Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
Results for ZNF274
Z-value: 0.53
Transcription factors associated with ZNF274
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZNF274
|
ENSG00000171606.13 | ZNF274 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZNF274 | hg19_v2_chr19_+_58694396_58694485 | 0.11 | 6.8e-01 | Click! |
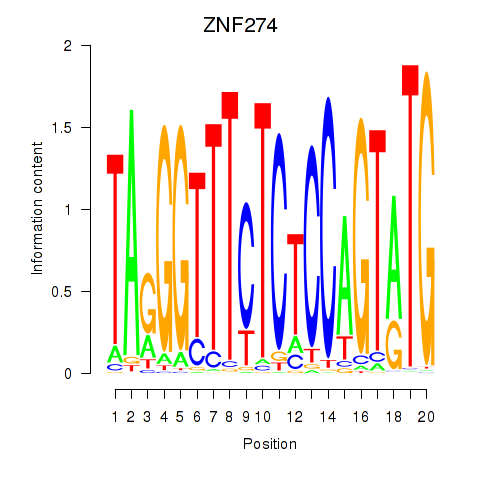
Activity profile of ZNF274 motif
Sorted Z-values of ZNF274 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZNF274
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_40195101 | 1.41 |
ENST00000360675.3 ENST00000601802.1 |
LGALS14 |
lectin, galactoside-binding, soluble, 14 |
| chr19_+_40194946 | 1.39 |
ENST00000392052.3 |
LGALS14 |
lectin, galactoside-binding, soluble, 14 |
| chr1_-_114414316 | 0.80 |
ENST00000528414.1 ENST00000538253.1 ENST00000460620.1 ENST00000420377.2 ENST00000525799.1 ENST00000359785.5 |
PTPN22 |
protein tyrosine phosphatase, non-receptor type 22 (lymphoid) |
| chr19_+_6887571 | 0.74 |
ENST00000250572.8 ENST00000381407.5 ENST00000312053.4 ENST00000450315.3 ENST00000381404.4 |
EMR1 |
egf-like module containing, mucin-like, hormone receptor-like 1 |
| chr1_+_45205478 | 0.69 |
ENST00000452259.1 ENST00000372224.4 |
KIF2C |
kinesin family member 2C |
| chr2_-_89292422 | 0.68 |
ENST00000495489.1 |
IGKV1-8 |
immunoglobulin kappa variable 1-8 |
| chr12_+_113344755 | 0.68 |
ENST00000550883.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr1_+_45205498 | 0.66 |
ENST00000372218.4 |
KIF2C |
kinesin family member 2C |
| chr6_-_32784687 | 0.61 |
ENST00000447394.1 ENST00000438763.2 |
HLA-DOB |
major histocompatibility complex, class II, DO beta |
| chr18_+_9103957 | 0.58 |
ENST00000400033.1 |
NDUFV2 |
NADH dehydrogenase (ubiquinone) flavoprotein 2, 24kDa |
| chr16_+_29819096 | 0.52 |
ENST00000568411.1 ENST00000563012.1 ENST00000562557.1 |
MAZ |
MYC-associated zinc finger protein (purine-binding transcription factor) |
| chr16_+_29819446 | 0.49 |
ENST00000568282.1 |
MAZ |
MYC-associated zinc finger protein (purine-binding transcription factor) |
| chr16_+_29819372 | 0.48 |
ENST00000568544.1 ENST00000569978.1 |
MAZ |
MYC-associated zinc finger protein (purine-binding transcription factor) |
| chr12_-_54653313 | 0.43 |
ENST00000550411.1 ENST00000439541.2 |
CBX5 |
chromobox homolog 5 |
| chr10_+_5488564 | 0.42 |
ENST00000449083.1 ENST00000380359.3 |
NET1 |
neuroepithelial cell transforming 1 |
| chr16_+_29818857 | 0.42 |
ENST00000567444.1 |
MAZ |
MYC-associated zinc finger protein (purine-binding transcription factor) |
| chr5_+_32531893 | 0.36 |
ENST00000512913.1 |
SUB1 |
SUB1 homolog (S. cerevisiae) |
| chr17_-_41132410 | 0.35 |
ENST00000409446.3 ENST00000453594.1 ENST00000409399.1 ENST00000421990.2 |
PTGES3L PTGES3L-AARSD1 |
prostaglandin E synthase 3 (cytosolic)-like PTGES3L-AARSD1 readthrough |
| chrX_+_119737806 | 0.31 |
ENST00000371317.5 |
MCTS1 |
malignant T cell amplified sequence 1 |
| chr2_-_96971259 | 0.28 |
ENST00000349783.5 |
SNRNP200 |
small nuclear ribonucleoprotein 200kDa (U5) |
| chr6_+_30687978 | 0.25 |
ENST00000327892.8 ENST00000435534.1 |
TUBB |
tubulin, beta class I |
| chr11_-_10590118 | 0.23 |
ENST00000529598.1 |
LYVE1 |
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr4_-_121993673 | 0.22 |
ENST00000379692.4 |
NDNF |
neuron-derived neurotrophic factor |
| chr11_+_16760161 | 0.19 |
ENST00000524439.1 ENST00000422258.2 ENST00000528634.1 ENST00000525684.1 |
C11orf58 |
chromosome 11 open reading frame 58 |
| chr15_+_89164520 | 0.15 |
ENST00000332810.3 |
AEN |
apoptosis enhancing nuclease |
| chr19_-_19774473 | 0.13 |
ENST00000357324.6 |
ATP13A1 |
ATPase type 13A1 |
| chr17_+_41132564 | 0.11 |
ENST00000361677.1 ENST00000589705.1 |
RUNDC1 |
RUN domain containing 1 |
| chr2_+_11864458 | 0.11 |
ENST00000396098.1 ENST00000396099.1 ENST00000425416.2 |
LPIN1 |
lipin 1 |
| chr6_+_31553978 | 0.09 |
ENST00000376096.1 ENST00000376099.1 ENST00000376110.3 |
LST1 |
leukocyte specific transcript 1 |
| chr6_+_31553901 | 0.07 |
ENST00000418507.2 ENST00000438075.2 ENST00000376100.3 ENST00000376111.4 |
LST1 |
leukocyte specific transcript 1 |
| chr19_+_6135646 | 0.06 |
ENST00000588304.1 ENST00000588485.1 ENST00000588722.1 ENST00000591403.1 ENST00000586696.1 ENST00000589401.1 ENST00000252669.5 |
ACSBG2 |
acyl-CoA synthetase bubblegum family member 2 |
| chr5_-_74162605 | 0.04 |
ENST00000389156.4 ENST00000510496.1 ENST00000380515.3 |
FAM169A |
family with sequence similarity 169, member A |
| chr17_-_3461092 | 0.04 |
ENST00000301365.4 ENST00000572519.1 |
TRPV3 |
transient receptor potential cation channel, subfamily V, member 3 |
| chr2_+_159313452 | 0.03 |
ENST00000389757.3 ENST00000389759.3 |
PKP4 |
plakophilin 4 |
| chr7_-_73038822 | 0.02 |
ENST00000414749.2 ENST00000429400.2 ENST00000434326.1 |
MLXIPL |
MLX interacting protein-like |
| chr22_-_43036607 | 0.01 |
ENST00000505920.1 |
ATP5L2 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit G2 |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.1 | 0.7 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.0 | 0.6 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.0 | 0.1 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.0 | 0.6 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.4 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 1.9 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.0 | 0.1 | GO:0005384 | manganese ion transmembrane transporter activity(GO:0005384) |
| 0.0 | 0.2 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.2 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:1902523 | positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 0.2 | 0.6 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 0.2 | 2.8 | GO:0070234 | positive regulation of T cell apoptotic process(GO:0070234) |
| 0.1 | 1.4 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.1 | 0.3 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.1 | 0.3 | GO:0000354 | cis assembly of pre-catalytic spliceosome(GO:0000354) |
| 0.0 | 0.4 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.0 | 1.9 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.0 | 0.2 | GO:0061042 | vascular wound healing(GO:0061042) |
| 0.0 | 0.1 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.0 | 0.1 | GO:0071421 | manganese ion transmembrane transport(GO:0071421) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0045272 | plasma membrane respiratory chain complex I(GO:0045272) |
| 0.0 | 1.4 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.0 | 0.4 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.6 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.0 | 0.2 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.8 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 2.8 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 0.2 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |