Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
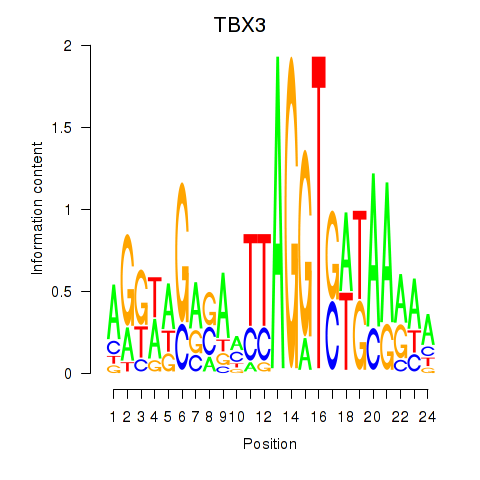
Results for TBX3
Z-value: 0.80
Transcription factors associated with TBX3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TBX3
|
ENSG00000135111.10 | TBX3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TBX3 | hg19_v2_chr12_-_115121962_115121987 | -0.33 | 2.1e-01 | Click! |
Activity profile of TBX3 motif
Sorted Z-values of TBX3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TBX3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr21_-_15918618 | 1.90 |
ENST00000400564.1 ENST00000400566.1 |
SAMSN1 |
SAM domain, SH3 domain and nuclear localization signals 1 |
| chr22_+_23237555 | 1.56 |
ENST00000390321.2 |
IGLC1 |
immunoglobulin lambda constant 1 (Mcg marker) |
| chr12_-_8765446 | 1.22 |
ENST00000537228.1 ENST00000229335.6 |
AICDA |
activation-induced cytidine deaminase |
| chr6_+_32821924 | 1.17 |
ENST00000374859.2 ENST00000453265.2 |
PSMB9 |
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr14_-_106471723 | 1.07 |
ENST00000390595.2 |
IGHV1-3 |
immunoglobulin heavy variable 1-3 |
| chr19_-_7766991 | 1.04 |
ENST00000597921.1 ENST00000346664.5 |
FCER2 |
Fc fragment of IgE, low affinity II, receptor for (CD23) |
| chr6_+_6588316 | 0.99 |
ENST00000379953.2 |
LY86 |
lymphocyte antigen 86 |
| chr19_+_1077393 | 0.95 |
ENST00000590577.1 |
HMHA1 |
histocompatibility (minor) HA-1 |
| chr11_-_73689037 | 0.85 |
ENST00000544615.1 |
UCP2 |
uncoupling protein 2 (mitochondrial, proton carrier) |
| chr7_+_150264365 | 0.78 |
ENST00000255945.2 ENST00000461940.1 |
GIMAP4 |
GTPase, IMAP family member 4 |
| chr3_+_133118839 | 0.75 |
ENST00000302334.2 |
BFSP2 |
beaded filament structural protein 2, phakinin |
| chr1_-_183560011 | 0.70 |
ENST00000367536.1 |
NCF2 |
neutrophil cytosolic factor 2 |
| chr4_-_84030996 | 0.66 |
ENST00000411416.2 |
PLAC8 |
placenta-specific 8 |
| chr14_-_106642049 | 0.65 |
ENST00000390605.2 |
IGHV1-18 |
immunoglobulin heavy variable 1-18 |
| chr6_-_32821599 | 0.57 |
ENST00000354258.4 |
TAP1 |
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr7_+_150434430 | 0.54 |
ENST00000358647.3 |
GIMAP5 |
GTPase, IMAP family member 5 |
| chr2_+_89184868 | 0.52 |
ENST00000390243.2 |
IGKV4-1 |
immunoglobulin kappa variable 4-1 |
| chr20_-_44539538 | 0.50 |
ENST00000372420.1 |
PLTP |
phospholipid transfer protein |
| chr11_+_19138670 | 0.49 |
ENST00000446113.2 ENST00000399351.3 |
ZDHHC13 |
zinc finger, DHHC-type containing 13 |
| chrX_-_70838306 | 0.47 |
ENST00000373691.4 ENST00000373693.3 |
CXCR3 |
chemokine (C-X-C motif) receptor 3 |
| chr11_+_7506713 | 0.46 |
ENST00000329293.3 ENST00000534244.1 |
OLFML1 |
olfactomedin-like 1 |
| chr5_-_40755987 | 0.41 |
ENST00000337702.4 |
TTC33 |
tetratricopeptide repeat domain 33 |
| chr7_+_22766766 | 0.41 |
ENST00000426291.1 ENST00000401651.1 ENST00000258743.5 ENST00000420258.2 ENST00000407492.1 ENST00000401630.3 ENST00000406575.1 |
IL6 |
interleukin 6 (interferon, beta 2) |
| chr7_+_98476095 | 0.41 |
ENST00000359863.4 ENST00000355540.3 |
TRRAP |
transformation/transcription domain-associated protein |
| chr2_-_24149977 | 0.39 |
ENST00000238789.5 |
ATAD2B |
ATPase family, AAA domain containing 2B |
| chr2_+_65215604 | 0.38 |
ENST00000531327.1 |
SLC1A4 |
solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 |
| chr14_-_21270561 | 0.38 |
ENST00000412779.2 |
RNASE1 |
ribonuclease, RNase A family, 1 (pancreatic) |
| chr4_+_178230985 | 0.38 |
ENST00000264596.3 |
NEIL3 |
nei endonuclease VIII-like 3 (E. coli) |
| chr19_+_56717283 | 0.37 |
ENST00000376267.1 |
ZSCAN5C |
zinc finger and SCAN domain containing 5C |
| chr8_-_81083341 | 0.37 |
ENST00000519303.2 |
TPD52 |
tumor protein D52 |
| chr14_-_74417096 | 0.36 |
ENST00000286544.3 |
FAM161B |
family with sequence similarity 161, member B |
| chr22_+_24891210 | 0.36 |
ENST00000382760.2 |
UPB1 |
ureidopropionase, beta |
| chr7_+_99971129 | 0.36 |
ENST00000394000.2 ENST00000350573.2 |
PILRA |
paired immunoglobin-like type 2 receptor alpha |
| chr9_-_130679257 | 0.35 |
ENST00000361444.3 ENST00000335791.5 ENST00000343609.2 |
ST6GALNAC4 |
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 4 |
| chr11_+_1718425 | 0.34 |
ENST00000382160.1 |
KRTAP5-6 |
keratin associated protein 5-6 |
| chr4_-_46911223 | 0.34 |
ENST00000396533.1 |
COX7B2 |
cytochrome c oxidase subunit VIIb2 |
| chr4_-_46911248 | 0.33 |
ENST00000355591.3 ENST00000505102.1 |
COX7B2 |
cytochrome c oxidase subunit VIIb2 |
| chr14_-_21270995 | 0.33 |
ENST00000555698.1 ENST00000397970.4 ENST00000340900.3 |
RNASE1 |
ribonuclease, RNase A family, 1 (pancreatic) |
| chr12_-_54652060 | 0.33 |
ENST00000552562.1 |
CBX5 |
chromobox homolog 5 |
| chr2_-_27545921 | 0.33 |
ENST00000402310.1 ENST00000405983.1 ENST00000403262.2 ENST00000428910.1 ENST00000402722.1 ENST00000399052.4 ENST00000380044.1 ENST00000405076.1 |
MPV17 |
MpV17 mitochondrial inner membrane protein |
| chr19_-_10491234 | 0.32 |
ENST00000524462.1 ENST00000531836.1 ENST00000525621.1 |
TYK2 |
tyrosine kinase 2 |
| chr9_+_114287433 | 0.30 |
ENST00000358151.4 ENST00000355824.3 ENST00000374374.3 ENST00000309235.5 |
ZNF483 |
zinc finger protein 483 |
| chr15_+_77712993 | 0.30 |
ENST00000336216.4 ENST00000381714.3 ENST00000558651.1 |
HMG20A |
high mobility group 20A |
| chr6_-_35888905 | 0.30 |
ENST00000510290.1 ENST00000423325.2 ENST00000373822.1 |
SRPK1 |
SRSF protein kinase 1 |
| chrX_+_51942963 | 0.30 |
ENST00000375625.3 |
RP11-363G10.2 |
RP11-363G10.2 |
| chr19_+_49403562 | 0.30 |
ENST00000407032.1 ENST00000452087.1 ENST00000411700.1 |
NUCB1 |
nucleobindin 1 |
| chr19_+_49404041 | 0.29 |
ENST00000263273.5 ENST00000424608.1 |
NUCB1 |
nucleobindin 1 |
| chr7_-_150675372 | 0.28 |
ENST00000262186.5 |
KCNH2 |
potassium voltage-gated channel, subfamily H (eag-related), member 2 |
| chrX_-_101726732 | 0.28 |
ENST00000457521.2 ENST00000412230.2 ENST00000453326.2 |
NXF2B TCP11X2 |
nuclear RNA export factor 2B t-complex 11 family, X-linked 2 |
| chrX_+_101470280 | 0.28 |
ENST00000395088.2 ENST00000330252.5 ENST00000333110.5 |
NXF2 TCP11X1 |
nuclear RNA export factor 2 t-complex 11 family, X-linked 1 |
| chr11_-_327537 | 0.27 |
ENST00000602735.1 |
IFITM3 |
interferon induced transmembrane protein 3 |
| chr10_-_13523073 | 0.27 |
ENST00000440282.1 |
BEND7 |
BEN domain containing 7 |
| chr19_+_58281014 | 0.27 |
ENST00000391702.3 ENST00000598885.1 ENST00000598183.1 ENST00000396154.2 ENST00000599802.1 ENST00000396150.4 |
ZNF586 |
zinc finger protein 586 |
| chr22_-_36031181 | 0.27 |
ENST00000594060.1 |
AL049747.1 |
AL049747.1 |
| chr16_-_68057770 | 0.26 |
ENST00000332395.5 |
DDX28 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 28 |
| chrX_+_65382381 | 0.26 |
ENST00000519389.1 |
HEPH |
hephaestin |
| chr2_-_89513402 | 0.25 |
ENST00000498435.1 |
IGKV1-27 |
immunoglobulin kappa variable 1-27 |
| chr16_+_84801852 | 0.25 |
ENST00000569925.1 ENST00000567526.1 |
USP10 |
ubiquitin specific peptidase 10 |
| chr6_-_131277510 | 0.25 |
ENST00000525193.1 ENST00000527659.1 |
EPB41L2 |
erythrocyte membrane protein band 4.1-like 2 |
| chr4_-_48908822 | 0.24 |
ENST00000508632.1 |
OCIAD2 |
OCIA domain containing 2 |
| chrX_+_65382433 | 0.24 |
ENST00000374727.3 |
HEPH |
hephaestin |
| chr3_-_61237050 | 0.24 |
ENST00000476844.1 ENST00000488467.1 ENST00000492590.1 ENST00000468189.1 |
FHIT |
fragile histidine triad |
| chr7_-_134896234 | 0.23 |
ENST00000354475.4 ENST00000344400.5 |
WDR91 |
WD repeat domain 91 |
| chr17_+_43861680 | 0.23 |
ENST00000314537.5 |
CRHR1 |
corticotropin releasing hormone receptor 1 |
| chr16_+_83932684 | 0.22 |
ENST00000262430.4 |
MLYCD |
malonyl-CoA decarboxylase |
| chr18_-_5544241 | 0.21 |
ENST00000341928.2 ENST00000540638.2 |
EPB41L3 |
erythrocyte membrane protein band 4.1-like 3 |
| chrX_-_131352152 | 0.21 |
ENST00000342983.2 |
RAP2C |
RAP2C, member of RAS oncogene family |
| chr1_-_231560790 | 0.21 |
ENST00000366641.3 |
EGLN1 |
egl-9 family hypoxia-inducible factor 1 |
| chr2_-_24270217 | 0.20 |
ENST00000295148.4 ENST00000406895.3 |
C2orf44 |
chromosome 2 open reading frame 44 |
| chr21_+_27011899 | 0.20 |
ENST00000425221.2 |
JAM2 |
junctional adhesion molecule 2 |
| chr11_+_71249071 | 0.20 |
ENST00000398534.3 |
KRTAP5-8 |
keratin associated protein 5-8 |
| chr11_-_10590118 | 0.20 |
ENST00000529598.1 |
LYVE1 |
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr19_-_42931567 | 0.20 |
ENST00000244289.4 |
LIPE |
lipase, hormone-sensitive |
| chr1_+_212208919 | 0.19 |
ENST00000366991.4 ENST00000542077.1 |
DTL |
denticleless E3 ubiquitin protein ligase homolog (Drosophila) |
| chr7_+_99971068 | 0.19 |
ENST00000198536.2 ENST00000453419.1 |
PILRA |
paired immunoglobin-like type 2 receptor alpha |
| chr1_-_11865351 | 0.19 |
ENST00000413656.1 ENST00000376585.1 ENST00000423400.1 ENST00000431243.1 |
MTHFR |
methylenetetrahydrofolate reductase (NAD(P)H) |
| chr11_-_10590238 | 0.19 |
ENST00000256178.3 |
LYVE1 |
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr2_+_90153696 | 0.18 |
ENST00000417279.2 |
IGKV3D-15 |
immunoglobulin kappa variable 3D-15 (gene/pseudogene) |
| chr14_-_107283278 | 0.18 |
ENST00000390639.2 |
IGHV7-81 |
immunoglobulin heavy variable 7-81 (non-functional) |
| chr3_-_81792780 | 0.18 |
ENST00000489715.1 |
GBE1 |
glucan (1,4-alpha-), branching enzyme 1 |
| chr2_-_217560248 | 0.18 |
ENST00000233813.4 |
IGFBP5 |
insulin-like growth factor binding protein 5 |
| chr10_+_47894023 | 0.18 |
ENST00000358474.5 |
FAM21B |
family with sequence similarity 21, member B |
| chr10_-_44144152 | 0.17 |
ENST00000395797.1 |
ZNF32 |
zinc finger protein 32 |
| chr9_-_95087604 | 0.17 |
ENST00000542613.1 ENST00000542053.1 ENST00000358855.4 ENST00000545558.1 ENST00000432670.2 ENST00000433029.2 ENST00000411621.2 |
NOL8 |
nucleolar protein 8 |
| chr11_-_1629693 | 0.17 |
ENST00000399685.1 |
KRTAP5-3 |
keratin associated protein 5-3 |
| chr11_+_71259466 | 0.16 |
ENST00000528743.2 |
KRTAP5-9 |
keratin associated protein 5-9 |
| chr17_-_41132410 | 0.16 |
ENST00000409446.3 ENST00000453594.1 ENST00000409399.1 ENST00000421990.2 |
PTGES3L PTGES3L-AARSD1 |
prostaglandin E synthase 3 (cytosolic)-like PTGES3L-AARSD1 readthrough |
| chr6_+_32146131 | 0.16 |
ENST00000375094.3 |
RNF5 |
ring finger protein 5, E3 ubiquitin protein ligase |
| chrX_-_57147748 | 0.15 |
ENST00000374910.3 |
SPIN2B |
spindlin family, member 2B |
| chr2_+_90192768 | 0.15 |
ENST00000390275.2 |
IGKV1D-13 |
immunoglobulin kappa variable 1D-13 |
| chr3_-_121264848 | 0.15 |
ENST00000264233.5 |
POLQ |
polymerase (DNA directed), theta |
| chr20_+_30865429 | 0.15 |
ENST00000375712.3 |
KIF3B |
kinesin family member 3B |
| chr13_+_92050928 | 0.15 |
ENST00000377067.3 |
GPC5 |
glypican 5 |
| chr19_-_55691472 | 0.15 |
ENST00000537500.1 |
SYT5 |
synaptotagmin V |
| chrX_+_149887090 | 0.15 |
ENST00000538506.1 |
MTMR1 |
myotubularin related protein 1 |
| chr1_+_212606219 | 0.15 |
ENST00000366988.3 |
NENF |
neudesin neurotrophic factor |
| chr4_+_15704573 | 0.14 |
ENST00000265016.4 |
BST1 |
bone marrow stromal cell antigen 1 |
| chrX_-_57147902 | 0.14 |
ENST00000275988.5 ENST00000434397.1 ENST00000333933.3 ENST00000374912.5 |
SPIN2B |
spindlin family, member 2B |
| chr19_-_14945933 | 0.14 |
ENST00000322301.3 |
OR7A5 |
olfactory receptor, family 7, subfamily A, member 5 |
| chr19_-_51538118 | 0.14 |
ENST00000529888.1 |
KLK12 |
kallikrein-related peptidase 12 |
| chr19_-_6333614 | 0.14 |
ENST00000301452.4 |
ACER1 |
alkaline ceramidase 1 |
| chr22_-_21581926 | 0.14 |
ENST00000401924.1 |
GGT2 |
gamma-glutamyltransferase 2 |
| chr2_+_178257372 | 0.14 |
ENST00000264167.4 ENST00000409888.1 |
AGPS |
alkylglycerone phosphate synthase |
| chr3_+_149192475 | 0.13 |
ENST00000465758.1 |
TM4SF4 |
transmembrane 4 L six family member 4 |
| chr14_+_74111578 | 0.13 |
ENST00000554113.1 ENST00000555631.2 ENST00000553645.2 ENST00000311089.3 ENST00000555919.3 ENST00000554339.1 ENST00000554871.1 |
DNAL1 |
dynein, axonemal, light chain 1 |
| chr22_+_24999114 | 0.13 |
ENST00000412658.1 ENST00000445029.1 ENST00000419133.1 ENST00000400382.1 ENST00000438643.2 ENST00000452551.1 ENST00000400383.1 ENST00000412898.1 ENST00000400380.1 ENST00000455483.1 ENST00000430289.1 |
GGT1 |
gamma-glutamyltransferase 1 |
| chr19_+_42724423 | 0.13 |
ENST00000301215.3 ENST00000597945.1 |
ZNF526 |
zinc finger protein 526 |
| chr13_-_46679144 | 0.13 |
ENST00000181383.4 |
CPB2 |
carboxypeptidase B2 (plasma) |
| chr3_+_108541608 | 0.13 |
ENST00000426646.1 |
TRAT1 |
T cell receptor associated transmembrane adaptor 1 |
| chr10_+_47894572 | 0.12 |
ENST00000355876.5 |
FAM21B |
family with sequence similarity 21, member B |
| chr2_+_162101247 | 0.12 |
ENST00000439050.1 ENST00000436506.1 |
AC009299.3 |
AC009299.3 |
| chrX_+_135570046 | 0.12 |
ENST00000370648.3 |
BRS3 |
bombesin-like receptor 3 |
| chr1_-_101491319 | 0.12 |
ENST00000342173.7 ENST00000488176.1 ENST00000370109.3 |
DPH5 |
diphthamide biosynthesis 5 |
| chr6_+_31514622 | 0.12 |
ENST00000376146.4 |
NFKBIL1 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor-like 1 |
| chr6_-_33771757 | 0.12 |
ENST00000507738.1 ENST00000266003.5 ENST00000430124.2 |
MLN |
motilin |
| chr17_-_61959202 | 0.12 |
ENST00000449787.2 ENST00000456543.2 ENST00000423893.2 ENST00000332800.7 |
GH2 |
growth hormone 2 |
| chr19_-_55791058 | 0.12 |
ENST00000587959.1 ENST00000585927.1 ENST00000587922.1 ENST00000585698.1 |
HSPBP1 |
HSPA (heat shock 70kDa) binding protein, cytoplasmic cochaperone 1 |
| chr12_-_7848364 | 0.11 |
ENST00000329913.3 |
GDF3 |
growth differentiation factor 3 |
| chr19_-_51538148 | 0.11 |
ENST00000319590.4 ENST00000250351.4 |
KLK12 |
kallikrein-related peptidase 12 |
| chr2_-_89266286 | 0.11 |
ENST00000464162.1 |
IGKV1-6 |
immunoglobulin kappa variable 1-6 |
| chr15_-_78592053 | 0.11 |
ENST00000267973.2 |
WDR61 |
WD repeat domain 61 |
| chr3_-_178976996 | 0.11 |
ENST00000485523.1 |
KCNMB3 |
potassium large conductance calcium-activated channel, subfamily M beta member 3 |
| chr10_+_51827648 | 0.11 |
ENST00000351071.6 ENST00000314664.7 ENST00000282633.5 |
FAM21A |
family with sequence similarity 21, member A |
| chr19_-_55691614 | 0.11 |
ENST00000592470.1 ENST00000354308.3 |
SYT5 |
synaptotagmin V |
| chr3_+_148709128 | 0.11 |
ENST00000345003.4 ENST00000296048.6 ENST00000483267.1 |
GYG1 |
glycogenin 1 |
| chr5_-_132362226 | 0.11 |
ENST00000509437.1 ENST00000355372.2 ENST00000513541.1 ENST00000509008.1 ENST00000513848.1 ENST00000504170.1 ENST00000324170.3 |
ZCCHC10 |
zinc finger, CCHC domain containing 10 |
| chr15_-_74501310 | 0.11 |
ENST00000423167.2 ENST00000432245.2 |
STRA6 |
stimulated by retinoic acid 6 |
| chr10_+_60028818 | 0.10 |
ENST00000333926.5 |
CISD1 |
CDGSH iron sulfur domain 1 |
| chr3_+_108541545 | 0.10 |
ENST00000295756.6 |
TRAT1 |
T cell receptor associated transmembrane adaptor 1 |
| chr5_+_150040403 | 0.10 |
ENST00000517768.1 ENST00000297130.4 |
MYOZ3 |
myozenin 3 |
| chr10_-_71176623 | 0.10 |
ENST00000373306.4 |
TACR2 |
tachykinin receptor 2 |
| chr1_-_182921119 | 0.09 |
ENST00000423786.1 |
SHCBP1L |
SHC SH2-domain binding protein 1-like |
| chr1_+_32608566 | 0.09 |
ENST00000545542.1 |
KPNA6 |
karyopherin alpha 6 (importin alpha 7) |
| chr3_+_52454971 | 0.09 |
ENST00000465863.1 |
PHF7 |
PHD finger protein 7 |
| chr1_+_36396791 | 0.09 |
ENST00000246314.6 |
AGO3 |
argonaute RISC catalytic component 3 |
| chr15_+_40861487 | 0.09 |
ENST00000315616.7 ENST00000559271.1 |
RPUSD2 |
RNA pseudouridylate synthase domain containing 2 |
| chr8_+_79578282 | 0.09 |
ENST00000263849.4 |
ZC2HC1A |
zinc finger, C2HC-type containing 1A |
| chr9_+_130186653 | 0.09 |
ENST00000342483.5 ENST00000543471.1 |
ZNF79 |
zinc finger protein 79 |
| chr9_+_5231413 | 0.08 |
ENST00000239316.4 |
INSL4 |
insulin-like 4 (placenta) |
| chr7_+_73106926 | 0.08 |
ENST00000453316.1 |
WBSCR22 |
Williams Beuren syndrome chromosome region 22 |
| chr1_+_36396677 | 0.08 |
ENST00000373191.4 ENST00000397828.2 |
AGO3 |
argonaute RISC catalytic component 3 |
| chr10_+_46222648 | 0.08 |
ENST00000336378.4 ENST00000540872.1 ENST00000537517.1 ENST00000374362.2 ENST00000359860.4 ENST00000420848.1 |
FAM21C |
family with sequence similarity 21, member C |
| chr17_-_26220366 | 0.08 |
ENST00000460380.2 ENST00000508862.1 ENST00000379102.3 ENST00000582441.1 |
LYRM9 RP1-66C13.4 |
LYR motif containing 9 Uncharacterized protein |
| chr10_+_52152766 | 0.08 |
ENST00000596442.1 |
AC069547.2 |
Uncharacterized protein |
| chr17_+_26800756 | 0.08 |
ENST00000537681.1 |
SLC13A2 |
solute carrier family 13 (sodium-dependent dicarboxylate transporter), member 2 |
| chr15_-_74504560 | 0.08 |
ENST00000449139.2 |
STRA6 |
stimulated by retinoic acid 6 |
| chr3_+_122103014 | 0.07 |
ENST00000232125.5 ENST00000477892.1 ENST00000469967.1 |
FAM162A |
family with sequence similarity 162, member A |
| chr19_-_47735918 | 0.07 |
ENST00000449228.1 ENST00000300880.7 ENST00000341983.4 |
BBC3 |
BCL2 binding component 3 |
| chr11_-_104972158 | 0.07 |
ENST00000598974.1 ENST00000593315.1 ENST00000594519.1 ENST00000415981.2 ENST00000525374.1 ENST00000375707.1 |
CASP1 CARD16 CARD17 |
caspase 1, apoptosis-related cysteine peptidase caspase recruitment domain family, member 16 caspase recruitment domain family, member 17 |
| chr19_+_21265028 | 0.07 |
ENST00000291770.7 |
ZNF714 |
zinc finger protein 714 |
| chr2_+_55459495 | 0.07 |
ENST00000272317.6 ENST00000449323.1 |
RPS27A |
ribosomal protein S27a |
| chr1_-_212208842 | 0.07 |
ENST00000366992.3 ENST00000366993.3 ENST00000440600.2 ENST00000366994.3 |
INTS7 |
integrator complex subunit 7 |
| chr10_+_71561704 | 0.06 |
ENST00000520267.1 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr11_-_8285405 | 0.06 |
ENST00000335790.3 ENST00000534484.1 |
LMO1 |
LIM domain only 1 (rhombotin 1) |
| chr15_+_31196053 | 0.06 |
ENST00000561594.1 ENST00000362065.4 |
FAN1 |
FANCD2/FANCI-associated nuclease 1 |
| chr12_-_88535842 | 0.06 |
ENST00000550962.1 ENST00000552810.1 |
CEP290 |
centrosomal protein 290kDa |
| chr19_-_33360647 | 0.06 |
ENST00000590341.1 ENST00000587772.1 ENST00000023064.4 |
SLC7A9 |
solute carrier family 7 (amino acid transporter light chain, bo,+ system), member 9 |
| chr1_+_36396313 | 0.06 |
ENST00000324350.5 |
AGO3 |
argonaute RISC catalytic component 3 |
| chr12_+_57849048 | 0.05 |
ENST00000266646.2 |
INHBE |
inhibin, beta E |
| chr19_+_41103063 | 0.05 |
ENST00000308370.7 |
LTBP4 |
latent transforming growth factor beta binding protein 4 |
| chr22_+_41487711 | 0.05 |
ENST00000263253.7 |
EP300 |
E1A binding protein p300 |
| chr14_-_50087312 | 0.05 |
ENST00000298289.6 |
RPL36AL |
ribosomal protein L36a-like |
| chr1_+_171810606 | 0.05 |
ENST00000358155.4 ENST00000367733.2 ENST00000355305.5 ENST00000367731.1 |
DNM3 |
dynamin 3 |
| chr17_-_1419914 | 0.05 |
ENST00000397335.3 ENST00000574561.1 |
INPP5K |
inositol polyphosphate-5-phosphatase K |
| chr7_-_76255444 | 0.04 |
ENST00000454397.1 |
POMZP3 |
POM121 and ZP3 fusion |
| chr11_+_71238313 | 0.04 |
ENST00000398536.4 |
KRTAP5-7 |
keratin associated protein 5-7 |
| chr5_+_54320078 | 0.04 |
ENST00000231009.2 |
GZMK |
granzyme K (granzyme 3; tryptase II) |
| chr17_+_38465441 | 0.04 |
ENST00000577646.1 ENST00000254066.5 |
RARA |
retinoic acid receptor, alpha |
| chr20_-_1472029 | 0.04 |
ENST00000359801.3 |
SIRPB2 |
signal-regulatory protein beta 2 |
| chr4_-_69215699 | 0.03 |
ENST00000510746.1 ENST00000344157.4 ENST00000355665.3 |
YTHDC1 |
YTH domain containing 1 |
| chr3_-_45838011 | 0.03 |
ENST00000358525.4 ENST00000413781.1 |
SLC6A20 |
solute carrier family 6 (proline IMINO transporter), member 20 |
| chr1_-_241799232 | 0.03 |
ENST00000366553.1 |
CHML |
choroideremia-like (Rab escort protein 2) |
| chr1_+_220267429 | 0.03 |
ENST00000366922.1 ENST00000302637.5 |
IARS2 |
isoleucyl-tRNA synthetase 2, mitochondrial |
| chr1_+_151020175 | 0.03 |
ENST00000368926.5 |
C1orf56 |
chromosome 1 open reading frame 56 |
| chr8_-_72268889 | 0.03 |
ENST00000388742.4 |
EYA1 |
eyes absent homolog 1 (Drosophila) |
| chr13_+_73356197 | 0.03 |
ENST00000326291.6 |
PIBF1 |
progesterone immunomodulatory binding factor 1 |
| chr7_-_99381798 | 0.03 |
ENST00000415003.1 ENST00000354593.2 |
CYP3A4 |
cytochrome P450, family 3, subfamily A, polypeptide 4 |
| chr1_-_154461642 | 0.03 |
ENST00000555188.1 |
SHE |
Src homology 2 domain containing E |
| chr22_-_42915788 | 0.02 |
ENST00000323013.6 |
RRP7A |
ribosomal RNA processing 7 homolog A (S. cerevisiae) |
| chr20_-_4229721 | 0.02 |
ENST00000379453.4 |
ADRA1D |
adrenoceptor alpha 1D |
| chr8_-_133123406 | 0.02 |
ENST00000434736.2 |
HHLA1 |
HERV-H LTR-associating 1 |
| chr4_+_156824840 | 0.02 |
ENST00000536354.2 |
TDO2 |
tryptophan 2,3-dioxygenase |
| chr18_+_55018044 | 0.02 |
ENST00000324000.3 |
ST8SIA3 |
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 |
| chr2_-_209119831 | 0.02 |
ENST00000345146.2 |
IDH1 |
isocitrate dehydrogenase 1 (NADP+), soluble |
| chr7_-_2883928 | 0.02 |
ENST00000275364.3 |
GNA12 |
guanine nucleotide binding protein (G protein) alpha 12 |
| chr11_-_122931881 | 0.02 |
ENST00000526110.1 ENST00000227378.3 |
HSPA8 |
heat shock 70kDa protein 8 |
| chr12_+_133757995 | 0.02 |
ENST00000536435.2 ENST00000228289.5 ENST00000541211.2 ENST00000500625.3 ENST00000539248.2 ENST00000542711.2 ENST00000536899.2 ENST00000542986.2 ENST00000537565.1 ENST00000541975.2 |
ZNF268 |
zinc finger protein 268 |
| chr12_+_133758115 | 0.02 |
ENST00000541009.2 ENST00000592241.1 |
ZNF268 |
zinc finger protein 268 |
| chr5_-_174871136 | 0.02 |
ENST00000393752.2 |
DRD1 |
dopamine receptor D1 |
| chr8_+_23386305 | 0.02 |
ENST00000519973.1 |
SLC25A37 |
solute carrier family 25 (mitochondrial iron transporter), member 37 |
| chr19_+_19322758 | 0.02 |
ENST00000252575.6 |
NCAN |
neurocan |
| chr12_-_88535747 | 0.02 |
ENST00000309041.7 |
CEP290 |
centrosomal protein 290kDa |
| chr19_+_496454 | 0.02 |
ENST00000346144.4 ENST00000215637.3 ENST00000382683.4 |
MADCAM1 |
mucosal vascular addressin cell adhesion molecule 1 |
| chrX_+_44703249 | 0.02 |
ENST00000339042.4 |
DUSP21 |
dual specificity phosphatase 21 |
| chr4_+_25314388 | 0.01 |
ENST00000302874.4 |
ZCCHC4 |
zinc finger, CCHC domain containing 4 |
| chr1_+_26869597 | 0.01 |
ENST00000530003.1 |
RPS6KA1 |
ribosomal protein S6 kinase, 90kDa, polypeptide 1 |
| chr8_-_72274095 | 0.01 |
ENST00000303824.7 |
EYA1 |
eyes absent homolog 1 (Drosophila) |
| chr2_+_55459808 | 0.01 |
ENST00000404735.1 |
RPS27A |
ribosomal protein S27a |
| chr19_+_41257084 | 0.01 |
ENST00000601393.1 |
SNRPA |
small nuclear ribonucleoprotein polypeptide A |
| chr22_+_23063100 | 0.01 |
ENST00000390309.2 |
IGLV3-19 |
immunoglobulin lambda variable 3-19 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 0.4 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.0 | 0.5 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.0 | 0.5 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 2.1 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 0.0 | 0.2 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.0 | 0.4 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.0 | 2.3 | PID IL4 2PATHWAY | IL4-mediated signaling events |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0090498 | extrinsic component of Golgi membrane(GO:0090498) |
| 0.1 | 1.2 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.1 | 0.4 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 0.1 | 0.6 | GO:0042825 | TAP complex(GO:0042825) |
| 0.1 | 0.7 | GO:0032010 | phagolysosome(GO:0032010) |
| 0.1 | 3.4 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 0.3 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.1 | 0.3 | GO:0042272 | nuclear RNA export factor complex(GO:0042272) |
| 0.1 | 0.3 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.1 | 0.2 | GO:0005873 | plus-end kinesin complex(GO:0005873) |
| 0.0 | 1.2 | GO:0000178 | exosome (RNase complex)(GO:0000178) |
| 0.0 | 0.4 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.0 | 0.7 | GO:0045277 | respiratory chain complex IV(GO:0045277) |
| 0.0 | 0.1 | GO:0055087 | Ski complex(GO:0055087) |
| 0.0 | 0.2 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.0 | 0.2 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.0 | 0.3 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.2 | GO:0008091 | spectrin(GO:0008091) |
| 0.0 | 0.1 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.0 | 0.1 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.0 | 0.7 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.2 | GO:0036513 | Derlin-1 retrotranslocation complex(GO:0036513) |
| 0.0 | 0.1 | GO:0097179 | protease inhibitor complex(GO:0097179) |
| 0.0 | 1.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.1 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.2 | GO:0033270 | paranode region of axon(GO:0033270) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.2 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.1 | 1.0 | GO:0019863 | IgE binding(GO:0019863) |
| 0.1 | 0.9 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 0.1 | 0.7 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.1 | 0.3 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.1 | 0.7 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.1 | 0.4 | GO:0015180 | L-alanine transmembrane transporter activity(GO:0015180) alanine transmembrane transporter activity(GO:0022858) |
| 0.1 | 0.6 | GO:0046979 | TAP2 binding(GO:0046979) |
| 0.1 | 0.2 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.1 | 3.4 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 0.5 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.0 | 0.1 | GO:0003953 | NAD+ nucleosidase activity(GO:0003953) NAD(P)+ nucleosidase activity(GO:0050135) |
| 0.0 | 0.4 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.0 | 0.3 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.0 | 0.4 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.0 | 0.2 | GO:0015056 | corticotrophin-releasing factor receptor activity(GO:0015056) |
| 0.0 | 0.5 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.0 | 1.2 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 0.2 | GO:0004489 | methylenetetrahydrofolate reductase (NAD(P)H) activity(GO:0004489) |
| 0.0 | 0.5 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.1 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.1 | GO:0051373 | telethonin binding(GO:0031433) FATZ binding(GO:0051373) |
| 0.0 | 1.9 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.3 | GO:0005131 | growth hormone receptor binding(GO:0005131) |
| 0.0 | 0.1 | GO:0008466 | glycogenin glucosyltransferase activity(GO:0008466) |
| 0.0 | 0.1 | GO:0015361 | low-affinity sodium:dicarboxylate symporter activity(GO:0015361) |
| 0.0 | 0.2 | GO:0042731 | PH domain binding(GO:0042731) |
| 0.0 | 0.7 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.7 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.2 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.0 | 0.1 | GO:0017108 | 5'-flap endonuclease activity(GO:0017108) |
| 0.0 | 0.1 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.0 | 0.3 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.0 | 0.2 | GO:0043142 | single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.0 | 0.6 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.6 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 0.2 | GO:0031545 | peptidyl-proline 4-dioxygenase activity(GO:0031545) |
| 0.0 | 0.2 | GO:0004551 | nucleotide diphosphatase activity(GO:0004551) |
| 0.0 | 0.1 | GO:0015184 | L-cystine transmembrane transporter activity(GO:0015184) |
| 0.0 | 0.1 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.2 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 0.2 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.0 | 0.2 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.0 | 0.0 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 0.0 | 0.2 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.0 | 0.1 | GO:0097157 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) pre-mRNA intronic binding(GO:0097157) |
| 0.0 | 0.4 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:0002925 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) |
| 0.2 | 1.2 | GO:0090308 | regulation of methylation-dependent chromatin silencing(GO:0090308) |
| 0.1 | 0.6 | GO:0046967 | cytosol to ER transport(GO:0046967) |
| 0.1 | 0.4 | GO:0002384 | hepatic immune response(GO:0002384) response to prolactin(GO:1990637) regulation of STAT protein import into nucleus(GO:2000364) positive regulation of STAT protein import into nucleus(GO:2000366) |
| 0.1 | 0.5 | GO:0042360 | vitamin E metabolic process(GO:0042360) |
| 0.1 | 0.4 | GO:0019482 | beta-alanine metabolic process(GO:0019482) |
| 0.1 | 1.0 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.1 | 0.4 | GO:0015808 | L-alanine transport(GO:0015808) |
| 0.1 | 0.3 | GO:1902303 | regulation of heart rate by hormone(GO:0003064) negative regulation of potassium ion export(GO:1902303) |
| 0.1 | 0.9 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.1 | 0.6 | GO:0072718 | response to cisplatin(GO:0072718) |
| 0.1 | 0.3 | GO:0061668 | mitochondrial ribosome assembly(GO:0061668) mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.1 | 0.8 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.0 | 0.4 | GO:0031936 | negative regulation of chromatin silencing(GO:0031936) |
| 0.0 | 3.4 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.1 | GO:1904204 | regulation of skeletal muscle hypertrophy(GO:1904204) |
| 0.0 | 0.2 | GO:0035279 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing by siRNA(GO:0090625) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.0 | 0.1 | GO:1901503 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) ether biosynthetic process(GO:1901503) |
| 0.0 | 1.9 | GO:0050869 | negative regulation of B cell activation(GO:0050869) |
| 0.0 | 0.2 | GO:0036115 | fatty-acyl-CoA catabolic process(GO:0036115) malonyl-CoA metabolic process(GO:2001293) |
| 0.0 | 0.2 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.0 | 0.2 | GO:0070829 | response to vitamin B2(GO:0033274) heterochromatin maintenance(GO:0070829) |
| 0.0 | 0.2 | GO:0061143 | alveolar primary septum development(GO:0061143) |
| 0.0 | 0.7 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.0 | 0.1 | GO:2001162 | regulation of histone H3-K79 methylation(GO:2001160) positive regulation of histone H3-K79 methylation(GO:2001162) |
| 0.0 | 0.5 | GO:1903830 | magnesium ion transmembrane transport(GO:1903830) |
| 0.0 | 0.2 | GO:0033121 | regulation of cyclic nucleotide catabolic process(GO:0030805) regulation of cAMP catabolic process(GO:0030820) regulation of purine nucleotide catabolic process(GO:0033121) regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051342) negative regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051344) |
| 0.0 | 0.1 | GO:0014057 | positive regulation of acetylcholine secretion, neurotransmission(GO:0014057) |
| 0.0 | 0.1 | GO:0003331 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.0 | 0.2 | GO:0051458 | corticotropin secretion(GO:0051458) |
| 0.0 | 0.2 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.3 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.0 | 0.1 | GO:0048859 | formation of anatomical boundary(GO:0048859) |
| 0.0 | 0.2 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.0 | 0.7 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.0 | 0.5 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 0.0 | 0.2 | GO:1902570 | protein localization to nucleolus(GO:1902570) |
| 0.0 | 0.5 | GO:0006825 | copper ion transport(GO:0006825) |
| 0.0 | 0.3 | GO:1901748 | leukotriene D4 metabolic process(GO:1901748) leukotriene D4 biosynthetic process(GO:1901750) |
| 0.0 | 0.2 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.0 | 0.1 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 0.0 | 0.2 | GO:1904778 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.0 | 0.1 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 0.0 | 0.1 | GO:0032099 | negative regulation of response to food(GO:0032096) negative regulation of appetite(GO:0032099) |
| 0.0 | 0.1 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.0 | 0.0 | GO:0016185 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) |
| 0.0 | 0.2 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) |
| 0.0 | 0.3 | GO:0033234 | negative regulation of protein sumoylation(GO:0033234) |
| 0.0 | 0.4 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.0 | 0.1 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.0 | 0.1 | GO:0015811 | L-cystine transport(GO:0015811) |
| 0.0 | 0.3 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.0 | 0.4 | GO:0097503 | sialylation(GO:0097503) |
| 0.0 | 0.2 | GO:0032486 | Rap protein signal transduction(GO:0032486) |
| 0.0 | 0.0 | GO:0045897 | positive regulation of transcription during mitosis(GO:0045897) |
| 0.0 | 0.2 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.0 | 0.1 | GO:1903322 | positive regulation of protein ubiquitination(GO:0031398) positive regulation of protein modification by small protein conjugation or removal(GO:1903322) |
| 0.0 | 0.1 | GO:0018076 | N-terminal peptidyl-lysine acetylation(GO:0018076) |
| 0.0 | 0.0 | GO:2001151 | regulation of renal water transport(GO:2001151) positive regulation of renal water transport(GO:2001153) |
| 0.0 | 0.1 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.0 | 0.7 | GO:0090502 | RNA phosphodiester bond hydrolysis, endonucleolytic(GO:0090502) |
| 0.0 | 0.0 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.0 | 0.3 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) |
| 0.0 | 0.5 | GO:0019985 | translesion synthesis(GO:0019985) |