Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
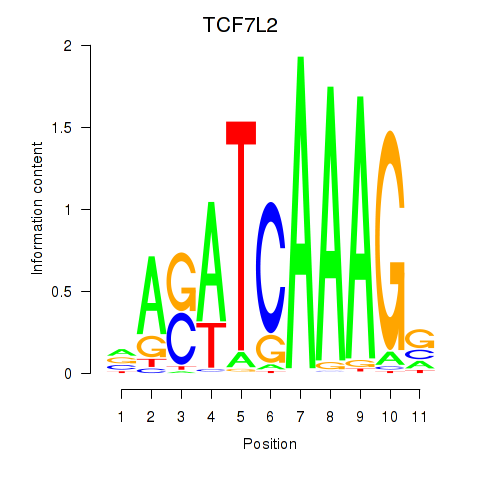
Results for TCF7L2
Z-value: 1.45
Transcription factors associated with TCF7L2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TCF7L2
|
ENSG00000148737.11 | TCF7L2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TCF7L2 | hg19_v2_chr10_+_114709999_114710031 | 0.87 | 1.2e-05 | Click! |
Activity profile of TCF7L2 motif
Sorted Z-values of TCF7L2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TCF7L2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_16936340 | 7.93 |
ENST00000507288.1 ENST00000513610.1 |
MYO10 |
myosin X |
| chr11_+_69455855 | 6.54 |
ENST00000227507.2 ENST00000536559.1 |
CCND1 |
cyclin D1 |
| chr11_+_114128522 | 5.98 |
ENST00000535401.1 |
NNMT |
nicotinamide N-methyltransferase |
| chr13_+_76334795 | 5.46 |
ENST00000526202.1 ENST00000465261.2 |
LMO7 |
LIM domain 7 |
| chr13_+_76334567 | 5.34 |
ENST00000321797.8 |
LMO7 |
LIM domain 7 |
| chr10_+_54074033 | 4.76 |
ENST00000373970.3 |
DKK1 |
dickkopf WNT signaling pathway inhibitor 1 |
| chr4_-_157892498 | 4.58 |
ENST00000502773.1 |
PDGFC |
platelet derived growth factor C |
| chr3_-_134093275 | 4.20 |
ENST00000513145.1 ENST00000422605.2 |
AMOTL2 |
angiomotin like 2 |
| chr3_-_134093395 | 3.85 |
ENST00000249883.5 |
AMOTL2 |
angiomotin like 2 |
| chr1_-_153522562 | 3.81 |
ENST00000368714.1 |
S100A4 |
S100 calcium binding protein A4 |
| chr13_+_102142296 | 3.75 |
ENST00000376162.3 |
ITGBL1 |
integrin, beta-like 1 (with EGF-like repeat domains) |
| chr8_-_119964434 | 3.49 |
ENST00000297350.4 |
TNFRSF11B |
tumor necrosis factor receptor superfamily, member 11b |
| chr1_-_110933663 | 3.09 |
ENST00000369781.4 ENST00000541986.1 ENST00000369779.4 |
SLC16A4 |
solute carrier family 16, member 4 |
| chrX_-_10588459 | 2.99 |
ENST00000380782.2 |
MID1 |
midline 1 (Opitz/BBB syndrome) |
| chr1_-_110933611 | 2.97 |
ENST00000472422.2 ENST00000437429.2 |
SLC16A4 |
solute carrier family 16, member 4 |
| chr5_+_102201509 | 2.82 |
ENST00000348126.2 ENST00000379787.4 |
PAM |
peptidylglycine alpha-amidating monooxygenase |
| chrX_-_10588595 | 2.81 |
ENST00000423614.1 ENST00000317552.4 |
MID1 |
midline 1 (Opitz/BBB syndrome) |
| chr17_+_1674982 | 2.81 |
ENST00000572048.1 ENST00000573763.1 |
SERPINF1 |
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr5_-_16742330 | 2.29 |
ENST00000505695.1 ENST00000427430.2 |
MYO10 |
myosin X |
| chr10_+_7745232 | 2.26 |
ENST00000358415.4 |
ITIH2 |
inter-alpha-trypsin inhibitor heavy chain 2 |
| chrX_+_54835493 | 2.22 |
ENST00000396224.1 |
MAGED2 |
melanoma antigen family D, 2 |
| chr10_+_114709999 | 2.14 |
ENST00000355995.4 ENST00000545257.1 ENST00000543371.1 ENST00000536810.1 ENST00000355717.4 ENST00000538897.1 ENST00000534894.1 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr3_-_149375783 | 1.99 |
ENST00000467467.1 ENST00000460517.1 ENST00000360632.3 |
WWTR1 |
WW domain containing transcription regulator 1 |
| chr10_+_7745303 | 1.93 |
ENST00000429820.1 ENST00000379587.4 |
ITIH2 |
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr5_+_102201430 | 1.91 |
ENST00000438793.3 ENST00000346918.2 |
PAM |
peptidylglycine alpha-amidating monooxygenase |
| chr10_+_114710211 | 1.83 |
ENST00000349937.2 ENST00000369397.4 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr2_+_69240415 | 1.81 |
ENST00000409829.3 |
ANTXR1 |
anthrax toxin receptor 1 |
| chr2_+_69240302 | 1.75 |
ENST00000303714.4 |
ANTXR1 |
anthrax toxin receptor 1 |
| chr2_+_69240511 | 1.71 |
ENST00000409349.3 |
ANTXR1 |
anthrax toxin receptor 1 |
| chr5_-_111093167 | 1.60 |
ENST00000446294.2 ENST00000419114.2 |
NREP |
neuronal regeneration related protein |
| chr1_-_94079648 | 1.59 |
ENST00000370247.3 |
BCAR3 |
breast cancer anti-estrogen resistance 3 |
| chr7_-_27213893 | 1.59 |
ENST00000283921.4 |
HOXA10 |
homeobox A10 |
| chr6_-_112575687 | 1.56 |
ENST00000521398.1 ENST00000424408.2 ENST00000243219.3 |
LAMA4 |
laminin, alpha 4 |
| chr2_+_228678550 | 1.51 |
ENST00000409189.3 ENST00000358813.4 |
CCL20 |
chemokine (C-C motif) ligand 20 |
| chr6_-_112575912 | 1.49 |
ENST00000522006.1 ENST00000230538.7 ENST00000519932.1 |
LAMA4 |
laminin, alpha 4 |
| chr10_+_114710425 | 1.46 |
ENST00000352065.5 ENST00000369395.1 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr12_+_1738363 | 1.40 |
ENST00000397196.2 |
WNT5B |
wingless-type MMTV integration site family, member 5B |
| chr1_-_109935819 | 1.20 |
ENST00000538502.1 |
SORT1 |
sortilin 1 |
| chr5_+_131593364 | 1.20 |
ENST00000253754.3 ENST00000379018.3 |
PDLIM4 |
PDZ and LIM domain 4 |
| chr10_+_114710516 | 1.19 |
ENST00000542695.1 ENST00000346198.4 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr5_+_140797296 | 1.18 |
ENST00000398594.2 |
PCDHGB7 |
protocadherin gamma subfamily B, 7 |
| chr5_+_140734570 | 1.08 |
ENST00000571252.1 |
PCDHGA4 |
protocadherin gamma subfamily A, 4 |
| chr6_-_32145861 | 1.02 |
ENST00000336984.6 |
AGPAT1 |
1-acylglycerol-3-phosphate O-acyltransferase 1 |
| chr5_-_148930960 | 0.97 |
ENST00000261798.5 ENST00000377843.2 |
CSNK1A1 |
casein kinase 1, alpha 1 |
| chr2_-_88427568 | 0.97 |
ENST00000393750.3 ENST00000295834.3 |
FABP1 |
fatty acid binding protein 1, liver |
| chr16_+_72088376 | 0.94 |
ENST00000570083.1 ENST00000355906.5 ENST00000398131.2 ENST00000569639.1 ENST00000564499.1 ENST00000357763.4 ENST00000562526.1 ENST00000565574.1 ENST00000568417.2 ENST00000356967.5 |
HP HPR |
haptoglobin haptoglobin-related protein |
| chr1_-_161993616 | 0.87 |
ENST00000294794.3 |
OLFML2B |
olfactomedin-like 2B |
| chr4_+_78078304 | 0.87 |
ENST00000316355.5 ENST00000354403.5 ENST00000502280.1 |
CCNG2 |
cyclin G2 |
| chr6_-_112575838 | 0.86 |
ENST00000455073.1 |
LAMA4 |
laminin, alpha 4 |
| chr12_+_9067123 | 0.83 |
ENST00000543824.1 |
PHC1 |
polyhomeotic homolog 1 (Drosophila) |
| chr18_+_46065393 | 0.79 |
ENST00000256413.3 |
CTIF |
CBP80/20-dependent translation initiation factor |
| chr8_+_32579341 | 0.77 |
ENST00000519240.1 ENST00000539990.1 |
NRG1 |
neuregulin 1 |
| chr10_+_101542462 | 0.76 |
ENST00000370449.4 ENST00000370434.1 |
ABCC2 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 2 |
| chrX_+_46937745 | 0.76 |
ENST00000397180.1 ENST00000457380.1 ENST00000352078.4 |
RGN |
regucalcin |
| chr14_-_61124977 | 0.74 |
ENST00000554986.1 |
SIX1 |
SIX homeobox 1 |
| chr17_-_38256973 | 0.74 |
ENST00000246672.3 |
NR1D1 |
nuclear receptor subfamily 1, group D, member 1 |
| chr13_-_36429763 | 0.73 |
ENST00000379893.1 |
DCLK1 |
doublecortin-like kinase 1 |
| chr7_+_80275621 | 0.70 |
ENST00000426978.1 ENST00000432207.1 |
CD36 |
CD36 molecule (thrombospondin receptor) |
| chr10_+_71561630 | 0.70 |
ENST00000398974.3 ENST00000398971.3 ENST00000398968.3 ENST00000398966.3 ENST00000398964.3 ENST00000398969.3 ENST00000356340.3 ENST00000398972.3 ENST00000398973.3 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr17_-_7297519 | 0.69 |
ENST00000576362.1 ENST00000571078.1 |
TMEM256-PLSCR3 |
TMEM256-PLSCR3 readthrough (NMD candidate) |
| chr7_+_80275752 | 0.67 |
ENST00000419819.2 |
CD36 |
CD36 molecule (thrombospondin receptor) |
| chr9_-_23826298 | 0.64 |
ENST00000380117.1 |
ELAVL2 |
ELAV like neuron-specific RNA binding protein 2 |
| chr17_-_39093672 | 0.62 |
ENST00000209718.3 ENST00000436344.3 ENST00000485751.1 |
KRT23 |
keratin 23 (histone deacetylase inducible) |
| chr6_-_112575758 | 0.61 |
ENST00000431543.2 ENST00000453937.2 ENST00000368638.4 ENST00000389463.4 |
LAMA4 |
laminin, alpha 4 |
| chr4_-_139163491 | 0.60 |
ENST00000280612.5 |
SLC7A11 |
solute carrier family 7 (anionic amino acid transporter light chain, xc- system), member 11 |
| chr17_-_56065484 | 0.58 |
ENST00000581208.1 |
VEZF1 |
vascular endothelial zinc finger 1 |
| chr5_-_58335281 | 0.57 |
ENST00000358923.6 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr10_+_71561649 | 0.56 |
ENST00000398978.3 ENST00000354547.3 ENST00000357811.3 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr17_-_7297833 | 0.55 |
ENST00000571802.1 ENST00000576201.1 ENST00000573213.1 ENST00000324822.11 |
TMEM256-PLSCR3 |
TMEM256-PLSCR3 readthrough (NMD candidate) |
| chr7_-_27219849 | 0.54 |
ENST00000396344.4 |
HOXA10 |
homeobox A10 |
| chr7_+_99613212 | 0.52 |
ENST00000426572.1 ENST00000535170.1 |
ZKSCAN1 |
zinc finger with KRAB and SCAN domains 1 |
| chrX_+_152760397 | 0.50 |
ENST00000331595.4 ENST00000431891.1 |
BGN |
biglycan |
| chr18_+_46065483 | 0.50 |
ENST00000382998.4 |
CTIF |
CBP80/20-dependent translation initiation factor |
| chr7_+_99613195 | 0.46 |
ENST00000324306.6 |
ZKSCAN1 |
zinc finger with KRAB and SCAN domains 1 |
| chr7_+_80275953 | 0.45 |
ENST00000538969.1 ENST00000544133.1 ENST00000433696.2 |
CD36 |
CD36 molecule (thrombospondin receptor) |
| chr19_-_40791302 | 0.44 |
ENST00000392038.2 ENST00000578123.1 |
AKT2 |
v-akt murine thymoma viral oncogene homolog 2 |
| chr7_-_148725733 | 0.43 |
ENST00000286091.4 |
PDIA4 |
protein disulfide isomerase family A, member 4 |
| chr14_-_65409502 | 0.43 |
ENST00000389614.5 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr4_+_30721968 | 0.41 |
ENST00000361762.2 |
PCDH7 |
protocadherin 7 |
| chr5_+_66124590 | 0.41 |
ENST00000490016.2 ENST00000403666.1 ENST00000450827.1 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr14_-_75179774 | 0.39 |
ENST00000555249.1 ENST00000556202.1 ENST00000356357.4 ENST00000338772.5 |
AREL1 AC007956.1 |
apoptosis resistant E3 ubiquitin protein ligase 1 Full-length cDNA 5-PRIME end of clone CS0CAP004YO05 of Thymus of Homo sapiens (human); Uncharacterized protein |
| chr13_-_67802549 | 0.39 |
ENST00000328454.5 ENST00000377865.2 |
PCDH9 |
protocadherin 9 |
| chr6_-_25930819 | 0.37 |
ENST00000360488.3 |
SLC17A2 |
solute carrier family 17, member 2 |
| chr12_+_122326630 | 0.34 |
ENST00000541212.1 ENST00000340175.5 |
PSMD9 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 9 |
| chr17_+_59529743 | 0.34 |
ENST00000589003.1 ENST00000393853.4 |
TBX4 |
T-box 4 |
| chr7_+_73868439 | 0.34 |
ENST00000424337.2 |
GTF2IRD1 |
GTF2I repeat domain containing 1 |
| chr13_-_72440901 | 0.33 |
ENST00000359684.2 |
DACH1 |
dachshund homolog 1 (Drosophila) |
| chr12_-_49582978 | 0.32 |
ENST00000301071.7 |
TUBA1A |
tubulin, alpha 1a |
| chr12_+_122326662 | 0.32 |
ENST00000261817.2 ENST00000538613.1 ENST00000542602.1 |
PSMD9 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 9 |
| chr16_-_73082274 | 0.31 |
ENST00000268489.5 |
ZFHX3 |
zinc finger homeobox 3 |
| chr19_-_40791211 | 0.31 |
ENST00000579047.1 |
AKT2 |
v-akt murine thymoma viral oncogene homolog 2 |
| chrX_+_47004599 | 0.31 |
ENST00000329236.7 |
RBM10 |
RNA binding motif protein 10 |
| chr2_-_55646957 | 0.31 |
ENST00000263630.8 |
CCDC88A |
coiled-coil domain containing 88A |
| chr9_-_27005686 | 0.29 |
ENST00000380055.5 |
LRRC19 |
leucine rich repeat containing 19 |
| chr6_-_25930904 | 0.29 |
ENST00000377850.3 |
SLC17A2 |
solute carrier family 17, member 2 |
| chr14_+_21236586 | 0.29 |
ENST00000326783.3 |
EDDM3B |
epididymal protein 3B |
| chr14_+_74815116 | 0.27 |
ENST00000256362.4 |
VRTN |
vertebrae development associated |
| chr11_-_76155618 | 0.25 |
ENST00000530759.1 |
RP11-111M22.3 |
RP11-111M22.3 |
| chrX_-_43832711 | 0.24 |
ENST00000378062.5 |
NDP |
Norrie disease (pseudoglioma) |
| chr5_+_170846640 | 0.22 |
ENST00000274625.5 |
FGF18 |
fibroblast growth factor 18 |
| chr2_-_55647057 | 0.21 |
ENST00000436346.1 |
CCDC88A |
coiled-coil domain containing 88A |
| chr5_-_157002775 | 0.21 |
ENST00000257527.4 |
ADAM19 |
ADAM metallopeptidase domain 19 |
| chr15_+_57511609 | 0.21 |
ENST00000543579.1 ENST00000537840.1 ENST00000343827.3 |
TCF12 |
transcription factor 12 |
| chr11_-_76155700 | 0.19 |
ENST00000572035.1 |
RP11-111M22.3 |
RP11-111M22.3 |
| chr7_+_110731062 | 0.19 |
ENST00000308478.5 ENST00000451085.1 ENST00000422987.3 ENST00000421101.1 |
LRRN3 |
leucine rich repeat neuronal 3 |
| chr11_+_65408273 | 0.18 |
ENST00000394227.3 |
SIPA1 |
signal-induced proliferation-associated 1 |
| chr2_+_71357434 | 0.18 |
ENST00000244230.2 |
MPHOSPH10 |
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr12_+_52056548 | 0.17 |
ENST00000545061.1 ENST00000355133.3 |
SCN8A |
sodium channel, voltage gated, type VIII, alpha subunit |
| chr22_+_19950060 | 0.16 |
ENST00000449653.1 |
COMT |
catechol-O-methyltransferase |
| chr6_-_11779403 | 0.16 |
ENST00000414691.3 |
ADTRP |
androgen-dependent TFPI-regulating protein |
| chr10_+_71561704 | 0.15 |
ENST00000520267.1 |
COL13A1 |
collagen, type XIII, alpha 1 |
| chr12_+_70760056 | 0.14 |
ENST00000258111.4 |
KCNMB4 |
potassium large conductance calcium-activated channel, subfamily M, beta member 4 |
| chr3_+_155838337 | 0.14 |
ENST00000490337.1 ENST00000389636.5 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr14_-_53258180 | 0.10 |
ENST00000554230.1 |
GNPNAT1 |
glucosamine-phosphate N-acetyltransferase 1 |
| chr1_+_16330723 | 0.09 |
ENST00000329454.2 |
C1orf64 |
chromosome 1 open reading frame 64 |
| chrX_-_122866874 | 0.09 |
ENST00000245838.8 ENST00000355725.4 |
THOC2 |
THO complex 2 |
| chrX_+_47004639 | 0.08 |
ENST00000345781.6 |
RBM10 |
RNA binding motif protein 10 |
| chr17_+_42923686 | 0.07 |
ENST00000591513.1 |
HIGD1B |
HIG1 hypoxia inducible domain family, member 1B |
| chr7_-_75452673 | 0.07 |
ENST00000416943.1 |
CCL24 |
chemokine (C-C motif) ligand 24 |
| chr2_+_71357744 | 0.07 |
ENST00000498451.2 |
MPHOSPH10 |
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr12_-_23737534 | 0.07 |
ENST00000396007.2 |
SOX5 |
SRY (sex determining region Y)-box 5 |
| chr6_+_64281906 | 0.06 |
ENST00000370651.3 |
PTP4A1 |
protein tyrosine phosphatase type IVA, member 1 |
| chr5_+_175288631 | 0.06 |
ENST00000509837.1 |
CPLX2 |
complexin 2 |
| chr4_-_69215467 | 0.05 |
ENST00000579690.1 |
YTHDC1 |
YTH domain containing 1 |
| chr22_+_41347363 | 0.05 |
ENST00000216225.8 |
RBX1 |
ring-box 1, E3 ubiquitin protein ligase |
| chr2_+_204732666 | 0.04 |
ENST00000295854.6 ENST00000472206.1 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
| chr5_-_157002749 | 0.04 |
ENST00000517905.1 ENST00000430702.2 ENST00000394020.1 |
ADAM19 |
ADAM metallopeptidase domain 19 |
| chr5_-_135290705 | 0.04 |
ENST00000274507.1 |
LECT2 |
leukocyte cell-derived chemotaxin 2 |
| chr18_-_52989217 | 0.03 |
ENST00000570287.2 |
TCF4 |
transcription factor 4 |
| chr20_-_17511962 | 0.02 |
ENST00000377873.3 |
BFSP1 |
beaded filament structural protein 1, filensin |
| chr4_-_109684120 | 0.01 |
ENST00000512646.1 ENST00000411864.2 ENST00000296486.3 ENST00000510706.1 |
ETNPPL |
ethanolamine-phosphate phospho-lyase |
| chr3_+_183353356 | 0.00 |
ENST00000242810.6 ENST00000493074.1 ENST00000437402.1 ENST00000454495.2 ENST00000473045.1 ENST00000468101.1 ENST00000427201.2 ENST00000482138.1 ENST00000454652.2 |
KLHL24 |
kelch-like family member 24 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 6.5 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.2 | 5.3 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.2 | 10.2 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.2 | 4.8 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.1 | 4.5 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.1 | 2.1 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.4 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 5.9 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.7 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.0 | 11.9 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 0.7 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 0.9 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.0 | 5.8 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.5 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 1.2 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 0.5 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 5.3 | GO:1901202 | negative regulation of extracellular matrix assembly(GO:1901202) |
| 1.6 | 4.8 | GO:0009996 | negative regulation of Wnt signaling pathway involved in heart development(GO:0003308) negative regulation of cell fate specification(GO:0009996) Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 1.2 | 4.7 | GO:0018032 | peptide amidation(GO:0001519) protein amidation(GO:0018032) peptide modification(GO:0031179) |
| 1.1 | 6.6 | GO:0010908 | regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010908) positive regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010909) canonical Wnt signaling pathway involved in positive regulation of epithelial to mesenchymal transition(GO:0044334) positive regulation of proteoglycan biosynthetic process(GO:1902730) |
| 0.7 | 6.5 | GO:0070141 | response to UV-A(GO:0070141) |
| 0.6 | 2.3 | GO:0042483 | negative regulation of odontogenesis(GO:0042483) |
| 0.6 | 2.8 | GO:0071279 | cellular response to cobalt ion(GO:0071279) |
| 0.5 | 1.5 | GO:0045362 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 0.5 | 6.0 | GO:0034356 | NAD biosynthesis via nicotinamide riboside salvage pathway(GO:0034356) |
| 0.4 | 5.8 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.4 | 0.8 | GO:1904833 | positive regulation of superoxide dismutase activity(GO:1901671) positive regulation of removal of superoxide radicals(GO:1904833) |
| 0.3 | 2.2 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.3 | 0.9 | GO:2000296 | negative regulation of hydrogen peroxide catabolic process(GO:2000296) |
| 0.3 | 1.8 | GO:2000334 | lipoprotein particle mediated signaling(GO:0055095) low-density lipoprotein particle mediated signaling(GO:0055096) blood microparticle formation(GO:0072564) regulation of blood microparticle formation(GO:2000332) positive regulation of blood microparticle formation(GO:2000334) |
| 0.3 | 0.8 | GO:0050787 | antibiotic metabolic process(GO:0016999) detoxification of mercury ion(GO:0050787) response to antineoplastic agent(GO:0097327) |
| 0.2 | 0.7 | GO:0061055 | myotome development(GO:0061055) positive regulation of mesenchymal cell proliferation involved in ureter development(GO:2000729) |
| 0.2 | 0.7 | GO:0060086 | circadian temperature homeostasis(GO:0060086) |
| 0.2 | 5.4 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 0.2 | 10.0 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.2 | 0.7 | GO:0010746 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) |
| 0.1 | 10.2 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.1 | 1.0 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 0.1 | 0.7 | GO:0070682 | proteasome regulatory particle assembly(GO:0070682) |
| 0.1 | 0.5 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.1 | 1.2 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.1 | 1.0 | GO:1904424 | regulation of GTP binding(GO:1904424) |
| 0.1 | 1.4 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.1 | 0.3 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.1 | 2.1 | GO:0009954 | proximal/distal pattern formation(GO:0009954) |
| 0.1 | 1.6 | GO:0002089 | lens morphogenesis in camera-type eye(GO:0002089) |
| 0.1 | 1.0 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) |
| 0.1 | 0.3 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 0.1 | 3.8 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.1 | 0.6 | GO:0086024 | adrenergic receptor signaling pathway involved in positive regulation of heart rate(GO:0086024) |
| 0.1 | 0.2 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) |
| 0.0 | 0.2 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) positive regulation of cell chemotaxis to fibroblast growth factor(GO:1904849) positive regulation of endothelial cell chemotaxis to fibroblast growth factor(GO:2000546) |
| 0.0 | 0.2 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.0 | 0.2 | GO:0016036 | cellular response to phosphate starvation(GO:0016036) positive regulation of sulfur amino acid metabolic process(GO:0031337) positive regulation of homocysteine metabolic process(GO:0050668) |
| 0.0 | 0.6 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 0.0 | 5.8 | GO:0015718 | monocarboxylic acid transport(GO:0015718) |
| 0.0 | 1.4 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 0.8 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.0 | 10.0 | GO:0000209 | protein polyubiquitination(GO:0000209) |
| 0.0 | 0.7 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.0 | 0.5 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 3.0 | GO:0045995 | regulation of embryonic development(GO:0045995) |
| 0.0 | 0.1 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.0 | 3.8 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.0 | 0.3 | GO:0032196 | transposition(GO:0032196) |
| 0.0 | 1.3 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.0 | 0.1 | GO:2000418 | positive regulation of eosinophil migration(GO:2000418) |
| 0.0 | 0.6 | GO:0001885 | endothelial cell development(GO:0001885) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 6.0 | GO:0008112 | nicotinamide N-methyltransferase activity(GO:0008112) pyridine N-methyltransferase activity(GO:0030760) |
| 1.2 | 4.7 | GO:0004598 | peptidylglycine monooxygenase activity(GO:0004504) peptidylamidoglycolate lyase activity(GO:0004598) |
| 1.1 | 10.2 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.5 | 1.5 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 0.4 | 4.8 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.3 | 1.8 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 0.3 | 5.2 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.2 | 1.2 | GO:0010465 | nerve growth factor receptor activity(GO:0010465) |
| 0.2 | 4.6 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.2 | 6.5 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.2 | 3.8 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.2 | 0.9 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.1 | 1.0 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.1 | 0.6 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) |
| 0.1 | 3.5 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 6.1 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.1 | 0.8 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.1 | 0.8 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.1 | 0.3 | GO:0004803 | transposase activity(GO:0004803) |
| 0.1 | 7.0 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 5.3 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 0.2 | GO:0005105 | type 1 fibroblast growth factor receptor binding(GO:0005105) |
| 0.1 | 0.7 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.1 | 5.0 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.7 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.0 | 1.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.6 | GO:0031698 | beta-2 adrenergic receptor binding(GO:0031698) |
| 0.0 | 4.3 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.0 | 0.7 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.0 | 0.5 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.0 | 0.2 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.0 | 0.4 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 8.4 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 2.1 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 0.1 | GO:0031728 | CCR3 chemokine receptor binding(GO:0031728) |
| 0.0 | 0.7 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.0 | 1.6 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 0.2 | GO:0035497 | cAMP response element binding(GO:0035497) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 6.6 | GO:0070369 | beta-catenin-TCF7L2 complex(GO:0070369) |
| 0.5 | 10.2 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.4 | 1.4 | GO:0030936 | collagen type XIII trimer(GO:0005600) transmembrane collagen trimer(GO:0030936) |
| 0.2 | 1.0 | GO:0045179 | apical cortex(GO:0045179) |
| 0.2 | 0.9 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.2 | 2.8 | GO:0043203 | axon hillock(GO:0043203) |
| 0.2 | 4.2 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.1 | 0.8 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 4.5 | GO:0005605 | basal lamina(GO:0005605) |
| 0.1 | 1.0 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.1 | 13.4 | GO:0005923 | bicellular tight junction(GO:0005923) |
| 0.1 | 2.2 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.1 | 4.7 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 0.8 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.0 | 0.2 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.0 | 10.5 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.0 | 0.7 | GO:0008540 | proteasome regulatory particle, base subcomplex(GO:0008540) |
| 0.0 | 1.2 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 4.9 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 10.4 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.8 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 2.2 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.0 | 3.6 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.2 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 0.1 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.0 | 3.6 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 0.6 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 10.0 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.3 | 6.5 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.2 | 4.6 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.2 | 10.2 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 0.7 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.0 | 5.6 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.0 | 0.8 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.7 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 2.8 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 0.8 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 1.5 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 1.4 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.5 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.0 | 0.5 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 1.2 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.6 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 0.6 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |