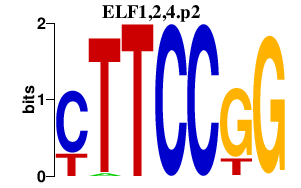
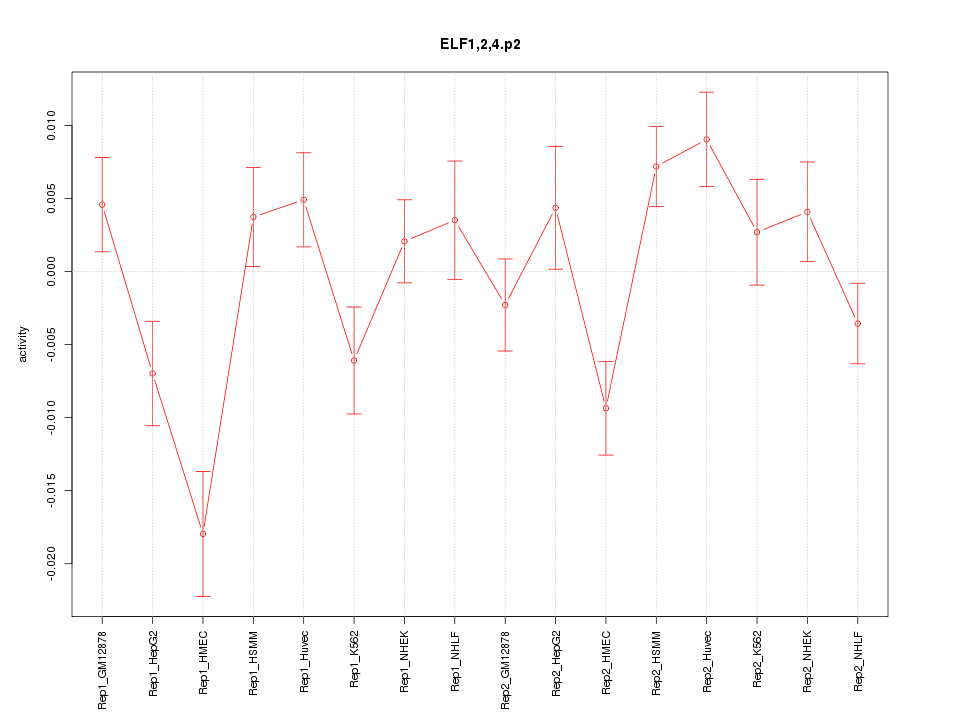
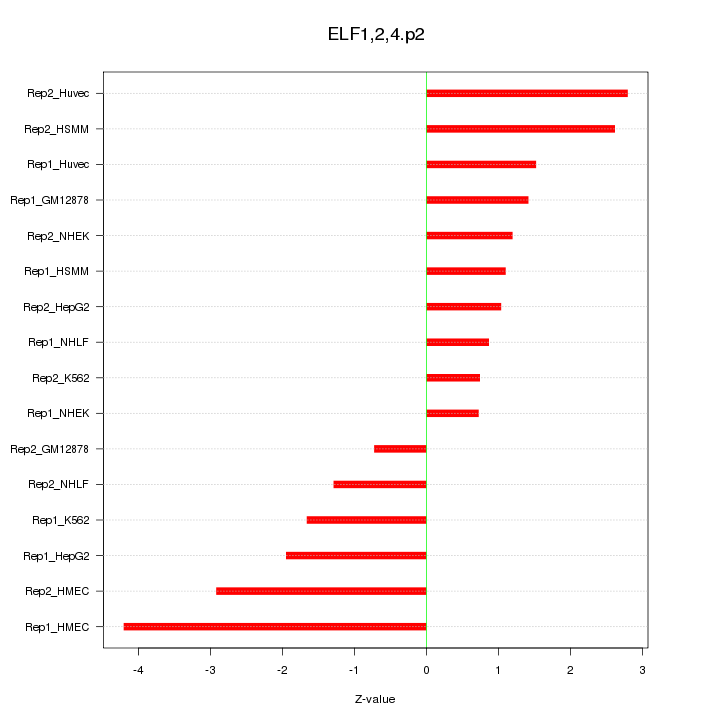
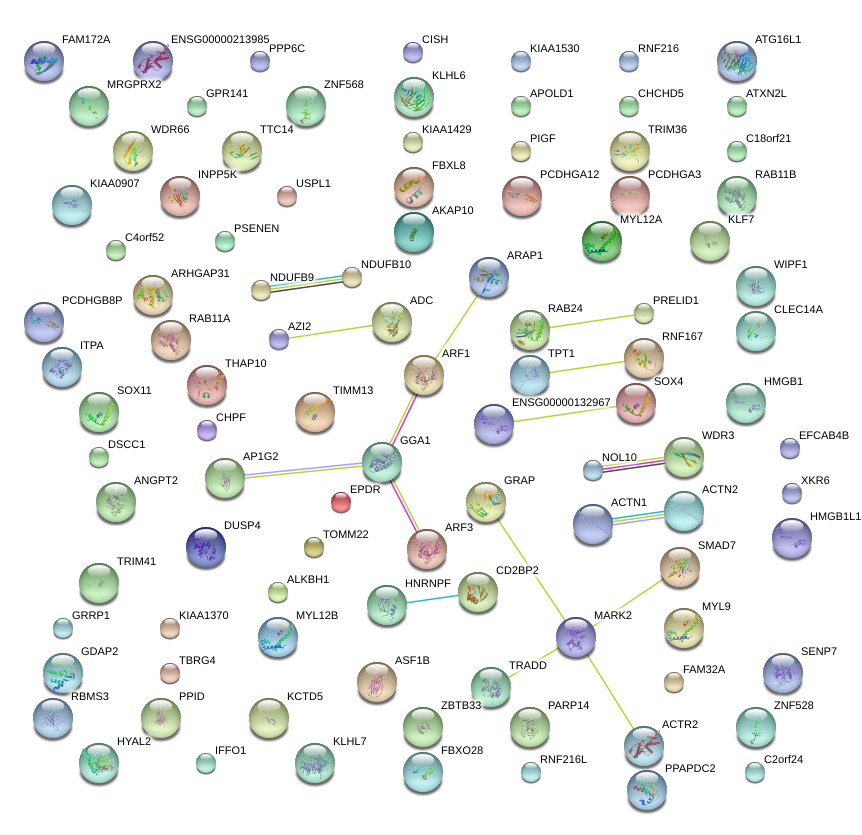
|
chr19_-_42098919
|
1.493
|
NM_001037232
|
ZNF829
|
zinc finger protein 829
|
|
chr19_-_42098758
|
1.353
|
NM_001171979
|
ZNF829
|
zinc finger protein 829
|
|
chr19_+_42099073
|
1.288
|
NM_198539
|
ZNF568
|
zinc finger protein 568
|
|
chr7_-_45117822
|
1.202
|
NM_004749
NM_030900
NM_199122
|
TBRG4
|
transforming growth factor beta regulator 4
|
|
chr17_+_4784524
|
1.148
|
|
RNF167
|
ring finger protein 167
|
|
chr7_-_45117797
|
1.130
|
|
TBRG4
|
transforming growth factor beta regulator 4
|
|
chr15_+_63948823
|
1.071
|
NM_004663
|
RAB11A
|
RAB11A, member RAS oncogene family
|
|
chr22_+_36334400
|
1.013
|
NM_001001560
NM_001001561
NM_001172687
NM_013365
|
GGA1
|
golgi-associated, gamma adaptin ear containing, ARF binding protein 1
|
|
chr12_+_120840829
|
1.011
|
NM_001178003
NM_144668
|
WDR66
|
WD repeat domain 66
|
|
chr3_-_184756101
|
0.995
|
NM_130446
|
KLHL6
|
kelch-like 6 (Drosophila)
|
|
chr1_+_45542138
|
0.980
|
|
LOC400752
|
hypothetical LOC400752
|
|
chr5_+_180582880
|
0.951
|
|
TRIM41
|
tripartite motif containing 41
|
|
chr7_-_5787776
|
0.915
|
NM_207111
NM_207116
|
RNF216
|
ring finger protein 216
|
|
chr19_+_16157198
|
0.909
|
NM_014077
|
FAM32A
|
family with sequence similarity 32, member A
|
|
chr1_+_226337425
|
0.905
|
NM_001024227
NM_001024228
|
ARF1
|
ADP-ribosylation factor 1
|
|
chr13_-_30089509
|
0.901
|
|
HMGB1
|
high-mobility group box 1
|
|
chr5_+_180582905
|
0.883
|
NM_033549
NM_201627
|
TRIM41
|
tripartite motif containing 41
|
|
chr3_+_120495906
|
0.875
|
NM_020754
|
ARHGAP31
|
Rho GTPase activating protein 31
|
|
chr8_-_6407882
|
0.852
|
NM_001118887
NM_001118888
NM_001147
|
ANGPT2
|
angiopoietin 2
|
|
chr16_-_30274064
|
0.852
|
|
CD2BP2
|
CD2 (cytoplasmic tail) binding protein 2
|
|
chr13_+_30089829
|
0.850
|
NM_005800
|
USPL1
|
ubiquitin specific peptidase like 1
|
|
chr11_-_72111027
|
0.848
|
NM_001135190
NM_015242
|
ARAP1
|
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1
|
|
chr2_+_113058642
|
0.844
|
|
CHCHD5
|
coiled-coil-helix-coiled-coil-helix domain containing 5
|
|
chr16_-_65751305
|
0.842
|
NM_003789
|
TRADD
|
TNFRSF1A-associated via death domain
|
|
chr16_+_65751329
|
0.824
|
NM_018378
|
FBXL8
|
F-box and leucine-rich repeat protein 8
|
|
chr12_+_12829807
|
0.811
|
NM_030817
|
APOLD1
|
apolipoprotein L domain containing 1
|
|
chr11_+_63362922
|
0.811
|
NM_001039469
NM_001163296
NM_001163297
NM_004954
|
MARK2
|
MAP/microtubule affinity-regulating kinase 2
|
|
chr19_-_57592778
|
0.811
|
|
LOC284373
|
hypothetical protein LOC284373
|
|
chr2_-_207738858
|
0.807
|
NM_003709
|
KLF7
|
Kruppel-like factor 7 (ubiquitous)
|
|
chr3_-_102714663
|
0.805
|
NM_001077203
NM_020654
|
SENP7
|
SUMO1/sentrin specific peptidase 7
|
|
chr1_-_154170798
|
0.803
|
|
KIAA0907
|
KIAA0907
|
|
chr16_-_65751263
|
0.794
|
|
TRADD
|
TNFRSF1A-associated via death domain
|
|
chr4_+_25524903
|
0.787
|
NM_001145432
|
C4orf52
|
chromosome 4 open reading frame 52
|
|
chr7_-_5787723
|
0.776
|
|
RNF216
|
ring finger protein 216
|
|
chr17_-_19821583
|
0.776
|
NM_007202
|
AKAP10
|
A kinase (PRKA) anchor protein 10
|
|
chr18_+_3252184
|
0.770
|
|
MYL12B
|
myosin, light chain 12B, regulatory
|
|
chr1_-_154170764
|
0.766
|
|
KIAA0907
|
KIAA0907
|
|
chr19_+_40928300
|
0.763
|
NM_172341
|
PSENEN
|
presenilin enhancer 2 homolog (C. elegans)
|
|
chr14_-_37795290
|
0.762
|
NM_175060
|
CLEC14A
|
C-type lectin domain family 14, member A
|
|
chr19_-_14108267
|
0.761
|
|
ASF1B
|
ASF1 anti-silencing function 1 homolog B (S. cerevisiae)
|
|
chr8_+_125620523
|
0.755
|
NM_005005
|
NDUFB9
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 9, 22kDa
|
|
chr21_+_25856311
|
0.753
|
|
MIR155HG
|
MIR155 host gene (non-protein coding)
|
|
chr19_+_57592913
|
0.731
|
NM_032423
|
ZNF528
|
zinc finger protein 528
|
|
chr19_-_14108400
|
0.726
|
NM_018154
|
ASF1B
|
ASF1 anti-silencing function 1 homolog B (S. cerevisiae)
|
|
chr1_+_222368413
|
0.721
|
NM_001136115
NM_015176
|
FBXO28
|
F-box protein 28
|
|
chr14_-_23107106
|
0.715
|
NM_003917
|
AP1G2
|
adaptor-related protein complex 1, gamma 2 subunit
|
|
chr1_+_26358097
|
0.714
|
NM_024869
|
GRRP1
|
glycine/arginine rich protein 1
|
|
chr14_-_37795001
|
0.712
|
|
CLEC14A
|
C-type lectin domain family 14, member A
|
|
chr7_+_7646840
|
0.705
|
|
LOC729852
|
hypothetical LOC729852
|
|
chr8_-_11096257
|
0.704
|
NM_173683
|
XKR6
|
XK, Kell blood group complex subunit-related family, member 6
|
|
chr2_-_175255717
|
0.703
|
NM_001077269
|
WIPF1
|
WAS/WASL interacting protein family, member 1
|
|
chr18_-_44729700
|
0.691
|
NM_001190822
|
SMAD7
|
SMAD family member 7
|
|
chr11_-_19038803
|
0.687
|
NM_054030
|
MRGPRX2
|
MAS-related GPR, member X2
|
|
chr8_-_120937215
|
0.674
|
NM_024094
|
DSCC1
|
defective in sister chromatid cohesion 1 homolog (S. cerevisiae)
|
|
chr15_-_50758100
|
0.673
|
NM_019600
|
KIAA1370
|
KIAA1370
|
|
chr19_-_2378530
|
0.665
|
|
TIMM13
|
translocase of inner mitochondrial membrane 13 homolog (yeast)
|
|
chr20_+_3138055
|
0.661
|
NM_033453
NM_181493
|
ITPA
|
inosine triphosphatase (nucleoside triphosphate pyrophosphatase)
|
|
chr22_+_37407921
|
0.659
|
|
TOMM22
|
translocase of outer mitochondrial membrane 22 homolog (yeast)
|
|
chr5_+_140719773
|
0.658
|
NM_018923
NM_032096
|
PCDHGB2
|
protocadherin gamma subfamily B, 2
|
|
chr3_+_123882355
|
0.657
|
NM_017554
|
PARP14
|
poly (ADP-ribose) polymerase family, member 14
|
|
chr8_-_95634845
|
0.653
|
NM_015496
NM_183009
|
KIAA1429
|
KIAA1429
|
|
chr22_+_37407925
|
0.650
|
|
TOMM22
|
translocase of outer mitochondrial membrane 22 homolog (yeast)
|
|
chr17_-_18890963
|
0.648
|
NM_006613
|
GRAP
|
GRB2-related adaptor protein
|
|
chr22_+_37407899
|
0.647
|
NM_020243
|
TOMM22
|
translocase of outer mitochondrial membrane 22 homolog (yeast)
|
|
chr16_-_30274182
|
0.644
|
NM_006110
|
CD2BP2
|
CD2 (cytoplasmic tail) binding protein 2
|
|
chr2_+_65308332
|
0.642
|
NM_001005386
NM_005722
|
ACTR2
|
ARP2 actin-related protein 2 homolog (yeast)
|
|
chr10_-_112668679
|
0.641
|
NM_001195306
|
BBIP1
|
BBSome interacting protein 1
|
|
chr2_-_219749793
|
0.640
|
NM_015680
|
C2orf24
|
chromosome 2 open reading frame 24
|
|
chr1_-_154170825
|
0.638
|
|
KIAA0907
|
KIAA0907
|
|
chr22_+_37407906
|
0.634
|
|
TOMM22
|
translocase of outer mitochondrial membrane 22 homolog (yeast)
|
|
chr2_-_46697646
|
0.632
|
|
PIGF
|
phosphatidylinositol glycan anchor biosynthesis, class F
|
|
chr1_+_118273887
|
0.628
|
NM_006784
|
WDR3
|
WD repeat domain 3
|
|
chr17_+_4784097
|
0.627
|
|
RNF167
|
ring finger protein 167
|
|
chr16_+_2672476
|
0.627
|
NM_018992
|
KCTD5
|
potassium channel tetramerisation domain containing 5
|
|
chr13_-_44813203
|
0.625
|
|
TPT1
|
tumor protein, translationally-controlled 1
|
|
chr3_+_181802592
|
0.624
|
NM_001042601
NM_133462
|
TTC14
|
tetratricopeptide repeat domain 14
|
|
chr4_+_1330957
|
0.623
|
NM_020894
|
KIAA1530
|
KIAA1530
|
|
chr10_-_43224312
|
0.621
|
NM_001098205
|
HNRNPF
|
heterogeneous nuclear ribonucleoprotein F
|
|
chr1_-_154170810
|
0.618
|
NM_014949
|
KIAA0907
|
KIAA0907
|
|
chr5_-_93472993
|
0.616
|
NM_001163417
NM_001163418
NM_032042
|
FAM172A
|
family with sequence similarity 172, member A
|
|
chr9_+_4652285
|
0.616
|
NM_203453
|
PPAPDC2
|
phosphatidic acid phosphatase type 2 domain containing 2
|
|
chr4_-_159863889
|
0.614
|
NM_005038
|
PPID
|
peptidylprolyl isomerase D
|
|
chr7_+_37926687
|
0.609
|
NM_017549
|
EPDR1
|
ependymin related protein 1 (zebrafish)
|
|
chr1_-_118273701
|
0.606
|
|
GDAP2
|
ganglioside induced differentiation associated protein 2
|
|
chr3_-_28365510
|
0.604
|
NM_001134432
NM_001134433
NM_022461
|
AZI2
|
5-azacytidine induced 2
|
|
chr12_-_6535481
|
0.603
|
NM_001039670
NM_001193457
NM_080730
|
IFFO1
|
intermediate filament family orphan 1
|
|
chr2_+_5750229
|
0.600
|
NM_003108
|
SOX11
|
SRY (sex determining region Y)-box 11
|
|
chr15_-_68971665
|
0.599
|
NM_020147
|
THAP10
|
THAP domain containing 10
|
|
chr14_-_77244073
|
0.594
|
NM_006020
|
ALKBH1
|
alkB, alkylation repair homolog 1 (E. coli)
|
|
chr1_-_154170812
|
0.593
|
|
KIAA0907
|
KIAA0907
|
|
chr5_-_176663265
|
0.591
|
NM_001031677
NM_130781
|
RAB24
|
RAB24, member RAS oncogene family
|
|
chr5_+_140780797
|
0.591
|
|
PCDHGA11
|
protocadherin gamma subfamily A, 11
|
|
chr2_-_220116062
|
0.591
|
NM_001195731
|
CHPF
|
chondroitin polymerizing factor
|
|
chr3_-_50335113
|
0.591
|
NM_003773
NM_033158
|
HYAL2
|
hyaluronoglucosaminidase 2
|
|
chr2_+_233825084
|
0.591
|
|
ATG16L1
|
ATG16 autophagy related 16-like 1 (S. cerevisiae)
|
|
chr8_-_29262240
|
0.579
|
NM_057158
|
DUSP4
|
dual specificity phosphatase 4
|
|
chr9_-_126991857
|
0.578
|
NM_001123355
NM_001123369
NM_002721
|
PPP6C
|
protein phosphatase 6, catalytic subunit
|
|
chr16_+_1949517
|
0.578
|
NM_004548
|
NDUFB10
|
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 10, 22kDa
|
|
chr5_+_180583315
|
0.575
|
|
TRIM41
|
tripartite motif containing 41
|
|
chr2_+_233824955
|
0.575
|
NM_001190266
NM_001190267
NM_017974
NM_030803
NM_198890
|
ATG16L1
|
ATG16 autophagy related 16-like 1 (S. cerevisiae)
|
|
chr2_+_233825113
|
0.573
|
|
ATG16L1
|
ATG16 autophagy related 16-like 1 (S. cerevisiae)
|
|
chr9_-_126991805
|
0.573
|
|
PPP6C
|
protein phosphatase 6, catalytic subunit
|
|
chr5_-_114543632
|
0.573
|
NM_001017397
NM_001017398
NM_018700
|
TRIM36
|
tripartite motif containing 36
|
|
chr12_-_3732614
|
0.572
|
NM_001144958
NM_001144959
NM_032680
|
EFCAB4B
|
EF-hand calcium binding domain 4B
|
|
chr14_-_68514723
|
0.571
|
|
ACTN1
|
actinin, alpha 1
|
|
chr17_-_1366927
|
0.570
|
NM_001135642
NM_016532
NM_130766
|
INPP5K
|
inositol polyphosphate-5-phosphatase K
|
|
chrX_+_119268518
|
0.569
|
NM_006777
NM_001184742
|
ZBTB33
|
zinc finger and BTB domain containing 33
|
|
chr10_-_112668892
|
0.566
|
NM_001195304
NM_001195305
NM_001195307
|
BBIP1
|
BBSome interacting protein 1
|
|
chr18_+_31806585
|
0.561
|
NM_031446
|
C18orf21
|
chromosome 18 open reading frame 21
|
|
chr3_-_50624114
|
0.557
|
NM_013324
NM_145071
|
CISH
|
cytokine inducible SH2-containing protein
|
|
chr5_+_176663385
|
0.555
|
|
PRELID1
|
PRELI domain containing 1
|
|
chr2_+_233825056
|
0.554
|
|
ATG16L1
|
ATG16 autophagy related 16-like 1 (S. cerevisiae)
|
|
chr3_+_29297806
|
0.551
|
NM_001003792
NM_001003793
NM_001177711
NM_001177712
NM_014483
|
RBMS3
|
RNA binding motif, single stranded interacting protein 3
|
|
chr19_-_2378581
|
0.549
|
|
TIMM13
|
translocase of inner mitochondrial membrane 13 homolog (yeast)
|
|
chr2_-_10747438
|
0.549
|
|
NOL10
|
nucleolar protein 10
|
|
chr1_-_224993498
|
0.545
|
NM_002221
|
ITPKB
|
inositol 1,4,5-trisphosphate 3-kinase B
|
|
chr19_-_54646837
|
0.545
|
NM_017916
|
PIH1D1
|
PIH1 domain containing 1
|
|
chr3_-_50624085
|
0.543
|
|
CISH
|
cytokine inducible SH2-containing protein
|
|
chr3_+_120496180
|
0.533
|
|
ARHGAP31
|
Rho GTPase activating protein 31
|
|
chr8_-_99126770
|
0.532
|
|
RPL30
|
ribosomal protein L30
|
|
chr9_+_126671321
|
0.531
|
|
ARPC5L
|
actin related protein 2/3 complex, subunit 5-like
|
|
chr4_-_47966570
|
0.530
|
NM_003215
|
TEC
|
tec protein tyrosine kinase
|
|
chr4_+_2783743
|
0.525
|
NM_001145855
|
SH3BP2
|
SH3-domain binding protein 2
|
|
chr7_-_134505923
|
0.523
|
|
C7orf49
|
chromosome 7 open reading frame 49
|
|
chr7_-_134506001
|
0.522
|
|
C7orf49
|
chromosome 7 open reading frame 49
|
|
chrX_-_1470862
|
0.520
|
NM_001636
|
SLC25A6
|
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 6
|
|
chr12_+_68265469
|
0.513
|
NM_006431
|
CCT2
|
chaperonin containing TCP1, subunit 2 (beta)
|
|
chr17_-_4783943
|
0.507
|
NM_001165417
NM_001165418
NM_003562
|
SLC25A11
|
solute carrier family 25 (mitochondrial carrier; oxoglutarate carrier), member 11
|
|
chr17_-_19821727
|
0.506
|
|
AKAP10
|
A kinase (PRKA) anchor protein 10
|
|
chr5_+_61910318
|
0.505
|
NM_181506
|
LRRC70
|
leucine rich repeat containing 70
|
|
chr7_-_134506012
|
0.504
|
|
C7orf49
|
chromosome 7 open reading frame 49
|
|
chr17_+_4784330
|
0.502
|
NM_015528
|
RNF167
|
ring finger protein 167
|
|
chr14_+_62740878
|
0.502
|
NM_020663
|
RHOJ
|
ras homolog gene family, member J
|
|
chr5_+_140772691
|
0.502
|
NM_018913
NM_032090
|
PCDHGA10
|
protocadherin gamma subfamily A, 10
|
|
chr17_+_3515217
|
0.501
|
|
RPL21
|
ribosomal protein L21
|
|
chr12_-_109372481
|
0.499
|
|
ARPC3
|
actin related protein 2/3 complex, subunit 3, 21kDa
|
|
chr11_-_73559507
|
0.497
|
NM_015531
|
C2CD3
|
C2 calcium-dependent domain containing 3
|
|
chr10_+_44816278
|
0.495
|
NM_006963
|
ZNF22
|
zinc finger protein 22 (KOX 15)
|
|
chr6_-_33375106
|
0.495
|
NM_004761
|
RGL2
|
ral guanine nucleotide dissociation stimulator-like 2
|
|
chr3_+_9379706
|
0.494
|
NM_001114092
NM_015453
|
THUMPD3
|
THUMP domain containing 3
|
|
chr11_-_77383359
|
0.494
|
NM_033547
|
INTS4
|
integrator complex subunit 4
|
|
chr1_-_153560532
|
0.491
|
NM_001039517
|
C1orf104
|
chromosome 1 open reading frame 104
|
|
chr1_-_1300680
|
0.490
|
NM_017900
|
AURKAIP1
|
aurora kinase A interacting protein 1
|
|
chr14_+_101345878
|
0.490
|
NM_002719
NM_178586
NM_178587
|
PPP2R5C
|
protein phosphatase 2, regulatory subunit B', gamma
|
|
chrX_+_69270489
|
0.487
|
|
IGBP1
|
immunoglobulin (CD79A) binding protein 1
|
|
chr3_+_157875408
|
0.486
|
NM_001184717
|
TIPARP
|
TCDD-inducible poly(ADP-ribose) polymerase
|
|
chr12_+_11694172
|
0.485
|
|
ETV6
|
ets variant 6
|
|
chr3_-_28365284
|
0.485
|
|
AZI2
|
5-azacytidine induced 2
|
|
chr12_-_52868855
|
0.481
|
NM_014311
|
SMUG1
|
single-strand-selective monofunctional uracil-DNA glycosylase 1
|
|
chr10_+_27483758
|
0.479
|
NM_001172303
NM_001172304
NM_032844
|
MASTL
|
microtubule associated serine/threonine kinase-like
|
|
chr12_-_368517
|
0.478
|
|
KDM5A
|
lysine (K)-specific demethylase 5A
|
|
chr16_+_2672527
|
0.478
|
|
KCTD5
|
potassium channel tetramerisation domain containing 5
|
|
chr5_+_67547357
|
0.476
|
|
PIK3R1
|
phosphoinositide-3-kinase, regulatory subunit 1 (alpha)
|
|
chr11_+_119712827
|
0.476
|
NM_001198665
NM_015313
|
ARHGEF12
|
Rho guanine nucleotide exchange factor (GEF) 12
|
|
chr8_-_78074851
|
0.475
|
NM_000318
NM_001079867
|
PEX2
|
peroxisomal biogenesis factor 2
|
|
chr9_-_35093144
|
0.472
|
NM_013442
|
STOML2
|
stomatin (EPB72)-like 2
|
|
chr19_-_57930102
|
0.472
|
NM_001161501
NM_001161499
NM_001161500
|
ZNF611
|
zinc finger protein 611
|
|
chr22_-_28279643
|
0.470
|
NM_001002878
NM_001002879
NM_001002877
NM_003678
|
THOC5
|
THO complex 5
|
|
chr15_-_80611897
|
0.470
|
NM_001021
|
RPS17
|
ribosomal protein S17
|
|
chr1_+_118273869
|
0.470
|
|
WDR3
|
WD repeat domain 3
|
|
chr14_+_77244177
|
0.470
|
NM_031210
|
C14orf156
|
chromosome 14 open reading frame 156
|
|
chr12_-_131915270
|
0.470
|
NM_005895
|
GOLGA3
|
golgin A3
|
|
chr5_-_159778621
|
0.469
|
NM_006425
|
SLU7
|
SLU7 splicing factor homolog (S. cerevisiae)
|
|
chr9_-_103289154
|
0.468
|
NM_032342
|
C9orf125
|
chromosome 9 open reading frame 125
|
|
chr1_-_226661089
|
0.467
|
NM_145214
|
TRIM11
|
tripartite motif containing 11
|
|
chr8_-_125620466
|
0.465
|
NM_001146160
NM_032026
|
TATDN1
|
TatD DNase domain containing 1
|
|
chr20_+_43028590
|
0.464
|
|
STK4
|
serine/threonine kinase 4
|
|
chr6_+_43097352
|
0.464
|
NM_033112
|
RRP36
|
ribosomal RNA processing 36 homolog (S. cerevisiae)
|
|
chr20_+_43028533
|
0.464
|
NM_006282
|
STK4
|
serine/threonine kinase 4
|
|
chr8_-_68136700
|
0.462
|
|
COPS5
|
COP9 constitutive photomorphogenic homolog subunit 5 (Arabidopsis)
|
|
chrX_+_119268713
|
0.462
|
|
|
|
|
chr22_+_25398690
|
0.462
|
|
LOC388889
|
hypothetical LOC388889
|
|
chr7_-_134506070
|
0.459
|
NM_024033
|
C7orf49
|
chromosome 7 open reading frame 49
|
|
chr9_+_19398924
|
0.458
|
NM_001010887
|
ACER2
|
alkaline ceramidase 2
|
|
chr7_+_44802834
|
0.456
|
|
PPIA
|
peptidylprolyl isomerase A (cyclophilin A)
|
|
chr6_-_44341222
|
0.456
|
|
NFKBIE
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, epsilon
|
|
chr19_+_39860398
|
0.456
|
NM_001012320
NM_018443
|
ZNF302
|
zinc finger protein 302
|
|
chr19_+_17391856
|
0.455
|
NM_138401
|
FAM125A
|
family with sequence similarity 125, member A
|
|
chr7_-_122314017
|
0.452
|
NM_001009571
NM_001167940
NM_017954
|
CADPS2
|
Ca++-dependent secretion activator 2
|
|
chr2_-_27147995
|
0.451
|
NM_001134693
|
OST4
|
oligosaccharyltransferase 4 homolog (S. cerevisiae)
|
|
chr9_-_35093189
|
0.450
|
|
STOML2
|
stomatin (EPB72)-like 2
|
|
chr15_+_73983250
|
0.448
|
NM_012170
NM_147188
|
FBXO22
|
F-box protein 22
|
|
chr3_-_170970296
|
0.448
|
NM_032487
|
ARPM1
|
actin related protein M1
|
|
chr12_-_131915476
|
0.448
|
NM_001172557
|
GOLGA3
|
golgin A3
|
|
chr11_+_63363174
|
0.447
|
|
MARK2
|
MAP/microtubule affinity-regulating kinase 2
|
|
chr2_-_65447370
|
0.444
|
NM_001128210
|
SPRED2
|
sprouty-related, EVH1 domain containing 2
|
|
chr1_+_155130146
|
0.443
|
NM_001080471
|
PEAR1
|
platelet endothelial aggregation receptor 1
|
|
chr18_-_70415788
|
0.443
|
|
LOC400657
|
hypothetical LOC400657
|
|
chr14_+_77244191
|
0.442
|
|
C14orf156
|
chromosome 14 open reading frame 156
|
|
chr14_-_23106779
|
0.438
|
|
AP1G2
|
adaptor-related protein complex 1, gamma 2 subunit
|
|
chr19_+_17391841
|
0.437
|
|
FAM125A
|
family with sequence similarity 125, member A
|
|
chr15_+_32304477
|
0.437
|
NM_016454
|
TMEM85
|
transmembrane protein 85
|
|
chr22_-_34566367
|
0.436
|
NM_001031695
NM_001082576
NM_001082577
NM_014309
|
RBFOX2
|
RNA binding protein, fox-1 homolog (C. elegans) 2
|
|
chr1_-_1300416
|
0.436
|
NM_001127230
NM_001127229
|
AURKAIP1
|
aurora kinase A interacting protein 1
|
|
chr9_+_130056513
|
0.434
|
|
|
|
|
chr11_-_65139277
|
0.433
|
|
MAP3K11
|
mitogen-activated protein kinase kinase kinase 11
|
|
chr1_+_45822246
|
0.433
|
NM_001195193
NM_002482
NM_152298
|
NASP
|
nuclear autoantigenic sperm protein (histone-binding)
|
|
chr15_+_68971835
|
0.432
|
NM_001199017
NM_017691
NM_001199018
|
LRRC49
|
leucine rich repeat containing 49
|
|
chr10_+_101282679
|
0.431
|
NM_145285
|
NKX2-3
|
NK2 transcription factor related, locus 3 (Drosophila)
|
|
chr6_-_136889302
|
0.430
|
|
MAP7
|
microtubule-associated protein 7
|