Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
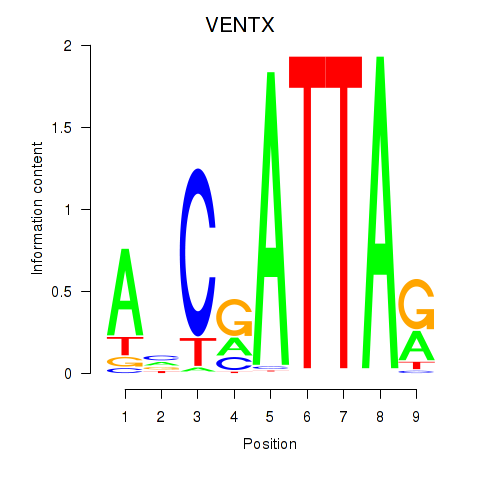
Results for VENTX
Z-value: 1.03
Transcription factors associated with VENTX
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
VENTX
|
ENSG00000151650.7 | VENTX |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| VENTX | hg19_v2_chr10_+_135050908_135050908 | 0.09 | 7.5e-01 | Click! |
Activity profile of VENTX motif
Sorted Z-values of VENTX motif
Network of associatons between targets according to the STRING database.
First level regulatory network of VENTX
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_84035905 | 3.81 |
ENST00000311507.4 |
PLAC8 |
placenta-specific 8 |
| chr4_-_84035868 | 3.74 |
ENST00000426923.2 ENST00000509973.1 |
PLAC8 |
placenta-specific 8 |
| chr4_-_155511887 | 2.07 |
ENST00000302053.3 ENST00000403106.3 |
FGA |
fibrinogen alpha chain |
| chr17_+_47448102 | 2.06 |
ENST00000576461.1 |
RP11-81K2.1 |
Uncharacterized protein |
| chr6_-_39693111 | 1.41 |
ENST00000373215.3 ENST00000538893.1 ENST00000287152.7 ENST00000373216.3 |
KIF6 |
kinesin family member 6 |
| chr12_+_25205568 | 1.34 |
ENST00000548766.1 ENST00000556887.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr7_-_22234381 | 1.34 |
ENST00000458533.1 |
RAPGEF5 |
Rap guanine nucleotide exchange factor (GEF) 5 |
| chr12_+_25205446 | 1.30 |
ENST00000557489.1 ENST00000354454.3 ENST00000536173.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr18_+_55888767 | 1.25 |
ENST00000431212.2 ENST00000586268.1 ENST00000587190.1 |
NEDD4L |
neural precursor cell expressed, developmentally down-regulated 4-like, E3 ubiquitin protein ligase |
| chr9_+_2159850 | 1.21 |
ENST00000416751.1 |
SMARCA2 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chrX_-_130423200 | 1.21 |
ENST00000361420.3 |
IGSF1 |
immunoglobulin superfamily, member 1 |
| chr1_+_226411319 | 1.19 |
ENST00000542034.1 ENST00000366810.5 |
MIXL1 |
Mix paired-like homeobox |
| chr18_+_29171689 | 1.18 |
ENST00000237014.3 |
TTR |
transthyretin |
| chr12_+_25205666 | 1.14 |
ENST00000547044.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr6_+_114178512 | 1.04 |
ENST00000368635.4 |
MARCKS |
myristoylated alanine-rich protein kinase C substrate |
| chr1_-_23886285 | 1.00 |
ENST00000374561.5 |
ID3 |
inhibitor of DNA binding 3, dominant negative helix-loop-helix protein |
| chr11_+_7618413 | 0.93 |
ENST00000528883.1 |
PPFIBP2 |
PTPRF interacting protein, binding protein 2 (liprin beta 2) |
| chr8_+_50824233 | 0.91 |
ENST00000522124.1 |
SNTG1 |
syntrophin, gamma 1 |
| chrX_-_130423240 | 0.91 |
ENST00000370910.1 ENST00000370901.4 |
IGSF1 |
immunoglobulin superfamily, member 1 |
| chr16_-_67517716 | 0.87 |
ENST00000290953.2 |
AGRP |
agouti related protein homolog (mouse) |
| chr12_+_56473939 | 0.86 |
ENST00000450146.2 |
ERBB3 |
v-erb-b2 avian erythroblastic leukemia viral oncogene homolog 3 |
| chr11_+_75526212 | 0.80 |
ENST00000356136.3 |
UVRAG |
UV radiation resistance associated |
| chr4_-_105416039 | 0.77 |
ENST00000394767.2 |
CXXC4 |
CXXC finger protein 4 |
| chr9_+_116225999 | 0.73 |
ENST00000317613.6 |
RGS3 |
regulator of G-protein signaling 3 |
| chr8_+_107738240 | 0.71 |
ENST00000449762.2 ENST00000297447.6 |
OXR1 |
oxidation resistance 1 |
| chr6_+_12290586 | 0.65 |
ENST00000379375.5 |
EDN1 |
endothelin 1 |
| chr14_-_20801427 | 0.64 |
ENST00000557665.1 ENST00000358932.4 ENST00000353689.4 |
CCNB1IP1 |
cyclin B1 interacting protein 1, E3 ubiquitin protein ligase |
| chr18_-_74207146 | 0.64 |
ENST00000443185.2 |
ZNF516 |
zinc finger protein 516 |
| chrX_+_133733457 | 0.63 |
ENST00000440614.1 |
RP11-308B5.2 |
RP11-308B5.2 |
| chr11_-_107729887 | 0.61 |
ENST00000525815.1 |
SLC35F2 |
solute carrier family 35, member F2 |
| chr18_+_59000815 | 0.55 |
ENST00000262717.4 |
CDH20 |
cadherin 20, type 2 |
| chr1_+_168250194 | 0.54 |
ENST00000367821.3 |
TBX19 |
T-box 19 |
| chr16_-_51185172 | 0.52 |
ENST00000251020.4 |
SALL1 |
spalt-like transcription factor 1 |
| chr12_+_56473910 | 0.50 |
ENST00000411731.2 |
ERBB3 |
v-erb-b2 avian erythroblastic leukemia viral oncogene homolog 3 |
| chr2_+_171034646 | 0.49 |
ENST00000409044.3 ENST00000408978.4 |
MYO3B |
myosin IIIB |
| chr12_-_6233828 | 0.48 |
ENST00000572068.1 ENST00000261405.5 |
VWF |
von Willebrand factor |
| chr6_+_153552455 | 0.48 |
ENST00000392385.2 |
AL590867.1 |
Uncharacterized protein; cDNA FLJ59044, highly similar to LINE-1 reverse transcriptase homolog |
| chr3_+_134514093 | 0.48 |
ENST00000398015.3 |
EPHB1 |
EPH receptor B1 |
| chr8_-_103425047 | 0.42 |
ENST00000520539.1 |
UBR5 |
ubiquitin protein ligase E3 component n-recognin 5 |
| chr12_+_81110684 | 0.42 |
ENST00000228644.3 |
MYF5 |
myogenic factor 5 |
| chr12_-_16758059 | 0.42 |
ENST00000261169.6 |
LMO3 |
LIM domain only 3 (rhombotin-like 2) |
| chr13_-_28545276 | 0.41 |
ENST00000381020.7 |
CDX2 |
caudal type homeobox 2 |
| chr4_-_26492076 | 0.40 |
ENST00000295589.3 |
CCKAR |
cholecystokinin A receptor |
| chr8_-_103424986 | 0.40 |
ENST00000521922.1 |
UBR5 |
ubiquitin protein ligase E3 component n-recognin 5 |
| chr18_-_53253323 | 0.40 |
ENST00000540999.1 ENST00000563888.2 |
TCF4 |
transcription factor 4 |
| chr8_-_103424916 | 0.39 |
ENST00000220959.4 |
UBR5 |
ubiquitin protein ligase E3 component n-recognin 5 |
| chr2_+_226265364 | 0.38 |
ENST00000272907.6 |
NYAP2 |
neuronal tyrosine-phosphorylated phosphoinositide-3-kinase adaptor 2 |
| chr2_-_203776864 | 0.38 |
ENST00000261015.4 |
WDR12 |
WD repeat domain 12 |
| chr6_-_76203345 | 0.37 |
ENST00000393004.2 |
FILIP1 |
filamin A interacting protein 1 |
| chr20_-_50722183 | 0.32 |
ENST00000371523.4 |
ZFP64 |
ZFP64 zinc finger protein |
| chr1_+_153747746 | 0.31 |
ENST00000368661.3 |
SLC27A3 |
solute carrier family 27 (fatty acid transporter), member 3 |
| chr2_-_88285309 | 0.30 |
ENST00000420840.2 |
RGPD2 |
RANBP2-like and GRIP domain containing 2 |
| chr18_-_53253112 | 0.29 |
ENST00000568673.1 ENST00000562847.1 ENST00000568147.1 |
TCF4 |
transcription factor 4 |
| chr3_-_57233966 | 0.28 |
ENST00000473921.1 ENST00000295934.3 |
HESX1 |
HESX homeobox 1 |
| chr17_-_47045949 | 0.28 |
ENST00000357424.2 |
GIP |
gastric inhibitory polypeptide |
| chrX_+_135730297 | 0.27 |
ENST00000370629.2 |
CD40LG |
CD40 ligand |
| chr22_-_40929812 | 0.26 |
ENST00000422851.1 |
MKL1 |
megakaryoblastic leukemia (translocation) 1 |
| chr17_-_202579 | 0.26 |
ENST00000577079.1 ENST00000331302.7 ENST00000536489.2 |
RPH3AL |
rabphilin 3A-like (without C2 domains) |
| chr1_+_62417957 | 0.26 |
ENST00000307297.7 ENST00000543708.1 |
INADL |
InaD-like (Drosophila) |
| chr4_-_168155169 | 0.25 |
ENST00000534949.1 ENST00000535728.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr12_-_101604185 | 0.24 |
ENST00000536262.2 |
SLC5A8 |
solute carrier family 5 (sodium/monocarboxylate cotransporter), member 8 |
| chr1_+_62439037 | 0.24 |
ENST00000545929.1 |
INADL |
InaD-like (Drosophila) |
| chr6_-_32157947 | 0.24 |
ENST00000375050.4 |
PBX2 |
pre-B-cell leukemia homeobox 2 |
| chr11_+_22359562 | 0.24 |
ENST00000263160.3 |
SLC17A6 |
solute carrier family 17 (vesicular glutamate transporter), member 6 |
| chrX_+_105937068 | 0.23 |
ENST00000324342.3 |
RNF128 |
ring finger protein 128, E3 ubiquitin protein ligase |
| chr13_-_67802549 | 0.22 |
ENST00000328454.5 ENST00000377865.2 |
PCDH9 |
protocadherin 9 |
| chr20_+_42136308 | 0.21 |
ENST00000434666.1 ENST00000427442.2 ENST00000439769.1 ENST00000418998.1 |
L3MBTL1 |
l(3)mbt-like 1 (Drosophila) |
| chr17_+_68100989 | 0.21 |
ENST00000585558.1 ENST00000392670.1 |
KCNJ16 |
potassium inwardly-rectifying channel, subfamily J, member 16 |
| chr2_+_166152283 | 0.21 |
ENST00000375427.2 |
SCN2A |
sodium channel, voltage-gated, type II, alpha subunit |
| chr9_+_133539981 | 0.21 |
ENST00000253008.2 |
PRDM12 |
PR domain containing 12 |
| chr2_-_163008903 | 0.20 |
ENST00000418842.2 ENST00000375497.3 |
GCG |
glucagon |
| chr11_+_46722368 | 0.19 |
ENST00000311764.2 |
ZNF408 |
zinc finger protein 408 |
| chr17_-_48785216 | 0.18 |
ENST00000285243.6 |
ANKRD40 |
ankyrin repeat domain 40 |
| chr12_+_52695617 | 0.18 |
ENST00000293525.5 |
KRT86 |
keratin 86 |
| chr5_-_82969405 | 0.18 |
ENST00000510978.1 |
HAPLN1 |
hyaluronan and proteoglycan link protein 1 |
| chr6_+_167704798 | 0.17 |
ENST00000230256.3 |
UNC93A |
unc-93 homolog A (C. elegans) |
| chr12_-_53994805 | 0.17 |
ENST00000328463.7 |
ATF7 |
activating transcription factor 7 |
| chr12_-_102591604 | 0.16 |
ENST00000329406.4 |
PMCH |
pro-melanin-concentrating hormone |
| chrX_+_135730373 | 0.16 |
ENST00000370628.2 |
CD40LG |
CD40 ligand |
| chr1_+_150337100 | 0.16 |
ENST00000401000.4 |
RPRD2 |
regulation of nuclear pre-mRNA domain containing 2 |
| chr9_+_117904097 | 0.15 |
ENST00000374016.1 |
DEC1 |
deleted in esophageal cancer 1 |
| chr9_+_136501478 | 0.15 |
ENST00000393056.2 ENST00000263611.2 |
DBH |
dopamine beta-hydroxylase (dopamine beta-monooxygenase) |
| chr3_+_171561127 | 0.15 |
ENST00000334567.5 ENST00000450693.1 |
TMEM212 |
transmembrane protein 212 |
| chr1_+_150337144 | 0.14 |
ENST00000539519.1 ENST00000369067.3 ENST00000369068.4 |
RPRD2 |
regulation of nuclear pre-mRNA domain containing 2 |
| chr5_-_79950775 | 0.14 |
ENST00000439211.2 |
DHFR |
dihydrofolate reductase |
| chr11_-_11374904 | 0.14 |
ENST00000528848.2 |
CSNK2A3 |
casein kinase 2, alpha 3 polypeptide |
| chr1_+_186265399 | 0.14 |
ENST00000367486.3 ENST00000367484.3 ENST00000533951.1 ENST00000367482.4 ENST00000367483.4 ENST00000367485.4 ENST00000445192.2 |
PRG4 |
proteoglycan 4 |
| chr2_+_87135076 | 0.13 |
ENST00000409776.2 |
RGPD1 |
RANBP2-like and GRIP domain containing 1 |
| chr6_+_167704838 | 0.13 |
ENST00000366829.2 |
UNC93A |
unc-93 homolog A (C. elegans) |
| chr2_-_165698662 | 0.13 |
ENST00000194871.6 ENST00000445474.2 |
COBLL1 |
cordon-bleu WH2 repeat protein-like 1 |
| chr6_-_99873145 | 0.13 |
ENST00000369239.5 ENST00000438806.1 |
PNISR |
PNN-interacting serine/arginine-rich protein |
| chr8_+_26150628 | 0.13 |
ENST00000523925.1 ENST00000315985.7 |
PPP2R2A |
protein phosphatase 2, regulatory subunit B, alpha |
| chr5_-_130500922 | 0.12 |
ENST00000513012.1 ENST00000508488.1 ENST00000506908.1 ENST00000304043.5 |
HINT1 |
histidine triad nucleotide binding protein 1 |
| chr6_-_111927062 | 0.12 |
ENST00000359831.4 |
TRAF3IP2 |
TRAF3 interacting protein 2 |
| chr1_-_28527152 | 0.11 |
ENST00000321830.5 |
AL353354.1 |
Uncharacterized protein |
| chr5_+_43121698 | 0.11 |
ENST00000505606.2 ENST00000509634.1 ENST00000509341.1 |
ZNF131 |
zinc finger protein 131 |
| chr14_+_65453432 | 0.10 |
ENST00000246166.2 |
FNTB |
farnesyltransferase, CAAX box, beta |
| chr12_-_51611477 | 0.10 |
ENST00000389243.4 |
POU6F1 |
POU class 6 homeobox 1 |
| chr5_-_156486120 | 0.10 |
ENST00000522693.1 |
HAVCR1 |
hepatitis A virus cellular receptor 1 |
| chr6_+_152011628 | 0.10 |
ENST00000404742.1 ENST00000440973.1 |
ESR1 |
estrogen receptor 1 |
| chr17_-_10372875 | 0.09 |
ENST00000255381.2 |
MYH4 |
myosin, heavy chain 4, skeletal muscle |
| chr14_+_32798462 | 0.09 |
ENST00000280979.4 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr17_-_39222131 | 0.09 |
ENST00000394015.2 |
KRTAP2-4 |
keratin associated protein 2-4 |
| chr3_+_9834227 | 0.08 |
ENST00000287613.7 ENST00000397261.3 |
ARPC4 |
actin related protein 2/3 complex, subunit 4, 20kDa |
| chr8_-_87755878 | 0.07 |
ENST00000320005.5 |
CNGB3 |
cyclic nucleotide gated channel beta 3 |
| chr7_+_139025875 | 0.07 |
ENST00000297534.6 |
C7orf55 |
chromosome 7 open reading frame 55 |
| chr1_-_190446759 | 0.06 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr22_+_39101728 | 0.06 |
ENST00000216044.5 ENST00000484657.1 |
GTPBP1 |
GTP binding protein 1 |
| chr19_-_15529790 | 0.05 |
ENST00000596195.1 ENST00000595067.1 ENST00000595465.2 ENST00000397410.5 ENST00000600247.1 |
AKAP8L |
A kinase (PRKA) anchor protein 8-like |
| chr7_-_104909435 | 0.05 |
ENST00000357311.3 |
SRPK2 |
SRSF protein kinase 2 |
| chr11_-_3859089 | 0.05 |
ENST00000396979.1 |
RHOG |
ras homolog family member G |
| chrX_+_78003204 | 0.04 |
ENST00000435339.3 ENST00000514744.1 |
LPAR4 |
lysophosphatidic acid receptor 4 |
| chr2_-_165698521 | 0.04 |
ENST00000409184.3 ENST00000392717.2 ENST00000456693.1 |
COBLL1 |
cordon-bleu WH2 repeat protein-like 1 |
| chr2_+_45168875 | 0.04 |
ENST00000260653.3 |
SIX3 |
SIX homeobox 3 |
| chr11_+_64018955 | 0.03 |
ENST00000279230.6 ENST00000540288.1 ENST00000325234.5 |
PLCB3 |
phospholipase C, beta 3 (phosphatidylinositol-specific) |
| chr6_-_90529418 | 0.03 |
ENST00000439638.1 ENST00000369393.3 ENST00000428876.1 |
MDN1 |
MDN1, midasin homolog (yeast) |
| chr4_+_37455536 | 0.01 |
ENST00000381980.4 ENST00000508175.1 |
C4orf19 |
chromosome 4 open reading frame 19 |
Gene Ontology Analysis
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 0.2 | 2.1 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.2 | 2.5 | GO:0043073 | germ cell nucleus(GO:0043073) |
| 0.1 | 0.9 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.1 | 8.7 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.1 | 0.4 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.1 | 0.8 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.1 | 1.2 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 3.8 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.0 | 0.5 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.0 | 1.3 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.0 | 0.1 | GO:0005965 | protein farnesyltransferase complex(GO:0005965) |
| 0.0 | 0.5 | GO:0010369 | chromocenter(GO:0010369) |
| 0.0 | 0.6 | GO:0000795 | synaptonemal complex(GO:0000795) |
| 0.0 | 0.1 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.0 | 1.3 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.0 | 0.2 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 7.5 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.3 | 0.8 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.3 | 0.8 | GO:0007352 | zygotic specification of dorsal/ventral axis(GO:0007352) |
| 0.3 | 1.3 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.2 | 0.7 | GO:0060585 | regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) |
| 0.2 | 2.1 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.2 | 1.2 | GO:2000382 | positive regulation of mesoderm development(GO:2000382) |
| 0.2 | 0.9 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.2 | 1.2 | GO:1900045 | negative regulation of histone ubiquitination(GO:0033183) negative regulation of protein K63-linked ubiquitination(GO:1900045) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.2 | 0.5 | GO:0072092 | olfactory bulb mitral cell layer development(GO:0061034) ureteric bud invasion(GO:0072092) |
| 0.2 | 0.5 | GO:0048170 | positive regulation of long-term neuronal synaptic plasticity(GO:0048170) trans-synaptic signaling by trans-synaptic complex(GO:0099545) |
| 0.1 | 0.4 | GO:0038188 | cholecystokinin signaling pathway(GO:0038188) |
| 0.1 | 0.4 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.1 | 1.2 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 0.6 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.1 | 0.4 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.1 | 0.2 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 0.1 | 1.0 | GO:0030903 | notochord development(GO:0030903) |
| 0.1 | 0.2 | GO:1900109 | regulation of histone H3-K9 dimethylation(GO:1900109) |
| 0.0 | 0.3 | GO:0070092 | regulation of glucagon secretion(GO:0070092) |
| 0.0 | 1.2 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.0 | 0.7 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.0 | 0.4 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.0 | 0.3 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.0 | 0.1 | GO:0046452 | dihydrofolate metabolic process(GO:0046452) |
| 0.0 | 0.4 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 3.8 | GO:0006903 | vesicle targeting(GO:0006903) |
| 0.0 | 0.2 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.2 | GO:0098700 | neurotransmitter loading into synaptic vesicle(GO:0098700) |
| 0.0 | 0.2 | GO:1904352 | positive regulation of protein catabolic process in the vacuole(GO:1904352) |
| 0.0 | 0.1 | GO:0042421 | norepinephrine biosynthetic process(GO:0042421) |
| 0.0 | 0.5 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 0.0 | 0.4 | GO:2000353 | positive regulation of endothelial cell apoptotic process(GO:2000353) |
| 0.0 | 0.2 | GO:0034356 | NAD biosynthesis via nicotinamide riboside salvage pathway(GO:0034356) |
| 0.0 | 0.4 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 0.0 | GO:0021798 | forebrain dorsal/ventral pattern formation(GO:0021798) |
| 0.0 | 0.6 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.0 | 0.2 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.0 | 0.1 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.0 | 0.2 | GO:0030916 | otic vesicle formation(GO:0030916) |
| 0.0 | 0.4 | GO:0000042 | protein targeting to Golgi(GO:0000042) |
| 0.0 | 0.1 | GO:0060737 | prostate epithelial cord elongation(GO:0060523) prostate gland morphogenetic growth(GO:0060737) epithelial cell proliferation involved in mammary gland duct elongation(GO:0060750) |
| 0.0 | 0.5 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.3 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 0.1 | GO:0010793 | regulation of mRNA export from nucleus(GO:0010793) |
| 0.0 | 0.1 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.0 | 1.2 | GO:0007286 | spermatid development(GO:0007286) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.1 | 1.1 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.0 | 1.1 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.9 | REACTOME INCRETIN SYNTHESIS SECRETION AND INACTIVATION | Genes involved in Incretin Synthesis, Secretion, and Inactivation |
| 0.0 | 0.5 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.0 | 1.1 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.5 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 1.2 | REACTOME AMYLOIDS | Genes involved in Amyloids |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.3 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.9 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 1.1 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 1.2 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 1.4 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 1.0 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.6 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.1 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.2 | 1.3 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.1 | 0.4 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.1 | 0.4 | GO:0004951 | cholecystokinin receptor activity(GO:0004951) |
| 0.1 | 0.7 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.1 | 1.2 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 1.4 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.1 | 1.3 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 1.2 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.1 | 0.5 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.1 | 0.2 | GO:0032093 | SAM domain binding(GO:0032093) |
| 0.1 | 0.7 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 2.7 | GO:0001190 | transcriptional activator activity, RNA polymerase II transcription factor binding(GO:0001190) transcriptional repressor activity, RNA polymerase II activating transcription factor binding(GO:0098811) |
| 0.0 | 0.1 | GO:0004660 | protein farnesyltransferase activity(GO:0004660) |
| 0.0 | 0.9 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.1 | GO:0051870 | methotrexate binding(GO:0051870) |
| 0.0 | 0.3 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 0.2 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.0 | 0.5 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.0 | 1.1 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 1.0 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.6 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 0.2 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 1.4 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 0.9 | GO:0001158 | enhancer sequence-specific DNA binding(GO:0001158) |
| 0.0 | 0.4 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.1 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.0 | 0.1 | GO:0001093 | TFIIB-class transcription factor binding(GO:0001093) |
| 0.0 | 0.2 | GO:0005313 | L-glutamate transmembrane transporter activity(GO:0005313) acidic amino acid transmembrane transporter activity(GO:0015172) |
| 0.0 | 6.9 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 0.1 | GO:0005223 | intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.0 | 0.3 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |