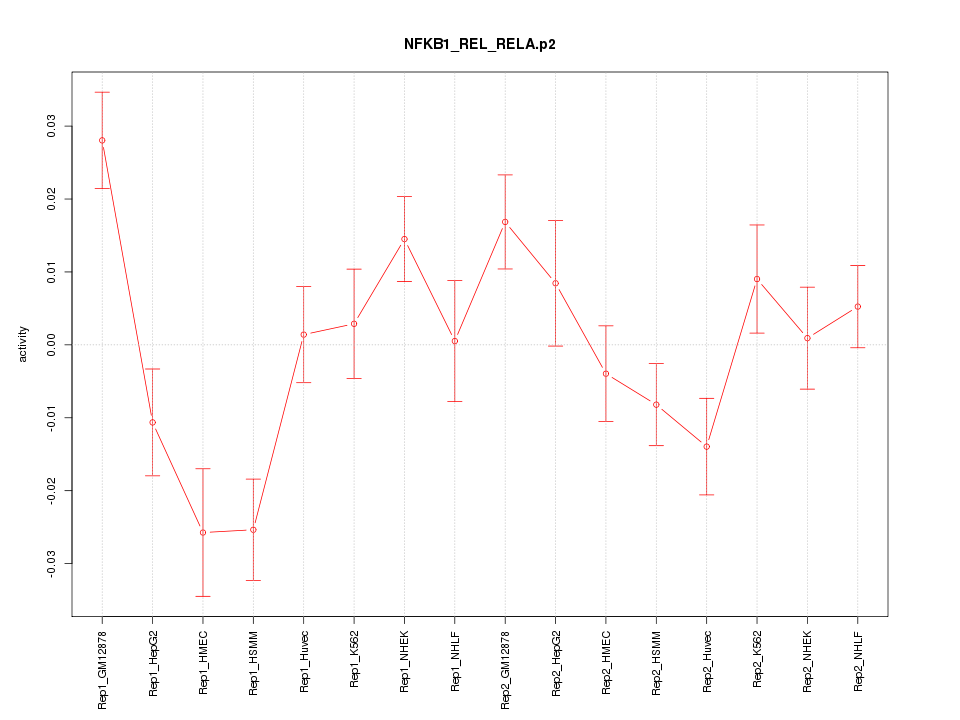
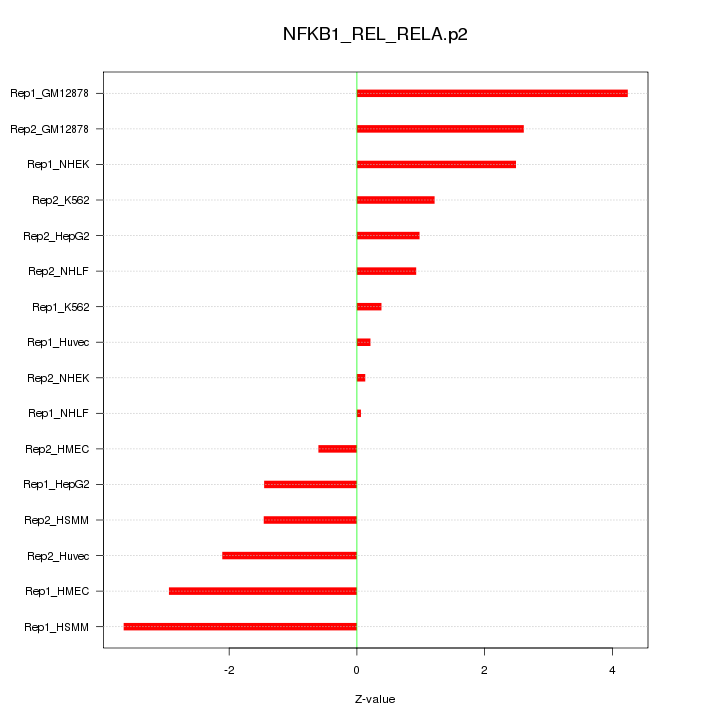
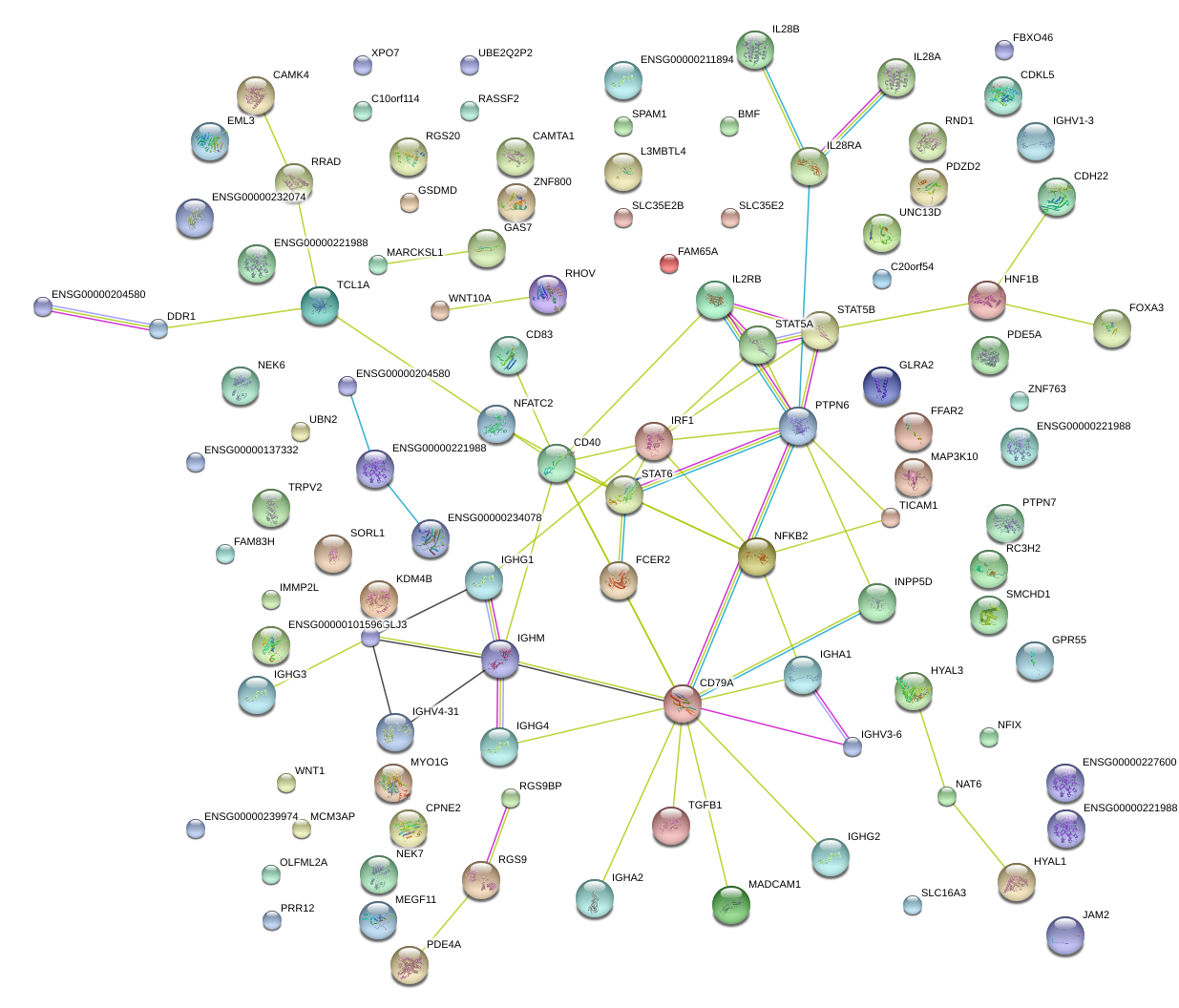
|
chr20_-_4743768
|
2.525
|
NM_170774
|
RASSF2
|
Ras association (RalGDS/AF-6) domain family member 2
|
|
chr20_-_697127
|
2.476
|
NM_033409
|
C20orf54
|
chromosome 20 open reading frame 54
|
|
chr15_-_38185578
|
2.403
|
NM_001003942
NM_001003943
|
BMF
|
Bcl2 modifying factor
|
|
chr1_-_200395741
|
2.084
|
NM_080588
|
PTPN7
|
protein tyrosine phosphatase, non-receptor type 7
|
|
chr14_-_106202262
|
1.668
|
|
IGLJ3
IGHA1
|
immunoglobulin lambda joining 3
immunoglobulin heavy constant alpha 1
|
|
chr9_+_126094388
|
1.657
|
NM_001166169
|
NEK6
|
NIMA (never in mitosis gene a)-related kinase 6
|
|
chr9_+_126094067
|
1.594
|
NM_001166170
NM_001166171
|
NEK6
|
NIMA (never in mitosis gene a)-related kinase 6
|
|
chr2_-_231498184
|
1.572
|
NM_005683
|
GPR55
|
G protein-coupled receptor 55
|
|
chr5_+_31675181
|
1.507
|
|
PDZD2
|
PDZ domain containing 2
|
|
chr19_-_4782711
|
1.490
|
NM_182919
|
TICAM1
|
toll-like receptor adaptor molecule 1
|
|
chr7_-_44985084
|
1.444
|
|
MYO1G
|
myosin IG
|
|
chr19_-_46551172
|
1.404
|
|
TGFB1
|
transforming growth factor, beta 1
|
|
chr19_+_13066956
|
1.399
|
|
NFIX
|
nuclear factor I/X (CCAAT-binding transcription factor)
|
|
chr17_+_16259636
|
1.348
|
|
TRPV2
|
transient receptor potential cation channel, subfamily V, member 2
|
|
chr16_-_65516919
|
1.342
|
NM_001128850
NM_004165
|
RRAD
|
Ras-related associated with diabetes
|
|
chr17_+_16259573
|
1.337
|
NM_016113
|
TRPV2
|
transient receptor potential cation channel, subfamily V, member 2
|
|
chr19_+_447453
|
1.335
|
NM_130760
NM_130762
|
MADCAM1
|
mucosal vascular addressin cell adhesion molecule 1
|
|
chr19_-_44427446
|
1.319
|
NM_172139
|
IL28B
|
interleukin 28B (interferon, lambda 3)
|
|
chr22_-_35875890
|
1.297
|
NM_000878
|
IL2RB
|
interleukin 2 receptor, beta
|
|
chr20_-_49592654
|
1.282
|
NM_012340
NM_173091
|
NFATC2
|
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 2
|
|
chr20_-_49592444
|
1.278
|
|
NFATC2
|
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 2
|
|
chr19_-_50975121
|
1.258
|
|
|
|
|
chr19_+_54786723
|
1.251
|
NM_020719
|
PRR12
|
proline rich 12
|
|
chr20_-_44370537
|
1.233
|
|
CDH22
|
cadherin 22, type 2
|
|
chr19_+_45389356
|
1.229
|
NM_002446
|
MAP3K10
|
mitogen-activated protein kinase kinase kinase 10
|
|
chr19_-_46551359
|
1.206
|
|
TGFB1
|
transforming growth factor, beta 1
|
|
chr7_-_44985202
|
1.150
|
NM_033054
|
MYO1G
|
myosin IG
|
|
chr12_-_55790355
|
1.144
|
NM_001178079
|
STAT6
|
signal transducer and activator of transcription 6, interleukin-4 induced
|
|
chr19_-_50925966
|
1.112
|
NM_001080469
|
FBXO46
|
F-box protein 46
|
|
chr17_-_33178987
|
1.103
|
|
HNF1B
|
HNF1 homeobox B
|
|
chr7_-_44985175
|
1.054
|
|
MYO1G
|
myosin IG
|
|
chr7_-_110989582
|
1.048
|
NM_032549
|
IMMP2L
|
IMP2 inner mitochondrial membrane peptidase-like (S. cerevisiae)
|
|
chr12_+_6926014
|
1.037
|
|
PTPN6
|
protein tyrosine phosphatase, non-receptor type 6
|
|
chr19_-_46551093
|
1.027
|
|
TGFB1
|
transforming growth factor, beta 1
|
|
chr9_+_126579203
|
1.025
|
NM_182487
|
OLFML2A
|
olfactomedin-like 2A
|
|
chr1_+_6767968
|
1.000
|
NM_001195563
NM_015215
|
CAMTA1
|
calmodulin binding transcription activator 1
|
|
chr3_-_50311666
|
0.997
|
NM_012191
NM_003549
|
NAT6
HYAL3
|
N-acetyltransferase 6 (GCN5-related)
hyaluronoglucosaminidase 3
|
|
chr4_-_120768563
|
0.991
|
NM_033437
|
PDE5A
|
phosphodiesterase 5A, cGMP-specific
|
|
chr1_-_200397300
|
0.990
|
|
PTPN7
|
protein tyrosine phosphatase, non-receptor type 7
|
|
chr16_+_66128843
|
0.966
|
NM_001193524
|
FAM65A
|
family with sequence similarity 65, member A
|
|
chr14_-_95250155
|
0.964
|
NM_001098725
NM_021966
|
TCL1A
|
T-cell leukemia/lymphoma 1A
|
|
chr21_+_25933459
|
0.954
|
NM_021219
|
JAM2
|
junctional adhesion molecule 2
|
|
chr8_+_144706699
|
0.950
|
NM_001166237
|
GSDMD
|
gasdermin D
|
|
chr10_+_104144218
|
0.941
|
NM_001077493
NM_001077494
|
NFKB2
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100)
|
|
chr19_+_47073029
|
0.937
|
NM_001783
NM_021601
|
CD79A
|
CD79a molecule, immunoglobulin-associated alpha
|
|
chr7_-_126819953
|
0.924
|
NM_176814
|
ZNF800
|
zinc finger protein 800
|
|
chr17_+_77783432
|
0.914
|
NM_001042423
|
SLC16A3
|
solute carrier family 16, member 3 (monocarboxylic acid transporter 4)
|
|
chr6_+_32229771
|
0.888
|
|
PPT2-EGFL8
PPT2
|
PPT2-EGFL8 read-through transcript
palmitoyl-protein thioesterase 2
|
|
chr9_-_124707314
|
0.867
|
NM_001100588
NM_018835
|
RC3H2
|
ring finger and CCCH-type domains 2
|
|
chr2_+_219453487
|
0.864
|
NM_025216
|
WNT10A
|
wingless-type MMTV integration site family, member 10A
|
|
chr19_+_10388448
|
0.863
|
|
PDE4A
|
phosphodiesterase 4A, cAMP-specific
|
|
chr6_+_14225932
|
0.862
|
|
CD83
|
CD83 molecule
|
|
chr1_-_32574185
|
0.858
|
NM_023009
|
MARCKSL1
|
MARCKS-like 1
|
|
chr19_+_40632456
|
0.851
|
NM_005306
|
FFAR2
|
free fatty acid receptor 2
|
|
chr12_+_6925997
|
0.845
|
NM_080548
|
PTPN6
|
protein tyrosine phosphatase, non-receptor type 6
|
|
chr19_+_11936868
|
0.835
|
NM_001012753
|
ZNF763
|
zinc finger protein 763
|
|
chr8_+_21833162
|
0.832
|
|
XPO7
|
exportin 7
|
|
chr17_-_71352017
|
0.832
|
|
UNC13D
|
unc-13 homolog D (C. elegans)
|
|
chr17_+_60563963
|
0.825
|
|
RGS9
|
regulator of G-protein signaling 9
|
|
chr19_+_44450993
|
0.824
|
NM_172138
|
IL28A
|
interleukin 28A (interferon, lambda 2)
|
|
chrX_+_18353623
|
0.817
|
NM_003159
|
CDKL5
|
cyclin-dependent kinase-like 5
|
|
chr19_-_46551670
|
0.807
|
NM_000660
|
TGFB1
|
transforming growth factor, beta 1
|
|
chr21_-_46529663
|
0.801
|
NM_003906
|
MCM3AP
|
minichromosome maintenance complex component 3 associated protein
|
|
chr8_+_54955962
|
0.800
|
NM_003702
|
RGS20
|
regulator of G-protein signaling 20
|
|
chr17_+_60563917
|
0.797
|
NM_001081955
NM_001165933
NM_003835
|
RGS9
|
regulator of G-protein signaling 9
|
|
chr17_+_37694007
|
0.791
|
|
STAT5A
|
signal transducer and activator of transcription 5A
|
|
chr5_-_131854325
|
0.785
|
NM_002198
|
IRF1
|
interferon regulatory factor 1
|
|
chr15_-_38953628
|
0.774
|
NM_133639
|
RHOV
|
ras homolog gene family, member V
|
|
chr12_-_47545816
|
0.766
|
NM_014470
|
RND1
|
Rho family GTPase 1
|
|
chr15_+_80434362
|
0.765
|
|
UBE2Q2P2
|
ubiquitin-conjugating enzyme E2Q family member 2 pseudogene 2
|
|
chr1_-_1613802
|
0.764
|
|
SLC35E2B
SLC35E2
|
solute carrier family 35, member E2B
solute carrier family 35, member E2
|
|
chr19_+_51059357
|
0.759
|
NM_004497
|
FOXA3
|
forkhead box A3
|
|
chr18_-_6404900
|
0.758
|
NM_173464
|
L3MBTL4
|
l(3)mbt-like 4 (Drosophila)
|
|
chr19_+_5082230
|
0.748
|
|
KDM4B
|
lysine (K)-specific demethylase 4B
|
|
chr8_-_144887876
|
0.745
|
NM_198488
|
FAM83H
|
family with sequence similarity 83, member H
|
|
chr1_-_200396354
|
0.740
|
NM_002832
|
PTPN7
|
protein tyrosine phosphatase, non-receptor type 7
|
|
chr15_-_64333048
|
0.736
|
NM_032445
|
MEGF11
|
multiple EGF-like-domains 11
|
|
chr1_-_200396328
|
0.735
|
|
PTPN7
|
protein tyrosine phosphatase, non-receptor type 7
|
|
chrX_+_14457340
|
0.720
|
NM_001118885
NM_001118886
NM_001171942
NM_002063
|
GLRA2
|
glycine receptor, alpha 2
|
|
chr19_-_7672978
|
0.720
|
NM_002002
|
FCER2
|
Fc fragment of IgE, low affinity II, receptor for (CD23)
|
|
chr20_+_44180296
|
0.716
|
NM_001250
NM_152854
|
CD40
|
CD40 molecule, TNF receptor superfamily member 5
|
|
chr10_-_21825879
|
0.709
|
|
C10orf114
|
chromosome 10 open reading frame 114
|
|
chr1_-_24386324
|
0.704
|
NM_170743
NM_173064
NM_173065
|
IL28RA
|
interleukin 28 receptor, alpha (interferon, lambda receptor)
|
|
chr12_+_47658502
|
0.700
|
NM_005430
|
WNT1
|
wingless-type MMTV integration site family, member 1
|
|
chr5_+_110587546
|
0.696
|
NM_001744
|
CAMK4
|
calcium/calmodulin-dependent protein kinase IV
|
|
chr6_+_32230172
|
0.696
|
|
PPT2
|
palmitoyl-protein thioesterase 2
|
|
chr2_-_104834344
|
0.688
|
|
LOC100506421
|
hypothetical LOC100506421
|
|
chr7_+_138566770
|
0.686
|
NM_173569
|
UBN2
|
ubinuclein 2
|
|
chr1_-_1667123
|
0.682
|
NM_182838
|
SLC35E2B
SLC35E2
|
solute carrier family 35, member E2B
solute carrier family 35, member E2
|
|
chr11_+_120828058
|
0.675
|
NM_003105
|
SORL1
|
sortilin-related receptor, L(DLR class) A repeats containing
|
|
chr16_+_55684031
|
0.675
|
|
CPNE2
|
copine II
|
|
chr11_-_62136778
|
0.672
|
NM_153265
|
EML3
|
echinoderm microtubule associated protein like 3
|
|
chr2_+_233633348
|
0.652
|
|
INPP5D
|
inositol polyphosphate-5-phosphatase, 145kDa
|
|
chr16_+_55683995
|
0.650
|
|
CPNE2
|
copine II
|
|
chr18_+_2645885
|
0.648
|
NM_015295
|
SMCHD1
|
structural maintenance of chromosomes flexible hinge domain containing 1
|
|
chr6_+_30958672
|
0.643
|
|
DDR1
|
discoidin domain receptor tyrosine kinase 1
|
|
chr16_+_55684030
|
0.636
|
|
CPNE2
|
copine II
|
|
chr15_+_80741827
|
0.636
|
|
LOC80154
|
hypothetical LOC80154
|
|
chr17_-_9803490
|
0.635
|
NM_003644
|
GAS7
|
growth arrest-specific 7
|
|
chr14_-_105644300
|
0.631
|
|
IGHV4-31
IGHA1
IGHA2
IGHG1
IGHG3
IGHM
|
immunoglobulin heavy variable 4-31
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant alpha 2 (A2m marker)
immunoglobulin heavy constant gamma 1 (G1m marker)
immunoglobulin heavy constant gamma 3 (G3m marker)
immunoglobulin heavy constant mu
|
|
chr17_+_35727998
|
0.627
|
NM_001145301
NM_001145302
|
RARA
|
retinoic acid receptor, alpha
|
|
chr17_+_22823151
|
0.627
|
NM_014238
|
KSR1
|
kinase suppressor of ras 1
|
|
chr16_+_87232501
|
0.625
|
NM_013278
|
IL17C
|
interleukin 17C
|
|
chr12_+_6745729
|
0.624
|
NM_002824
|
PTMS
|
parathymosin
|
|
chr12_+_6745835
|
0.620
|
|
PTMS
|
parathymosin
|
|
chr17_-_31231421
|
0.618
|
NM_002985
|
CCL5
|
chemokine (C-C motif) ligand 5
|
|
chr2_-_127679812
|
0.618
|
NM_001001665
|
CYP27C1
|
cytochrome P450, family 27, subfamily C, polypeptide 1
|
|
chr3_-_13089547
|
0.610
|
NM_001134382
|
IQSEC1
|
IQ motif and Sec7 domain 1
|
|
chr6_+_149680747
|
0.610
|
|
TAB2
|
TGF-beta activated kinase 1/MAP3K7 binding protein 2
|
|
chr2_+_233633279
|
0.608
|
NM_001017915
NM_005541
|
INPP5D
|
inositol polyphosphate-5-phosphatase, 145kDa
|
|
chr7_+_100584397
|
0.607
|
NM_001283
|
AP1S1
|
adaptor-related protein complex 1, sigma 1 subunit
|
|
chr2_+_233093416
|
0.603
|
NM_001195129
|
LOC646960
|
trypsin serine protease-like
|
|
chr17_+_3515217
|
0.603
|
|
RPL21
|
ribosomal protein L21
|
|
chr19_-_4289750
|
0.601
|
NM_001013841
NM_017720
|
STAP2
|
signal transducing adaptor family member 2
|
|
chr17_-_33179118
|
0.601
|
NM_000458
NM_001165923
|
HNF1B
|
HNF1 homeobox B
|
|
chr1_-_1614005
|
0.598
|
|
SLC35E2B
|
solute carrier family 35, member E2B
|
|
chr11_-_76800525
|
0.592
|
|
PAK1
|
p21 protein (Cdc42/Rac)-activated kinase 1
|
|
chr22_-_16864183
|
0.590
|
|
MICAL3
|
microtubule associated monoxygenase, calponin and LIM domain containing 3
|
|
chr19_-_46551607
|
0.590
|
|
TGFB1
|
transforming growth factor, beta 1
|
|
chr11_+_65740789
|
0.589
|
|
PACS1
|
phosphofurin acidic cluster sorting protein 1
|
|
chr10_+_102746819
|
0.586
|
NM_032429
|
LZTS2
|
leucine zipper, putative tumor suppressor 2
|
|
chr16_+_55683940
|
0.586
|
NM_152727
|
CPNE2
|
copine II
|
|
chr17_-_71352182
|
0.584
|
NM_199242
|
UNC13D
|
unc-13 homolog D (C. elegans)
|
|
chr19_-_10537689
|
0.583
|
NM_023008
|
KRI1
|
KRI1 homolog (S. cerevisiae)
|
|
chr8_-_11361594
|
0.582
|
NM_053279
|
FAM167A
|
family with sequence similarity 167, member A
|
|
chr5_+_177473071
|
0.580
|
NM_015111
|
N4BP3
|
NEDD4 binding protein 3
|
|
chr15_+_71131877
|
0.580
|
NM_001172623
NM_001172624
NM_002499
|
NEO1
|
neogenin 1
|
|
chr17_+_35728039
|
0.580
|
|
RARA
|
retinoic acid receptor, alpha
|
|
chr7_+_8440541
|
0.577
|
|
NXPH1
|
neurexophilin 1
|
|
chr16_-_73292188
|
0.577
|
NM_001142497
NM_152649
|
MLKL
|
mixed lineage kinase domain-like
|
|
chr11_+_62136788
|
0.577
|
NM_000327
|
ROM1
|
retinal outer segment membrane protein 1
|
|
chr20_-_24986501
|
0.575
|
|
ACSS1
|
acyl-CoA synthetase short-chain family member 1
|
|
chr11_+_58666749
|
0.572
|
NM_001142519
NM_001142520
NM_001142521
|
FAM111A
|
family with sequence similarity 111, member A
|
|
chr11_+_61352348
|
0.570
|
|
FADS2
|
fatty acid desaturase 2
|
|
chr1_-_27834210
|
0.569
|
NM_005248
|
FGR
|
Gardner-Rasheed feline sarcoma viral (v-fgr) oncogene homolog
|
|
chr6_+_32229599
|
0.566
|
NM_005155
|
PPT2-EGFL8
PPT2
|
PPT2-EGFL8 read-through transcript
palmitoyl-protein thioesterase 2
|
|
chr1_-_164402700
|
0.566
|
|
FAM78B
|
family with sequence similarity 78, member B
|
|
chr19_-_52426055
|
0.564
|
NM_014417
|
BBC3
|
BCL2 binding component 3
|
|
chr19_-_50964336
|
0.562
|
NM_175875
|
SIX5
|
SIX homeobox 5
|
|
chr1_-_47428310
|
0.561
|
NM_005764
|
PDZK1IP1
|
PDZK1 interacting protein 1
|
|
chr1_-_1614040
|
0.560
|
NM_001110781
|
SLC35E2B
|
solute carrier family 35, member E2B
|
|
chr19_+_55614006
|
0.559
|
NM_003121
|
SPIB
|
Spi-B transcription factor (Spi-1/PU.1 related)
|
|
chr12_+_6745947
|
0.557
|
|
PTMS
|
parathymosin
|
|
chr19_-_17819814
|
0.548
|
NM_000215
|
JAK3
|
Janus kinase 3
|
|
chr17_+_35277942
|
0.544
|
NM_198844
NM_199321
|
ZPBP2
|
zona pellucida binding protein 2
|
|
chr1_-_210939929
|
0.541
|
NM_018664
|
BATF3
|
basic leucine zipper transcription factor, ATF-like 3
|
|
chr7_-_129379947
|
0.538
|
NM_003344
NM_182697
|
UBE2H
|
ubiquitin-conjugating enzyme E2H (UBC8 homolog, yeast)
|
|
chr17_-_36346240
|
0.537
|
|
KRT23
|
keratin 23 (histone deacetylase inducible)
|
|
chr11_-_74740298
|
0.533
|
NM_004041
NM_020251
|
ARRB1
|
arrestin, beta 1
|
|
chr20_-_44034194
|
0.532
|
NM_022095
|
ZNF335
|
zinc finger protein 335
|
|
chr4_-_56107725
|
0.531
|
NM_004898
|
CLOCK
|
clock homolog (mouse)
|
|
chr3_-_71715431
|
0.531
|
NM_001012505
NM_032682
|
FOXP1
|
forkhead box P1
|
|
chr8_+_144888275
|
0.530
|
|
LOC100128338
|
hypothetical LOC100128338
|
|
chr1_+_44185005
|
0.528
|
NM_014652
|
IPO13
|
importin 13
|
|
chr15_+_80741876
|
0.528
|
|
LOC80154
|
hypothetical LOC80154
|
|
chr2_+_242679507
|
0.528
|
|
LOC728323
|
hypothetical LOC728323
|
|
chr1_+_44185169
|
0.524
|
|
IPO13
|
importin 13
|
|
chr12_+_92488932
|
0.524
|
|
SOCS2
|
suppressor of cytokine signaling 2
|
|
chr19_+_1199548
|
0.521
|
NM_177401
|
MIDN
|
midnolin
|
|
chr1_+_44185227
|
0.520
|
|
IPO13
|
importin 13
|
|
chr12_-_92488767
|
0.518
|
|
|
|
|
chr10_-_134940315
|
0.518
|
NM_001109
NM_001164489
NM_001164490
|
ADAM8
|
ADAM metallopeptidase domain 8
|
|
chr17_-_4831388
|
0.518
|
NM_001171167
NM_001171168
|
CAMTA2
|
calmodulin binding transcription activator 2
|
|
chr1_+_35104120
|
0.517
|
|
|
|
|
chr1_-_75854281
|
0.516
|
|
SLC44A5
|
solute carrier family 44, member 5
|
|
chr10_+_12431477
|
0.516
|
NM_020397
NM_153498
|
CAMK1D
|
calcium/calmodulin-dependent protein kinase ID
|
|
chr16_+_705080
|
0.515
|
NM_024042
|
METRN
|
meteorin, glial cell differentiation regulator
|
|
chr19_-_48976852
|
0.513
|
|
KCNN4
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 4
|
|
chr1_+_23990228
|
0.509
|
NM_007260
|
LYPLA2
|
lysophospholipase II
|
|
chr8_-_103737058
|
0.509
|
NM_005655
|
KLF10
|
Kruppel-like factor 10
|
|
chr11_-_65070625
|
0.508
|
|
LTBP3
|
latent transforming growth factor beta binding protein 3
|
|
chr8_-_95343716
|
0.507
|
NM_005261
NM_181702
|
GEM
|
GTP binding protein overexpressed in skeletal muscle
|
|
chr17_+_35728070
|
0.506
|
|
RARA
|
retinoic acid receptor, alpha
|
|
chr13_+_107719977
|
0.505
|
NM_001145645
NM_006573
|
TNFSF13B
|
tumor necrosis factor (ligand) superfamily, member 13b
|
|
chr14_-_69725342
|
0.505
|
NM_033262
NM_058240
NM_182932
NM_183002
|
SLC8A3
|
solute carrier family 8 (sodium/calcium exchanger), member 3
|
|
chr17_+_62804036
|
0.503
|
|
PITPNC1
|
phosphatidylinositol transfer protein, cytoplasmic 1
|
|
chr15_+_54998264
|
0.501
|
|
TCF12
|
transcription factor 12
|
|
chr12_+_50731457
|
0.500
|
NM_002135
NM_173157
|
NR4A1
|
nuclear receptor subfamily 4, group A, member 1
|
|
chr19_-_10537646
|
0.496
|
|
KRI1
|
KRI1 homolog (S. cerevisiae)
|
|
chr1_+_24614831
|
0.494
|
NM_020448
|
NIPAL3
|
NIPA-like domain containing 3
|
|
chr14_-_73296643
|
0.494
|
NM_194278
|
C14orf43
|
chromosome 14 open reading frame 43
|
|
chr19_+_44595589
|
0.492
|
NM_022835
|
PLEKHG2
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 2
|
|
chr1_+_31855873
|
0.491
|
NM_001525
|
HCRTR1
|
hypocretin (orexin) receptor 1
|
|
chr2_+_231968578
|
0.491
|
NM_145236
|
B3GNT7
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 7
|
|
chr17_-_4831644
|
0.490
|
NM_001171166
NM_015099
|
CAMTA2
|
calmodulin binding transcription activator 2
|
|
chr12_-_56312696
|
0.489
|
|
B4GALNT1
|
beta-1,4-N-acetyl-galactosaminyl transferase 1
|
|
chr7_+_149042960
|
0.488
|
NM_032534
|
KRBA1
|
KRAB-A domain containing 1
|
|
chr18_+_2836979
|
0.486
|
NM_032048
|
EMILIN2
|
elastin microfibril interfacer 2
|
|
chr11_+_66004454
|
0.485
|
NM_005700
NM_130443
|
DPP3
|
dipeptidyl-peptidase 3
|
|
chr6_+_36272506
|
0.485
|
NM_015695
|
BRPF3
|
bromodomain and PHD finger containing, 3
|
|
chr4_-_122304925
|
0.484
|
NM_024873
|
TNIP3
|
TNFAIP3 interacting protein 3
|
|
chr1_-_59020875
|
0.480
|
|
JUN
|
jun proto-oncogene
|
|
chr1_+_205694237
|
0.480
|
NM_001006658
NM_001877
|
CR2
|
complement component (3d/Epstein Barr virus) receptor 2
|
|
chr11_+_120828275
|
0.476
|
|
SORL1
|
sortilin-related receptor, L(DLR class) A repeats containing
|
|
chr20_+_2972267
|
0.475
|
NM_001501
NM_178331
NM_178332
|
GNRH2
|
gonadotropin-releasing hormone 2
|
|
chr4_-_47966570
|
0.474
|
NM_003215
|
TEC
|
tec protein tyrosine kinase
|
|
chr7_-_156125876
|
0.472
|
|
C7orf13
|
chromosome 7 open reading frame 13
|
|
chr5_+_96297101
|
0.471
|
NM_005575
|
LNPEP
|
leucyl/cystinyl aminopeptidase
|
|
chr16_+_29735208
|
0.470
|
|
C16orf53
|
chromosome 16 open reading frame 53
|
|
chr1_-_158098962
|
0.470
|
NM_001013661
|
VSIG8
|
V-set and immunoglobulin domain containing 8
|