Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
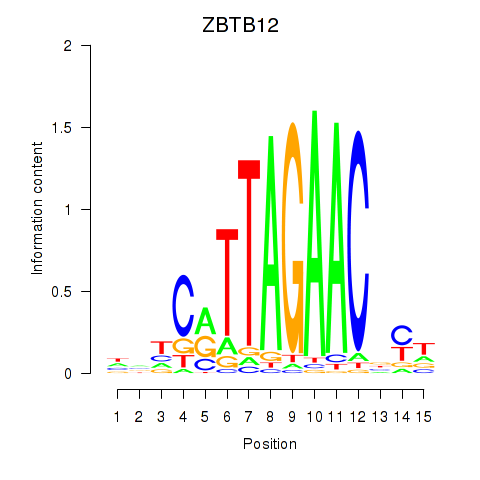
Results for ZBTB12
Z-value: 1.02
Transcription factors associated with ZBTB12
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZBTB12
|
ENSG00000204366.3 | ZBTB12 |
Activity profile of ZBTB12 motif
Sorted Z-values of ZBTB12 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZBTB12
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_26644441 | 2.30 |
ENST00000374213.2 |
CD52 |
CD52 molecule |
| chr1_-_117210290 | 2.28 |
ENST00000369483.1 ENST00000369486.3 |
IGSF3 |
immunoglobulin superfamily, member 3 |
| chr17_-_39677971 | 2.25 |
ENST00000393976.2 |
KRT15 |
keratin 15 |
| chrX_-_73072534 | 2.06 |
ENST00000429829.1 |
XIST |
X inactive specific transcript (non-protein coding) |
| chr22_+_22930626 | 2.02 |
ENST00000390302.2 |
IGLV2-33 |
immunoglobulin lambda variable 2-33 (non-functional) |
| chr13_-_46716969 | 1.79 |
ENST00000435666.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chr1_+_158979792 | 1.79 |
ENST00000359709.3 ENST00000430894.2 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr11_-_104972158 | 1.79 |
ENST00000598974.1 ENST00000593315.1 ENST00000594519.1 ENST00000415981.2 ENST00000525374.1 ENST00000375707.1 |
CASP1 CARD16 CARD17 |
caspase 1, apoptosis-related cysteine peptidase caspase recruitment domain family, member 16 caspase recruitment domain family, member 17 |
| chr1_+_158979680 | 1.57 |
ENST00000368131.4 ENST00000340979.6 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr1_+_158979686 | 1.54 |
ENST00000368132.3 ENST00000295809.7 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr15_-_55562479 | 1.29 |
ENST00000564609.1 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr7_-_93520191 | 1.29 |
ENST00000545378.1 |
TFPI2 |
tissue factor pathway inhibitor 2 |
| chr4_-_36246060 | 1.27 |
ENST00000303965.4 |
ARAP2 |
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 2 |
| chr15_-_55563072 | 1.21 |
ENST00000567380.1 ENST00000565972.1 ENST00000569493.1 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr19_-_11688500 | 1.17 |
ENST00000433365.2 |
ACP5 |
acid phosphatase 5, tartrate resistant |
| chr1_-_200992827 | 1.10 |
ENST00000332129.2 ENST00000422435.2 |
KIF21B |
kinesin family member 21B |
| chr1_+_35220613 | 1.07 |
ENST00000338513.1 |
GJB5 |
gap junction protein, beta 5, 31.1kDa |
| chr7_-_93520259 | 1.03 |
ENST00000222543.5 |
TFPI2 |
tissue factor pathway inhibitor 2 |
| chr14_-_50154921 | 0.88 |
ENST00000553805.2 ENST00000554396.1 ENST00000216367.5 ENST00000539565.2 |
POLE2 |
polymerase (DNA directed), epsilon 2, accessory subunit |
| chr11_-_104905840 | 0.87 |
ENST00000526568.1 ENST00000393136.4 ENST00000531166.1 ENST00000534497.1 ENST00000527979.1 ENST00000446369.1 ENST00000353247.5 ENST00000528974.1 ENST00000533400.1 ENST00000525825.1 ENST00000436863.3 |
CASP1 |
caspase 1, apoptosis-related cysteine peptidase |
| chr15_-_55562582 | 0.82 |
ENST00000396307.2 |
RAB27A |
RAB27A, member RAS oncogene family |
| chr16_-_67969888 | 0.72 |
ENST00000574576.2 |
PSMB10 |
proteasome (prosome, macropain) subunit, beta type, 10 |
| chr1_+_32757668 | 0.70 |
ENST00000373548.3 |
HDAC1 |
histone deacetylase 1 |
| chr6_+_26158343 | 0.69 |
ENST00000377777.4 ENST00000289316.2 |
HIST1H2BD |
histone cluster 1, H2bd |
| chrX_+_149861836 | 0.68 |
ENST00000542156.1 ENST00000370390.3 ENST00000490316.2 ENST00000445323.2 ENST00000544228.1 ENST00000451863.2 |
MTMR1 |
myotubularin related protein 1 |
| chr8_+_38758737 | 0.63 |
ENST00000521746.1 ENST00000420274.1 |
PLEKHA2 |
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 2 |
| chr2_+_234545092 | 0.63 |
ENST00000344644.5 |
UGT1A10 |
UDP glucuronosyltransferase 1 family, polypeptide A10 |
| chr8_+_104892639 | 0.62 |
ENST00000436393.2 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr15_+_81475047 | 0.61 |
ENST00000559388.1 |
IL16 |
interleukin 16 |
| chr11_+_74303575 | 0.59 |
ENST00000263681.2 |
POLD3 |
polymerase (DNA-directed), delta 3, accessory subunit |
| chr1_+_45212051 | 0.58 |
ENST00000372222.3 |
KIF2C |
kinesin family member 2C |
| chr19_-_6767431 | 0.58 |
ENST00000437152.3 ENST00000597687.1 |
SH2D3A |
SH2 domain containing 3A |
| chr15_+_66797627 | 0.57 |
ENST00000565627.1 ENST00000564179.1 |
ZWILCH |
zwilch kinetochore protein |
| chr10_+_52152766 | 0.56 |
ENST00000596442.1 |
AC069547.2 |
Uncharacterized protein |
| chr10_-_12084770 | 0.55 |
ENST00000357604.5 |
UPF2 |
UPF2 regulator of nonsense transcripts homolog (yeast) |
| chr15_+_66797455 | 0.54 |
ENST00000446801.2 |
ZWILCH |
zwilch kinetochore protein |
| chr17_+_46184911 | 0.52 |
ENST00000580219.1 ENST00000452859.2 ENST00000393405.2 ENST00000439357.2 ENST00000359238.2 |
SNX11 |
sorting nexin 11 |
| chr19_-_46148820 | 0.49 |
ENST00000587152.1 |
EML2 |
echinoderm microtubule associated protein like 2 |
| chr19_-_51531272 | 0.48 |
ENST00000319720.7 |
KLK11 |
kallikrein-related peptidase 11 |
| chr22_+_38035459 | 0.46 |
ENST00000357436.4 |
SH3BP1 |
SH3-domain binding protein 1 |
| chr19_-_6767516 | 0.46 |
ENST00000245908.6 |
SH2D3A |
SH2 domain containing 3A |
| chr1_+_45212074 | 0.45 |
ENST00000372217.1 |
KIF2C |
kinesin family member 2C |
| chr19_-_51530916 | 0.43 |
ENST00000594768.1 |
KLK11 |
kallikrein-related peptidase 11 |
| chr17_-_61781750 | 0.43 |
ENST00000582026.1 |
STRADA |
STE20-related kinase adaptor alpha |
| chr4_-_99851766 | 0.41 |
ENST00000450253.2 |
EIF4E |
eukaryotic translation initiation factor 4E |
| chr22_+_51039098 | 0.39 |
ENST00000399912.1 ENST00000329492.3 ENST00000442429.2 ENST00000341339.4 |
MAPK8IP2 |
mitogen-activated protein kinase 8 interacting protein 2 |
| chr14_-_91526462 | 0.35 |
ENST00000536315.2 |
RPS6KA5 |
ribosomal protein S6 kinase, 90kDa, polypeptide 5 |
| chr18_-_53177984 | 0.33 |
ENST00000543082.1 |
TCF4 |
transcription factor 4 |
| chr4_-_87028478 | 0.32 |
ENST00000515400.1 ENST00000395157.3 |
MAPK10 |
mitogen-activated protein kinase 10 |
| chr19_-_51531210 | 0.31 |
ENST00000391804.3 |
KLK11 |
kallikrein-related peptidase 11 |
| chr6_+_36097992 | 0.31 |
ENST00000211287.4 |
MAPK13 |
mitogen-activated protein kinase 13 |
| chr19_+_44716678 | 0.28 |
ENST00000586228.1 ENST00000588219.1 ENST00000313040.7 ENST00000589707.1 ENST00000588394.1 ENST00000589005.1 |
ZNF227 |
zinc finger protein 227 |
| chr5_-_96518907 | 0.27 |
ENST00000508447.1 ENST00000283109.3 |
RIOK2 |
RIO kinase 2 |
| chr2_+_89999259 | 0.27 |
ENST00000558026.1 |
IGKV2D-28 |
immunoglobulin kappa variable 2D-28 |
| chr20_+_36373032 | 0.27 |
ENST00000373473.1 |
CTNNBL1 |
catenin, beta like 1 |
| chr3_-_50336826 | 0.26 |
ENST00000443842.1 ENST00000354862.4 ENST00000443094.2 ENST00000415204.1 ENST00000336307.1 |
NAT6 HYAL3 |
N-acetyltransferase 6 (GCN5-related) hyaluronoglucosaminidase 3 |
| chr19_-_13227534 | 0.25 |
ENST00000588229.1 ENST00000357720.4 |
TRMT1 |
tRNA methyltransferase 1 homolog (S. cerevisiae) |
| chr14_+_76618242 | 0.25 |
ENST00000557542.1 ENST00000557263.1 ENST00000557207.1 ENST00000312858.5 ENST00000261530.7 |
GPATCH2L |
G patch domain containing 2-like |
| chr12_-_7077125 | 0.25 |
ENST00000545555.2 |
PHB2 |
prohibitin 2 |
| chr5_-_88179302 | 0.24 |
ENST00000504921.2 |
MEF2C |
myocyte enhancer factor 2C |
| chr1_+_200993071 | 0.24 |
ENST00000446333.1 ENST00000458003.1 |
RP11-168O16.1 |
RP11-168O16.1 |
| chr1_-_94344754 | 0.23 |
ENST00000436063.2 |
DNTTIP2 |
deoxynucleotidyltransferase, terminal, interacting protein 2 |
| chr17_-_5487768 | 0.22 |
ENST00000269280.4 ENST00000345221.3 ENST00000262467.5 |
NLRP1 |
NLR family, pyrin domain containing 1 |
| chr17_+_73257945 | 0.21 |
ENST00000579002.1 |
MRPS7 |
mitochondrial ribosomal protein S7 |
| chr15_+_40453204 | 0.20 |
ENST00000287598.6 ENST00000412359.3 |
BUB1B |
BUB1 mitotic checkpoint serine/threonine kinase B |
| chr2_+_166152283 | 0.20 |
ENST00000375427.2 |
SCN2A |
sodium channel, voltage-gated, type II, alpha subunit |
| chr4_-_85419603 | 0.20 |
ENST00000295886.4 |
NKX6-1 |
NK6 homeobox 1 |
| chr13_+_45694583 | 0.20 |
ENST00000340473.6 |
GTF2F2 |
general transcription factor IIF, polypeptide 2, 30kDa |
| chr14_-_70826438 | 0.20 |
ENST00000389912.6 |
COX16 |
COX16 cytochrome c oxidase assembly homolog (S. cerevisiae) |
| chr5_-_88178964 | 0.19 |
ENST00000513252.1 ENST00000508569.1 ENST00000510942.1 ENST00000506554.1 |
MEF2C |
myocyte enhancer factor 2C |
| chr18_-_24445664 | 0.19 |
ENST00000578776.1 |
AQP4 |
aquaporin 4 |
| chr5_-_88179017 | 0.19 |
ENST00000514028.1 ENST00000514015.1 ENST00000503075.1 ENST00000437473.2 |
MEF2C |
myocyte enhancer factor 2C |
| chr7_+_92158083 | 0.18 |
ENST00000265732.5 ENST00000481551.1 ENST00000496410.1 |
RBM48 |
RNA binding motif protein 48 |
| chr5_-_35089722 | 0.16 |
ENST00000511486.1 ENST00000310101.5 ENST00000231423.3 ENST00000513753.1 ENST00000348262.3 ENST00000397391.3 ENST00000542609.1 |
PRLR |
prolactin receptor |
| chr10_+_76969909 | 0.15 |
ENST00000298468.5 ENST00000543351.1 |
VDAC2 |
voltage-dependent anion channel 2 |
| chr8_+_1993173 | 0.15 |
ENST00000523438.1 |
MYOM2 |
myomesin 2 |
| chr2_+_173600514 | 0.15 |
ENST00000264111.6 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr11_+_64323156 | 0.15 |
ENST00000377585.3 |
SLC22A11 |
solute carrier family 22 (organic anion/urate transporter), member 11 |
| chr14_-_23877474 | 0.15 |
ENST00000405093.3 |
MYH6 |
myosin, heavy chain 6, cardiac muscle, alpha |
| chr3_-_52804872 | 0.15 |
ENST00000535191.1 ENST00000461689.1 ENST00000383721.4 ENST00000233027.5 |
NEK4 |
NIMA-related kinase 4 |
| chrX_+_101478829 | 0.14 |
ENST00000372763.1 ENST00000372758.1 |
NXF2 |
nuclear RNA export factor 2 |
| chr10_-_98273668 | 0.14 |
ENST00000357947.3 |
TLL2 |
tolloid-like 2 |
| chr17_+_11501748 | 0.14 |
ENST00000262442.4 ENST00000579828.1 |
DNAH9 |
dynein, axonemal, heavy chain 9 |
| chr8_-_141728760 | 0.14 |
ENST00000430260.2 |
PTK2 |
protein tyrosine kinase 2 |
| chr20_-_33543546 | 0.13 |
ENST00000216951.2 |
GSS |
glutathione synthetase |
| chr14_-_21270995 | 0.12 |
ENST00000555698.1 ENST00000397970.4 ENST00000340900.3 |
RNASE1 |
ribonuclease, RNase A family, 1 (pancreatic) |
| chrX_+_140096761 | 0.12 |
ENST00000370530.1 |
SPANXB1 |
SPANX family, member B1 |
| chr1_+_46049706 | 0.11 |
ENST00000527470.1 ENST00000525515.1 ENST00000537798.1 ENST00000402363.3 ENST00000528238.1 ENST00000350030.3 ENST00000470768.1 ENST00000372052.4 ENST00000351223.3 |
NASP |
nuclear autoantigenic sperm protein (histone-binding) |
| chr18_-_47018769 | 0.11 |
ENST00000583637.1 ENST00000578528.1 ENST00000578532.1 ENST00000580387.1 ENST00000579248.1 ENST00000581373.1 |
RPL17 |
ribosomal protein L17 |
| chr12_+_123459127 | 0.10 |
ENST00000397389.2 ENST00000538755.1 ENST00000536150.1 ENST00000545056.1 ENST00000545612.1 ENST00000538628.1 ENST00000545317.1 |
OGFOD2 |
2-oxoglutarate and iron-dependent oxygenase domain containing 2 |
| chr4_-_168155417 | 0.10 |
ENST00000511269.1 ENST00000506697.1 ENST00000512042.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr14_-_74959994 | 0.10 |
ENST00000238633.2 ENST00000434013.2 |
NPC2 |
Niemann-Pick disease, type C2 |
| chr3_+_89156674 | 0.09 |
ENST00000336596.2 |
EPHA3 |
EPH receptor A3 |
| chr2_+_190722119 | 0.09 |
ENST00000452382.1 |
PMS1 |
PMS1 postmeiotic segregation increased 1 (S. cerevisiae) |
| chr4_-_10118348 | 0.09 |
ENST00000502702.1 |
WDR1 |
WD repeat domain 1 |
| chr1_+_46016703 | 0.08 |
ENST00000481885.1 ENST00000351829.4 ENST00000471651.1 |
AKR1A1 |
aldo-keto reductase family 1, member A1 (aldehyde reductase) |
| chr18_-_24445729 | 0.08 |
ENST00000383168.4 |
AQP4 |
aquaporin 4 |
| chr14_-_74960030 | 0.08 |
ENST00000553490.1 ENST00000557510.1 |
NPC2 |
Niemann-Pick disease, type C2 |
| chr8_-_63998590 | 0.08 |
ENST00000260116.4 |
TTPA |
tocopherol (alpha) transfer protein |
| chr8_+_1993152 | 0.08 |
ENST00000262113.4 |
MYOM2 |
myomesin 2 |
| chr1_+_22778337 | 0.08 |
ENST00000404138.1 ENST00000400239.2 ENST00000375647.4 ENST00000374651.4 |
ZBTB40 |
zinc finger and BTB domain containing 40 |
| chr20_-_8000426 | 0.07 |
ENST00000527925.1 ENST00000246024.2 |
TMX4 |
thioredoxin-related transmembrane protein 4 |
| chr1_+_171107241 | 0.07 |
ENST00000236166.3 |
FMO6P |
flavin containing monooxygenase 6 pseudogene |
| chr5_-_176778803 | 0.07 |
ENST00000303127.7 |
LMAN2 |
lectin, mannose-binding 2 |
| chr19_-_54618650 | 0.07 |
ENST00000391757.1 |
TFPT |
TCF3 (E2A) fusion partner (in childhood Leukemia) |
| chr8_+_110098850 | 0.06 |
ENST00000518632.1 |
TRHR |
thyrotropin-releasing hormone receptor |
| chrX_-_24690771 | 0.06 |
ENST00000379145.1 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr3_-_151047327 | 0.06 |
ENST00000325602.5 |
P2RY13 |
purinergic receptor P2Y, G-protein coupled, 13 |
| chr11_+_64323098 | 0.05 |
ENST00000301891.4 |
SLC22A11 |
solute carrier family 22 (organic anion/urate transporter), member 11 |
| chr10_+_57358750 | 0.04 |
ENST00000512524.2 |
MTRNR2L5 |
MT-RNR2-like 5 |
| chr19_+_55417499 | 0.04 |
ENST00000291890.4 ENST00000447255.1 ENST00000598576.1 ENST00000594765.1 |
NCR1 |
natural cytotoxicity triggering receptor 1 |
| chr19_+_55417530 | 0.04 |
ENST00000350790.5 ENST00000338835.5 ENST00000357397.5 |
NCR1 |
natural cytotoxicity triggering receptor 1 |
| chr19_+_52264449 | 0.04 |
ENST00000599326.1 ENST00000598953.1 |
FPR2 |
formyl peptide receptor 2 |
| chr11_-_101000445 | 0.03 |
ENST00000534013.1 |
PGR |
progesterone receptor |
| chr4_-_96470350 | 0.03 |
ENST00000504962.1 ENST00000453304.1 ENST00000506749.1 |
UNC5C |
unc-5 homolog C (C. elegans) |
| chr1_+_155583012 | 0.03 |
ENST00000462250.2 |
MSTO1 |
misato 1, mitochondrial distribution and morphology regulator |
| chr8_-_30670384 | 0.03 |
ENST00000221138.4 ENST00000518243.1 |
PPP2CB |
protein phosphatase 2, catalytic subunit, beta isozyme |
| chr19_+_11485333 | 0.02 |
ENST00000312423.2 |
SWSAP1 |
SWIM-type zinc finger 7 associated protein 1 |
| chr22_-_50524298 | 0.02 |
ENST00000311597.5 |
MLC1 |
megalencephalic leukoencephalopathy with subcortical cysts 1 |
| chr20_-_36793663 | 0.01 |
ENST00000536701.1 ENST00000536724.1 |
TGM2 |
transglutaminase 2 |
| chr5_-_148929848 | 0.01 |
ENST00000504676.1 ENST00000515435.1 |
CSNK1A1 |
casein kinase 1, alpha 1 |
| chr15_-_34880646 | 0.00 |
ENST00000543376.1 |
GOLGA8A |
golgin A8 family, member A |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.7 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 3.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 1.5 | REACTOME REPAIR SYNTHESIS FOR GAP FILLING BY DNA POL IN TC NER | Genes involved in Repair synthesis for gap-filling by DNA polymerase in TC-NER |
| 0.1 | 1.1 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 1.0 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 0.3 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.0 | 0.7 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 0.3 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.0 | 0.4 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.2 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.0 | 0.4 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.0 | 1.0 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.2 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.0 | 0.3 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.0 | 1.3 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.3 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.0 | 0.7 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 0.3 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.7 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.0 | 1.3 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 1.4 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 1.0 | PID AURORA B PATHWAY | Aurora B signaling |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.7 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 0.6 | 3.3 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.3 | 1.1 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.2 | 1.2 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.2 | 2.3 | GO:0032808 | lacrimal gland development(GO:0032808) |
| 0.2 | 0.6 | GO:0003185 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.1 | 0.6 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 0.1 | 1.8 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 1.0 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.1 | 5.1 | GO:0032731 | positive regulation of interleukin-1 beta production(GO:0032731) |
| 0.1 | 0.3 | GO:0090070 | positive regulation of ribosome biogenesis(GO:0090070) positive regulation of rRNA processing(GO:2000234) |
| 0.1 | 2.3 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.1 | 0.4 | GO:0043988 | histone H3-S28 phosphorylation(GO:0043988) histone H2A phosphorylation(GO:1990164) |
| 0.1 | 0.6 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.1 | 0.2 | GO:0072560 | glandular epithelial cell maturation(GO:0002071) regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification(GO:0021913) type B pancreatic cell maturation(GO:0072560) |
| 0.1 | 2.3 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.1 | 0.6 | GO:0051552 | flavone metabolic process(GO:0051552) |
| 0.1 | 0.2 | GO:0051754 | meiotic sister chromatid cohesion, centromeric(GO:0051754) |
| 0.0 | 1.5 | GO:0032201 | telomere maintenance via semi-conservative replication(GO:0032201) |
| 0.0 | 0.2 | GO:0042695 | thelarche(GO:0042695) mammary gland branching involved in thelarche(GO:0060744) regulation of branching involved in mammary gland duct morphogenesis(GO:0060762) |
| 0.0 | 0.5 | GO:0010968 | regulation of microtubule nucleation(GO:0010968) |
| 0.0 | 0.2 | GO:0019747 | regulation of isoprenoid metabolic process(GO:0019747) |
| 0.0 | 0.3 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.0 | 0.4 | GO:0046958 | nonassociative learning(GO:0046958) |
| 0.0 | 0.1 | GO:0048631 | regulation of skeletal muscle tissue growth(GO:0048631) |
| 0.0 | 0.6 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.0 | 0.2 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.0 | 0.2 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.0 | 0.1 | GO:0019853 | L-ascorbic acid biosynthetic process(GO:0019853) |
| 0.0 | 0.1 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.0 | 0.7 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 0.2 | GO:0015747 | urate transport(GO:0015747) |
| 0.0 | 0.1 | GO:0016333 | morphogenesis of follicular epithelium(GO:0016333) establishment or maintenance of polarity of follicular epithelium(GO:0016334) establishment of planar polarity of follicular epithelium(GO:0042247) |
| 0.0 | 0.1 | GO:0090212 | vitamin E metabolic process(GO:0042360) regulation of establishment of blood-brain barrier(GO:0090210) negative regulation of establishment of blood-brain barrier(GO:0090212) |
| 0.0 | 2.3 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 0.2 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 0.0 | 0.2 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.0 | 0.1 | GO:0001302 | replicative cell aging(GO:0001302) |
| 0.0 | 0.2 | GO:0032968 | positive regulation of transcription elongation from RNA polymerase II promoter(GO:0032968) |
| 0.0 | 2.3 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.2 | GO:0071493 | cellular response to UV-B(GO:0071493) |
| 0.0 | 0.1 | GO:0006335 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.0 | 0.3 | GO:0006833 | water transport(GO:0006833) |
| 0.0 | 0.0 | GO:0060748 | tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.7 | GO:0097179 | protease inhibitor complex(GO:0097179) |
| 0.3 | 1.1 | GO:1990423 | RZZ complex(GO:1990423) |
| 0.3 | 3.3 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.2 | 0.9 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.1 | 0.6 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.1 | 0.7 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.1 | 1.8 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 0.2 | GO:0005674 | transcription factor TFIIF complex(GO:0005674) |
| 0.1 | 1.1 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 0.2 | GO:0072558 | NLRP1 inflammasome complex(GO:0072558) |
| 0.0 | 0.4 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 2.1 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.3 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.0 | 0.1 | GO:0042272 | nuclear RNA export factor complex(GO:0042272) |
| 0.0 | 0.6 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.0 | 0.5 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.0 | 0.2 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.0 | 2.3 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.1 | GO:0042643 | actomyosin, actin portion(GO:0042643) |
| 0.0 | 2.3 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.6 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.1 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.3 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.0 | 0.2 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.0 | 5.6 | GO:0016607 | nuclear speck(GO:0016607) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.7 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.1 | 3.3 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 1.0 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.1 | 1.2 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.1 | 0.3 | GO:0004809 | tRNA (guanine-N2-)-methyltransferase activity(GO:0004809) |
| 0.1 | 0.3 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.1 | 0.7 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.1 | 0.4 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.1 | 1.5 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.6 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.2 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.0 | 1.3 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.0 | 0.6 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.0 | 0.2 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.0 | 0.4 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.0 | 0.7 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.0 | 0.6 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.0 | 0.3 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 2.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 3.8 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.2 | GO:0000990 | transcription factor activity, core RNA polymerase binding(GO:0000990) |
| 0.0 | 0.3 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.6 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 1.8 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 0.2 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.0 | 0.1 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.0 | 0.3 | GO:0015250 | water channel activity(GO:0015250) |
| 0.0 | 0.2 | GO:0015288 | porin activity(GO:0015288) |
| 0.0 | 0.6 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 1.2 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 0.1 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.0 | 0.5 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 0.1 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.0 | 0.2 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.0 | 0.2 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |