST00000261333.6 ENST00000355632.4 ENST00000585800.1 |
ZNF397 |
zinc finger protein 397 |
| chr1_+_10509971 | 0.14 |
ENST00000320498.4 |
CORT |
cortistatin |
| chr5_+_175511859 | 0.12 |
ENST00000503724.2 ENST00000253490.4 |
FAM153B |
family with sequence similarity 153, member B |
| chr12_-_58027138 | 0.11 |
ENST00000341156.4 |
B4GALNT1 |
beta-1,4-N-acetyl-galactosaminyl transferase 1 |
| chr9_-_70465758 | 0.10 |
ENST00000489273.1 |
CBWD5 |
COBW domain containing 5 |
| chr10_+_96162242 | 0.10 |
ENST00000225235.4 |
TBC1D12 |
TBC1 domain family, member 12 |
| chr17_-_66951474 | 0.10 |
ENST00000269080.2 |
ABCA8 |
ATP-binding cassette, sub-family A (ABC1), member 8 |
| chr8_+_12803176 | 0.09 |
ENST00000524591.2 |
KIAA1456 |
KIAA1456 |
| chr4_+_76995855 | 0.08 |
ENST00000355810.4 ENST00000349321.3 |
ART3 |
ADP-ribosyltransferase 3 |
| chr6_+_150690028 | 0.08 |
ENST00000229447.5 ENST00000344419.3 |
IYD |
iodotyrosine deiodinase |
| chr6_+_150690133 | 0.07 |
ENST00000392255.3 ENST00000500320.3 |
IYD |
iodotyrosine deiodinase |
| chr15_+_59439899 | 0.07 |
ENST00000599727.1 |
C15ORF31 |
C15ORF31 |
| chr6_+_29555683 | 0.06 |
ENST00000383640.2 |
OR2H2 |
olfactory receptor, family 2, subfamily H, member 2 |
| chr9_+_105757590 | 0.05 |
ENST00000374798.3 ENST00000487798.1 |
CYLC2 |
cylicin, basic protein of sperm head cytoskeleton 2 |
| chr8_-_19459993 | 0.05 |
ENST00000454498.2 ENST00000520003.1 |
CSGALNACT1 |
chondroitin sulfate N-acetylgalactosaminyltransferase 1 |
| chr1_+_11724167 | 0.05 |
ENST00000376753.4 |
FBXO6 |
F-box protein 6 |
| chr1_-_54303934 | 0.05 |
ENST00000537333.1 |
NDC1 |
NDC1 transmembrane nucleoporin |
| chr4_-_47983519 | 0.03 |
ENST00000358519.4 ENST00000544810.1 ENST00000402813.3 |
CNGA1 |
cyclic nucleotide gated channel alpha 1 |
| chr5_+_161494521 | 0.03 |
ENST00000356592.3 |
GABRG2 |
gamma-aminobutyric acid (GABA) A receptor, gamma 2 |
| chr2_-_88285309 | 0.02 |
ENST00000420840.2 |
RGPD2 |
RANBP2-like and GRIP domain containing 2 |
| chr6_-_46048116 | 0.02 |
ENST00000185206.6 |
CLIC5 |
chloride intracellular channel 5 |
| chr6_-_132910877 | 0.01 |
ENST00000258034.2 |
TAAR5 |
trace amine associated receptor 5 |
| chr14_+_50291993 | 0.00 |
ENST00000595378.1 |
AL627171.2 |
HCG1786899; PRO2610; Uncharacterized protein |
| chr16_-_20364030 | 0.00 |
ENST00000396134.2 ENST00000573567.1 ENST00000570757.1 ENST00000424589.1 ENST00000302509.4 ENST00000571174.1 ENST00000576688.1 |
UMOD |
uromodulin |
| chr5_+_161494770 | 0.00 |
ENST00000414552.2 ENST00000361925.4 |
GABRG2 |
gamma-aminobutyric acid (GABA) A receptor, gamma 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.1 | 2.5 | GO:0045838 | positive regulation of membrane potential(GO:0045838) |
| 0.1 | 0.5 | GO:0009753 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 0.1 | 0.3 | GO:0090467 | L-arginine import(GO:0043091) arginine import(GO:0090467) L-arginine transport(GO:1902023) |
| 0.0 | 0.1 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.0 | 0.2 | GO:0071233 | cellular response to leucine(GO:0071233) |
| 0.0 | 2.9 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.0 | 0.5 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 0.5 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.0 | 0.7 | GO:0050869 | negative regulation of B cell activation(GO:0050869) |
| 0.0 | 0.2 | GO:0098870 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.0 | 0.0 | GO:0070632 | spindle pole body duplication(GO:0030474) spindle pole body organization(GO:0051300) spindle pole body localization(GO:0070631) establishment of spindle pole body localization(GO:0070632) spindle pole body localization to nuclear envelope(GO:0071789) establishment of spindle pole body localization to nuclear envelope(GO:0071790) |
| 0.0 | 0.1 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.0 | 0.2 | GO:0006570 | tyrosine metabolic process(GO:0006570) |
| 0.0 | 0.0 | GO:0019276 | UDP-N-acetylgalactosamine metabolic process(GO:0019276) chondroitin sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0050653) |
| 0.0 | 0.2 | GO:0060632 | regulation of microtubule-based movement(GO:0060632) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.5 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.5 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 0.5 | GO:0042575 | DNA polymerase complex(GO:0042575) |
| 0.0 | 0.5 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.0 | 0.1 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 0.1 | GO:0033150 | perinuclear theca(GO:0033011) cytoskeletal calyx(GO:0033150) |
| 0.0 | 0.0 | GO:0070762 | nuclear pore transmembrane ring(GO:0070762) |
| 0.0 | 0.2 | GO:0043194 | voltage-gated sodium channel complex(GO:0001518) axon initial segment(GO:0043194) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.5 | GO:0015038 | glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.1 | 0.5 | GO:0047086 | phenanthrene 9,10-monooxygenase activity(GO:0018636) ketosteroid monooxygenase activity(GO:0047086) trans-1,2-dihydrobenzene-1,2-diol dehydrogenase activity(GO:0047115) |
| 0.0 | 0.2 | GO:0070728 | leucine binding(GO:0070728) |
| 0.0 | 0.9 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.2 | GO:0004447 | iodide peroxidase activity(GO:0004447) |
| 0.0 | 0.5 | GO:0008175 | tRNA methyltransferase activity(GO:0008175) |
| 0.0 | 0.5 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.2 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.0 | 0.1 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.0 | 0.0 | GO:0008955 | peptidoglycan glycosyltransferase activity(GO:0008955) |
| 0.0 | 0.5 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 0.1 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.0 | 2.9 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 0.7 | GO:0001784 | phosphotyrosine binding(GO:0001784) |