Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
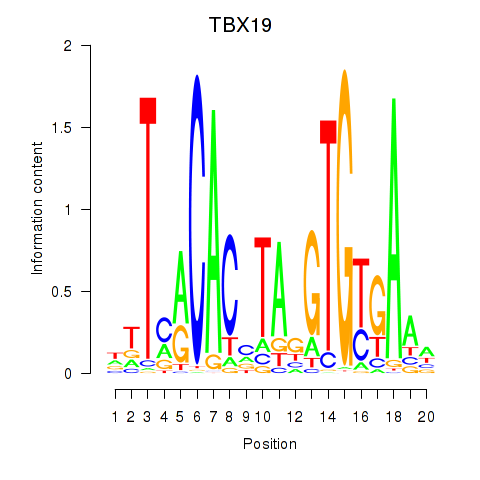
Results for TBX19
Z-value: 0.73
Transcription factors associated with TBX19
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TBX19
|
ENSG00000143178.8 | TBX19 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TBX19 | hg19_v2_chr1_+_168250194_168250278 | 0.04 | 9.0e-01 | Click! |
Activity profile of TBX19 motif
Sorted Z-values of TBX19 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TBX19
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_113594279 | 1.83 |
ENST00000416750.1 ENST00000418817.1 ENST00000263341.2 |
IL1B |
interleukin 1, beta |
| chr18_+_61554932 | 1.62 |
ENST00000299502.4 ENST00000457692.1 ENST00000413956.1 |
SERPINB2 |
serpin peptidase inhibitor, clade B (ovalbumin), member 2 |
| chr1_+_46640750 | 1.55 |
ENST00000372003.1 |
TSPAN1 |
tetraspanin 1 |
| chr19_+_38755042 | 1.31 |
ENST00000301244.7 |
SPINT2 |
serine peptidase inhibitor, Kunitz type, 2 |
| chr6_-_2842087 | 1.14 |
ENST00000537185.1 |
SERPINB1 |
serpin peptidase inhibitor, clade B (ovalbumin), member 1 |
| chr11_-_115375107 | 1.13 |
ENST00000545380.1 ENST00000452722.3 ENST00000537058.1 ENST00000536727.1 ENST00000542447.2 ENST00000331581.6 |
CADM1 |
cell adhesion molecule 1 |
| chr5_+_96211643 | 1.12 |
ENST00000437043.3 ENST00000510373.1 |
ERAP2 |
endoplasmic reticulum aminopeptidase 2 |
| chr6_-_2842219 | 1.11 |
ENST00000380739.5 |
SERPINB1 |
serpin peptidase inhibitor, clade B (ovalbumin), member 1 |
| chr5_+_96212185 | 1.08 |
ENST00000379904.4 |
ERAP2 |
endoplasmic reticulum aminopeptidase 2 |
| chr10_-_95241951 | 0.92 |
ENST00000358334.5 ENST00000359263.4 ENST00000371488.3 |
MYOF |
myoferlin |
| chr10_-_95242044 | 0.88 |
ENST00000371501.4 ENST00000371502.4 ENST00000371489.1 |
MYOF |
myoferlin |
| chr7_-_76255444 | 0.87 |
ENST00000454397.1 |
POMZP3 |
POM121 and ZP3 fusion |
| chr3_+_118905564 | 0.86 |
ENST00000460625.1 |
UPK1B |
uroplakin 1B |
| chr11_-_119993734 | 0.79 |
ENST00000533302.1 |
TRIM29 |
tripartite motif containing 29 |
| chr5_+_125695805 | 0.74 |
ENST00000513040.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr21_+_39628655 | 0.73 |
ENST00000398925.1 ENST00000398928.1 ENST00000328656.4 ENST00000443341.1 |
KCNJ15 |
potassium inwardly-rectifying channel, subfamily J, member 15 |
| chr4_-_156298028 | 0.67 |
ENST00000433024.1 ENST00000379248.2 |
MAP9 |
microtubule-associated protein 9 |
| chr20_+_44036620 | 0.66 |
ENST00000372710.3 |
DBNDD2 |
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr2_+_62900986 | 0.64 |
ENST00000405015.3 ENST00000413434.1 ENST00000426940.1 ENST00000449820.1 |
EHBP1 |
EH domain binding protein 1 |
| chr10_+_90750378 | 0.57 |
ENST00000355740.2 ENST00000352159.4 |
FAS |
Fas cell surface death receptor |
| chr4_-_57976544 | 0.54 |
ENST00000295666.4 ENST00000537922.1 |
IGFBP7 |
insulin-like growth factor binding protein 7 |
| chr1_+_19923454 | 0.53 |
ENST00000602662.1 ENST00000602293.1 ENST00000322753.6 |
MINOS1-NBL1 MINOS1 |
MINOS1-NBL1 readthrough mitochondrial inner membrane organizing system 1 |
| chr22_+_31477296 | 0.50 |
ENST00000426927.1 ENST00000440425.1 ENST00000358743.1 ENST00000347557.2 ENST00000333137.7 |
SMTN |
smoothelin |
| chr6_+_123110465 | 0.44 |
ENST00000539041.1 |
SMPDL3A |
sphingomyelin phosphodiesterase, acid-like 3A |
| chr14_+_32546274 | 0.44 |
ENST00000396582.2 |
ARHGAP5 |
Rho GTPase activating protein 5 |
| chr10_+_90750493 | 0.43 |
ENST00000357339.2 ENST00000355279.2 |
FAS |
Fas cell surface death receptor |
| chr2_-_9563575 | 0.35 |
ENST00000488451.1 ENST00000238091.4 ENST00000355346.4 |
ITGB1BP1 |
integrin beta 1 binding protein 1 |
| chr14_+_69658194 | 0.34 |
ENST00000409018.3 ENST00000409014.1 ENST00000409675.1 |
EXD2 |
exonuclease 3'-5' domain containing 2 |
| chr16_-_66583701 | 0.32 |
ENST00000527800.1 ENST00000525974.1 ENST00000563369.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr12_-_10962767 | 0.31 |
ENST00000240691.2 |
TAS2R9 |
taste receptor, type 2, member 9 |
| chr2_-_9563319 | 0.30 |
ENST00000497105.1 ENST00000360635.3 ENST00000359712.3 |
ITGB1BP1 |
integrin beta 1 binding protein 1 |
| chr2_-_85636928 | 0.30 |
ENST00000449030.1 |
CAPG |
capping protein (actin filament), gelsolin-like |
| chr15_-_60771128 | 0.26 |
ENST00000558512.1 ENST00000561114.1 |
NARG2 |
NMDA receptor regulated 2 |
| chr7_-_43946175 | 0.23 |
ENST00000603700.1 ENST00000402306.3 ENST00000443736.1 ENST00000446958.1 |
URGCP-MRPS24 URGCP |
URGCP-MRPS24 readthrough upregulator of cell proliferation |
| chrX_-_15402498 | 0.22 |
ENST00000297904.3 |
FIGF |
c-fos induced growth factor (vascular endothelial growth factor D) |
| chr8_-_19540266 | 0.22 |
ENST00000311540.4 |
CSGALNACT1 |
chondroitin sulfate N-acetylgalactosaminyltransferase 1 |
| chr1_-_167522982 | 0.22 |
ENST00000370509.4 |
CREG1 |
cellular repressor of E1A-stimulated genes 1 |
| chr2_-_9563469 | 0.22 |
ENST00000484735.1 ENST00000456913.2 |
ITGB1BP1 |
integrin beta 1 binding protein 1 |
| chr8_-_19540086 | 0.22 |
ENST00000332246.6 |
CSGALNACT1 |
chondroitin sulfate N-acetylgalactosaminyltransferase 1 |
| chr20_-_3762087 | 0.21 |
ENST00000379756.3 |
SPEF1 |
sperm flagellar 1 |
| chr6_-_27880174 | 0.20 |
ENST00000303324.2 |
OR2B2 |
olfactory receptor, family 2, subfamily B, member 2 |
| chr5_+_140165876 | 0.20 |
ENST00000504120.2 ENST00000394633.3 ENST00000378133.3 |
PCDHA1 |
protocadherin alpha 1 |
| chr1_-_146040968 | 0.19 |
ENST00000401010.3 |
NBPF11 |
neuroblastoma breakpoint family, member 11 |
| chr2_+_201754135 | 0.19 |
ENST00000409357.1 ENST00000409129.2 |
NIF3L1 |
NIF3 NGG1 interacting factor 3-like 1 (S. cerevisiae) |
| chr9_+_125137565 | 0.19 |
ENST00000373698.5 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr2_+_152214098 | 0.19 |
ENST00000243347.3 |
TNFAIP6 |
tumor necrosis factor, alpha-induced protein 6 |
| chr14_+_24458123 | 0.18 |
ENST00000545240.1 ENST00000382755.4 |
DHRS4L2 |
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr2_+_201754050 | 0.17 |
ENST00000426253.1 ENST00000416651.1 ENST00000454952.1 ENST00000409020.1 ENST00000359683.4 |
NIF3L1 |
NIF3 NGG1 interacting factor 3-like 1 (S. cerevisiae) |
| chr20_-_7921090 | 0.17 |
ENST00000378789.3 |
HAO1 |
hydroxyacid oxidase (glycolate oxidase) 1 |
| chr11_+_46740730 | 0.17 |
ENST00000311907.5 ENST00000530231.1 ENST00000442468.1 |
F2 |
coagulation factor II (thrombin) |
| chrX_+_150866779 | 0.17 |
ENST00000370353.3 |
PRRG3 |
proline rich Gla (G-carboxyglutamic acid) 3 (transmembrane) |
| chr15_-_65809581 | 0.16 |
ENST00000341861.5 |
DPP8 |
dipeptidyl-peptidase 8 |
| chr20_-_22565101 | 0.15 |
ENST00000419308.2 |
FOXA2 |
forkhead box A2 |
| chr2_+_228337079 | 0.14 |
ENST00000409315.1 ENST00000373671.3 ENST00000409171.1 |
AGFG1 |
ArfGAP with FG repeats 1 |
| chr14_+_24458021 | 0.13 |
ENST00000397071.1 ENST00000559411.1 ENST00000335125.6 |
DHRS4L2 |
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr4_+_69962185 | 0.13 |
ENST00000305231.7 |
UGT2B7 |
UDP glucuronosyltransferase 2 family, polypeptide B7 |
| chr2_-_211341411 | 0.13 |
ENST00000233714.4 ENST00000443314.1 ENST00000441020.3 ENST00000450366.2 ENST00000431941.2 |
LANCL1 |
LanC lantibiotic synthetase component C-like 1 (bacterial) |
| chr19_-_46526304 | 0.13 |
ENST00000008938.4 |
PGLYRP1 |
peptidoglycan recognition protein 1 |
| chr14_+_24458093 | 0.11 |
ENST00000558753.1 ENST00000537912.1 |
DHRS4L2 |
dehydrogenase/reductase (SDR family) member 4 like 2 |
| chr19_+_852291 | 0.11 |
ENST00000263621.1 |
ELANE |
elastase, neutrophil expressed |
| chr12_-_10875831 | 0.10 |
ENST00000279550.7 ENST00000228251.4 |
YBX3 |
Y box binding protein 3 |
| chr3_+_132316081 | 0.10 |
ENST00000249887.2 |
ACKR4 |
atypical chemokine receptor 4 |
| chr17_+_4736627 | 0.10 |
ENST00000355280.6 ENST00000347992.7 |
MINK1 |
misshapen-like kinase 1 |
| chr1_+_153963227 | 0.10 |
ENST00000368567.4 ENST00000392558.4 |
RPS27 |
ribosomal protein S27 |
| chr10_+_103113802 | 0.10 |
ENST00000370187.3 |
BTRC |
beta-transducin repeat containing E3 ubiquitin protein ligase |
| chr8_-_134115118 | 0.09 |
ENST00000395352.3 ENST00000338087.5 |
SLA |
Src-like-adaptor |
| chr9_+_78505581 | 0.09 |
ENST00000376767.3 ENST00000376752.4 |
PCSK5 |
proprotein convertase subtilisin/kexin type 5 |
| chr1_-_23521222 | 0.09 |
ENST00000374619.1 |
HTR1D |
5-hydroxytryptamine (serotonin) receptor 1D, G protein-coupled |
| chr3_+_32726774 | 0.09 |
ENST00000538368.1 |
CNOT10 |
CCR4-NOT transcription complex, subunit 10 |
| chr19_+_4247070 | 0.09 |
ENST00000262962.7 |
CCDC94 |
coiled-coil domain containing 94 |
| chr9_+_78505554 | 0.09 |
ENST00000545128.1 |
PCSK5 |
proprotein convertase subtilisin/kexin type 5 |
| chr18_-_3874752 | 0.08 |
ENST00000534970.1 |
DLGAP1 |
discs, large (Drosophila) homolog-associated protein 1 |
| chr1_-_23520755 | 0.08 |
ENST00000314113.3 |
HTR1D |
5-hydroxytryptamine (serotonin) receptor 1D, G protein-coupled |
| chr12_+_110011571 | 0.08 |
ENST00000539696.1 ENST00000228510.3 ENST00000392727.3 |
MVK |
mevalonate kinase |
| chr20_-_50385138 | 0.08 |
ENST00000338821.5 |
ATP9A |
ATPase, class II, type 9A |
| chr5_-_39270725 | 0.08 |
ENST00000512138.1 ENST00000512982.1 ENST00000540520.1 |
FYB |
FYN binding protein |
| chr17_+_41561317 | 0.08 |
ENST00000540306.1 ENST00000262415.3 ENST00000605777.1 |
DHX8 |
DEAH (Asp-Glu-Ala-His) box polypeptide 8 |
| chr1_+_16090914 | 0.07 |
ENST00000441801.2 |
FBLIM1 |
filamin binding LIM protein 1 |
| chr2_-_162931052 | 0.07 |
ENST00000360534.3 |
DPP4 |
dipeptidyl-peptidase 4 |
| chr10_+_43916052 | 0.07 |
ENST00000442526.2 |
RP11-517P14.2 |
RP11-517P14.2 |
| chr17_-_32690239 | 0.07 |
ENST00000225842.3 |
CCL1 |
chemokine (C-C motif) ligand 1 |
| chr3_-_49466686 | 0.06 |
ENST00000273598.3 ENST00000436744.2 |
NICN1 |
nicolin 1 |
| chr5_-_78281603 | 0.06 |
ENST00000264914.4 |
ARSB |
arylsulfatase B |
| chr1_+_117297007 | 0.06 |
ENST00000369478.3 ENST00000369477.1 |
CD2 |
CD2 molecule |
| chr17_+_3118915 | 0.06 |
ENST00000304094.1 |
OR1A1 |
olfactory receptor, family 1, subfamily A, member 1 |
| chr19_-_39421377 | 0.06 |
ENST00000430193.3 ENST00000600042.1 ENST00000221431.6 |
SARS2 |
seryl-tRNA synthetase 2, mitochondrial |
| chr4_+_26321284 | 0.06 |
ENST00000506956.1 ENST00000512671.1 ENST00000345843.3 ENST00000342295.1 |
RBPJ |
recombination signal binding protein for immunoglobulin kappa J region |
| chr6_+_42847649 | 0.06 |
ENST00000424341.2 ENST00000602561.1 |
RPL7L1 |
ribosomal protein L7-like 1 |
| chr6_+_153552455 | 0.06 |
ENST00000392385.2 |
AL590867.1 |
Uncharacterized protein; cDNA FLJ59044, highly similar to LINE-1 reverse transcriptase homolog |
| chr7_+_117251671 | 0.06 |
ENST00000468795.1 |
CFTR |
cystic fibrosis transmembrane conductance regulator (ATP-binding cassette sub-family C, member 7) |
| chr10_+_106034884 | 0.05 |
ENST00000369707.2 ENST00000429569.2 |
GSTO2 |
glutathione S-transferase omega 2 |
| chr16_+_66995121 | 0.05 |
ENST00000303334.4 |
CES3 |
carboxylesterase 3 |
| chr12_-_9360966 | 0.05 |
ENST00000261336.2 |
PZP |
pregnancy-zone protein |
| chr16_+_66995144 | 0.05 |
ENST00000394037.1 |
CES3 |
carboxylesterase 3 |
| chr6_+_22569784 | 0.05 |
ENST00000510882.2 |
HDGFL1 |
hepatoma derived growth factor-like 1 |
| chrX_+_107288239 | 0.05 |
ENST00000217957.5 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr19_-_58514129 | 0.04 |
ENST00000552184.1 ENST00000546715.1 ENST00000536132.1 ENST00000547828.1 ENST00000547121.1 ENST00000551380.1 |
ZNF606 |
zinc finger protein 606 |
| chr12_+_120740119 | 0.04 |
ENST00000536460.1 ENST00000202967.4 |
SIRT4 |
sirtuin 4 |
| chr10_+_12171636 | 0.04 |
ENST00000379051.1 ENST00000379033.3 ENST00000441368.1 ENST00000298428.9 ENST00000304267.8 |
SEC61A2 |
Sec61 alpha 2 subunit (S. cerevisiae) |
| chr15_-_20170354 | 0.03 |
ENST00000338912.5 |
IGHV1OR15-9 |
immunoglobulin heavy variable 1/OR15-9 (non-functional) |
| chr9_-_100684845 | 0.03 |
ENST00000375119.3 |
C9orf156 |
chromosome 9 open reading frame 156 |
| chr3_+_119499331 | 0.03 |
ENST00000393716.2 ENST00000466380.1 |
NR1I2 |
nuclear receptor subfamily 1, group I, member 2 |
| chr11_-_59383617 | 0.03 |
ENST00000263847.1 |
OSBP |
oxysterol binding protein |
| chr3_-_150264272 | 0.03 |
ENST00000491660.1 ENST00000487153.1 ENST00000239944.2 |
SERP1 |
stress-associated endoplasmic reticulum protein 1 |
| chr3_+_46204959 | 0.03 |
ENST00000357422.2 |
CCR3 |
chemokine (C-C motif) receptor 3 |
| chr13_+_21141208 | 0.03 |
ENST00000351808.5 |
IFT88 |
intraflagellar transport 88 homolog (Chlamydomonas) |
| chrX_+_107288197 | 0.02 |
ENST00000415430.3 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr10_-_97200772 | 0.02 |
ENST00000371241.1 ENST00000354106.3 ENST00000371239.1 ENST00000361941.3 ENST00000277982.5 ENST00000371245.3 |
SORBS1 |
sorbin and SH3 domain containing 1 |
| chr20_+_43835638 | 0.02 |
ENST00000372781.3 ENST00000244069.6 |
SEMG1 |
semenogelin I |
| chr17_+_55162453 | 0.02 |
ENST00000575322.1 ENST00000337714.3 ENST00000314126.3 |
AKAP1 |
A kinase (PRKA) anchor protein 1 |
| chr5_+_54320078 | 0.01 |
ENST00000231009.2 |
GZMK |
granzyme K (granzyme 3; tryptase II) |
| chr12_-_11036844 | 0.01 |
ENST00000428168.2 |
PRH1 |
proline-rich protein HaeIII subfamily 1 |
| chr12_+_8276433 | 0.01 |
ENST00000345999.3 ENST00000352620.3 ENST00000360500.3 |
CLEC4A |
C-type lectin domain family 4, member A |
| chr16_-_20339123 | 0.01 |
ENST00000381360.5 |
GP2 |
glycoprotein 2 (zymogen granule membrane) |
| chr14_-_58894332 | 0.01 |
ENST00000395159.2 |
TIMM9 |
translocase of inner mitochondrial membrane 9 homolog (yeast) |
| chr7_+_93535817 | 0.01 |
ENST00000248572.5 |
GNGT1 |
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr19_+_58111241 | 0.01 |
ENST00000597700.1 ENST00000332854.6 ENST00000597864.1 |
ZNF530 |
zinc finger protein 530 |
| chr12_-_49319265 | 0.01 |
ENST00000552878.1 ENST00000453172.2 |
FKBP11 |
FK506 binding protein 11, 19 kDa |
| chr8_-_6795823 | 0.01 |
ENST00000297435.2 |
DEFA4 |
defensin, alpha 4, corticostatin |
| chr21_-_35884573 | 0.00 |
ENST00000399286.2 |
KCNE1 |
potassium voltage-gated channel, Isk-related family, member 1 |
| chr11_+_27062502 | 0.00 |
ENST00000263182.3 |
BBOX1 |
butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.8 | GO:0046136 | positive regulation of vitamin metabolic process(GO:0046136) positive regulation of vitamin D biosynthetic process(GO:0060557) positive regulation of calcidiol 1-monooxygenase activity(GO:0060559) |
| 0.3 | 1.3 | GO:0060671 | epithelial cell differentiation involved in embryonic placenta development(GO:0060671) epithelial cell morphogenesis involved in placental branching(GO:0060672) |
| 0.3 | 1.1 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.2 | 2.2 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.2 | 1.0 | GO:0097527 | Fas signaling pathway(GO:0036337) necroptotic signaling pathway(GO:0097527) |
| 0.1 | 1.8 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.1 | 0.9 | GO:0035803 | egg coat formation(GO:0035803) |
| 0.1 | 0.4 | GO:0019276 | UDP-N-acetylgalactosamine metabolic process(GO:0019276) chondroitin sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0050653) |
| 0.1 | 0.9 | GO:0006933 | negative regulation of cell adhesion involved in substrate-bound cell migration(GO:0006933) |
| 0.1 | 0.3 | GO:0046125 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.1 | 0.5 | GO:0051414 | response to cortisol(GO:0051414) |
| 0.1 | 0.7 | GO:1902412 | regulation of mitotic cytokinesis(GO:1902412) |
| 0.1 | 1.8 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.0 | 0.1 | GO:0045013 | carbon catabolite repression of transcription(GO:0045013) negative regulation of transcription by glucose(GO:0045014) |
| 0.0 | 0.4 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 0.1 | GO:0051710 | regulation of cytolysis in other organism(GO:0051710) |
| 0.0 | 0.2 | GO:0032455 | nerve growth factor processing(GO:0032455) |
| 0.0 | 0.1 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.0 | 0.1 | GO:0010716 | negative regulation of extracellular matrix disassembly(GO:0010716) |
| 0.0 | 0.1 | GO:0070945 | neutrophil mediated killing of gram-negative bacterium(GO:0070945) |
| 0.0 | 0.2 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 0.0 | 0.2 | GO:0014827 | intestine smooth muscle contraction(GO:0014827) |
| 0.0 | 0.1 | GO:0006434 | seryl-tRNA aminoacylation(GO:0006434) selenocysteinyl-tRNA(Sec) biosynthetic process(GO:0097056) |
| 0.0 | 0.1 | GO:1905068 | arterial endothelial cell fate commitment(GO:0060844) blood vessel lumenization(GO:0072554) positive regulation of ephrin receptor signaling pathway(GO:1901189) positive regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901297) positive regulation of canonical Wnt signaling pathway involved in heart development(GO:1905068) |
| 0.0 | 0.1 | GO:1902161 | positive regulation of cyclic nucleotide-gated ion channel activity(GO:1902161) |
| 0.0 | 0.1 | GO:0001675 | acrosome assembly(GO:0001675) |
| 0.0 | 0.7 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.0 | 0.3 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.0 | 0.1 | GO:1902219 | negative regulation of intrinsic apoptotic signaling pathway in response to osmotic stress(GO:1902219) |
| 0.0 | 0.3 | GO:0000729 | DNA double-strand break processing(GO:0000729) |
| 0.0 | 0.0 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.0 | 0.1 | GO:0030885 | regulation of myeloid dendritic cell activation(GO:0030885) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.1 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 1.0 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 1.9 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.7 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 0.4 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 0.2 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 0.2 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.2 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 0.3 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0000235 | astral microtubule(GO:0000235) aster(GO:0005818) |
| 0.1 | 1.0 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 0.0 | 0.2 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 0.3 | GO:0090543 | Flemming body(GO:0090543) |
| 0.0 | 0.1 | GO:0097013 | phagocytic vesicle lumen(GO:0097013) |
| 0.0 | 1.9 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 1.8 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 0.1 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.0 | 0.4 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.1 | GO:0031205 | endoplasmic reticulum Sec complex(GO:0031205) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.0 | 1.0 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 1.0 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.0 | 4.3 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.8 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 0.1 | 0.4 | GO:0008955 | peptidoglycan glycosyltransferase activity(GO:0008955) |
| 0.1 | 0.3 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) thymidine kinase activity(GO:0004797) |
| 0.1 | 1.8 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.1 | 0.9 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.1 | 2.2 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.1 | 0.2 | GO:0016899 | oxidoreductase activity, acting on the CH-OH group of donors, oxygen as acceptor(GO:0016899) |
| 0.1 | 0.3 | GO:0008310 | single-stranded DNA 3'-5' exodeoxyribonuclease activity(GO:0008310) |
| 0.0 | 5.2 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.2 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.0 | 0.4 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.1 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 0.0 | 0.7 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.0 | 0.9 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.0 | 0.8 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.2 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 0.1 | GO:0004828 | serine-tRNA ligase activity(GO:0004828) |
| 0.0 | 0.1 | GO:0005260 | channel-conductance-controlling ATPase activity(GO:0005260) |
| 0.0 | 0.1 | GO:0045174 | oxidoreductase activity, acting on a sulfur group of donors, quinone or similar compound as acceptor(GO:0016672) glutathione dehydrogenase (ascorbate) activity(GO:0045174) methylarsonate reductase activity(GO:0050610) |
| 0.0 | 0.2 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.0 | 0.5 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 0.2 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 1.1 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.2 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |