Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
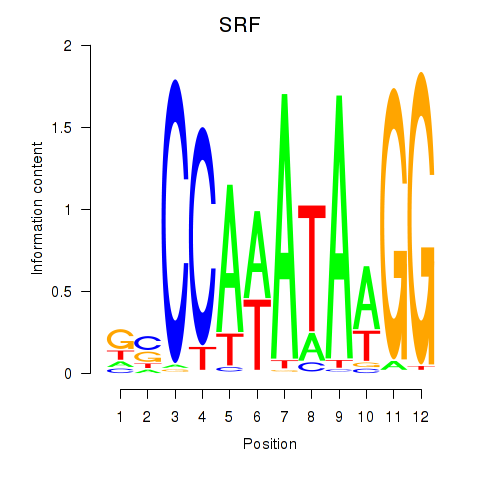
Results for SRF
Z-value: 4.00
Transcription factors associated with SRF
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SRF
|
ENSG00000112658.6 | SRF |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SRF | hg19_v2_chr6_+_43139037_43139094 | -0.23 | 3.9e-01 | Click! |
Activity profile of SRF motif
Sorted Z-values of SRF motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SRF
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_+_35169885 | 31.81 |
ENST00000279022.2 ENST00000346786.2 |
MYL9 |
myosin, light chain 9, regulatory |
| chr15_-_35088340 | 22.47 |
ENST00000290378.4 |
ACTC1 |
actin, alpha, cardiac muscle 1 |
| chr9_-_35691017 | 17.27 |
ENST00000378292.3 |
TPM2 |
tropomyosin 2 (beta) |
| chr2_+_74120094 | 15.97 |
ENST00000409731.3 ENST00000345517.3 ENST00000409918.1 ENST00000442912.1 ENST00000409624.1 |
ACTG2 |
actin, gamma 2, smooth muscle, enteric |
| chr12_+_75874460 | 13.20 |
ENST00000266659.3 |
GLIPR1 |
GLI pathogenesis-related 1 |
| chr1_-_86043921 | 13.20 |
ENST00000535924.2 |
DDAH1 |
dimethylarginine dimethylaminohydrolase 1 |
| chr12_+_75874580 | 12.83 |
ENST00000456650.3 |
GLIPR1 |
GLI pathogenesis-related 1 |
| chr6_+_151561506 | 12.32 |
ENST00000253332.1 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr6_+_151561085 | 12.27 |
ENST00000402676.2 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr11_+_117070037 | 10.35 |
ENST00000392951.4 ENST00000525531.1 ENST00000278968.6 |
TAGLN |
transgelin |
| chr2_-_106054952 | 9.91 |
ENST00000336660.5 ENST00000393352.3 ENST00000607522.1 |
FHL2 |
four and a half LIM domains 2 |
| chr3_-_123339418 | 9.49 |
ENST00000583087.1 |
MYLK |
myosin light chain kinase |
| chr7_-_94285402 | 9.25 |
ENST00000428696.2 ENST00000445866.2 |
SGCE |
sarcoglycan, epsilon |
| chr7_-_94285472 | 9.20 |
ENST00000437425.2 ENST00000447873.1 ENST00000415788.2 |
SGCE |
sarcoglycan, epsilon |
| chr3_-_123339343 | 9.15 |
ENST00000578202.1 |
MYLK |
myosin light chain kinase |
| chr7_-_94285511 | 8.97 |
ENST00000265735.7 |
SGCE |
sarcoglycan, epsilon |
| chr10_-_90712520 | 8.87 |
ENST00000224784.6 |
ACTA2 |
actin, alpha 2, smooth muscle, aorta |
| chr2_-_106015527 | 8.49 |
ENST00000344213.4 ENST00000358129.4 |
FHL2 |
four and a half LIM domains 2 |
| chr15_+_74218787 | 8.39 |
ENST00000261921.7 |
LOXL1 |
lysyl oxidase-like 1 |
| chr2_-_106015491 | 8.38 |
ENST00000408995.1 ENST00000393353.3 ENST00000322142.8 |
FHL2 |
four and a half LIM domains 2 |
| chr3_-_99594948 | 8.16 |
ENST00000471562.1 ENST00000495625.2 |
FILIP1L |
filamin A interacting protein 1-like |
| chr3_-_99595037 | 8.12 |
ENST00000383694.2 |
FILIP1L |
filamin A interacting protein 1-like |
| chr1_-_89531041 | 7.84 |
ENST00000370473.4 |
GBP1 |
guanylate binding protein 1, interferon-inducible |
| chr16_+_30383613 | 6.91 |
ENST00000568749.1 |
MYLPF |
myosin light chain, phosphorylatable, fast skeletal muscle |
| chr9_-_79520989 | 6.87 |
ENST00000376713.3 ENST00000376718.3 ENST00000428286.1 |
PRUNE2 |
prune homolog 2 (Drosophila) |
| chr9_+_124088860 | 6.76 |
ENST00000373806.1 |
GSN |
gelsolin |
| chr1_-_229569834 | 6.62 |
ENST00000366684.3 ENST00000366683.2 |
ACTA1 |
actin, alpha 1, skeletal muscle |
| chr10_-_17659357 | 6.57 |
ENST00000326961.6 ENST00000361271.3 |
PTPLA |
protein tyrosine phosphatase-like (proline instead of catalytic arginine), member A |
| chr2_-_161264385 | 6.46 |
ENST00000409972.1 |
RBMS1 |
RNA binding motif, single stranded interacting protein 1 |
| chr8_+_97597148 | 6.42 |
ENST00000521590.1 |
SDC2 |
syndecan 2 |
| chr19_+_16178317 | 6.32 |
ENST00000344824.6 ENST00000538887.1 |
TPM4 |
tropomyosin 4 |
| chr1_+_223889285 | 6.18 |
ENST00000433674.2 |
CAPN2 |
calpain 2, (m/II) large subunit |
| chr17_-_10560619 | 6.00 |
ENST00000583535.1 |
MYH3 |
myosin, heavy chain 3, skeletal muscle, embryonic |
| chr17_+_42634844 | 5.97 |
ENST00000315323.3 |
FZD2 |
frizzled family receptor 2 |
| chr19_-_45826125 | 5.89 |
ENST00000221476.3 |
CKM |
creatine kinase, muscle |
| chr7_-_107643674 | 5.38 |
ENST00000222399.6 |
LAMB1 |
laminin, beta 1 |
| chr10_+_31610064 | 5.26 |
ENST00000446923.2 ENST00000559476.1 |
ZEB1 |
zinc finger E-box binding homeobox 1 |
| chr5_-_111093167 | 5.22 |
ENST00000446294.2 ENST00000419114.2 |
NREP |
neuronal regeneration related protein |
| chr3_-_52486841 | 5.17 |
ENST00000496590.1 |
TNNC1 |
troponin C type 1 (slow) |
| chr5_+_92919043 | 5.09 |
ENST00000327111.3 |
NR2F1 |
nuclear receptor subfamily 2, group F, member 1 |
| chr3_+_187930719 | 4.63 |
ENST00000312675.4 |
LPP |
LIM domain containing preferred translocation partner in lipoma |
| chr9_+_92219919 | 4.45 |
ENST00000252506.6 ENST00000375769.1 |
GADD45G |
growth arrest and DNA-damage-inducible, gamma |
| chr3_+_49027308 | 4.43 |
ENST00000383729.4 ENST00000343546.4 |
P4HTM |
prolyl 4-hydroxylase, transmembrane (endoplasmic reticulum) |
| chr10_+_123922941 | 4.43 |
ENST00000360561.3 |
TACC2 |
transforming, acidic coiled-coil containing protein 2 |
| chr22_-_36784035 | 4.42 |
ENST00000216181.5 |
MYH9 |
myosin, heavy chain 9, non-muscle |
| chr19_+_2476116 | 4.39 |
ENST00000215631.4 ENST00000587345.1 |
GADD45B |
growth arrest and DNA-damage-inducible, beta |
| chr5_+_40679584 | 4.27 |
ENST00000302472.3 |
PTGER4 |
prostaglandin E receptor 4 (subtype EP4) |
| chr17_-_46671323 | 4.22 |
ENST00000239151.5 |
HOXB5 |
homeobox B5 |
| chr22_-_36357671 | 4.22 |
ENST00000408983.2 |
RBFOX2 |
RNA binding protein, fox-1 homolog (C. elegans) 2 |
| chr14_+_74034310 | 4.16 |
ENST00000538782.1 |
ACOT2 |
acyl-CoA thioesterase 2 |
| chr10_-_17659234 | 4.05 |
ENST00000466335.1 |
PTPLA |
protein tyrosine phosphatase-like (proline instead of catalytic arginine), member A |
| chr7_-_27183263 | 3.72 |
ENST00000222726.3 |
HOXA5 |
homeobox A5 |
| chr15_+_83776324 | 3.71 |
ENST00000379390.6 ENST00000379386.4 ENST00000565774.1 ENST00000565982.1 |
TM6SF1 |
transmembrane 6 superfamily member 1 |
| chr19_-_36247910 | 3.65 |
ENST00000587965.1 ENST00000004982.3 |
HSPB6 |
heat shock protein, alpha-crystallin-related, B6 |
| chr16_+_56965960 | 3.50 |
ENST00000439977.2 ENST00000344114.4 ENST00000300302.5 ENST00000379792.2 |
HERPUD1 |
homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 |
| chr7_+_143078652 | 3.40 |
ENST00000354434.4 ENST00000449423.2 |
ZYX |
zyxin |
| chr11_-_65667997 | 3.37 |
ENST00000312562.2 ENST00000534222.1 |
FOSL1 |
FOS-like antigen 1 |
| chr1_-_89591749 | 3.25 |
ENST00000370466.3 |
GBP2 |
guanylate binding protein 2, interferon-inducible |
| chr12_-_50616382 | 3.11 |
ENST00000552783.1 |
LIMA1 |
LIM domain and actin binding 1 |
| chr2_-_231989808 | 3.07 |
ENST00000258400.3 |
HTR2B |
5-hydroxytryptamine (serotonin) receptor 2B, G protein-coupled |
| chr14_+_75745477 | 3.01 |
ENST00000303562.4 ENST00000554617.1 ENST00000554212.1 ENST00000535987.1 ENST00000555242.1 |
FOS |
FBJ murine osteosarcoma viral oncogene homolog |
| chr12_-_50616122 | 3.01 |
ENST00000552823.1 ENST00000552909.1 |
LIMA1 |
LIM domain and actin binding 1 |
| chr11_-_65667884 | 3.00 |
ENST00000448083.2 ENST00000531493.1 ENST00000532401.1 |
FOSL1 |
FOS-like antigen 1 |
| chr11_-_65626753 | 2.98 |
ENST00000526975.1 ENST00000531413.1 |
CFL1 |
cofilin 1 (non-muscle) |
| chr10_-_16859442 | 2.89 |
ENST00000602389.1 ENST00000345264.5 |
RSU1 |
Ras suppressor protein 1 |
| chr10_-_16859361 | 2.84 |
ENST00000377921.3 |
RSU1 |
Ras suppressor protein 1 |
| chr2_-_128432639 | 2.74 |
ENST00000545738.2 ENST00000355119.4 ENST00000409808.2 |
LIMS2 |
LIM and senescent cell antigen-like domains 2 |
| chr10_+_75757863 | 2.55 |
ENST00000372755.3 ENST00000211998.4 ENST00000417648.2 |
VCL |
vinculin |
| chr11_-_2162468 | 2.32 |
ENST00000434045.2 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr4_-_10118469 | 2.29 |
ENST00000499869.2 |
WDR1 |
WD repeat domain 1 |
| chr5_-_142000883 | 2.29 |
ENST00000359370.6 |
FGF1 |
fibroblast growth factor 1 (acidic) |
| chr17_-_79481666 | 2.27 |
ENST00000575659.1 |
ACTG1 |
actin, gamma 1 |
| chr4_-_10118573 | 2.25 |
ENST00000382452.2 ENST00000382451.2 |
WDR1 |
WD repeat domain 1 |
| chr5_-_176923803 | 2.24 |
ENST00000506161.1 |
PDLIM7 |
PDZ and LIM domain 7 (enigma) |
| chr15_+_96873921 | 2.24 |
ENST00000394166.3 |
NR2F2 |
nuclear receptor subfamily 2, group F, member 2 |
| chr5_-_176923846 | 2.23 |
ENST00000506537.1 |
PDLIM7 |
PDZ and LIM domain 7 (enigma) |
| chr18_+_54318566 | 2.11 |
ENST00000589935.1 ENST00000357574.3 |
WDR7 |
WD repeat domain 7 |
| chr2_-_208031943 | 2.10 |
ENST00000421199.1 ENST00000457962.1 |
KLF7 |
Kruppel-like factor 7 (ubiquitous) |
| chrX_-_153602991 | 2.07 |
ENST00000369850.3 ENST00000422373.1 |
FLNA |
filamin A, alpha |
| chr4_-_54930790 | 1.92 |
ENST00000263921.3 |
CHIC2 |
cysteine-rich hydrophobic domain 2 |
| chr7_+_94285637 | 1.88 |
ENST00000482108.1 ENST00000488574.1 |
PEG10 |
paternally expressed 10 |
| chr10_+_112257596 | 1.84 |
ENST00000369583.3 |
DUSP5 |
dual specificity phosphatase 5 |
| chr1_+_28099683 | 1.80 |
ENST00000373943.4 |
STX12 |
syntaxin 12 |
| chr18_+_54318616 | 1.79 |
ENST00000254442.3 |
WDR7 |
WD repeat domain 7 |
| chr10_+_88428206 | 1.78 |
ENST00000429277.2 ENST00000458213.2 ENST00000352360.5 |
LDB3 |
LIM domain binding 3 |
| chr11_-_65626797 | 1.77 |
ENST00000525451.2 |
CFL1 |
cofilin 1 (non-muscle) |
| chr2_+_191792376 | 1.72 |
ENST00000409428.1 ENST00000409215.1 |
GLS |
glutaminase |
| chr19_+_1026566 | 1.61 |
ENST00000348419.3 ENST00000565096.2 ENST00000562958.2 ENST00000562075.2 ENST00000607102.1 |
CNN2 |
calponin 2 |
| chr10_+_96162242 | 1.55 |
ENST00000225235.4 |
TBC1D12 |
TBC1 domain family, member 12 |
| chr1_+_11994715 | 1.54 |
ENST00000449038.1 ENST00000376369.3 ENST00000429000.2 ENST00000196061.4 |
PLOD1 |
procollagen-lysine, 2-oxoglutarate 5-dioxygenase 1 |
| chr11_-_2162162 | 1.52 |
ENST00000381389.1 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr12_-_6772303 | 1.46 |
ENST00000396807.4 ENST00000446105.2 ENST00000341550.4 |
ING4 |
inhibitor of growth family, member 4 |
| chr6_-_111804905 | 1.44 |
ENST00000358835.3 ENST00000435970.1 |
REV3L |
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr12_+_59989791 | 1.40 |
ENST00000552432.1 |
SLC16A7 |
solute carrier family 16 (monocarboxylate transporter), member 7 |
| chr19_+_1026298 | 1.28 |
ENST00000263097.4 |
CNN2 |
calponin 2 |
| chr3_+_149192475 | 1.24 |
ENST00000465758.1 |
TM4SF4 |
transmembrane 4 L six family member 4 |
| chr17_-_79479789 | 1.03 |
ENST00000571691.1 ENST00000571721.1 ENST00000573283.1 ENST00000575842.1 ENST00000575087.1 ENST00000570382.1 ENST00000331925.2 |
ACTG1 |
actin, gamma 1 |
| chr7_-_5570229 | 0.89 |
ENST00000331789.5 |
ACTB |
actin, beta |
| chr18_-_54318353 | 0.88 |
ENST00000590954.1 ENST00000540155.1 |
TXNL1 |
thioredoxin-like 1 |
| chr19_-_15344243 | 0.85 |
ENST00000602233.1 |
EPHX3 |
epoxide hydrolase 3 |
| chr19_+_13135386 | 0.83 |
ENST00000360105.4 ENST00000588228.1 ENST00000591028.1 |
NFIX |
nuclear factor I/X (CCAAT-binding transcription factor) |
| chr17_-_4890919 | 0.81 |
ENST00000572543.1 ENST00000381311.5 ENST00000348066.3 ENST00000358183.4 |
CAMTA2 |
calmodulin binding transcription activator 2 |
| chr13_+_27825706 | 0.81 |
ENST00000272274.4 ENST00000319826.4 ENST00000326092.4 |
RPL21 |
ribosomal protein L21 |
| chr17_-_4890649 | 0.75 |
ENST00000361571.5 |
CAMTA2 |
calmodulin binding transcription activator 2 |
| chr3_+_9834179 | 0.72 |
ENST00000498623.2 |
ARPC4 |
actin related protein 2/3 complex, subunit 4, 20kDa |
| chr3_-_9834375 | 0.64 |
ENST00000343450.2 ENST00000301964.2 |
TADA3 |
transcriptional adaptor 3 |
| chr12_-_125398850 | 0.61 |
ENST00000535859.1 ENST00000546271.1 ENST00000540700.1 ENST00000546120.1 |
UBC |
ubiquitin C |
| chr3_+_9834758 | 0.60 |
ENST00000485273.1 ENST00000433034.1 ENST00000397256.1 |
ARPC4 ARPC4-TTLL3 |
actin related protein 2/3 complex, subunit 4, 20kDa ARPC4-TTLL3 readthrough |
| chr12_+_59989918 | 0.60 |
ENST00000547379.1 ENST00000549465.1 |
SLC16A7 |
solute carrier family 16 (monocarboxylate transporter), member 7 |
| chr1_-_208084729 | 0.59 |
ENST00000310833.7 ENST00000356522.4 |
CD34 |
CD34 molecule |
| chr13_+_27825446 | 0.55 |
ENST00000311549.6 |
RPL21 |
ribosomal protein L21 |
| chr17_+_7487146 | 0.54 |
ENST00000396501.4 ENST00000584378.1 ENST00000423172.2 ENST00000579445.1 ENST00000585217.1 ENST00000581380.1 |
MPDU1 |
mannose-P-dolichol utilization defect 1 |
| chr1_-_168106536 | 0.53 |
ENST00000537209.1 ENST00000361697.2 ENST00000546300.1 ENST00000367835.1 |
GPR161 |
G protein-coupled receptor 161 |
| chr19_+_45971246 | 0.47 |
ENST00000585836.1 ENST00000417353.2 ENST00000353609.3 ENST00000591858.1 ENST00000443841.2 ENST00000590335.1 |
FOSB |
FBJ murine osteosarcoma viral oncogene homolog B |
| chr3_+_9834227 | 0.40 |
ENST00000287613.7 ENST00000397261.3 |
ARPC4 |
actin related protein 2/3 complex, subunit 4, 20kDa |
| chr17_+_73452695 | 0.38 |
ENST00000582186.1 ENST00000582455.1 ENST00000581252.1 ENST00000579208.1 |
KIAA0195 |
KIAA0195 |
| chr5_+_137774706 | 0.38 |
ENST00000378339.2 ENST00000254901.5 ENST00000506158.1 |
REEP2 |
receptor accessory protein 2 |
| chr8_+_1993152 | 0.36 |
ENST00000262113.4 |
MYOM2 |
myomesin 2 |
| chr20_-_62258394 | 0.35 |
ENST00000370077.1 |
GMEB2 |
glucocorticoid modulatory element binding protein 2 |
| chr22_+_38004832 | 0.30 |
ENST00000405147.3 ENST00000429218.1 ENST00000325180.8 ENST00000337437.4 |
GGA1 |
golgi-associated, gamma adaptin ear containing, ARF binding protein 1 |
| chr22_+_38004473 | 0.27 |
ENST00000414350.3 ENST00000343632.4 |
GGA1 |
golgi-associated, gamma adaptin ear containing, ARF binding protein 1 |
| chr2_-_209118974 | 0.26 |
ENST00000415913.1 ENST00000415282.1 ENST00000446179.1 |
IDH1 |
isocitrate dehydrogenase 1 (NADP+), soluble |
| chr10_+_88428370 | 0.24 |
ENST00000372066.3 ENST00000263066.6 ENST00000372056.4 ENST00000310944.6 ENST00000361373.4 ENST00000542786.1 |
LDB3 |
LIM domain binding 3 |
| chr19_-_15343773 | 0.21 |
ENST00000435261.1 ENST00000594042.1 |
EPHX3 |
epoxide hydrolase 3 |
| chr17_-_4871085 | 0.20 |
ENST00000575142.1 ENST00000206020.3 |
SPAG7 |
sperm associated antigen 7 |
| chr8_+_1993173 | 0.15 |
ENST00000523438.1 |
MYOM2 |
myomesin 2 |
| chr17_+_3379284 | 0.14 |
ENST00000263080.2 |
ASPA |
aspartoacylase |
| chr11_+_34073195 | 0.13 |
ENST00000341394.4 |
CAPRIN1 |
cell cycle associated protein 1 |
| chr5_+_66254698 | 0.11 |
ENST00000405643.1 ENST00000407621.1 ENST00000432426.1 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr11_-_71823266 | 0.02 |
ENST00000538919.1 ENST00000539395.1 ENST00000542531.1 |
ANAPC15 |
anaphase promoting complex subunit 15 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 101.4 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.4 | 4.3 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.4 | 52.8 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.2 | 6.4 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.2 | 3.1 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.2 | 6.8 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.2 | 2.3 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.2 | 4.3 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 3.8 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 3.5 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.1 | 2.0 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.1 | 11.1 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.1 | 5.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 4.4 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 3.1 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.1 | 1.8 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.1 | 1.7 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 1.8 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 5.4 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 4.2 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 4.0 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.9 | REACTOME PREFOLDIN MEDIATED TRANSFER OF SUBSTRATE TO CCT TRIC | Genes involved in Prefoldin mediated transfer of substrate to CCT/TriC |
| 0.0 | 1.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 1.8 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 4.9 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.7 | 18.6 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 2.6 | 13.2 | GO:0016403 | dimethylargininase activity(GO:0016403) |
| 1.7 | 27.6 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 1.5 | 24.6 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.8 | 6.8 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.7 | 5.9 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.7 | 4.4 | GO:0017018 | myosin phosphatase activity(GO:0017018) |
| 0.6 | 5.1 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.5 | 4.3 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.5 | 8.4 | GO:0016641 | oxidoreductase activity, acting on the CH-NH2 group of donors, oxygen as acceptor(GO:0016641) |
| 0.5 | 2.1 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 0.5 | 1.5 | GO:0033823 | procollagen-lysine 5-dioxygenase activity(GO:0008475) procollagen glucosyltransferase activity(GO:0033823) |
| 0.5 | 2.0 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.5 | 5.2 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.4 | 4.2 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.4 | 26.8 | GO:0050681 | androgen receptor binding(GO:0050681) |
| 0.3 | 1.7 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.3 | 22.9 | GO:0017022 | myosin binding(GO:0017022) |
| 0.3 | 27.7 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.3 | 6.0 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.3 | 3.1 | GO:0051378 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.2 | 4.4 | GO:0043495 | protein anchor(GO:0043495) |
| 0.2 | 2.6 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.2 | 4.4 | GO:0031418 | L-ascorbic acid binding(GO:0031418) |
| 0.2 | 2.0 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.2 | 7.8 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.2 | 6.1 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.2 | 0.6 | GO:0070735 | protein-glycine ligase activity(GO:0070735) |
| 0.1 | 0.9 | GO:0047134 | protein-disulfide reductase activity(GO:0047134) |
| 0.1 | 3.8 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 3.7 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 0.9 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.1 | 3.8 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.1 | 16.8 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.1 | 5.3 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 0.3 | GO:0004450 | isocitrate dehydrogenase (NADP+) activity(GO:0004450) |
| 0.1 | 4.2 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.1 | 2.3 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.1 | 3.0 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 6.4 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.1 | 2.2 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 0.6 | GO:0043199 | sulfate binding(GO:0043199) |
| 0.1 | 5.4 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 1.4 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 6.5 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 0.6 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.0 | 1.8 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.6 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 3.3 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 3.9 | GO:0035257 | nuclear hormone receptor binding(GO:0035257) |
| 0.0 | 1.5 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.1 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.0 | 1.2 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 4.2 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 2.2 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 0.2 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 0.5 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 3.9 | GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding(GO:0000978) |
| 0.0 | 17.6 | GO:0005524 | ATP binding(GO:0005524) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 26.8 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.8 | 44.7 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.5 | 41.5 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.2 | 5.4 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.2 | 8.8 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.2 | 11.9 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.2 | 6.0 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.2 | 10.7 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.2 | 6.8 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.2 | 4.5 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.2 | 3.8 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 6.4 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.1 | 6.4 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 19.3 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.1 | 3.8 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.1 | 2.3 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 1.8 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 2.2 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 7.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.9 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.2 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.0 | 53.9 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 5.4 | 26.8 | GO:0055014 | atrial cardiac muscle cell differentiation(GO:0055011) atrial cardiac muscle cell development(GO:0055014) |
| 2.3 | 18.6 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 1.9 | 11.2 | GO:1900756 | protein processing in phagocytic vesicle(GO:1900756) regulation of protein processing in phagocytic vesicle(GO:1903921) positive regulation of protein processing in phagocytic vesicle(GO:1903923) |
| 1.7 | 8.4 | GO:1900041 | negative regulation of interleukin-2 secretion(GO:1900041) |
| 1.6 | 24.6 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 1.4 | 4.3 | GO:2000417 | negative regulation of eosinophil migration(GO:2000417) |
| 1.3 | 5.4 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 1.3 | 5.2 | GO:0032972 | diaphragm contraction(GO:0002086) regulation of muscle filament sliding speed(GO:0032972) |
| 1.2 | 3.7 | GO:0060535 | trachea cartilage morphogenesis(GO:0060535) |
| 1.1 | 4.5 | GO:0042247 | morphogenesis of follicular epithelium(GO:0016333) establishment or maintenance of polarity of follicular epithelium(GO:0016334) establishment of planar polarity of follicular epithelium(GO:0042247) |
| 1.1 | 13.2 | GO:1900038 | negative regulation of cellular response to hypoxia(GO:1900038) |
| 1.1 | 6.4 | GO:0007296 | vitellogenesis(GO:0007296) |
| 1.0 | 4.8 | GO:2000771 | regulation of unidimensional cell growth(GO:0051510) negative regulation of unidimensional cell growth(GO:0051511) establishment of cell polarity regulating cell shape(GO:0071964) regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000769) positive regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000771) regulation of establishment of cell polarity regulating cell shape(GO:2000782) positive regulation of establishment of cell polarity regulating cell shape(GO:2000784) regulation of barbed-end actin filament capping(GO:2000812) positive regulation of barbed-end actin filament capping(GO:2000814) |
| 0.9 | 6.4 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.7 | 6.0 | GO:0003150 | muscular septum morphogenesis(GO:0003150) |
| 0.7 | 4.2 | GO:0060318 | regulation of definitive erythrocyte differentiation(GO:0010724) definitive erythrocyte differentiation(GO:0060318) |
| 0.6 | 3.1 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.6 | 5.3 | GO:0045602 | negative regulation of endothelial cell differentiation(GO:0045602) |
| 0.6 | 2.3 | GO:0060681 | branch elongation involved in ureteric bud branching(GO:0060681) |
| 0.6 | 2.2 | GO:0060849 | radial pattern formation(GO:0009956) regulation of transcription involved in lymphatic endothelial cell fate commitment(GO:0060849) |
| 0.5 | 5.9 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.5 | 1.5 | GO:0046946 | hydroxylysine metabolic process(GO:0046946) hydroxylysine biosynthetic process(GO:0046947) |
| 0.5 | 3.8 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.4 | 31.8 | GO:0045652 | regulation of megakaryocyte differentiation(GO:0045652) |
| 0.4 | 8.8 | GO:1900745 | positive regulation of p38MAPK cascade(GO:1900745) |
| 0.4 | 3.0 | GO:0001661 | conditioned taste aversion(GO:0001661) |
| 0.4 | 1.5 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.3 | 2.1 | GO:0016479 | negative regulation of transcription from RNA polymerase I promoter(GO:0016479) |
| 0.3 | 1.7 | GO:0006543 | glutamine catabolic process(GO:0006543) |
| 0.3 | 23.7 | GO:0030049 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.2 | 2.0 | GO:0006848 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.2 | 3.5 | GO:1903071 | positive regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903071) |
| 0.2 | 2.7 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.2 | 3.8 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.2 | 0.6 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 0.2 | 2.6 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.1 | 8.1 | GO:0035904 | aorta development(GO:0035904) |
| 0.1 | 3.7 | GO:0010667 | negative regulation of cardiac muscle cell apoptotic process(GO:0010667) |
| 0.1 | 0.4 | GO:0050916 | sensory perception of sweet taste(GO:0050916) |
| 0.1 | 1.8 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.1 | 4.2 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.1 | 2.0 | GO:0045214 | sarcomere organization(GO:0045214) |
| 0.1 | 6.1 | GO:0031529 | ruffle organization(GO:0031529) |
| 0.1 | 0.6 | GO:0090043 | regulation of tubulin deacetylation(GO:0090043) |
| 0.1 | 0.3 | GO:0006097 | glyoxylate cycle(GO:0006097) |
| 0.1 | 4.4 | GO:0045646 | regulation of erythrocyte differentiation(GO:0045646) |
| 0.1 | 23.7 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.1 | 4.1 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.1 | 2.1 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.1 | 4.2 | GO:0070527 | platelet aggregation(GO:0070527) |
| 0.1 | 4.5 | GO:0045669 | positive regulation of osteoblast differentiation(GO:0045669) |
| 0.1 | 0.5 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.1 | 1.6 | GO:2000785 | regulation of autophagosome assembly(GO:2000785) |
| 0.0 | 4.2 | GO:0048704 | embryonic skeletal system morphogenesis(GO:0048704) |
| 0.0 | 0.1 | GO:0006533 | aspartate catabolic process(GO:0006533) |
| 0.0 | 2.9 | GO:0035722 | interleukin-12-mediated signaling pathway(GO:0035722) cellular response to interleukin-12(GO:0071349) |
| 0.0 | 1.9 | GO:0006893 | Golgi to plasma membrane transport(GO:0006893) |
| 0.0 | 1.9 | GO:1903845 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.0 | 14.7 | GO:0019216 | regulation of lipid metabolic process(GO:0019216) |
| 0.0 | 0.5 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 5.1 | GO:0010977 | negative regulation of neuron projection development(GO:0010977) |
| 0.0 | 1.8 | GO:0000045 | autophagosome assembly(GO:0000045) |
| 0.0 | 3.2 | GO:0060337 | type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 0.0 | 0.5 | GO:0051412 | response to corticosterone(GO:0051412) |
| 0.0 | 1.1 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 3.4 | GO:0021987 | cerebral cortex development(GO:0021987) |
| 0.0 | 1.2 | GO:0042246 | tissue regeneration(GO:0042246) |
| 0.0 | 2.2 | GO:0048813 | dendrite morphogenesis(GO:0048813) |
| 0.0 | 0.6 | GO:1901998 | toxin transport(GO:1901998) |
| 0.0 | 5.3 | GO:0006260 | DNA replication(GO:0006260) |
| 0.0 | 2.0 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.0 | 0.9 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 3.5 | GO:0007517 | muscle organ development(GO:0007517) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.4 | 27.0 | GO:0042643 | actomyosin, actin portion(GO:0042643) |
| 3.4 | 27.4 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 2.7 | 16.0 | GO:0032982 | myosin filament(GO:0032982) |
| 2.2 | 8.9 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 1.8 | 5.4 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 1.5 | 4.4 | GO:0005826 | actomyosin contractile ring(GO:0005826) |
| 1.2 | 23.6 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 1.0 | 36.0 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.9 | 5.2 | GO:1990584 | cardiac Troponin complex(GO:1990584) |
| 0.7 | 2.1 | GO:0031523 | Myb complex(GO:0031523) |
| 0.7 | 26.4 | GO:0031430 | M band(GO:0031430) |
| 0.6 | 3.5 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.5 | 6.8 | GO:0030478 | actin cap(GO:0030478) |
| 0.4 | 40.7 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.4 | 3.0 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.3 | 26.0 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.3 | 4.2 | GO:0097433 | dense body(GO:0097433) |
| 0.3 | 5.9 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.2 | 2.6 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.1 | 4.8 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.1 | 1.4 | GO:0042575 | DNA polymerase complex(GO:0042575) |
| 0.1 | 1.8 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.1 | 1.7 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 6.0 | GO:0030669 | clathrin-coated endocytic vesicle membrane(GO:0030669) |
| 0.1 | 25.2 | GO:0005938 | cell cortex(GO:0005938) |
| 0.1 | 8.4 | GO:0005604 | basement membrane(GO:0005604) |
| 0.1 | 6.4 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.1 | 1.5 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.1 | 6.4 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 3.8 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.0 | 0.6 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 0.6 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 0.6 | GO:0046930 | pore complex(GO:0046930) |
| 0.0 | 1.9 | GO:0005798 | Golgi-associated vesicle(GO:0005798) |
| 0.0 | 4.2 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 3.9 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 1.4 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 10.4 | GO:0005925 | cell-substrate adherens junction(GO:0005924) focal adhesion(GO:0005925) |
| 0.0 | 0.6 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.0 | 3.7 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 1.6 | GO:0005776 | autophagosome(GO:0005776) |
| 0.0 | 1.5 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.0 | 11.1 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 4.0 | GO:0005667 | transcription factor complex(GO:0005667) |