Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
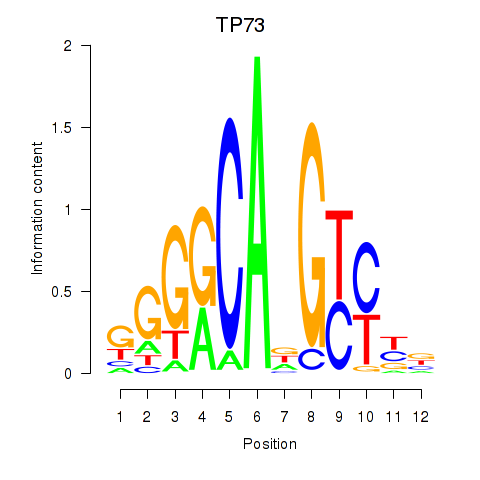
Results for TP73
Z-value: 0.63
Transcription factors associated with TP73
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TP73
|
ENSG00000078900.10 | TP73 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TP73 | hg19_v2_chr1_+_3614591_3614622 | -0.29 | 2.7e-01 | Click! |
Activity profile of TP73 motif
Sorted Z-values of TP73 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TP73
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_230135937 | 1.57 |
ENST00000392054.3 ENST00000409462.1 ENST00000392055.3 |
PID1 |
phosphotyrosine interaction domain containing 1 |
| chr5_+_125758865 | 1.48 |
ENST00000542322.1 ENST00000544396.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr10_+_47746929 | 1.47 |
ENST00000340243.6 ENST00000374277.5 ENST00000449464.2 ENST00000538825.1 ENST00000335083.5 |
ANXA8L2 AL603965.1 |
annexin A8-like 2 Protein LOC100996760 |
| chr5_+_125758813 | 1.47 |
ENST00000285689.3 ENST00000515200.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr10_-_47173994 | 1.37 |
ENST00000414655.2 ENST00000545298.1 ENST00000359178.4 ENST00000358140.4 ENST00000503031.1 |
ANXA8L1 LINC00842 |
annexin A8-like 1 long intergenic non-protein coding RNA 842 |
| chr1_-_68698222 | 1.13 |
ENST00000370976.3 ENST00000354777.2 ENST00000262348.4 ENST00000540432.1 |
WLS |
wntless Wnt ligand secretion mediator |
| chr17_+_73750699 | 1.04 |
ENST00000584939.1 |
ITGB4 |
integrin, beta 4 |
| chr8_+_54764346 | 0.98 |
ENST00000297313.3 ENST00000344277.6 |
RGS20 |
regulator of G-protein signaling 20 |
| chr8_-_145016692 | 0.97 |
ENST00000357649.2 |
PLEC |
plectin |
| chr12_-_91573249 | 0.96 |
ENST00000550099.1 ENST00000546391.1 ENST00000551354.1 |
DCN |
decorin |
| chr3_-_185538849 | 0.90 |
ENST00000421047.2 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr11_+_65686952 | 0.85 |
ENST00000527119.1 |
DRAP1 |
DR1-associated protein 1 (negative cofactor 2 alpha) |
| chr14_+_75745477 | 0.80 |
ENST00000303562.4 ENST00000554617.1 ENST00000554212.1 ENST00000535987.1 ENST00000555242.1 |
FOS |
FBJ murine osteosarcoma viral oncogene homolog |
| chr5_+_119799927 | 0.72 |
ENST00000407149.2 ENST00000379551.2 |
PRR16 |
proline rich 16 |
| chr20_+_48807351 | 0.62 |
ENST00000303004.3 |
CEBPB |
CCAAT/enhancer binding protein (C/EBP), beta |
| chr14_+_75746340 | 0.57 |
ENST00000555686.1 ENST00000555672.1 |
FOS |
FBJ murine osteosarcoma viral oncogene homolog |
| chr11_+_65687158 | 0.57 |
ENST00000532933.1 |
DRAP1 |
DR1-associated protein 1 (negative cofactor 2 alpha) |
| chr6_-_46703069 | 0.56 |
ENST00000538237.1 ENST00000274793.7 |
PLA2G7 |
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
| chr1_-_68698197 | 0.43 |
ENST00000370973.2 ENST00000370971.1 |
WLS |
wntless Wnt ligand secretion mediator |
| chr8_+_37654693 | 0.43 |
ENST00000412232.2 |
GPR124 |
G protein-coupled receptor 124 |
| chr10_-_79686284 | 0.41 |
ENST00000372391.2 ENST00000372388.2 |
DLG5 |
discs, large homolog 5 (Drosophila) |
| chr19_+_45281118 | 0.40 |
ENST00000270279.3 ENST00000341505.4 |
CBLC |
Cbl proto-oncogene C, E3 ubiquitin protein ligase |
| chr15_+_41136216 | 0.38 |
ENST00000562057.1 ENST00000344051.4 |
SPINT1 |
serine peptidase inhibitor, Kunitz type 1 |
| chr15_+_41136586 | 0.36 |
ENST00000431806.1 |
SPINT1 |
serine peptidase inhibitor, Kunitz type 1 |
| chr6_-_46703430 | 0.35 |
ENST00000537365.1 |
PLA2G7 |
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
| chr15_+_74466744 | 0.34 |
ENST00000560862.1 ENST00000395118.1 |
ISLR |
immunoglobulin superfamily containing leucine-rich repeat |
| chr11_-_65686496 | 0.32 |
ENST00000449692.3 |
C11orf68 |
chromosome 11 open reading frame 68 |
| chr7_-_2883928 | 0.30 |
ENST00000275364.3 |
GNA12 |
guanine nucleotide binding protein (G protein) alpha 12 |
| chr1_-_203055129 | 0.24 |
ENST00000241651.4 |
MYOG |
myogenin (myogenic factor 4) |
| chr6_-_122792919 | 0.24 |
ENST00000339697.4 |
SERINC1 |
serine incorporator 1 |
| chr17_+_21191341 | 0.21 |
ENST00000526076.2 ENST00000361818.5 ENST00000316920.6 |
MAP2K3 |
mitogen-activated protein kinase kinase 3 |
| chr14_-_25519317 | 0.19 |
ENST00000323944.5 |
STXBP6 |
syntaxin binding protein 6 (amisyn) |
| chr1_-_214724566 | 0.18 |
ENST00000366956.5 |
PTPN14 |
protein tyrosine phosphatase, non-receptor type 14 |
| chr8_+_37654424 | 0.18 |
ENST00000315215.7 |
GPR124 |
G protein-coupled receptor 124 |
| chr16_+_8891670 | 0.18 |
ENST00000268261.4 ENST00000539622.1 ENST00000569958.1 ENST00000537352.1 |
PMM2 |
phosphomannomutase 2 |
| chr8_+_120885949 | 0.18 |
ENST00000523492.1 ENST00000286234.5 |
DEPTOR |
DEP domain containing MTOR-interacting protein |
| chr17_-_79818354 | 0.17 |
ENST00000576541.1 ENST00000576380.1 ENST00000571617.1 ENST00000576052.1 ENST00000576390.1 ENST00000573778.2 ENST00000439918.2 ENST00000574914.1 ENST00000331483.4 |
P4HB |
prolyl 4-hydroxylase, beta polypeptide |
| chr2_+_220283091 | 0.17 |
ENST00000373960.3 |
DES |
desmin |
| chr14_-_25519095 | 0.17 |
ENST00000419632.2 ENST00000358326.2 ENST00000396700.1 ENST00000548724.1 |
STXBP6 |
syntaxin binding protein 6 (amisyn) |
| chr1_-_52344471 | 0.16 |
ENST00000352171.7 ENST00000354831.7 |
NRD1 |
nardilysin (N-arginine dibasic convertase) |
| chr9_-_128003606 | 0.15 |
ENST00000324460.6 |
HSPA5 |
heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) |
| chr12_-_57505121 | 0.15 |
ENST00000538913.2 ENST00000537215.2 ENST00000454075.3 ENST00000554825.1 ENST00000553275.1 ENST00000300134.3 |
STAT6 |
signal transducer and activator of transcription 6, interleukin-4 induced |
| chr22_+_39966758 | 0.14 |
ENST00000407673.1 ENST00000401624.1 ENST00000404898.1 ENST00000402142.3 ENST00000336649.4 ENST00000400164.3 |
CACNA1I |
calcium channel, voltage-dependent, T type, alpha 1I subunit |
| chr16_+_84178874 | 0.11 |
ENST00000378553.5 |
DNAAF1 |
dynein, axonemal, assembly factor 1 |
| chr18_-_28681950 | 0.11 |
ENST00000251081.6 |
DSC2 |
desmocollin 2 |
| chr19_+_17326521 | 0.10 |
ENST00000593597.1 |
USE1 |
unconventional SNARE in the ER 1 homolog (S. cerevisiae) |
| chr6_-_72130472 | 0.10 |
ENST00000426635.2 |
LINC00472 |
long intergenic non-protein coding RNA 472 |
| chr15_+_64680003 | 0.10 |
ENST00000261884.3 |
TRIP4 |
thyroid hormone receptor interactor 4 |
| chr14_-_105531759 | 0.09 |
ENST00000329797.3 ENST00000539291.2 ENST00000392585.2 |
GPR132 |
G protein-coupled receptor 132 |
| chr3_+_48481658 | 0.08 |
ENST00000438607.2 |
TMA7 |
translation machinery associated 7 homolog (S. cerevisiae) |
| chr10_+_70661014 | 0.08 |
ENST00000373585.3 |
DDX50 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 50 |
| chr3_-_100120223 | 0.08 |
ENST00000284320.5 |
TOMM70A |
translocase of outer mitochondrial membrane 70 homolog A (S. cerevisiae) |
| chr16_-_2581409 | 0.06 |
ENST00000567119.1 ENST00000565480.1 ENST00000382350.1 |
CEMP1 |
cementum protein 1 |
| chr11_+_60197040 | 0.06 |
ENST00000300190.2 |
MS4A5 |
membrane-spanning 4-domains, subfamily A, member 5 |
| chr11_+_60197069 | 0.05 |
ENST00000528905.1 ENST00000528093.1 |
MS4A5 |
membrane-spanning 4-domains, subfamily A, member 5 |
| chr19_+_45254529 | 0.05 |
ENST00000444487.1 |
BCL3 |
B-cell CLL/lymphoma 3 |
| chr20_-_52790055 | 0.04 |
ENST00000395955.3 |
CYP24A1 |
cytochrome P450, family 24, subfamily A, polypeptide 1 |
| chr16_+_5008290 | 0.04 |
ENST00000251170.7 |
SEC14L5 |
SEC14-like 5 (S. cerevisiae) |
| chr10_-_105212059 | 0.03 |
ENST00000260743.5 |
CALHM2 |
calcium homeostasis modulator 2 |
| chr3_-_48936272 | 0.03 |
ENST00000544097.1 ENST00000430379.1 ENST00000319017.4 |
SLC25A20 |
solute carrier family 25 (carnitine/acylcarnitine translocase), member 20 |
| chr17_+_4835580 | 0.03 |
ENST00000329125.5 |
GP1BA |
glycoprotein Ib (platelet), alpha polypeptide |
| chr11_+_65154070 | 0.03 |
ENST00000317568.5 ENST00000531296.1 ENST00000533782.1 ENST00000355991.5 ENST00000416776.2 ENST00000526201.1 |
FRMD8 |
FERM domain containing 8 |
| chr19_+_41620335 | 0.03 |
ENST00000331105.2 |
CYP2F1 |
cytochrome P450, family 2, subfamily F, polypeptide 1 |
| chr19_+_19627026 | 0.02 |
ENST00000608404.1 ENST00000555938.1 ENST00000503283.1 ENST00000512771.3 ENST00000428459.2 |
YJEFN3 CTC-260F20.3 NDUFA13 |
YjeF N-terminal domain containing 3 Uncharacterized protein NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 13 |
| chr17_-_2614927 | 0.02 |
ENST00000435359.1 |
CLUH |
clustered mitochondria (cluA/CLU1) homolog |
| chr18_-_4455260 | 0.02 |
ENST00000581527.1 |
DLGAP1 |
discs, large (Drosophila) homolog-associated protein 1 |
| chr11_+_236959 | 0.02 |
ENST00000431206.2 ENST00000528906.1 |
PSMD13 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 13 |
| chr18_+_77623668 | 0.01 |
ENST00000316249.3 |
KCNG2 |
potassium voltage-gated channel, subfamily G, member 2 |
| chr9_+_36169380 | 0.01 |
ENST00000335119.2 |
CCIN |
calicin |
| chr7_+_116502527 | 0.01 |
ENST00000361183.3 |
CAPZA2 |
capping protein (actin filament) muscle Z-line, alpha 2 |
| chr16_+_67197288 | 0.01 |
ENST00000264009.8 ENST00000421453.1 |
HSF4 |
heat shock transcription factor 4 |
| chr12_+_44152740 | 0.00 |
ENST00000440781.2 ENST00000431837.1 ENST00000550616.1 ENST00000448290.2 ENST00000551736.1 |
IRAK4 |
interleukin-1 receptor-associated kinase 4 |
| chr11_+_237016 | 0.00 |
ENST00000352303.5 |
PSMD13 |
proteasome (prosome, macropain) 26S subunit, non-ATPase, 13 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.6 | GO:1901859 | negative regulation of mitochondrial DNA replication(GO:0090298) negative regulation of mitochondrial DNA metabolic process(GO:1901859) |
| 0.5 | 1.6 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.2 | 1.4 | GO:0001661 | conditioned taste aversion(GO:0001661) |
| 0.2 | 0.9 | GO:0034441 | plasma lipoprotein particle oxidation(GO:0034441) |
| 0.1 | 0.2 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) |
| 0.1 | 1.0 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.1 | 1.0 | GO:0035878 | nail development(GO:0035878) |
| 0.1 | 0.8 | GO:1990440 | positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.1 | 0.4 | GO:0071896 | negative regulation of hippo signaling(GO:0035331) protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.2 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 0.0 | 1.0 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.7 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.0 | 0.4 | GO:0035542 | regulation of SNARE complex assembly(GO:0035542) |
| 0.0 | 0.1 | GO:0002296 | T-helper 1 cell lineage commitment(GO:0002296) |
| 0.0 | 0.5 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.0 | 0.4 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.0 | 0.2 | GO:0035897 | proteolysis in other organism(GO:0035897) |
| 0.0 | 0.2 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.0 | 0.2 | GO:0015825 | L-serine transport(GO:0015825) |
| 0.0 | 0.1 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.0 | 0.1 | GO:0086073 | cardiac muscle cell-cardiac muscle cell adhesion(GO:0086042) bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 0.0 | GO:0045082 | positive regulation of interleukin-10 biosynthetic process(GO:0045082) |
| 0.0 | 0.2 | GO:0045792 | negative regulation of cell size(GO:0045792) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.4 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.1 | 2.0 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.1 | 1.0 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.1 | 0.2 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.1 | 1.6 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.0 | 0.9 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.0 | 0.4 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 0.2 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.0 | 0.0 | GO:0033257 | Bcl3/NF-kappaB2 complex(GO:0033257) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 1.0 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 0.3 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.0 | 0.9 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 1.0 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.0 | ST G ALPHA I PATHWAY | G alpha i Pathway |
| 0.0 | 0.6 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.0 | 0.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 1.4 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.9 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.0 | 1.0 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 1.0 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 1.0 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 0.2 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 0.2 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.1 | 0.9 | GO:0003847 | 1-alkyl-2-acetylglycerophosphocholine esterase activity(GO:0003847) |
| 0.1 | 1.0 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.0 | 0.1 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.0 | 0.2 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) procollagen-proline dioxygenase activity(GO:0019798) |
| 0.0 | 1.0 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.6 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.0 | 1.4 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 1.4 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 0.9 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 0.4 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.2 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.0 | 0.3 | GO:0050780 | dopamine receptor binding(GO:0050780) |
| 0.0 | 0.2 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.0 | 0.1 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 0.0 | GO:0015199 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |