Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
Results for UAAGACU
Z-value: 0.29
miRNA associated with seed UAAGACU
| Name | miRBASE accession |
|---|---|
|
hsa-miR-499a-5p
|
MIMAT0002870 |
Activity profile of UAAGACU motif
Sorted Z-values of UAAGACU motif
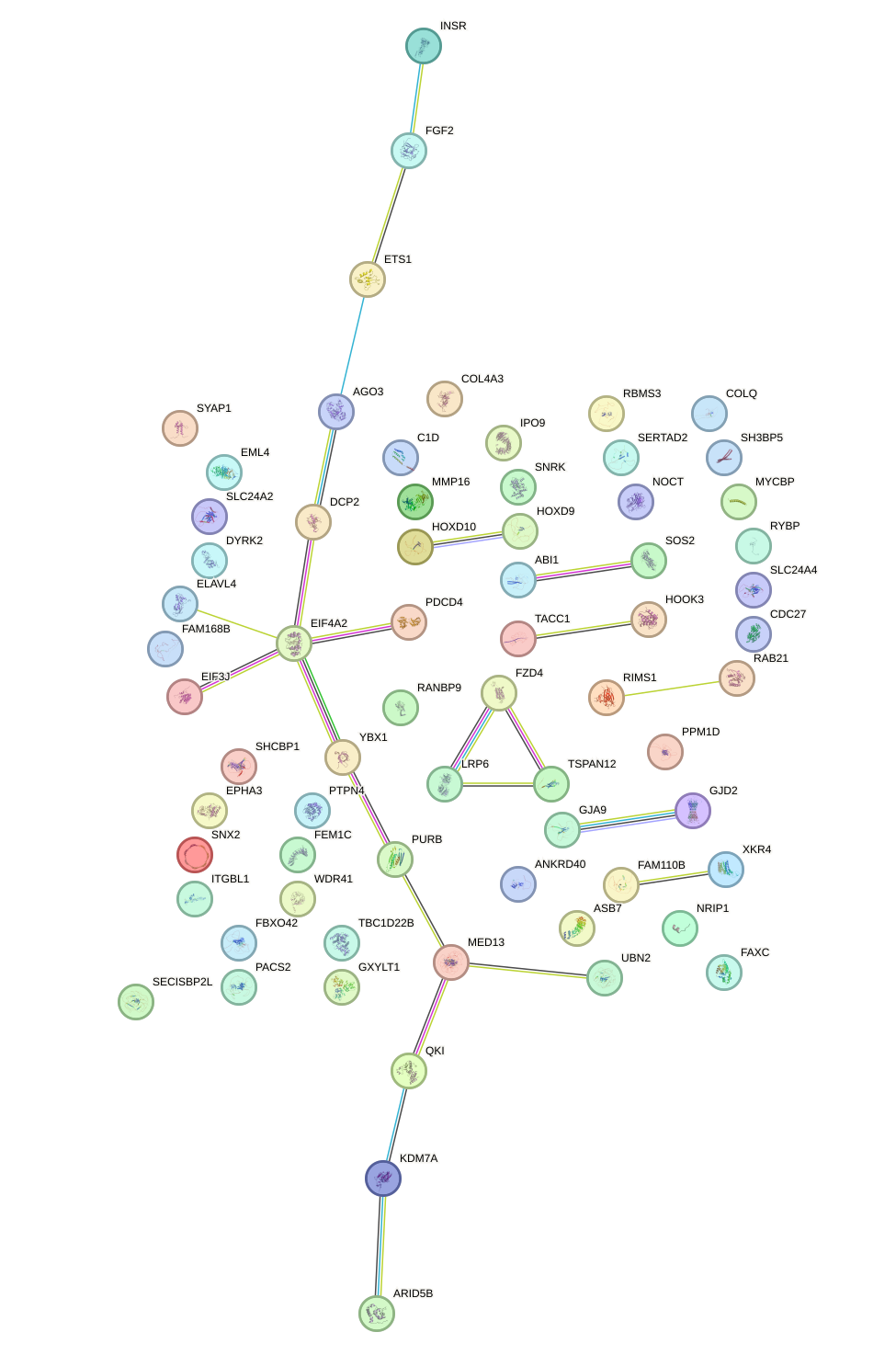
Network of associatons between targets according to the STRING database.
First level regulatory network of UAAGACU
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_38614807 | 0.46 |
ENST00000330691.6 ENST00000348567.4 |
TACC1 |
transforming, acidic coiled-coil containing protein 1 |
| chr19_-_7293942 | 0.26 |
ENST00000341500.5 ENST00000302850.5 |
INSR |
insulin receptor |
| chr1_-_154842741 | 0.21 |
ENST00000271915.4 |
KCNN3 |
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 |
| chr8_+_56014949 | 0.21 |
ENST00000327381.6 |
XKR4 |
XK, Kell blood group complex subunit-related family, member 4 |
| chr10_+_112631547 | 0.19 |
ENST00000280154.7 ENST00000393104.2 |
PDCD4 |
programmed cell death 4 (neoplastic transformation inhibitor) |
| chr7_-_120498357 | 0.18 |
ENST00000415871.1 ENST00000222747.3 ENST00000430985.1 |
TSPAN12 |
tetraspanin 12 |
| chr5_+_112312416 | 0.17 |
ENST00000389063.2 |
DCP2 |
decapping mRNA 2 |
| chr17_+_58677539 | 0.17 |
ENST00000305921.3 |
PPM1D |
protein phosphatase, Mg2+/Mn2+ dependent, 1D |
| chr2_-_64881018 | 0.17 |
ENST00000313349.3 |
SERTAD2 |
SERTA domain containing 2 |
| chr3_+_186501336 | 0.16 |
ENST00000323963.5 ENST00000440191.2 ENST00000356531.5 |
EIF4A2 |
eukaryotic translation initiation factor 4A2 |
| chr6_-_99797522 | 0.15 |
ENST00000389677.5 |
FAXC |
failed axon connections homolog (Drosophila) |
| chr16_-_46655538 | 0.14 |
ENST00000303383.3 |
SHCBP1 |
SHC SH2-domain binding protein 1 |
| chrY_+_15016725 | 0.14 |
ENST00000336079.3 |
DDX3Y |
DEAD (Asp-Glu-Ala-Asp) box helicase 3, Y-linked |
| chr3_-_72496035 | 0.11 |
ENST00000477973.2 |
RYBP |
RING1 and YY1 binding protein |
| chr7_-_139876812 | 0.11 |
ENST00000397560.2 |
JHDM1D |
lysine (K)-specific demethylase 7A |
| chr10_+_63661053 | 0.10 |
ENST00000279873.7 |
ARID5B |
AT rich interactive domain 5B (MRF1-like) |
| chr3_+_43328004 | 0.08 |
ENST00000454177.1 ENST00000429705.2 ENST00000296088.7 ENST00000437827.1 |
SNRK |
SNF related kinase |
| chr5_-_76788317 | 0.08 |
ENST00000296679.4 |
WDR41 |
WD repeat domain 41 |
| chr1_+_145611010 | 0.08 |
ENST00000369291.5 |
RNF115 |
ring finger protein 115 |
| chr1_-_16678914 | 0.08 |
ENST00000375592.3 |
FBXO42 |
F-box protein 42 |
| chr2_+_228029281 | 0.08 |
ENST00000396578.3 |
COL4A3 |
collagen, type IV, alpha 3 (Goodpasture antigen) |
| chr2_-_86564776 | 0.08 |
ENST00000165698.5 ENST00000541910.1 ENST00000535845.1 |
REEP1 |
receptor accessory protein 1 |
| chr8_+_58907104 | 0.08 |
ENST00000361488.3 |
FAM110B |
family with sequence similarity 110, member B |
| chr12_+_72148614 | 0.07 |
ENST00000261263.3 |
RAB21 |
RAB21, member RAS oncogene family |
| chr17_-_60142609 | 0.07 |
ENST00000397786.2 |
MED13 |
mediator complex subunit 13 |
| chr3_-_15374033 | 0.07 |
ENST00000253688.5 ENST00000383791.3 |
SH3BP5 |
SH3-domain binding protein 5 (BTK-associated) |
| chr2_+_42396472 | 0.07 |
ENST00000318522.5 ENST00000402711.2 |
EML4 |
echinoderm microtubule associated protein like 4 |
| chr15_-_49338748 | 0.06 |
ENST00000559471.1 |
SECISBP2L |
SECIS binding protein 2-like |
| chr10_-_27149792 | 0.06 |
ENST00000376140.3 ENST00000376170.4 |
ABI1 |
abl-interactor 1 |
| chr11_-_86666427 | 0.06 |
ENST00000531380.1 |
FZD4 |
frizzled family receptor 4 |
| chr12_+_68042495 | 0.06 |
ENST00000344096.3 |
DYRK2 |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 2 |
| chr7_+_90225796 | 0.06 |
ENST00000380050.3 |
CDK14 |
cyclin-dependent kinase 14 |
| chr12_-_12419703 | 0.06 |
ENST00000543091.1 ENST00000261349.4 |
LRP6 |
low density lipoprotein receptor-related protein 6 |
| chr1_+_97187318 | 0.06 |
ENST00000609116.1 ENST00000370198.1 ENST00000370197.1 ENST00000426398.2 ENST00000394184.3 |
PTBP2 |
polypyrimidine tract binding protein 2 |
| chr8_+_42752053 | 0.06 |
ENST00000307602.4 |
HOOK3 |
hook microtubule-tethering protein 3 |
| chr2_-_68290106 | 0.05 |
ENST00000407324.1 ENST00000355848.3 ENST00000409302.1 ENST00000410067.3 |
C1D |
C1D nuclear receptor corepressor |
| chr6_+_72596604 | 0.05 |
ENST00000348717.5 ENST00000517960.1 ENST00000518273.1 ENST00000522291.1 ENST00000521978.1 ENST00000520567.1 ENST00000264839.7 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr17_+_29718642 | 0.05 |
ENST00000325874.8 |
RAB11FIP4 |
RAB11 family interacting protein 4 (class II) |
| chr14_-_50698276 | 0.05 |
ENST00000216373.5 |
SOS2 |
son of sevenless homolog 2 (Drosophila) |
| chr8_+_11561660 | 0.05 |
ENST00000526716.1 ENST00000335135.4 ENST00000528027.1 |
GATA4 |
GATA binding protein 4 |
| chr17_+_66508537 | 0.05 |
ENST00000392711.1 ENST00000585427.1 ENST00000589228.1 ENST00000536854.2 ENST00000588702.1 ENST00000589309.1 |
PRKAR1A |
protein kinase, cAMP-dependent, regulatory, type I, alpha |
| chr1_+_36396677 | 0.05 |
ENST00000373191.4 ENST00000397828.2 |
AGO3 |
argonaute RISC catalytic component 3 |
| chr22_+_21771656 | 0.05 |
ENST00000407464.2 |
HIC2 |
hypermethylated in cancer 2 |
| chr1_-_39339777 | 0.05 |
ENST00000397572.2 |
MYCBP |
MYC binding protein |
| chr5_+_161274685 | 0.05 |
ENST00000428797.2 |
GABRA1 |
gamma-aminobutyric acid (GABA) A receptor, alpha 1 |
| chr5_+_122110691 | 0.05 |
ENST00000379516.2 ENST00000505934.1 ENST00000514949.1 |
SNX2 |
sorting nexin 2 |
| chr2_+_120517174 | 0.05 |
ENST00000263708.2 |
PTPN4 |
protein tyrosine phosphatase, non-receptor type 4 (megakaryocyte) |
| chr15_+_57210818 | 0.04 |
ENST00000438423.2 ENST00000267811.5 ENST00000452095.2 ENST00000559609.1 ENST00000333725.5 |
TCF12 |
transcription factor 12 |
| chr15_-_64648273 | 0.04 |
ENST00000607537.1 ENST00000303052.7 ENST00000303032.6 |
CSNK1G1 |
casein kinase 1, gamma 1 |
| chr1_+_50574585 | 0.04 |
ENST00000371824.1 ENST00000371823.4 |
ELAVL4 |
ELAV like neuron-specific RNA binding protein 4 |
| chr6_-_13711773 | 0.04 |
ENST00000011619.3 |
RANBP9 |
RAN binding protein 9 |
| chr3_+_29322803 | 0.04 |
ENST00000396583.3 ENST00000383767.2 |
RBMS3 |
RNA binding motif, single stranded interacting protein 3 |
| chr17_-_45266542 | 0.04 |
ENST00000531206.1 ENST00000527547.1 ENST00000446365.2 ENST00000575483.1 ENST00000066544.3 |
CDC27 |
cell division cycle 27 |
| chrX_+_16737718 | 0.04 |
ENST00000380155.3 |
SYAP1 |
synapse associated protein 1 |
| chr5_-_114880533 | 0.03 |
ENST00000274457.3 |
FEM1C |
fem-1 homolog c (C. elegans) |
| chrX_+_41192595 | 0.03 |
ENST00000399959.2 |
DDX3X |
DEAD (Asp-Glu-Ala-Asp) box helicase 3, X-linked |
| chr11_-_128392085 | 0.03 |
ENST00000526145.2 ENST00000531611.1 ENST00000319397.6 ENST00000345075.4 ENST00000535549.1 |
ETS1 |
v-ets avian erythroblastosis virus E26 oncogene homolog 1 |
| chr12_-_42538657 | 0.03 |
ENST00000398675.3 |
GXYLT1 |
glucoside xylosyltransferase 1 |
| chr9_-_19786926 | 0.03 |
ENST00000341998.2 ENST00000286344.3 |
SLC24A2 |
solute carrier family 24 (sodium/potassium/calcium exchanger), member 2 |
| chr6_-_166075557 | 0.03 |
ENST00000539869.2 ENST00000366882.1 |
PDE10A |
phosphodiesterase 10A |
| chr15_+_101142722 | 0.02 |
ENST00000332783.7 ENST00000558747.1 ENST00000343276.4 |
ASB7 |
ankyrin repeat and SOCS box containing 7 |
| chr15_+_44829255 | 0.02 |
ENST00000261868.5 ENST00000424492.3 |
EIF3J |
eukaryotic translation initiation factor 3, subunit J |
| chr17_-_48785216 | 0.02 |
ENST00000285243.6 |
ANKRD40 |
ankyrin repeat domain 40 |
| chr7_+_65338230 | 0.02 |
ENST00000360768.3 |
VKORC1L1 |
vitamin K epoxide reductase complex, subunit 1-like 1 |
| chr1_-_39347255 | 0.02 |
ENST00000454994.2 ENST00000357771.3 |
GJA9 |
gap junction protein, alpha 9, 59kDa |
| chr2_+_176981307 | 0.02 |
ENST00000249501.4 |
HOXD10 |
homeobox D10 |
| chr13_+_49550015 | 0.02 |
ENST00000492622.2 |
FNDC3A |
fibronectin type III domain containing 3A |
| chr9_-_23821273 | 0.02 |
ENST00000380110.4 |
ELAVL2 |
ELAV like neuron-specific RNA binding protein 2 |
| chr7_+_138916231 | 0.02 |
ENST00000473989.3 ENST00000288561.8 |
UBN2 |
ubinuclein 2 |
| chr1_-_21113105 | 0.02 |
ENST00000375000.1 ENST00000419490.1 ENST00000414993.1 ENST00000443615.1 ENST00000312239.5 |
HP1BP3 |
heterochromatin protein 1, binding protein 3 |
| chr6_+_163835669 | 0.02 |
ENST00000453779.2 ENST00000275262.7 ENST00000392127.2 ENST00000361752.3 |
QKI |
QKI, KH domain containing, RNA binding |
| chr13_+_102104980 | 0.02 |
ENST00000545560.2 |
ITGBL1 |
integrin, beta-like 1 (with EGF-like repeat domains) |
| chr14_+_105781048 | 0.02 |
ENST00000458164.2 ENST00000447393.1 |
PACS2 |
phosphofurin acidic cluster sorting protein 2 |
| chr7_-_44924939 | 0.02 |
ENST00000395699.2 |
PURB |
purine-rich element binding protein B |
| chr6_+_133562472 | 0.02 |
ENST00000430974.2 ENST00000367895.5 ENST00000355167.3 ENST00000355286.6 |
EYA4 |
eyes absent homolog 4 (Drosophila) |
| chr4_+_139936905 | 0.01 |
ENST00000280614.2 |
CCRN4L |
CCR4 carbon catabolite repression 4-like (S. cerevisiae) |
| chr3_-_15563229 | 0.01 |
ENST00000383786.5 ENST00000383787.2 ENST00000383785.2 ENST00000383788.5 ENST00000603808.1 |
COLQ |
collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase |
| chr9_+_82186872 | 0.01 |
ENST00000376544.3 ENST00000376520.4 |
TLE4 |
transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) |
| chr13_+_50571155 | 0.01 |
ENST00000420995.2 ENST00000378182.3 ENST00000356017.4 ENST00000457662.2 |
TRIM13 |
tripartite motif containing 13 |
| chr8_-_89339705 | 0.01 |
ENST00000286614.6 |
MMP16 |
matrix metallopeptidase 16 (membrane-inserted) |
| chr3_+_89156674 | 0.01 |
ENST00000336596.2 |
EPHA3 |
EPH receptor A3 |
| chr1_+_43148059 | 0.01 |
ENST00000321358.7 ENST00000332220.6 |
YBX1 |
Y box binding protein 1 |
| chr6_+_37225540 | 0.01 |
ENST00000373491.3 |
TBC1D22B |
TBC1 domain family, member 22B |
| chr2_+_176987088 | 0.00 |
ENST00000249499.6 |
HOXD9 |
homeobox D9 |
| chr5_-_79287060 | 0.00 |
ENST00000512560.1 ENST00000509852.1 ENST00000512528.1 |
MTX3 |
metaxin 3 |
| chr4_+_123747834 | 0.00 |
ENST00000264498.3 |
FGF2 |
fibroblast growth factor 2 (basic) |
| chr2_-_131850951 | 0.00 |
ENST00000409185.1 ENST00000389915.3 |
FAM168B |
family with sequence similarity 168, member B |
| chr21_-_16437255 | 0.00 |
ENST00000400199.1 ENST00000400202.1 |
NRIP1 |
nuclear receptor interacting protein 1 |
| chr15_-_56535464 | 0.00 |
ENST00000559447.2 ENST00000422057.1 ENST00000317318.6 ENST00000423270.1 |
RFX7 |
regulatory factor X, 7 |
| chr14_-_61116168 | 0.00 |
ENST00000247182.6 |
SIX1 |
SIX homeobox 1 |
Gene Ontology Analysis
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.0 | 0.1 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.0 | 0.0 | GO:1990851 | Wnt-Frizzled-LRP5/6 complex(GO:1990851) |
| 0.0 | 0.1 | GO:0070695 | FHF complex(GO:0070695) |
| 0.0 | 0.2 | GO:0031332 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.0 | 0.1 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.0 | 0.2 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:1990535 | transformation of host cell by virus(GO:0019087) neuron projection maintenance(GO:1990535) |
| 0.1 | 0.2 | GO:1905064 | epithelial to mesenchymal transition involved in cardiac fibroblast development(GO:0060940) negative regulation of vascular smooth muscle cell differentiation(GO:1905064) |
| 0.0 | 0.2 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.0 | 0.1 | GO:0090244 | trachea cartilage morphogenesis(GO:0060535) Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 0.0 | 0.0 | GO:0003285 | septum secundum development(GO:0003285) |
| 0.0 | 0.1 | GO:0061304 | extracellular matrix-cell signaling(GO:0035426) retinal blood vessel morphogenesis(GO:0061304) |
| 0.0 | 0.1 | GO:0022027 | interkinetic nuclear migration(GO:0022027) |
| 0.0 | 0.1 | GO:0051534 | negative regulation of NFAT protein import into nucleus(GO:0051534) |
| 0.0 | 0.2 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.0 | 0.3 | GO:0043559 | insulin binding(GO:0043559) |
| 0.0 | 0.2 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.0 | 0.3 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.1 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |