|
chr13_-_40454188
|
1.920
|
|
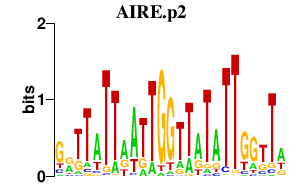
ELF1
|
E74-like factor 1 (ets domain transcription factor)
|
|
chr5_+_75734759
|
1.830
|
NM_006633
|
IQGAP2
|
IQ motif containing GTPase activating protein 2
|
|
chr22_-_34886727
|
1.555
|
NM_145640
|
APOL3
|
apolipoprotein L, 3
|
|
chr13_-_40454360
|
1.543
|
NM_001145353
|
ELF1
|
E74-like factor 1 (ets domain transcription factor)
|
|
chr19_-_53444733
|
1.434
|
NM_001184901
NM_001184903
NM_014959
|
CARD8
|
caspase recruitment domain family, member 8
|
|
chr16_+_20328355
|
1.300
|
NM_017888
|
ACSM5
|
acyl-CoA synthetase medium-chain family member 5
|
|
chr4_+_69716298
|
1.235
|
NM_001075
NM_001144767
|
UGT2B10
|
UDP glucuronosyltransferase 2 family, polypeptide B10
|
|
chr19_+_55627971
|
1.169
|
NM_004533
|
MYBPC2
|
myosin binding protein C, fast type
|
|
chr11_-_73371531
|
1.128
|
NM_003355
|
UCP2
|
uncoupling protein 2 (mitochondrial, proton carrier)
|
|
chr13_-_40491491
|
1.118
|
NM_172373
|
ELF1
|
E74-like factor 1 (ets domain transcription factor)
|
|
chr19_+_18069596
|
1.104
|
NM_015016
|
MAST3
|
microtubule associated serine/threonine kinase 3
|
|
chr4_-_139382672
|
1.016
|
NM_014331
|
SLC7A11
|
solute carrier family 7, (cationic amino acid transporter, y+ system) member 11
|
|
chr16_-_3567354
|
1.008
|
NM_178844
|
NLRC3
|
NLR family, CARD domain containing 3
|
|
chr7_-_149960612
|
0.863
|
NM_024711
|
GIMAP6
|
GTPase, IMAP family member 6
|
|
chr19_-_6621078
|
0.858
|
|
TNFSF14
|
tumor necrosis factor (ligand) superfamily, member 14
|
|
chr2_-_190336168
|
0.852
|
NM_022353
|
OSGEPL1
|
O-sialoglycoprotein endopeptidase-like 1
|
|
chr6_+_10629550
|
0.848
|
NM_145649
|
GCNT2
|
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme (I blood group)
|
|
chr13_-_40491405
|
0.846
|
|
ELF1
|
E74-like factor 1 (ets domain transcription factor)
|
|
chr1_+_153222464
|
0.824
|
|
FLAD1
|
FAD1 flavin adenine dinucleotide synthetase homolog (S. cerevisiae)
|
|
chr19_-_44518516
|
0.817
|
NM_004877
|
GMFG
|
glia maturation factor, gamma
|
|
chr6_+_6533932
|
0.815
|
NM_004271
|
LY86
|
lymphocyte antigen 86
|
|
chr7_+_116238309
|
0.812
|
|
CAPZA2
|
capping protein (actin filament) muscle Z-line, alpha 2
|
|
chr3_-_17757402
|
0.811
|
NM_014744
|
TBC1D5
|
TBC1 domain family, member 5
|
|
chr1_-_89437098
|
0.780
|
NM_052941
|
GBP4
|
guanylate binding protein 4
|
|
chr17_-_59437782
|
0.764
|
NM_000873
|
ICAM2
|
intercellular adhesion molecule 2
|
|
chr9_+_116413443
|
0.754
|
NM_153045
|
C9orf91
|
chromosome 9 open reading frame 91
|
|
chr4_+_88187215
|
0.738
|
|
AFF1
|
AF4/FMR2 family, member 1
|
|
chr17_-_31231421
|
0.730
|
NM_002985
|
CCL5
|
chemokine (C-C motif) ligand 5
|
|
chr19_-_53450953
|
0.718
|
NM_001184902
NM_001184904
|
CARD8
|
caspase recruitment domain family, member 8
|
|
chr11_-_62195587
|
0.696
|
|
C11orf48
|
chromosome 11 open reading frame 48
|
|
chr1_+_158975662
|
0.695
|
NM_021181
|
SLAMF7
|
SLAM family member 7
|
|
chr5_-_149772563
|
0.681
|
|
CD74
|
CD74 molecule, major histocompatibility complex, class II invariant chain
|
|
chr7_-_64104441
|
0.668
|
NM_001007253
|
ERV3
ZNF117
|
endogenous retroviral sequence 3
zinc finger protein 117
|
|
chr11_+_64694200
|
0.653
|
NM_001008778
|
SPDYC
|
speedy homolog C (Xenopus laevis)
|
|
chr2_-_231498184
|
0.649
|
NM_005683
|
GPR55
|
G protein-coupled receptor 55
|
|
chr11_-_104274606
|
0.648
|
NM_001191016
|
CASP12
|
caspase 12 (gene/pseudogene)
|
|
chr8_-_42742729
|
0.640
|
NM_004198
|
CHRNA6
|
cholinergic receptor, nicotinic, alpha 6
|
|
chr15_-_97607108
|
0.640
|
NM_001040655
NM_001040656
NM_001040657
NM_001040658
NM_001040659
NM_001040660
NM_022905
|
TTC23
|
tetratricopeptide repeat domain 23
|
|
chr10_-_32707549
|
0.629
|
|
EPC1
|
enhancer of polycomb homolog 1 (Drosophila)
|
|
chr19_-_6542012
|
0.618
|
|
CD70
|
CD70 molecule
|
|
chr1_+_157067729
|
0.618
|
NM_002432
|
MNDA
|
myeloid cell nuclear differentiation antigen
|
|
chrX_-_100193747
|
0.608
|
NM_001167970
NM_001167971
NM_001167972
NM_024917
|
TRMT2B
|
TRM2 tRNA methyltransferase 2 homolog B (S. cerevisiae)
|
|
chrX_-_100193698
|
0.602
|
|
TRMT2B
|
TRM2 tRNA methyltransferase 2 homolog B (S. cerevisiae)
|
|
chr15_+_39363442
|
0.602
|
|
LOC729082
|
hypothetical LOC729082
|
|
chr2_-_188021117
|
0.599
|
NM_005795
|
CALCRL
|
calcitonin receptor-like
|
|
chr18_+_27425727
|
0.585
|
NM_000371
|
TTR
|
transthyretin
|
|
chr22_+_34979118
|
0.576
|
|
APOL1
|
apolipoprotein L, 1
|
|
chr2_+_215885053
|
0.561
|
|
ATIC
|
5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase
|
|
chr8_-_30704984
|
0.553
|
NM_000637
NM_001195102
NM_001195103
NM_001195104
|
GSR
|
glutathione reductase
|
|
chr19_-_12695628
|
0.551
|
NM_001136195
|
TNPO2
|
transportin 2
|
|
chr1_+_153222393
|
0.549
|
NM_001184891
NM_001184892
NM_025207
NM_201398
|
FLAD1
|
FAD1 flavin adenine dinucleotide synthetase homolog (S. cerevisiae)
|
|
chr22_+_34979062
|
0.533
|
NM_001136540
NM_001136541
NM_003661
NM_145343
|
APOL1
|
apolipoprotein L, 1
|
|
chr6_-_33085329
|
0.521
|
NM_002119
|
HLA-DOA
|
major histocompatibility complex, class II, DO alpha
|
|
chr14_+_101345878
|
0.514
|
NM_002719
NM_178586
NM_178587
|
PPP2R5C
|
protein phosphatase 2, regulatory subunit B', gamma
|
|
chr19_-_10474767
|
0.513
|
NM_203500
|
KEAP1
|
kelch-like ECH-associated protein 1
|
|
chr19_-_44618375
|
0.512
|
NM_001020
|
RPS16
|
ribosomal protein S16
|
|
chr1_-_167947301
|
0.509
|
NM_000655
|
SELL
|
selectin L
|
|
chr2_+_215884923
|
0.504
|
NM_004044
|
ATIC
|
5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase
|
|
chr16_-_11282595
|
0.493
|
NM_002761
|
PRM1
|
protamine 1
|
|
chr4_+_17187936
|
0.476
|
NM_015907
|
LAP3
|
leucine aminopeptidase 3
|
|
chr1_-_32157845
|
0.466
|
NM_001195100
NM_001195101
|
PTP4A2
|
protein tyrosine phosphatase type IVA, member 2
|
|
chr15_+_53398697
|
0.465
|
|
PIGB
|
phosphatidylinositol glycan anchor biosynthesis, class B
|
|
chr2_-_170138558
|
0.461
|
NM_024622
|
FASTKD1
|
FAST kinase domains 1
|
|
chr1_-_9052207
|
0.458
|
|
SLC2A5
|
solute carrier family 2 (facilitated glucose/fructose transporter), member 5
|
|
chr4_-_123761615
|
0.457
|
NM_021803
|
IL21
|
interleukin 21
|
|
chr12_+_49605010
|
0.456
|
|
METTL7A
|
methyltransferase like 7A
|
|
chr12_+_45896318
|
0.453
|
NM_138371
|
FAM113B
|
family with sequence similarity 113, member B
|
|
chr6_-_37575599
|
0.449
|
|
C6orf129
|
chromosome 6 open reading frame 129
|
|
chr4_-_38710376
|
0.439
|
NM_024943
|
TMEM156
|
transmembrane protein 156
|
|
chr1_-_114855281
|
0.438
|
NM_015906
NM_033020
|
TRIM33
|
tripartite motif containing 33
|
|
chr6_+_43005214
|
0.431
|
|
CNPY3
|
canopy 3 homolog (zebrafish)
|
|
chr2_+_215885063
|
0.431
|
|
ATIC
|
5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase
|
|
chr16_+_80370400
|
0.430
|
|
PLCG2
|
phospholipase C, gamma 2 (phosphatidylinositol-specific)
|
|
chr6_+_53043729
|
0.425
|
NM_012347
|
FBXO9
|
F-box protein 9
|
|
chr2_-_176656935
|
0.417
|
NM_001080458
|
EVX2
|
even-skipped homeobox 2
|
|
chr12_-_30740072
|
0.409
|
NM_006390
|
IPO8
|
importin 8
|
|
chr1_-_143708467
|
0.402
|
NM_001195260
|
PDE4DIP
|
phosphodiesterase 4D interacting protein
|
|
chr6_-_99957253
|
0.400
|
|
SFRS18
|
splicing factor, arginine/serine-rich 18
|
|
chr4_+_53986759
|
0.394
|
|
FIP1L1
|
FIP1 like 1 (S. cerevisiae)
|
|
chr3_+_173954940
|
0.392
|
NM_018098
|
ECT2
|
epithelial cell transforming sequence 2 oncogene
|
|
chr21_-_42219849
|
0.392
|
NM_199050
|
C2CD2
|
C2 calcium-dependent domain containing 2
|
|
chr10_+_7785360
|
0.387
|
|
ITIH2
|
inter-alpha (globulin) inhibitor H2
|
|
chr13_-_98708549
|
0.387
|
NM_001098200
NM_005292
|
GPR18
|
G protein-coupled receptor 18
|
|
chr12_-_55405209
|
0.382
|
NM_001113201
NM_005594
|
NACA
|
nascent polypeptide-associated complex alpha subunit
|
|
chr16_+_28993770
|
0.374
|
|
RRN3P2
|
RNA polymerase I transcription factor homolog (S. cerevisiae) pseudogene 2
|
|
chr16_-_727889
|
0.371
|
|
NARFL
|
nuclear prelamin A recognition factor-like
|
|
chr2_+_113911737
|
0.358
|
NM_172003
|
CBWD2
CBWD1
|
COBW domain containing 2
COBW domain containing 1
|
|
chr17_+_39778191
|
0.357
|
|
GRN
|
granulin
|
|
chr8_+_7340777
|
0.357
|
NM_001037668
NM_001040705
|
DEFB107A
DEFB107B
|
defensin, beta 107A
defensin, beta 107B
|
|
chr9_-_85761353
|
0.350
|
|
C9orf64
|
chromosome 9 open reading frame 64
|
|
chr6_+_31910623
|
0.350
|
NM_001040437
NM_001040438
|
C6orf48
|
chromosome 6 open reading frame 48
|
|
chr10_+_7785347
|
0.349
|
|
ITIH2
|
inter-alpha (globulin) inhibitor H2
|
|
chr21_-_18697829
|
0.347
|
NM_002772
|
TMPRSS15
|
transmembrane protease, serine 15
|
|
chr6_-_32892705
|
0.345
|
NM_002120
|
HLA-DOB
|
major histocompatibility complex, class II, DO beta
|
|
chr4_+_39874921
|
0.337
|
NM_004310
|
RHOH
|
ras homolog gene family, member H
|
|
chr15_+_64372876
|
0.335
|
NM_001143688
|
DIS3L
|
DIS3 mitotic control homolog (S. cerevisiae)-like
|
|
chr10_+_7785331
|
0.334
|
|
ITIH2
|
inter-alpha (globulin) inhibitor H2
|
|
chr22_+_24174039
|
0.333
|
|
CRYBB2P1
|
crystallin, beta B2 pseudogene 1
|
|
chr6_-_149847809
|
0.330
|
NM_207360
|
ZC3H12D
|
zinc finger CCCH-type containing 12D
|
|
chr12_+_70119785
|
0.329
|
|
LGR5
|
leucine-rich repeat containing G protein-coupled receptor 5
|
|
chr22_-_16080265
|
0.324
|
|
CECR1
|
cat eye syndrome chromosome region, candidate 1
|
|
chr9_-_124329392
|
0.322
|
NM_012363
|
OR1N1
|
olfactory receptor, family 1, subfamily N, member 1
|
|
chr19_+_17499064
|
0.316
|
|
FAM129C
|
family with sequence similarity 129, member C
|
|
chr6_-_31668476
|
0.316
|
NM_001145466
NM_001145467
NM_147130
|
NCR3
|
natural cytotoxicity triggering receptor 3
|
|
chr10_+_7785290
|
0.313
|
|
ITIH2
|
inter-alpha (globulin) inhibitor H2
|
|
chr6_+_32240357
|
0.309
|
NM_030652
|
EGFL8
|
EGF-like-domain, multiple 8
|
|
chr6_-_11490487
|
0.297
|
NM_001142393
|
NEDD9
|
neural precursor cell expressed, developmentally down-regulated 9
|
|
chrX_-_2892288
|
0.294
|
NM_000047
|
ARSE
|
arylsulfatase E (chondrodysplasia punctata 1)
|
|
chr22_+_34126294
|
0.292
|
|
MCM5
|
minichromosome maintenance complex component 5
|
|
chr13_+_48968097
|
0.292
|
|
PHF11
|
PHD finger protein 11
|
|
chr11_-_73371452
|
0.289
|
|
UCP2
|
uncoupling protein 2 (mitochondrial, proton carrier)
|
|
chr6_+_43005126
|
0.286
|
|
CNPY3
|
canopy 3 homolog (zebrafish)
|
|
chr17_-_18207416
|
0.283
|
NM_004169
NM_148918
|
SHMT1
|
serine hydroxymethyltransferase 1 (soluble)
|
|
chr19_-_16111594
|
0.280
|
|
|
|
|
chrX_+_153310313
|
0.279
|
|
ATP6AP1
|
ATPase, H+ transporting, lysosomal accessory protein 1
|
|
chr2_+_201806410
|
0.277
|
NM_001080124
NM_001228
NM_033355
NM_033358
|
CASP8
|
caspase 8, apoptosis-related cysteine peptidase
|
|
chr6_+_26473376
|
0.276
|
NM_001197247
NM_001197249
NM_007047
NM_001197248
|
BTN3A2
|
butyrophilin, subfamily 3, member A2
|
|
chr8_-_102034777
|
0.274
|
NM_003406
|
YWHAZ
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide
|
|
chr2_-_85490997
|
0.270
|
NM_001747
|
CAPG
|
capping protein (actin filament), gelsolin-like
|
|
chr19_-_14646626
|
0.269
|
NM_032571
|
EMR3
|
egf-like module containing, mucin-like, hormone receptor-like 3
|
|
chr13_-_45640679
|
0.266
|
|
LCP1
|
lymphocyte cytosolic protein 1 (L-plastin)
|
|
chr2_+_68445821
|
0.266
|
NM_002664
|
PLEK
|
pleckstrin
|
|
chr1_+_239762179
|
0.265
|
NM_003679
|
KMO
|
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase)
|
|
chr1_-_9052253
|
0.263
|
NM_001135585
NM_003039
|
SLC2A5
|
solute carrier family 2 (facilitated glucose/fructose transporter), member 5
|
|
chr19_-_8314120
|
0.261
|
NM_198471
|
KANK3
|
KN motif and ankyrin repeat domains 3
|
|
chr7_-_149960218
|
0.261
|
|
GIMAP6
|
GTPase, IMAP family member 6
|
|
chr16_-_73258218
|
0.260
|
|
RFWD3
|
ring finger and WD repeat domain 3
|
|
chr2_-_61098867
|
0.256
|
NM_144709
|
PUS10
|
pseudouridylate synthase 10
|
|
chr6_-_159386007
|
0.255
|
NM_054114
NM_138810
NM_152133
|
TAGAP
|
T-cell activation RhoGTPase activating protein
|
|
chr7_+_2565238
|
0.253
|
|
IQCE
|
IQ motif containing E
|
|
chr19_-_16111695
|
0.253
|
|
|
|
|
chr9_-_98580148
|
0.251
|
NM_014930
|
ZNF510
|
zinc finger protein 510
|
|
chr17_+_38259620
|
0.246
|
|
AOC3
|
amine oxidase, copper containing 3 (vascular adhesion protein 1)
|
|
chr15_-_43458247
|
0.246
|
NM_001482
|
GATM
|
glycine amidinotransferase (L-arginine:glycine amidinotransferase)
|
|
chr16_-_28528780
|
0.244
|
NM_001055
NM_177529
|
SULT1A1
|
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1
|
|
chr17_-_30414789
|
0.244
|
NM_001017368
|
RFFL
|
ring finger and FYVE-like domain containing 1
|
|
chr19_+_59866237
|
0.242
|
|
LILRB4
|
leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 4
|
|
chr19_-_42355407
|
0.241
|
NM_152655
|
ZNF585A
|
zinc finger protein 585A
|
|
chr2_+_232354675
|
0.240
|
|
COPS7B
|
COP9 constitutive photomorphogenic homolog subunit 7B (Arabidopsis)
|
|
chr15_+_20287609
|
0.238
|
NM_001001413
|
GOLGA6L1
|
golgin A6 family-like 1
|
|
chr1_+_84382562
|
0.238
|
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chr4_-_84254912
|
0.237
|
|
PLAC8
|
placenta-specific 8
|
|
chr5_-_156502363
|
0.237
|
NM_001100816
NM_004270
|
MED7
|
mediator complex subunit 7
|
|
chr9_+_70840298
|
0.236
|
NM_000144
NM_001161706
NM_181425
|
FXN
|
frataxin
|
|
chr12_+_49604782
|
0.234
|
NM_014033
|
METTL7A
|
methyltransferase like 7A
|
|
chr19_+_21056792
|
0.233
|
NM_182515
|
ZNF714
|
zinc finger protein 714
|
|
chr8_+_30072463
|
0.233
|
NM_001128208
NM_015344
|
LEPROTL1
|
leptin receptor overlapping transcript-like 1
|
|
chr1_+_51208499
|
0.233
|
|
CDKN2C
|
cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4)
|
|
chr6_-_90119169
|
0.232
|
|
UBE2J1
|
ubiquitin-conjugating enzyme E2, J1 (UBC6 homolog, yeast)
|
|
chr6_-_37575666
|
0.231
|
NM_138493
|
C6orf129
|
chromosome 6 open reading frame 129
|
|
chr6_+_4835224
|
0.231
|
NM_001143971
|
CDYL
|
chromodomain protein, Y-like
|
|
chr1_-_21249974
|
0.230
|
|
EIF4G3
|
eukaryotic translation initiation factor 4 gamma, 3
|
|
chr20_-_18425811
|
0.226
|
NM_006606
|
RBBP9
|
retinoblastoma binding protein 9
|
|
chr12_-_7979896
|
0.226
|
NM_006931
|
SLC2A3
|
solute carrier family 2 (facilitated glucose transporter), member 3
|
|
chr11_+_129484540
|
0.222
|
|
APLP2
|
amyloid beta (A4) precursor-like protein 2
|
|
chr22_+_21359175
|
0.219
|
|
IGLV3-25
|
immunoglobulin lambda variable 3-25
|
|
chr19_-_10475083
|
0.216
|
|
KEAP1
|
kelch-like ECH-associated protein 1
|
|
chr19_+_10223598
|
0.215
|
NM_015956
NM_146387
NM_146388
|
MRPL4
|
mitochondrial ribosomal protein L4
|
|
chr13_-_35327798
|
0.215
|
NM_001195415
NM_001195416
NM_001195430
|
DCLK1
|
doublecortin-like kinase 1
|
|
chr11_-_75057299
|
0.214
|
|
MAP6
|
microtubule-associated protein 6
|
|
chr16_+_20682812
|
0.212
|
NM_005622
NM_202000
|
ACSM3
|
acyl-CoA synthetase medium-chain family member 3
|
|
chr2_+_68445936
|
0.212
|
|
PLEK
|
pleckstrin
|
|
chr4_+_39084867
|
0.211
|
NM_175737
|
KLB
|
klotho beta
|
|
chr16_-_56886380
|
0.210
|
NM_001080492
|
PRSS54
|
protease, serine, 54
|
|
chr2_+_231829021
|
0.209
|
|
|
|
|
chr19_-_6542136
|
0.208
|
NM_001252
|
CD70
|
CD70 molecule
|
|
chr13_+_49100435
|
0.207
|
NM_138450
|
ARL11
|
ADP-ribosylation factor-like 11
|
|
chr14_+_57964462
|
0.206
|
NM_014749
|
KIAA0586
|
KIAA0586
|
|
chr6_-_90119329
|
0.206
|
NM_016021
|
UBE2J1
|
ubiquitin-conjugating enzyme E2, J1 (UBC6 homolog, yeast)
|
|
chr20_-_33658823
|
0.205
|
|
FER1L4
|
fer-1-like 4 (C. elegans) pseudogene
|
|
chr22_-_23281842
|
0.203
|
|
C22orf13
|
chromosome 22 open reading frame 13
|
|
chr17_+_39778153
|
0.203
|
|
GRN
|
granulin
|
|
chr10_-_101831579
|
0.202
|
NM_001308
|
CPN1
|
carboxypeptidase N, polypeptide 1
|
|
chr20_-_31452964
|
0.201
|
|
CDK5RAP1
|
CDK5 regulatory subunit associated protein 1
|
|
chr20_-_10360569
|
0.200
|
NM_018848
|
MKKS
|
McKusick-Kaufman syndrome
|
|
chr4_+_170817787
|
0.197
|
|
CLCN3
|
chloride channel 3
|
|
chr10_+_135057597
|
0.194
|
NM_138384
|
MTG1
|
mitochondrial GTPase 1 homolog (S. cerevisiae)
|
|
chr9_-_129740583
|
0.194
|
NM_003863
|
DPM2
|
dolichyl-phosphate mannosyltransferase polypeptide 2, regulatory subunit
|
|
chr3_-_152403636
|
0.193
|
NM_013308
|
GPR171
|
G protein-coupled receptor 171
|
|
chr17_+_64922420
|
0.192
|
NM_002758
|
MAP2K6
|
mitogen-activated protein kinase kinase 6
|
|
chr4_-_84254873
|
0.192
|
|
PLAC8
|
placenta-specific 8
|
|
chr15_+_57067442
|
0.192
|
|
RNF111
|
ring finger protein 111
|
|
chr18_+_2896178
|
0.191
|
|
EMILIN2
|
elastin microfibril interfacer 2
|
|
chr13_+_96726197
|
0.191
|
|
MBNL2
|
muscleblind-like 2 (Drosophila)
|
|
chr1_+_246635918
|
0.190
|
NM_030904
|
OR2T1
|
olfactory receptor, family 2, subfamily T, member 1
|
|
chr1_-_159543847
|
0.190
|
|
MPZ
|
myelin protein zero
|
|
chr16_+_65144019
|
0.190
|
|
CKLF
|
chemokine-like factor
|
|
chr12_+_108035424
|
0.190
|
|
|
|
|
chr4_-_84254932
|
0.188
|
NM_001130715
NM_016619
|
PLAC8
|
placenta-specific 8
|
|
chr6_+_33352886
|
0.188
|
NM_003782
|
B3GALT4
|
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 4
|
|
chr1_-_89364273
|
0.186
|
NM_004120
|
GBP2
|
guanylate binding protein 2, interferon-inducible
|
|
chr10_+_22645304
|
0.185
|
NM_012071
|
COMMD3-BMI1
COMMD3
|
COMMD3-BMI1 read-through transcript
COMM domain containing 3
|
|
chr17_+_39778180
|
0.184
|
|
GRN
|
granulin
|
|
chr13_-_36471474
|
0.183
|
NM_001142364
NM_013338
|
ALG5
|
asparagine-linked glycosylation 5, dolichyl-phosphate beta-glucosyltransferase homolog (S. cerevisiae)
|
|
chr17_+_63766874
|
0.181
|
NM_014960
|
ARSG
|
arylsulfatase G
|
|
chr19_+_46489987
|
0.178
|
|
HNRNPUL1
|
heterogeneous nuclear ribonucleoprotein U-like 1
|
|
chr9_-_85761451
|
0.178
|
|
C9orf64
|
chromosome 9 open reading frame 64
|
|
chr6_+_74228331
|
0.177
|
|
MTO1
|
mitochondrial translation optimization 1 homolog (S. cerevisiae)
|
|
chrX_-_47815426
|
0.177
|
NM_001037735
|
ZNF630
|
zinc finger protein 630
|
|
chr17_+_63367052
|
0.177
|
|
BPTF
|
bromodomain PHD finger transcription factor
|