Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
Results for ZNF8
Z-value: 0.33
Transcription factors associated with ZNF8
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZNF8
|
ENSG00000083842.8 | ZNF8 |
|
ZNF8
|
ENSG00000273439.1 | ZNF8 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZNF8 | hg19_v2_chr19_+_58790314_58790358 | 0.09 | 7.4e-01 | Click! |
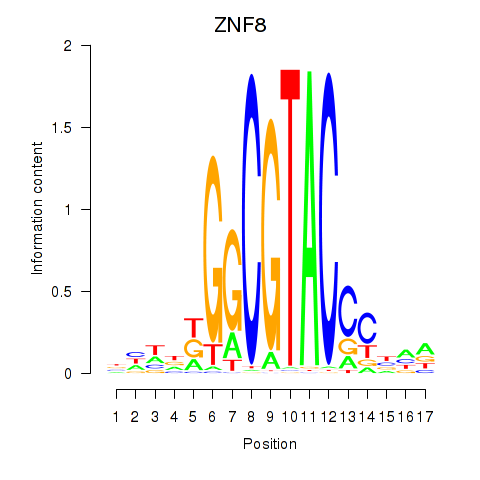
Activity profile of ZNF8 motif
Sorted Z-values of ZNF8 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZNF8
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_106959718 | 0.60 |
ENST00000369066.3 |
AIM1 |
absent in melanoma 1 |
| chrX_+_128913906 | 0.49 |
ENST00000356892.3 |
SASH3 |
SAM and SH3 domain containing 3 |
| chr6_+_29691198 | 0.45 |
ENST00000440587.2 ENST00000434407.2 |
HLA-F |
major histocompatibility complex, class I, F |
| chr6_+_29691056 | 0.44 |
ENST00000414333.1 ENST00000334668.4 ENST00000259951.7 |
HLA-F |
major histocompatibility complex, class I, F |
| chr12_-_46662772 | 0.35 |
ENST00000549049.1 ENST00000439706.1 ENST00000398637.5 |
SLC38A1 |
solute carrier family 38, member 1 |
| chr2_-_172290482 | 0.34 |
ENST00000442541.1 ENST00000392599.2 ENST00000375258.4 |
METTL8 |
methyltransferase like 8 |
| chr19_+_35739782 | 0.27 |
ENST00000347609.4 |
LSR |
lipolysis stimulated lipoprotein receptor |
| chr19_+_35739897 | 0.27 |
ENST00000605618.1 ENST00000427250.1 ENST00000601623.1 |
LSR |
lipolysis stimulated lipoprotein receptor |
| chr17_+_75181292 | 0.25 |
ENST00000431431.2 |
SEC14L1 |
SEC14-like 1 (S. cerevisiae) |
| chr6_+_53659746 | 0.24 |
ENST00000370888.1 |
LRRC1 |
leucine rich repeat containing 1 |
| chr22_+_37415676 | 0.22 |
ENST00000401419.3 |
MPST |
mercaptopyruvate sulfurtransferase |
| chr11_-_58345569 | 0.22 |
ENST00000528954.1 ENST00000528489.1 |
LPXN |
leupaxin |
| chr1_+_214161272 | 0.21 |
ENST00000498508.2 ENST00000366958.4 |
PROX1 |
prospero homeobox 1 |
| chr22_+_37415728 | 0.20 |
ENST00000404802.3 |
MPST |
mercaptopyruvate sulfurtransferase |
| chr22_+_37415700 | 0.19 |
ENST00000397129.1 |
MPST |
mercaptopyruvate sulfurtransferase |
| chr15_+_85523671 | 0.18 |
ENST00000310298.4 ENST00000557957.1 |
PDE8A |
phosphodiesterase 8A |
| chr22_+_37415776 | 0.18 |
ENST00000341116.3 ENST00000429360.2 ENST00000404393.1 |
MPST |
mercaptopyruvate sulfurtransferase |
| chr3_-_9994021 | 0.16 |
ENST00000411976.2 ENST00000412055.1 |
PRRT3 |
proline-rich transmembrane protein 3 |
| chr6_-_26056695 | 0.15 |
ENST00000343677.2 |
HIST1H1C |
histone cluster 1, H1c |
| chr7_+_117824086 | 0.15 |
ENST00000249299.2 ENST00000424702.1 |
NAA38 |
N(alpha)-acetyltransferase 38, NatC auxiliary subunit |
| chr5_-_137911049 | 0.13 |
ENST00000297185.3 |
HSPA9 |
heat shock 70kDa protein 9 (mortalin) |
| chr13_+_53226963 | 0.12 |
ENST00000343788.6 ENST00000535397.1 ENST00000310528.8 |
SUGT1 |
SGT1, suppressor of G2 allele of SKP1 (S. cerevisiae) |
| chr10_+_135192695 | 0.11 |
ENST00000368539.4 ENST00000278060.5 ENST00000357296.3 |
PAOX |
polyamine oxidase (exo-N4-amino) |
| chr2_+_169926047 | 0.10 |
ENST00000428522.1 ENST00000450153.1 ENST00000421653.1 |
DHRS9 |
dehydrogenase/reductase (SDR family) member 9 |
| chr1_-_235292250 | 0.10 |
ENST00000366607.4 |
TOMM20 |
translocase of outer mitochondrial membrane 20 homolog (yeast) |
| chr11_-_46638720 | 0.10 |
ENST00000326737.3 |
HARBI1 |
harbinger transposase derived 1 |
| chr7_+_100303676 | 0.10 |
ENST00000303151.4 |
POP7 |
processing of precursor 7, ribonuclease P/MRP subunit (S. cerevisiae) |
| chr17_+_48624450 | 0.10 |
ENST00000006658.6 ENST00000356488.4 ENST00000393244.3 |
SPATA20 |
spermatogenesis associated 20 |
| chr9_-_32552551 | 0.08 |
ENST00000360538.2 ENST00000379858.1 |
TOPORS |
topoisomerase I binding, arginine/serine-rich, E3 ubiquitin protein ligase |
| chr13_+_21750780 | 0.08 |
ENST00000309594.4 |
MRP63 |
mitochondrial ribosomal protein 63 |
| chr10_-_1034237 | 0.08 |
ENST00000381466.1 |
AL359878.1 |
Uncharacterized protein |
| chrX_+_129535937 | 0.07 |
ENST00000305536.6 ENST00000370947.1 |
RBMX2 |
RNA binding motif protein, X-linked 2 |
| chr19_+_37998031 | 0.07 |
ENST00000586138.1 ENST00000588578.1 ENST00000587986.1 |
ZNF793 |
zinc finger protein 793 |
| chr2_+_172290707 | 0.06 |
ENST00000375255.3 ENST00000539783.1 |
DCAF17 |
DDB1 and CUL4 associated factor 17 |
| chr8_-_109260897 | 0.06 |
ENST00000521297.1 ENST00000519030.1 ENST00000521440.1 ENST00000518345.1 ENST00000519627.1 ENST00000220849.5 |
EIF3E |
eukaryotic translation initiation factor 3, subunit E |
| chr2_+_87144738 | 0.06 |
ENST00000559485.1 |
RGPD1 |
RANBP2-like and GRIP domain containing 1 |
| chr2_+_198365122 | 0.05 |
ENST00000604458.1 |
HSPE1-MOB4 |
HSPE1-MOB4 readthrough |
| chr16_+_21964662 | 0.04 |
ENST00000561553.1 ENST00000565331.1 |
UQCRC2 |
ubiquinol-cytochrome c reductase core protein II |
| chr22_+_30163340 | 0.04 |
ENST00000330029.6 ENST00000401406.3 |
UQCR10 |
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr11_+_46638805 | 0.04 |
ENST00000434074.1 ENST00000312040.4 ENST00000451945.1 |
ATG13 |
autophagy related 13 |
| chr4_-_77069573 | 0.04 |
ENST00000264883.3 |
NUP54 |
nucleoporin 54kDa |
| chr19_-_18392422 | 0.03 |
ENST00000252818.3 |
JUND |
jun D proto-oncogene |
| chr1_-_100715372 | 0.03 |
ENST00000370131.3 ENST00000370132.4 |
DBT |
dihydrolipoamide branched chain transacylase E2 |
| chr2_-_240322643 | 0.02 |
ENST00000345617.3 |
HDAC4 |
histone deacetylase 4 |
| chr20_+_37377085 | 0.02 |
ENST00000243903.4 |
ACTR5 |
ARP5 actin-related protein 5 homolog (yeast) |
| chr19_-_5903714 | 0.02 |
ENST00000586349.1 ENST00000585661.1 ENST00000308961.4 ENST00000592634.1 ENST00000418389.2 ENST00000252675.5 |
AC024592.12 NDUFA11 FUT5 |
Uncharacterized protein NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 11, 14.7kDa fucosyltransferase 5 (alpha (1,3) fucosyltransferase) |
| chr8_+_145064233 | 0.01 |
ENST00000529301.1 ENST00000395068.4 |
GRINA |
glutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1 (glutamate binding) |
| chr14_+_51706886 | 0.01 |
ENST00000457354.2 |
TMX1 |
thioredoxin-related transmembrane protein 1 |
| chr5_+_10353780 | 0.01 |
ENST00000449913.2 ENST00000503788.1 ENST00000274140.5 |
MARCH6 |
membrane-associated ring finger (C3HC4) 6, E3 ubiquitin protein ligase |
| chr7_-_99698338 | 0.01 |
ENST00000354230.3 ENST00000425308.1 |
MCM7 |
minichromosome maintenance complex component 7 |
| chr1_-_70671216 | 0.00 |
ENST00000370952.3 |
LRRC40 |
leucine rich repeat containing 40 |
| chr1_+_70671363 | 0.00 |
ENST00000370951.1 ENST00000370950.3 ENST00000405432.1 ENST00000454435.2 |
SRSF11 |
serine/arginine-rich splicing factor 11 |
Gene Ontology Analysis
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.1 | 0.9 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.0 | 0.1 | GO:0031417 | NatC complex(GO:0031417) |
| 0.0 | 0.1 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.1 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0016784 | 3-mercaptopyruvate sulfurtransferase activity(GO:0016784) |
| 0.1 | 0.9 | GO:0046979 | TAP2 binding(GO:0046979) |
| 0.0 | 0.3 | GO:0039552 | RIG-I binding(GO:0039552) |
| 0.0 | 0.2 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.0 | 0.1 | GO:0046592 | polyamine oxidase activity(GO:0046592) spermine:oxygen oxidoreductase (spermidine-forming) activity(GO:0052901) |
| 0.0 | 0.4 | GO:0005283 | sodium:amino acid symporter activity(GO:0005283) |
| 0.0 | 0.1 | GO:0044547 | DNA topoisomerase binding(GO:0044547) |
| 0.0 | 0.1 | GO:0004022 | alcohol dehydrogenase (NAD) activity(GO:0004022) |
| 0.0 | 0.1 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0009440 | cyanate metabolic process(GO:0009439) cyanate catabolic process(GO:0009440) |
| 0.1 | 0.5 | GO:1904274 | tricellular tight junction assembly(GO:1904274) |
| 0.1 | 0.9 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 0.1 | 0.2 | GO:0046619 | optic placode formation involved in camera-type eye formation(GO:0046619) |
| 0.0 | 0.1 | GO:0009447 | putrescine catabolic process(GO:0009447) |
| 0.0 | 0.1 | GO:0035927 | RNA import into mitochondrion(GO:0035927) |
| 0.0 | 0.2 | GO:0050859 | negative regulation of B cell receptor signaling pathway(GO:0050859) |
| 0.0 | 0.5 | GO:0002726 | positive regulation of T cell cytokine production(GO:0002726) |
| 0.0 | 0.3 | GO:0039536 | negative regulation of RIG-I signaling pathway(GO:0039536) |
| 0.0 | 0.1 | GO:0052055 | modulation by symbiont of host molecular function(GO:0052055) modulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052203) modulation by host of symbiont catalytic activity(GO:0052422) |
| 0.0 | 0.2 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.0 | 0.1 | GO:1902037 | negative regulation of hematopoietic stem cell differentiation(GO:1902037) |
| 0.0 | 0.1 | GO:0046549 | retinal cone cell differentiation(GO:0042670) retinal cone cell development(GO:0046549) |
| 0.0 | 0.0 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |