Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
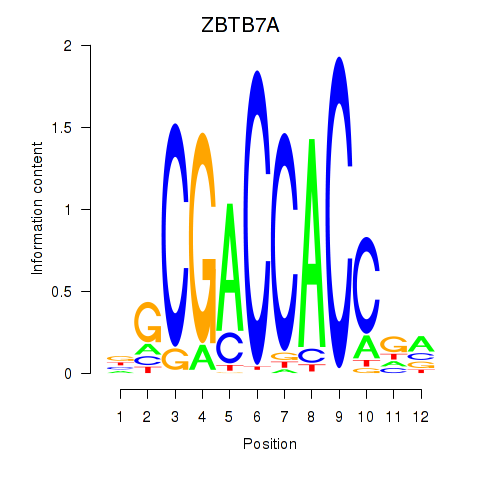
Results for ZBTB7A_ZBTB7C
Z-value: 0.83
Transcription factors associated with ZBTB7A_ZBTB7C
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZBTB7A
|
ENSG00000178951.4 | ZBTB7A |
|
ZBTB7C
|
ENSG00000184828.5 | ZBTB7C |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZBTB7A | hg19_v2_chr19_-_4066890_4066943 | 0.59 | 1.6e-02 | Click! |
| ZBTB7C | hg19_v2_chr18_-_45663666_45663732, hg19_v2_chr18_-_45935663_45935793 | 0.31 | 2.5e-01 | Click! |
Activity profile of ZBTB7A_ZBTB7C motif
Sorted Z-values of ZBTB7A_ZBTB7C motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZBTB7A_ZBTB7C
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_235405168 | 3.88 |
ENST00000339728.3 |
ARL4C |
ADP-ribosylation factor-like 4C |
| chr19_-_50143452 | 2.87 |
ENST00000246792.3 |
RRAS |
related RAS viral (r-ras) oncogene homolog |
| chr14_-_51411194 | 1.63 |
ENST00000544180.2 |
PYGL |
phosphorylase, glycogen, liver |
| chr14_-_51411146 | 1.56 |
ENST00000532462.1 |
PYGL |
phosphorylase, glycogen, liver |
| chr7_-_94285472 | 1.56 |
ENST00000437425.2 ENST00000447873.1 ENST00000415788.2 |
SGCE |
sarcoglycan, epsilon |
| chr7_-_94285402 | 1.52 |
ENST00000428696.2 ENST00000445866.2 |
SGCE |
sarcoglycan, epsilon |
| chr7_-_94285511 | 1.50 |
ENST00000265735.7 |
SGCE |
sarcoglycan, epsilon |
| chr1_-_95007193 | 1.49 |
ENST00000370207.4 ENST00000334047.7 |
F3 |
coagulation factor III (thromboplastin, tissue factor) |
| chr3_-_185542761 | 1.33 |
ENST00000457616.2 ENST00000346192.3 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr3_-_185542817 | 1.31 |
ENST00000382199.2 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr1_+_182992545 | 1.27 |
ENST00000258341.4 |
LAMC1 |
laminin, gamma 1 (formerly LAMB2) |
| chr17_-_39674668 | 1.24 |
ENST00000393981.3 |
KRT15 |
keratin 15 |
| chr14_-_102771462 | 1.23 |
ENST00000522874.1 |
MOK |
MOK protein kinase |
| chr12_-_106641728 | 1.22 |
ENST00000378026.4 |
CKAP4 |
cytoskeleton-associated protein 4 |
| chr12_-_52887034 | 1.16 |
ENST00000330722.6 |
KRT6A |
keratin 6A |
| chr19_+_38755042 | 1.15 |
ENST00000301244.7 |
SPINT2 |
serine peptidase inhibitor, Kunitz type, 2 |
| chr12_-_52867569 | 1.14 |
ENST00000252250.6 |
KRT6C |
keratin 6C |
| chr3_-_136471204 | 1.14 |
ENST00000480733.1 ENST00000383202.2 ENST00000236698.5 ENST00000434713.2 |
STAG1 |
stromal antigen 1 |
| chr19_+_38755203 | 1.14 |
ENST00000587090.1 ENST00000454580.3 |
SPINT2 |
serine peptidase inhibitor, Kunitz type, 2 |
| chr14_-_102771516 | 1.07 |
ENST00000524214.1 ENST00000193029.6 ENST00000361847.2 |
MOK |
MOK protein kinase |
| chr10_-_126849068 | 1.05 |
ENST00000494626.2 ENST00000337195.5 |
CTBP2 |
C-terminal binding protein 2 |
| chr1_+_169075554 | 0.93 |
ENST00000367815.4 |
ATP1B1 |
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr17_-_39769005 | 0.93 |
ENST00000301653.4 ENST00000593067.1 |
KRT16 |
keratin 16 |
| chr11_+_133938955 | 0.91 |
ENST00000534549.1 ENST00000441717.3 |
JAM3 |
junctional adhesion molecule 3 |
| chr11_+_133938820 | 0.90 |
ENST00000299106.4 ENST00000529443.2 |
JAM3 |
junctional adhesion molecule 3 |
| chrX_-_119603138 | 0.83 |
ENST00000200639.4 ENST00000371335.4 ENST00000538785.1 ENST00000434600.2 |
LAMP2 |
lysosomal-associated membrane protein 2 |
| chrX_-_51812268 | 0.78 |
ENST00000486010.1 ENST00000497164.1 ENST00000360134.6 ENST00000485287.1 ENST00000335504.5 ENST00000431659.1 |
MAGED4B |
melanoma antigen family D, 4B |
| chr13_+_110959598 | 0.76 |
ENST00000360467.5 |
COL4A2 |
collagen, type IV, alpha 2 |
| chr6_+_139456226 | 0.66 |
ENST00000367658.2 |
HECA |
headcase homolog (Drosophila) |
| chr15_+_92396920 | 0.63 |
ENST00000318445.6 |
SLCO3A1 |
solute carrier organic anion transporter family, member 3A1 |
| chr11_-_1330834 | 0.61 |
ENST00000525159.1 ENST00000317204.6 ENST00000542915.1 ENST00000527938.1 ENST00000530541.1 ENST00000263646.7 |
TOLLIP |
toll interacting protein |
| chr1_+_15480197 | 0.61 |
ENST00000400796.3 ENST00000434578.2 ENST00000376008.2 |
TMEM51 |
transmembrane protein 51 |
| chr9_+_137533615 | 0.49 |
ENST00000371817.3 |
COL5A1 |
collagen, type V, alpha 1 |
| chr12_+_13197218 | 0.48 |
ENST00000197268.8 |
KIAA1467 |
KIAA1467 |
| chr20_+_44034804 | 0.45 |
ENST00000357275.2 ENST00000372720.3 |
DBNDD2 |
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr20_+_44034676 | 0.45 |
ENST00000372723.3 ENST00000372722.3 |
DBNDD2 |
dysbindin (dystrobrevin binding protein 1) domain containing 2 |
| chr7_+_116312411 | 0.44 |
ENST00000456159.1 ENST00000397752.3 ENST00000318493.6 |
MET |
met proto-oncogene |
| chr17_-_8534031 | 0.42 |
ENST00000411957.1 ENST00000396239.1 ENST00000379980.4 |
MYH10 |
myosin, heavy chain 10, non-muscle |
| chr14_-_24584138 | 0.42 |
ENST00000558280.1 ENST00000561028.1 |
NRL |
neural retina leucine zipper |
| chrX_-_153141302 | 0.42 |
ENST00000361699.4 ENST00000543994.1 ENST00000370057.3 ENST00000538883.1 ENST00000361981.3 |
L1CAM |
L1 cell adhesion molecule |
| chr10_-_16859361 | 0.42 |
ENST00000377921.3 |
RSU1 |
Ras suppressor protein 1 |
| chr21_-_45079341 | 0.41 |
ENST00000443485.1 ENST00000291560.2 |
HSF2BP |
heat shock transcription factor 2 binding protein |
| chr4_+_39184024 | 0.40 |
ENST00000399820.3 ENST00000509560.1 ENST00000512112.1 ENST00000288634.7 ENST00000506503.1 |
WDR19 |
WD repeat domain 19 |
| chr14_+_59655369 | 0.39 |
ENST00000360909.3 ENST00000351081.1 ENST00000556135.1 |
DAAM1 |
dishevelled associated activator of morphogenesis 1 |
| chr17_+_37844331 | 0.38 |
ENST00000578199.1 ENST00000406381.2 |
ERBB2 |
v-erb-b2 avian erythroblastic leukemia viral oncogene homolog 2 |
| chr20_-_17662705 | 0.38 |
ENST00000455029.2 |
RRBP1 |
ribosome binding protein 1 |
| chr9_+_132934835 | 0.37 |
ENST00000372398.3 |
NCS1 |
neuronal calcium sensor 1 |
| chr6_-_122792919 | 0.36 |
ENST00000339697.4 |
SERINC1 |
serine incorporator 1 |
| chr11_-_129872712 | 0.36 |
ENST00000358825.5 ENST00000360871.3 ENST00000528746.1 |
PRDM10 |
PR domain containing 10 |
| chr16_+_15528332 | 0.35 |
ENST00000566490.1 |
C16orf45 |
chromosome 16 open reading frame 45 |
| chr17_-_8534067 | 0.35 |
ENST00000360416.3 ENST00000269243.4 |
MYH10 |
myosin, heavy chain 10, non-muscle |
| chr2_-_97405775 | 0.34 |
ENST00000264963.4 ENST00000537039.1 ENST00000377079.4 ENST00000426463.2 ENST00000534882.1 |
LMAN2L |
lectin, mannose-binding 2-like |
| chr4_-_146101304 | 0.33 |
ENST00000447906.2 |
OTUD4 |
OTU domain containing 4 |
| chr1_+_89246647 | 0.33 |
ENST00000544045.1 |
PKN2 |
protein kinase N2 |
| chr1_+_167691185 | 0.32 |
ENST00000359523.2 |
MPZL1 |
myelin protein zero-like 1 |
| chr2_+_242255297 | 0.30 |
ENST00000401990.1 ENST00000407971.1 ENST00000436795.1 ENST00000411484.1 ENST00000434955.1 ENST00000402092.2 ENST00000441533.1 ENST00000443492.1 ENST00000437066.1 ENST00000429791.1 |
SEPT2 |
septin 2 |
| chr6_-_82462425 | 0.29 |
ENST00000369754.3 ENST00000320172.6 ENST00000369756.3 |
FAM46A |
family with sequence similarity 46, member A |
| chr9_-_131872928 | 0.28 |
ENST00000455830.2 ENST00000393384.3 ENST00000318080.2 |
CRAT |
carnitine O-acetyltransferase |
| chr20_-_36156293 | 0.28 |
ENST00000373537.2 ENST00000414542.2 |
BLCAP |
bladder cancer associated protein |
| chr14_-_54955721 | 0.28 |
ENST00000554908.1 |
GMFB |
glia maturation factor, beta |
| chr4_-_16085314 | 0.28 |
ENST00000510224.1 |
PROM1 |
prominin 1 |
| chr2_-_242255117 | 0.28 |
ENST00000420451.1 ENST00000417540.1 ENST00000310931.4 |
HDLBP |
high density lipoprotein binding protein |
| chr8_-_134584092 | 0.28 |
ENST00000522652.1 |
ST3GAL1 |
ST3 beta-galactoside alpha-2,3-sialyltransferase 1 |
| chr9_+_131873227 | 0.28 |
ENST00000358994.4 ENST00000455292.1 |
PPP2R4 |
protein phosphatase 2A activator, regulatory subunit 4 |
| chr20_+_36149602 | 0.28 |
ENST00000062104.2 ENST00000346199.2 |
NNAT |
neuronatin |
| chr5_-_132112907 | 0.27 |
ENST00000458488.2 |
SEPT8 |
septin 8 |
| chr8_-_134584152 | 0.27 |
ENST00000521180.1 ENST00000517668.1 ENST00000319914.5 |
ST3GAL1 |
ST3 beta-galactoside alpha-2,3-sialyltransferase 1 |
| chr6_+_125474939 | 0.26 |
ENST00000527711.1 |
TPD52L1 |
tumor protein D52-like 1 |
| chr5_+_178286925 | 0.26 |
ENST00000322434.3 |
ZNF354B |
zinc finger protein 354B |
| chr9_-_123476612 | 0.26 |
ENST00000426959.1 |
MEGF9 |
multiple EGF-like-domains 9 |
| chr9_+_35161998 | 0.25 |
ENST00000396787.1 ENST00000378495.3 ENST00000378496.4 |
UNC13B |
unc-13 homolog B (C. elegans) |
| chr4_-_16085340 | 0.25 |
ENST00000508167.1 |
PROM1 |
prominin 1 |
| chr8_-_101964265 | 0.25 |
ENST00000395958.2 |
YWHAZ |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta |
| chr16_+_66400533 | 0.24 |
ENST00000341529.3 |
CDH5 |
cadherin 5, type 2 (vascular endothelium) |
| chr20_-_36156125 | 0.24 |
ENST00000397135.1 ENST00000397137.1 |
BLCAP |
bladder cancer associated protein |
| chr7_+_77167376 | 0.24 |
ENST00000435495.2 |
PTPN12 |
protein tyrosine phosphatase, non-receptor type 12 |
| chr4_-_119274121 | 0.24 |
ENST00000296498.3 |
PRSS12 |
protease, serine, 12 (neurotrypsin, motopsin) |
| chr14_-_61116168 | 0.24 |
ENST00000247182.6 |
SIX1 |
SIX homeobox 1 |
| chr8_+_9413410 | 0.24 |
ENST00000520408.1 ENST00000310430.6 ENST00000522110.1 |
TNKS |
tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase |
| chr4_-_5021164 | 0.24 |
ENST00000506508.1 ENST00000509419.1 ENST00000307746.4 |
CYTL1 |
cytokine-like 1 |
| chr8_+_26240414 | 0.23 |
ENST00000380629.2 |
BNIP3L |
BCL2/adenovirus E1B 19kDa interacting protein 3-like |
| chr22_+_38004832 | 0.23 |
ENST00000405147.3 ENST00000429218.1 ENST00000325180.8 ENST00000337437.4 |
GGA1 |
golgi-associated, gamma adaptin ear containing, ARF binding protein 1 |
| chr21_-_47648665 | 0.22 |
ENST00000450351.1 ENST00000522411.1 ENST00000356396.4 ENST00000397728.3 ENST00000457828.2 |
LSS |
lanosterol synthase (2,3-oxidosqualene-lanosterol cyclase) |
| chr17_-_7080801 | 0.22 |
ENST00000572879.1 |
ASGR1 |
asialoglycoprotein receptor 1 |
| chr20_-_61493115 | 0.22 |
ENST00000335351.3 ENST00000217162.5 |
TCFL5 |
transcription factor-like 5 (basic helix-loop-helix) |
| chr8_-_101964231 | 0.21 |
ENST00000521309.1 |
YWHAZ |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta |
| chr19_+_45971246 | 0.21 |
ENST00000585836.1 ENST00000417353.2 ENST00000353609.3 ENST00000591858.1 ENST00000443841.2 ENST00000590335.1 |
FOSB |
FBJ murine osteosarcoma viral oncogene homolog B |
| chr2_+_242255275 | 0.21 |
ENST00000391971.2 |
SEPT2 |
septin 2 |
| chr6_+_125475335 | 0.20 |
ENST00000532429.1 ENST00000534199.1 |
TPD52L1 |
tumor protein D52-like 1 |
| chr6_-_41715128 | 0.20 |
ENST00000356667.4 ENST00000373025.3 ENST00000425343.2 |
PGC |
progastricsin (pepsinogen C) |
| chr14_+_24583836 | 0.20 |
ENST00000559115.1 ENST00000558215.1 ENST00000557810.1 ENST00000561375.1 ENST00000446197.3 ENST00000559796.1 ENST00000560713.1 ENST00000560901.1 ENST00000559382.1 |
DCAF11 |
DDB1 and CUL4 associated factor 11 |
| chr5_-_137878887 | 0.19 |
ENST00000507939.1 ENST00000572514.1 ENST00000499810.2 ENST00000360541.5 |
ETF1 |
eukaryotic translation termination factor 1 |
| chr14_-_68283291 | 0.19 |
ENST00000555452.1 ENST00000347230.4 |
ZFYVE26 |
zinc finger, FYVE domain containing 26 |
| chr5_-_132112921 | 0.18 |
ENST00000378721.4 ENST00000378701.1 |
SEPT8 |
septin 8 |
| chr20_-_58508702 | 0.17 |
ENST00000357552.3 ENST00000425931.1 |
SYCP2 |
synaptonemal complex protein 2 |
| chr17_+_4736627 | 0.17 |
ENST00000355280.6 ENST00000347992.7 |
MINK1 |
misshapen-like kinase 1 |
| chr5_-_176924562 | 0.17 |
ENST00000359895.2 ENST00000355572.2 ENST00000355841.2 ENST00000393551.1 ENST00000505074.1 ENST00000356618.4 ENST00000393546.4 |
PDLIM7 |
PDZ and LIM domain 7 (enigma) |
| chr17_-_37844267 | 0.16 |
ENST00000579146.1 ENST00000378011.4 ENST00000429199.2 ENST00000300658.4 |
PGAP3 |
post-GPI attachment to proteins 3 |
| chr11_-_66115032 | 0.16 |
ENST00000311181.4 |
B3GNT1 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 1 |
| chr17_-_8868991 | 0.16 |
ENST00000447110.1 |
PIK3R5 |
phosphoinositide-3-kinase, regulatory subunit 5 |
| chr6_+_125474992 | 0.16 |
ENST00000528193.1 |
TPD52L1 |
tumor protein D52-like 1 |
| chr6_-_46459675 | 0.16 |
ENST00000306764.7 |
RCAN2 |
regulator of calcineurin 2 |
| chr12_-_107487604 | 0.16 |
ENST00000008527.5 |
CRY1 |
cryptochrome 1 (photolyase-like) |
| chr17_+_40834580 | 0.16 |
ENST00000264638.4 |
CNTNAP1 |
contactin associated protein 1 |
| chr12_-_56122426 | 0.15 |
ENST00000551173.1 |
CD63 |
CD63 molecule |
| chr5_-_443239 | 0.15 |
ENST00000408966.2 |
C5orf55 |
chromosome 5 open reading frame 55 |
| chr19_-_56988677 | 0.15 |
ENST00000504904.3 ENST00000292069.6 |
ZNF667 |
zinc finger protein 667 |
| chr2_-_152118352 | 0.13 |
ENST00000331426.5 |
RBM43 |
RNA binding motif protein 43 |
| chr22_+_38005033 | 0.13 |
ENST00000447515.1 ENST00000406772.1 ENST00000431745.1 |
GGA1 |
golgi-associated, gamma adaptin ear containing, ARF binding protein 1 |
| chr1_+_167691191 | 0.12 |
ENST00000392121.3 ENST00000474859.1 |
MPZL1 |
myelin protein zero-like 1 |
| chr5_+_175792459 | 0.11 |
ENST00000310389.5 |
ARL10 |
ADP-ribosylation factor-like 10 |
| chr7_-_51384451 | 0.11 |
ENST00000441453.1 ENST00000265136.7 ENST00000395542.2 ENST00000395540.2 |
COBL |
cordon-bleu WH2 repeat protein |
| chr14_+_70233810 | 0.11 |
ENST00000394366.2 ENST00000553548.1 ENST00000553369.1 ENST00000557154.1 ENST00000451983.2 ENST00000553635.1 |
SRSF5 |
serine/arginine-rich splicing factor 5 |
| chr7_+_43152191 | 0.11 |
ENST00000395891.2 |
HECW1 |
HECT, C2 and WW domain containing E3 ubiquitin protein ligase 1 |
| chr5_-_72861484 | 0.11 |
ENST00000296785.3 |
ANKRA2 |
ankyrin repeat, family A (RFXANK-like), 2 |
| chr11_-_14380664 | 0.11 |
ENST00000545643.1 ENST00000256196.4 |
RRAS2 |
related RAS viral (r-ras) oncogene homolog 2 |
| chr1_-_21616901 | 0.11 |
ENST00000436918.2 ENST00000374893.6 |
ECE1 |
endothelin converting enzyme 1 |
| chr3_+_15468862 | 0.10 |
ENST00000396842.2 |
EAF1 |
ELL associated factor 1 |
| chr20_+_37590942 | 0.10 |
ENST00000373325.2 ENST00000252011.3 ENST00000373323.4 |
DHX35 |
DEAH (Asp-Glu-Ala-His) box polypeptide 35 |
| chr17_-_58469474 | 0.10 |
ENST00000300896.4 |
USP32 |
ubiquitin specific peptidase 32 |
| chr19_+_5904866 | 0.10 |
ENST00000339485.3 |
VMAC |
vimentin-type intermediate filament associated coiled-coil protein |
| chr3_+_50126341 | 0.09 |
ENST00000347869.3 ENST00000469838.1 ENST00000404526.2 ENST00000441305.1 |
RBM5 |
RNA binding motif protein 5 |
| chr19_-_49622348 | 0.09 |
ENST00000408991.2 |
C19orf73 |
chromosome 19 open reading frame 73 |
| chr9_+_74526384 | 0.09 |
ENST00000334731.2 ENST00000377031.3 |
C9orf85 |
chromosome 9 open reading frame 85 |
| chr16_-_89724064 | 0.09 |
ENST00000535997.2 ENST00000253475.5 ENST00000397901.3 |
CHMP1A |
charged multivesicular body protein 1A |
| chr5_-_140070897 | 0.09 |
ENST00000448240.1 ENST00000438307.2 ENST00000415192.2 ENST00000457527.2 ENST00000307633.3 ENST00000507746.1 ENST00000431330.2 |
HARS |
histidyl-tRNA synthetase |
| chr5_+_140071011 | 0.09 |
ENST00000230771.3 ENST00000509299.1 ENST00000503873.1 ENST00000435019.2 ENST00000437649.2 ENST00000432671.2 |
HARS2 |
histidyl-tRNA synthetase 2, mitochondrial |
| chr19_-_46296011 | 0.09 |
ENST00000377735.3 ENST00000270223.6 |
DMWD |
dystrophia myotonica, WD repeat containing |
| chr20_-_61557821 | 0.08 |
ENST00000354665.4 ENST00000370368.1 ENST00000395343.1 ENST00000395340.1 |
DIDO1 |
death inducer-obliterator 1 |
| chr17_-_3704500 | 0.08 |
ENST00000263087.4 |
ITGAE |
integrin, alpha E (antigen CD103, human mucosal lymphocyte antigen 1; alpha polypeptide) |
| chrX_+_133507327 | 0.08 |
ENST00000332070.3 ENST00000394292.1 ENST00000370799.1 ENST00000416404.2 |
PHF6 |
PHD finger protein 6 |
| chr10_+_70091847 | 0.08 |
ENST00000441000.2 ENST00000354695.5 |
HNRNPH3 |
heterogeneous nuclear ribonucleoprotein H3 (2H9) |
| chr3_+_46618727 | 0.07 |
ENST00000296145.5 |
TDGF1 |
teratocarcinoma-derived growth factor 1 |
| chr19_-_14228541 | 0.07 |
ENST00000590853.1 ENST00000308677.4 |
PRKACA |
protein kinase, cAMP-dependent, catalytic, alpha |
| chr4_-_76598326 | 0.06 |
ENST00000503660.1 |
G3BP2 |
GTPase activating protein (SH3 domain) binding protein 2 |
| chr10_-_105615164 | 0.06 |
ENST00000355946.2 ENST00000369774.4 |
SH3PXD2A |
SH3 and PX domains 2A |
| chr4_+_41258786 | 0.06 |
ENST00000503431.1 ENST00000284440.4 ENST00000508768.1 ENST00000512788.1 |
UCHL1 |
ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) |
| chr10_-_101380121 | 0.06 |
ENST00000370495.4 |
SLC25A28 |
solute carrier family 25 (mitochondrial iron transporter), member 28 |
| chr6_-_36953833 | 0.05 |
ENST00000538808.1 ENST00000460219.1 ENST00000373616.5 ENST00000373627.5 |
MTCH1 |
mitochondrial carrier 1 |
| chr4_+_74718906 | 0.05 |
ENST00000226524.3 |
PF4V1 |
platelet factor 4 variant 1 |
| chr8_-_143957213 | 0.05 |
ENST00000519285.1 |
CYP11B1 |
cytochrome P450, family 11, subfamily B, polypeptide 1 |
| chr2_+_25015968 | 0.05 |
ENST00000380834.2 ENST00000473706.1 |
CENPO |
centromere protein O |
| chr4_+_156588350 | 0.05 |
ENST00000296518.7 |
GUCY1A3 |
guanylate cyclase 1, soluble, alpha 3 |
| chr1_+_25664408 | 0.05 |
ENST00000374358.4 |
TMEM50A |
transmembrane protein 50A |
| chr6_-_34113856 | 0.05 |
ENST00000538487.2 |
GRM4 |
glutamate receptor, metabotropic 4 |
| chr4_-_74847800 | 0.05 |
ENST00000296029.3 |
PF4 |
platelet factor 4 |
| chr17_-_4890919 | 0.04 |
ENST00000572543.1 ENST00000381311.5 ENST00000348066.3 ENST00000358183.4 |
CAMTA2 |
calmodulin binding transcription activator 2 |
| chr17_-_7137857 | 0.04 |
ENST00000005340.5 |
DVL2 |
dishevelled segment polarity protein 2 |
| chr9_-_139094988 | 0.04 |
ENST00000371746.3 |
LHX3 |
LIM homeobox 3 |
| chr22_+_21369316 | 0.04 |
ENST00000413302.2 ENST00000402329.3 ENST00000336296.2 ENST00000401443.1 ENST00000443995.3 |
P2RX6 |
purinergic receptor P2X, ligand-gated ion channel, 6 |
| chr11_-_117747607 | 0.04 |
ENST00000540359.1 ENST00000539526.1 |
FXYD6 |
FXYD domain containing ion transport regulator 6 |
| chr6_+_30689401 | 0.04 |
ENST00000396389.1 ENST00000396384.1 |
TUBB |
tubulin, beta class I |
| chr10_+_49514698 | 0.04 |
ENST00000432379.1 ENST00000429041.1 ENST00000374189.1 |
MAPK8 |
mitogen-activated protein kinase 8 |
| chr4_+_95373037 | 0.04 |
ENST00000359265.4 ENST00000437932.1 ENST00000380180.3 ENST00000318007.5 ENST00000450793.1 ENST00000538141.1 ENST00000317968.4 ENST00000512274.1 ENST00000503974.1 ENST00000504489.1 ENST00000542407.1 |
PDLIM5 |
PDZ and LIM domain 5 |
| chr7_+_77167343 | 0.04 |
ENST00000433369.2 ENST00000415482.2 |
PTPN12 |
protein tyrosine phosphatase, non-receptor type 12 |
| chr22_-_46931191 | 0.04 |
ENST00000454637.1 |
CELSR1 |
cadherin, EGF LAG seven-pass G-type receptor 1 |
| chr11_-_64013663 | 0.03 |
ENST00000392210.2 |
PPP1R14B |
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr22_+_38035459 | 0.03 |
ENST00000357436.4 |
SH3BP1 |
SH3-domain binding protein 1 |
| chr19_+_45394477 | 0.03 |
ENST00000252487.5 ENST00000405636.2 ENST00000592434.1 ENST00000426677.2 ENST00000589649.1 |
TOMM40 |
translocase of outer mitochondrial membrane 40 homolog (yeast) |
| chr1_-_32110467 | 0.03 |
ENST00000440872.2 ENST00000373703.4 |
PEF1 |
penta-EF-hand domain containing 1 |
| chr8_-_74884511 | 0.03 |
ENST00000518127.1 |
TCEB1 |
transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) |
| chr10_+_70091812 | 0.03 |
ENST00000265866.7 |
HNRNPH3 |
heterogeneous nuclear ribonucleoprotein H3 (2H9) |
| chr10_+_101419187 | 0.03 |
ENST00000370489.4 |
ENTPD7 |
ectonucleoside triphosphate diphosphohydrolase 7 |
| chr2_+_25016282 | 0.02 |
ENST00000260662.1 |
CENPO |
centromere protein O |
| chr22_+_41487711 | 0.02 |
ENST00000263253.7 |
EP300 |
E1A binding protein p300 |
| chr8_-_101734170 | 0.01 |
ENST00000522387.1 ENST00000518196.1 |
PABPC1 |
poly(A) binding protein, cytoplasmic 1 |
| chr16_-_49315731 | 0.01 |
ENST00000219197.6 |
CBLN1 |
cerebellin 1 precursor |
| chr8_-_74884482 | 0.01 |
ENST00000520242.1 ENST00000519082.1 |
TCEB1 |
transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) |
| chr8_-_74884399 | 0.01 |
ENST00000520210.1 ENST00000602840.1 |
TCEB1 |
transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) |
| chr6_+_64282447 | 0.01 |
ENST00000370650.2 ENST00000578299.1 |
PTP4A1 |
protein tyrosine phosphatase type IVA, member 1 |
| chr5_+_78407602 | 0.00 |
ENST00000274353.5 ENST00000524080.1 |
BHMT |
betaine--homocysteine S-methyltransferase |
| chr3_+_23986748 | 0.00 |
ENST00000312521.4 |
NR1D2 |
nuclear receptor subfamily 1, group D, member 2 |
| chr1_-_156721502 | 0.00 |
ENST00000357325.5 |
HDGF |
hepatoma-derived growth factor |
| chr14_+_39644387 | 0.00 |
ENST00000553331.1 ENST00000216832.4 |
PNN |
pinin, desmosome associated protein |
| chr3_-_149470229 | 0.00 |
ENST00000473414.1 |
COMMD2 |
COMM domain containing 2 |
| chr1_+_167906056 | 0.00 |
ENST00000367840.3 |
DCAF6 |
DDB1 and CUL4 associated factor 6 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 3.2 | GO:0006015 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 0.6 | 2.3 | GO:0060671 | epithelial cell differentiation involved in embryonic placenta development(GO:0060671) epithelial cell morphogenesis involved in placental branching(GO:0060672) |
| 0.5 | 1.5 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 0.4 | 1.2 | GO:0051710 | regulation of cytolysis in other organism(GO:0051710) |
| 0.3 | 0.8 | GO:0021592 | fourth ventricle development(GO:0021592) third ventricle development(GO:0021678) |
| 0.2 | 0.9 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) positive regulation of potassium ion import(GO:1903288) |
| 0.2 | 0.6 | GO:0061055 | myotome development(GO:0061055) |
| 0.2 | 1.8 | GO:0002318 | myeloid progenitor cell differentiation(GO:0002318) |
| 0.1 | 0.5 | GO:2000768 | glomerular parietal epithelial cell differentiation(GO:0072139) positive regulation of nephron tubule epithelial cell differentiation(GO:2000768) |
| 0.1 | 0.5 | GO:1903224 | regulation of endodermal cell differentiation(GO:1903224) |
| 0.1 | 1.0 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.1 | 0.4 | GO:0045872 | positive regulation of rhodopsin gene expression(GO:0045872) |
| 0.1 | 0.3 | GO:0019254 | carnitine metabolic process, CoA-linked(GO:0019254) |
| 0.1 | 0.4 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) |
| 0.1 | 2.9 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.1 | 3.9 | GO:0032456 | endocytic recycling(GO:0032456) |
| 0.1 | 0.2 | GO:2000302 | positive regulation of synaptic vesicle exocytosis(GO:2000302) |
| 0.1 | 0.2 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.1 | 0.4 | GO:1903976 | negative regulation of glial cell migration(GO:1903976) |
| 0.1 | 0.8 | GO:1905146 | protein targeting to lysosome involved in chaperone-mediated autophagy(GO:0061740) lysosomal protein catabolic process(GO:1905146) |
| 0.1 | 0.8 | GO:0038063 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) |
| 0.1 | 0.9 | GO:0051546 | keratinocyte migration(GO:0051546) |
| 0.1 | 0.2 | GO:0006427 | histidyl-tRNA aminoacylation(GO:0006427) |
| 0.0 | 0.4 | GO:0033088 | negative regulation of immature T cell proliferation in thymus(GO:0033088) |
| 0.0 | 1.3 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.6 | GO:0033235 | positive regulation of protein sumoylation(GO:0033235) |
| 0.0 | 0.3 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) |
| 0.0 | 0.2 | GO:0035694 | mitochondrial protein catabolic process(GO:0035694) |
| 0.0 | 0.3 | GO:2000587 | negative regulation of platelet-derived growth factor receptor-beta signaling pathway(GO:2000587) |
| 0.0 | 0.2 | GO:0002760 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antimicrobial humoral response(GO:0002760) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.0 | 0.6 | GO:0015732 | prostaglandin transport(GO:0015732) |
| 0.0 | 0.1 | GO:0045014 | carbon catabolite repression of transcription(GO:0045013) negative regulation of transcription by glucose(GO:0045014) |
| 0.0 | 0.4 | GO:0015825 | L-serine transport(GO:0015825) |
| 0.0 | 0.2 | GO:1901552 | positive regulation of endothelial cell development(GO:1901552) positive regulation of establishment of endothelial barrier(GO:1903142) |
| 0.0 | 0.1 | GO:0001757 | somite specification(GO:0001757) |
| 0.0 | 0.3 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.0 | 0.1 | GO:0010816 | substance P catabolic process(GO:0010814) calcitonin catabolic process(GO:0010816) endothelin maturation(GO:0034959) |
| 0.0 | 0.2 | GO:2000323 | negative regulation of glucocorticoid receptor signaling pathway(GO:2000323) |
| 0.0 | 0.2 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.9 | GO:0045104 | intermediate filament cytoskeleton organization(GO:0045104) |
| 0.0 | 0.6 | GO:0097503 | sialylation(GO:0097503) |
| 0.0 | 0.1 | GO:0007412 | axon target recognition(GO:0007412) |
| 0.0 | 0.1 | GO:0048250 | mitochondrial iron ion transport(GO:0048250) |
| 0.0 | 0.1 | GO:0072674 | multinuclear osteoclast differentiation(GO:0072674) osteoclast fusion(GO:0072675) |
| 0.0 | 0.3 | GO:0034384 | high-density lipoprotein particle clearance(GO:0034384) |
| 0.0 | 0.2 | GO:0035112 | genitalia morphogenesis(GO:0035112) |
| 0.0 | 0.3 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.0 | 0.2 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 0.0 | 0.0 | GO:1902595 | regulation of DNA replication origin binding(GO:1902595) |
| 0.0 | 4.1 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.0 | 1.2 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 0.0 | GO:0060448 | dichotomous subdivision of terminal units involved in lung branching(GO:0060448) |
| 0.0 | 0.5 | GO:0030262 | apoptotic nuclear changes(GO:0030262) |
| 0.0 | 0.0 | GO:0045918 | negative regulation of cytolysis(GO:0045918) |
| 0.0 | 2.2 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.0 | 0.1 | GO:0090168 | Golgi reassembly(GO:0090168) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.9 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.1 | 3.2 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 1.6 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 1.6 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.0 | 1.4 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.6 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 0.6 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.0 | 3.0 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.4 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 1.4 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 0.1 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.0 | 2.0 | REACTOME DIABETES PATHWAYS | Genes involved in Diabetes pathways |
| 0.0 | 0.2 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 3.2 | GO:0008184 | purine nucleobase binding(GO:0002060) glycogen phosphorylase activity(GO:0008184) |
| 0.2 | 0.6 | GO:0005150 | interleukin-1, Type I receptor binding(GO:0005150) |
| 0.1 | 1.0 | GO:0030267 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.1 | 3.9 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 2.6 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.1 | 0.9 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.1 | 0.2 | GO:0004821 | histidine-tRNA ligase activity(GO:0004821) |
| 0.0 | 0.5 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.0 | 0.6 | GO:0003836 | beta-galactoside (CMP) alpha-2,3-sialyltransferase activity(GO:0003836) |
| 0.0 | 2.3 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.0 | 2.9 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 0.3 | GO:0016406 | carnitine O-acyltransferase activity(GO:0016406) |
| 0.0 | 0.3 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.0 | 0.4 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.0 | 0.2 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.4 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.0 | 0.2 | GO:0004873 | asialoglycoprotein receptor activity(GO:0004873) |
| 0.0 | 0.2 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.0 | 0.4 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 0.4 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.0 | 2.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 2.0 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.3 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.0 | 3.1 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.3 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 0.6 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.1 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.0 | 0.2 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 0.2 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 0.3 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.0 | 0.0 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.0 | 0.1 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 0.2 | GO:0009881 | photoreceptor activity(GO:0009881) |
| 0.0 | 0.2 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.9 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 1.3 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 1.3 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 1.8 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.5 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.7 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 0.5 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 0.4 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 4.6 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.3 | 1.3 | GO:0043259 | laminin-1 complex(GO:0005606) laminin-10 complex(GO:0043259) laminin-11 complex(GO:0043260) |
| 0.2 | 0.8 | GO:0031166 | integral component of vacuolar membrane(GO:0031166) |
| 0.2 | 0.5 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.2 | 2.0 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.2 | 0.8 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.1 | 0.6 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 0.1 | 1.3 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.1 | 0.8 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.1 | 0.4 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.1 | 1.5 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.1 | 2.4 | GO:0097546 | ciliary base(GO:0097546) |
| 0.1 | 1.2 | GO:0042599 | lamellar body(GO:0042599) |
| 0.1 | 0.2 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.1 | 2.3 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.9 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.0 | 0.5 | GO:0097227 | sperm annulus(GO:0097227) |
| 0.0 | 0.5 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 0.5 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 3.9 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 0.1 | GO:0031310 | intrinsic component of vacuolar membrane(GO:0031310) |
| 0.0 | 2.0 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.4 | GO:0031045 | dense core granule(GO:0031045) |
| 0.0 | 0.2 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.0 | 0.1 | GO:1990393 | 3M complex(GO:1990393) |
| 0.0 | 0.2 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.0 | 3.2 | GO:1904813 | ficolin-1-rich granule(GO:0101002) ficolin-1-rich granule lumen(GO:1904813) |
| 0.0 | 0.0 | GO:0097679 | symbiont-containing vacuole(GO:0020003) symbiont-containing vacuole membrane(GO:0020005) other organism cytoplasm(GO:0097679) |
| 0.0 | 0.2 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.0 | 0.2 | GO:0000800 | lateral element(GO:0000800) |
| 0.0 | 0.2 | GO:0097487 | multivesicular body, internal vesicle(GO:0097487) |
| 0.0 | 0.4 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 0.9 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |