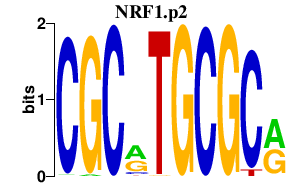
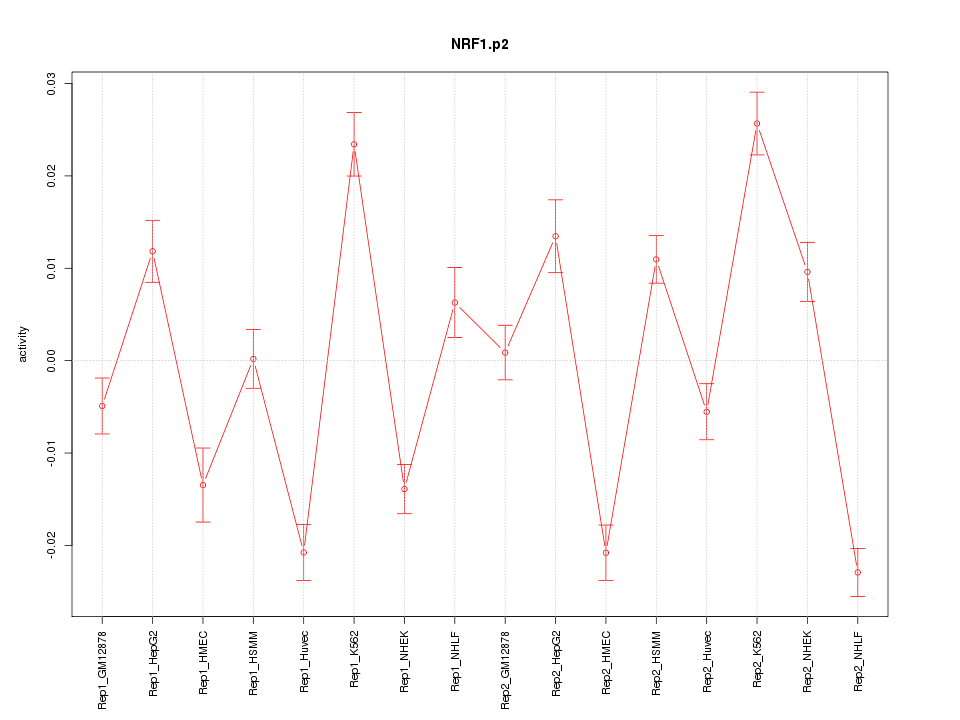
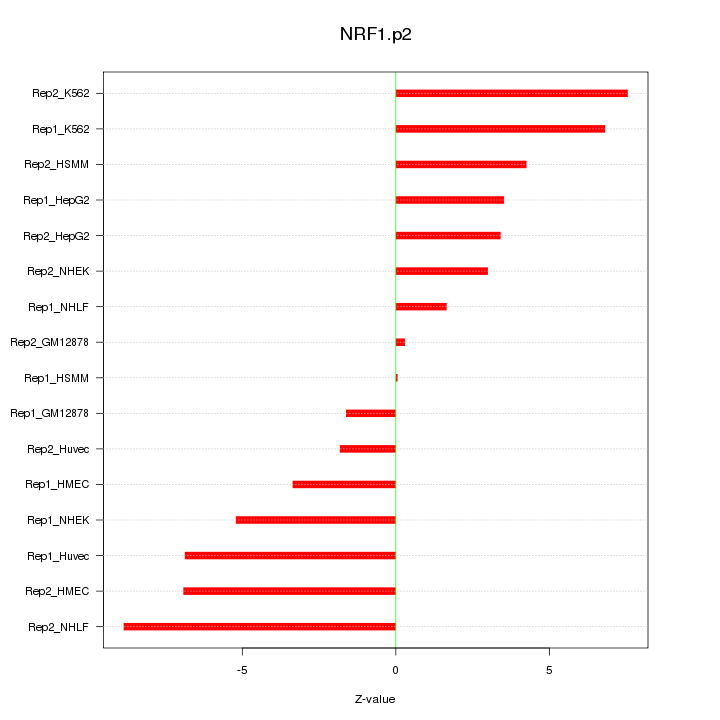
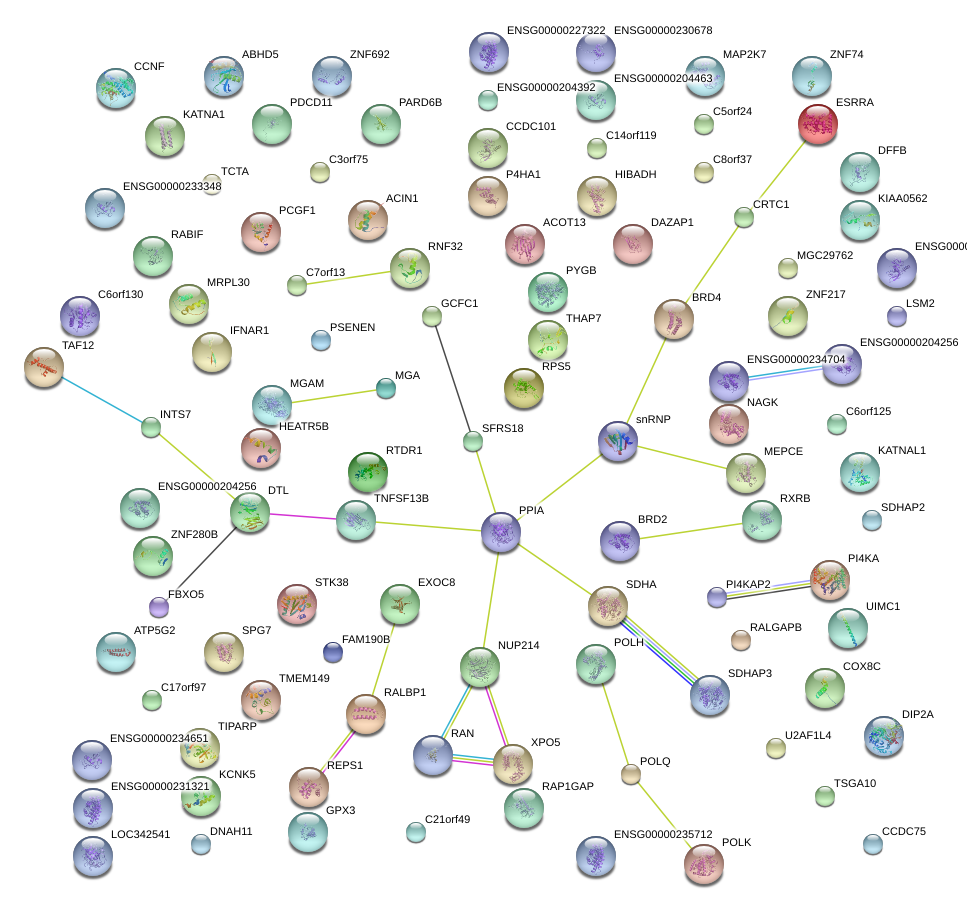
|
chr17_+_260408
|
10.177
|
NM_001013672
|
C17orf97
|
chromosome 17 open reading frame 97
|
|
chr1_-_3763480
|
4.896
|
|
KIAA0562
|
KIAA0562
|
|
chr1_+_3763704
|
4.140
|
NM_004402
|
DFFB
|
DNA fragmentation factor, 40kDa, beta polypeptide (caspase-activated DNase)
|
|
chr6_+_33044353
|
3.937
|
|
|
|
|
chr21_+_33066281
|
3.744
|
|
C21orf49
|
chromosome 21 open reading frame 49
|
|
chr21_-_33065849
|
3.661
|
NM_013329
NM_016631
|
GCFC1
|
GC-rich sequence DNA-binding factor 1
|
|
chr20_+_48781457
|
3.396
|
NM_032521
|
PARD6B
|
par-6 partitioning defective 6 homolog beta (C. elegans)
|
|
chr19_-_7874426
|
3.007
|
|
|
|
|
chr14_+_22633728
|
3.002
|
|
C14orf119
|
chromosome 14 open reading frame 119
|
|
chr6_+_33044378
|
2.999
|
NM_005104
|
BRD2
|
bromodomain containing 2
|
|
chr1_-_3763620
|
2.987
|
NM_014704
|
KIAA0562
|
KIAA0562
|
|
chr12_-_52356375
|
2.977
|
NM_005176
|
ATP5G2
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C2 (subunit 9)
|
|
chr6_-_31728121
|
2.899
|
NM_001098534
|
BAG6
|
BCL2-associated athanogene 6
|
|
chr22_-_21814208
|
2.783
|
NM_014433
|
RTDR1
|
rhabdoid tumor deletion region gene 1
|
|
chr6_-_31728363
|
2.589
|
|
BAG6
|
BCL2-associated athanogene 6
|
|
chr7_+_99865340
|
2.579
|
|
MEPCE
|
methylphosphate capping enzyme
|
|
chr6_-_31728405
|
2.560
|
|
BAG6
|
BCL2-associated athanogene 6
|
|
chr19_-_40928131
|
2.544
|
NM_001040425
NM_144987
|
U2AF1L4
|
U2 small nuclear RNA auxiliary factor 1-like 4
|
|
chr6_-_41148061
|
2.538
|
NM_145063
|
C6orf130
|
chromosome 6 open reading frame 130
|
|
chr6_-_39305203
|
2.517
|
NM_003740
|
KCNK5
|
potassium channel, subfamily K, member 5
|
|
chr6_-_31728418
|
2.513
|
|
BAG6
|
BCL2-associated athanogene 6
|
|
chr19_+_40928300
|
2.484
|
NM_172341
|
PSENEN
|
presenilin enhancer 2 homolog (C. elegans)
|
|
chr20_+_36534871
|
2.449
|
NM_020336
|
RALGAPB
|
Ral GTPase activating protein, beta subunit (non-catalytic)
|
|
chr22_+_19078397
|
2.428
|
NM_003426
|
ZNF74
|
zinc finger protein 74
|
|
chr1_+_210275541
|
2.406
|
NM_016448
|
DTL
|
denticleless homolog (Drosophila)
|
|
chr12_-_52356123
|
2.367
|
|
ATP5G2
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C2 (subunit 9)
|
|
chrX_-_73429101
|
2.361
|
|
MIR374AHG
|
mir-374a-545 cluster host gene (non-protein coding)
|
|
chr20_-_51644187
|
2.353
|
|
ZNF217
|
zinc finger protein 217
|
|
chr12_+_129922510
|
2.337
|
NM_006325
|
RAN
|
RAN, member RAS oncogene family
|
|
chr21_+_33619147
|
2.290
|
|
IFNAR1
|
interferon (alpha, beta and omega) receptor 1
|
|
chr1_-_210275402
|
2.261
|
NM_015434
|
INTS7
|
integrator complex subunit 7
|
|
chr9_+_132990760
|
2.244
|
NM_005085
|
NUP214
|
nucleoporin 214kDa
|
|
chr22_-_19686351
|
2.229
|
NM_001008695
NM_030573
|
THAP7
|
THAP domain containing 7
|
|
chr3_+_157875063
|
2.228
|
|
TIPARP
|
TCDD-inducible poly(ADP-ribose) polymerase
|
|
chr11_+_63829616
|
2.164
|
NM_004451
|
ESRRA
|
estrogen-related receptor alpha
|
|
chr10_+_86078136
|
2.141
|
NM_018999
|
FAM190B
|
family with sequence similarity 190, member B
|
|
chr9_+_132990834
|
2.079
|
|
NUP214
|
nucleoporin 214kDa
|
|
chr14_-_22634134
|
2.066
|
|
ACIN1
|
apoptotic chromatin condensation inducer 1
|
|
chr3_-_197201497
|
2.048
|
|
SDHAP1
SDHAP2
|
succinate dehydrogenase complex, subunit A, flavoprotein pseudogene 1
succinate dehydrogenase complex, subunit A, flavoprotein pseudogene 2
|
|
chr1_-_201124848
|
2.004
|
NM_002871
|
RABIF
|
RAB interacting factor
|
|
chr19_+_1358664
|
1.998
|
|
DAZAP1
|
DAZ associated protein 1
|
|
chr19_+_7874701
|
1.995
|
|
MAP2K7
|
mitogen-activated protein kinase kinase 7
|
|
chr6_-_36623223
|
1.994
|
NM_007271
|
STK38
|
serine/threonine kinase 38
|
|
chr3_+_43707362
|
1.991
|
NM_016006
|
ABHD5
|
abhydrolase domain containing 5
|
|
chr3_+_49424757
|
1.965
|
|
TCTA
|
T-cell leukemia translocation altered gene
|
|
chr19_+_7874774
|
1.954
|
|
MAP2K7
|
mitogen-activated protein kinase kinase 7
|
|
chr6_-_163754903
|
1.939
|
|
LOC100526820
|
hypothetical LOC100526820
|
|
chr6_-_33787413
|
1.891
|
|
C6orf125
|
chromosome 6 open reading frame 125
|
|
chr19_+_7874765
|
1.891
|
|
MAP2K7
|
mitogen-activated protein kinase kinase 7
|
|
chr2_-_74588006
|
1.873
|
|
PCGF1
|
polycomb group ring finger 1
|
|
chr5_-_176365971
|
1.869
|
NM_016290
|
UIMC1
|
ubiquitin interaction motif containing 1
|
|
chr6_-_36623038
|
1.861
|
|
STK38
|
serine/threonine kinase 38
|
|
chr1_-_28842103
|
1.859
|
NM_001135218
NM_005644
|
TAF12
|
TAF12 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 20kDa
|
|
chr6_-_43651578
|
1.839
|
|
XPO5
|
exportin 5
|
|
chr22_-_21193037
|
1.814
|
|
ZNF280B
|
zinc finger protein 280B
|
|
chr3_-_47530083
|
1.811
|
NM_001031703
|
C3orf75
|
chromosome 3 open reading frame 75
|
|
chr3_-_47529810
|
1.804
|
|
C3orf75
|
chromosome 3 open reading frame 75
|
|
chr19_+_18655483
|
1.780
|
|
CRTC1
|
CREB regulated transcription coactivator 1
|
|
chr6_-_33787479
|
1.767
|
NM_032340
|
C6orf125
|
chromosome 6 open reading frame 125
|
|
chr7_-_156125876
|
1.759
|
|
C7orf13
|
chromosome 7 open reading frame 13
|
|
chr19_+_7874764
|
1.756
|
NM_145185
|
MAP2K7
|
mitogen-activated protein kinase kinase 7
|
|
chr21_+_46703192
|
1.747
|
NM_001146114
NM_001146115
NM_001146116
NM_015151
NM_206889
NM_206890
NM_206891
|
DIP2A
|
DIP2 disco-interacting protein 2 homolog A (Drosophila)
|
|
chr22_-_21193504
|
1.747
|
NM_080764
|
ZNF280B
|
zinc finger protein 280B
|
|
chr16_+_88102305
|
1.737
|
NM_003119
NM_199367
|
SPG7
|
spastic paraplegia 7 (pure and complicated autosomal recessive)
|
|
chr2_-_74588293
|
1.713
|
NM_032673
|
PCGF1
|
polycomb group ring finger 1
|
|
chr2_+_71148915
|
1.708
|
NM_017567
|
NAGK
|
N-acetylglucosamine kinase
|
|
chr10_-_74526662
|
1.685
|
NM_000917
NM_001017962
NM_001142595
NM_001142596
|
P4HA1
|
prolyl 4-hydroxylase, alpha polypeptide I
|
|
chr6_-_43651727
|
1.684
|
|
XPO5
|
exportin 5
|
|
chr6_-_153346283
|
1.674
|
NM_001142522
|
FBXO5
|
F-box protein 5
|
|
chr18_+_9465528
|
1.670
|
NM_006788
|
RALBP1
|
ralA binding protein 1
|
|
chr5_+_271428
|
1.665
|
|
SDHA
|
succinate dehydrogenase complex, subunit A, flavoprotein (Fp)
|
|
chr6_-_31882558
|
1.655
|
|
LSM2
|
LSM2 homolog, U6 small nuclear RNA associated (S. cerevisiae)
|
|
chr22_-_19685998
|
1.649
|
|
THAP7
|
THAP domain containing 7
|
|
chr7_-_149201696
|
1.645
|
|
LOC401431
|
hypothetical LOC401431
|
|
chr19_+_7874762
|
1.627
|
|
MAP2K7
|
mitogen-activated protein kinase kinase 7
|
|
chr7_+_44802834
|
1.617
|
|
PPIA
|
peptidylprolyl isomerase A (cyclophilin A)
|
|
chr20_+_25176698
|
1.588
|
NM_002862
|
PYGB
|
phosphorylase, glycogen; brain
|
|
chr14_+_92883289
|
1.584
|
NM_182971
|
COX8C
|
cytochrome c oxidase subunit 8C
|
|
chr6_-_43651745
|
1.575
|
NM_020750
|
XPO5
|
exportin 5
|
|
chr21_+_33619067
|
1.565
|
NM_000629
|
IFNAR1
|
interferon (alpha, beta and omega) receptor 1
|
|
chr21_-_46703020
|
1.560
|
|
|
|
|
chr19_+_18655391
|
1.558
|
NM_001098482
NM_015321
|
CRTC1
|
CREB regulated transcription coactivator 1
|
|
chr6_-_31728447
|
1.545
|
NM_004639
NM_080702
NM_080703
|
BAG6
|
BCL2-associated athanogene 6
|
|
chr6_-_99979850
|
1.540
|
NM_015491
NM_032870
|
SFRS18
|
splicing factor, arginine/serine-rich 18
|
|
chr2_-_37164919
|
1.537
|
|
HEATR5B
|
HEAT repeat containing 5B
|
|
chr1_-_247119516
|
1.535
|
NM_001136036
|
ZNF692
|
zinc finger protein 692
|
|
chr6_-_31882704
|
1.535
|
NM_021177
|
LSM2
|
LSM2 homolog, U6 small nuclear RNA associated (S. cerevisiae)
|
|
chr15_+_39739864
|
1.532
|
NM_001080541
NM_001164273
|
MGA
|
MAX gene associated
|
|
chr8_-_96350575
|
1.531
|
NM_177965
|
C8orf37
|
chromosome 8 open reading frame 37
|
|
chr7_+_99865189
|
1.522
|
NM_019606
|
MEPCE
|
methylphosphate capping enzyme
|
|
chr2_+_99124379
|
1.516
|
NM_144706
|
C2orf15
|
chromosome 2 open reading frame 15
|
|
chr2_-_99124466
|
1.514
|
NM_182911
|
TSGA10
|
testis specific, 10
|
|
chr5_+_271355
|
1.499
|
NM_004168
|
SDHA
|
succinate dehydrogenase complex, subunit A, flavoprotein (Fp)
|
|
chr7_-_27669029
|
1.487
|
NM_152740
|
HIBADH
|
3-hydroxyisobutyrate dehydrogenase
|
|
chr19_+_18655487
|
1.485
|
|
CRTC1
|
CREB regulated transcription coactivator 1
|
|
chr16_+_2419388
|
1.484
|
NM_001761
|
CCNF
|
cyclin F
|
|
chr15_+_39739933
|
1.475
|
|
MGA
|
MAX gene associated
|
|
chr10_-_74526596
|
1.475
|
|
P4HA1
|
prolyl 4-hydroxylase, alpha polypeptide I
|
|
chr16_+_28472780
|
1.467
|
|
CCDC101
|
coiled-coil domain containing 101
|
|
chr22_-_19543069
|
1.466
|
NM_058004
|
PI4KA
|
phosphatidylinositol 4-kinase, catalytic, alpha
|
|
chr19_+_7874820
|
1.463
|
|
MAP2K7
|
mitogen-activated protein kinase kinase 7
|
|
chr6_-_150011450
|
1.460
|
|
KATNA1
|
katanin p60 (ATPase containing) subunit A 1
|
|
chr1_-_229540101
|
1.458
|
NM_175876
|
EXOC8
|
exocyst complex component 8
|
|
chr19_+_63590446
|
1.452
|
NM_001009
|
RPS5
|
ribosomal protein S5
|
|
chr6_+_24775241
|
1.429
|
NM_001160094
NM_018473
|
ACOT13
|
acyl-CoA thioesterase 13
|
|
chr2_+_37165097
|
1.409
|
NM_174931
|
CCDC75
|
coiled-coil domain containing 75
|
|
chr1_+_210275735
|
1.407
|
|
DTL
|
denticleless homolog (Drosophila)
|
|
chr1_-_21868380
|
1.392
|
NM_001145657
NM_002885
|
RAP1GAP
|
RAP1 GTPase activating protein
|
|
chr5_+_134209628
|
1.386
|
|
C5orf24
|
chromosome 5 open reading frame 24
|
|
chr7_+_156126157
|
1.386
|
NM_030936
NM_001184996
|
RNF32
|
ring finger protein 32
|
|
chr6_+_43651855
|
1.376
|
NM_006502
|
POLH
|
polymerase (DNA directed), eta
|
|
chr2_-_37164962
|
1.362
|
NM_019024
|
HEATR5B
|
HEAT repeat containing 5B
|
|
chr6_-_33276416
|
1.354
|
|
RXRB
|
retinoid X receptor, beta
|
|
chr6_-_24774797
|
1.346
|
NM_016614
|
TDP2
|
tyrosyl-DNA phosphodiesterase 2
|
|
chr17_-_24302597
|
1.345
|
NM_001033561
NM_020889
|
PHF12
|
PHD finger protein 12
|
|
chr7_+_107171514
|
1.344
|
NM_024814
|
CBLL1
|
Cas-Br-M (murine) ecotropic retroviral transforming sequence-like 1
|
|
chr6_+_43651899
|
1.340
|
|
POLH
|
polymerase (DNA directed), eta
|
|
chr17_-_18526296
|
1.339
|
NM_001145045
|
ZNF286B
|
zinc finger protein 286B
|
|
chr19_+_45996881
|
1.333
|
NM_080732
|
EGLN2
|
egl nine homolog 2 (C. elegans)
|
|
chr19_-_18294933
|
1.310
|
NM_012321
|
LSM4
|
LSM4 homolog, U6 small nuclear RNA associated (S. cerevisiae)
|
|
chr10_-_73280964
|
1.302
|
|
PSAP
|
prosaposin
|
|
chr10_-_71663076
|
1.300
|
NM_021129
|
PPA1
|
pyrophosphatase (inorganic) 1
|
|
chr4_+_26471410
|
1.298
|
NM_001169117
NM_001169118
NM_020860
|
STIM2
|
stromal interaction molecule 2
|
|
chr6_-_33276347
|
1.297
|
NM_021976
|
RXRB
|
retinoid X receptor, beta
|
|
chr6_+_237033
|
1.297
|
NM_020185
|
DUSP22
|
dual specificity phosphatase 22
|
|
chr1_-_25446509
|
1.297
|
|
C1orf63
|
chromosome 1 open reading frame 63
|
|
chr2_+_71149193
|
1.296
|
|
NAGK
|
N-acetylglucosamine kinase
|
|
chr6_-_33276426
|
1.294
|
|
RXRB
|
retinoid X receptor, beta
|
|
chr14_+_22305570
|
1.292
|
NM_005015
|
OXA1L
|
oxidase (cytochrome c) assembly 1-like
|
|
chr5_+_134209444
|
1.287
|
NM_001135586
|
C5orf24
|
chromosome 5 open reading frame 24
|
|
chr16_+_27122796
|
1.284
|
NM_001145348
|
JMJD5
|
jumonji domain containing 5
|
|
chr5_+_271447
|
1.283
|
|
SDHA
|
succinate dehydrogenase complex, subunit A, flavoprotein (Fp)
|
|
chr20_+_25124333
|
1.282
|
NM_001114089
NM_001247
|
ENTPD6
|
ectonucleoside triphosphate diphosphohydrolase 6 (putative)
|
|
chr1_+_32251988
|
1.277
|
NM_006559
|
KHDRBS1
|
KH domain containing, RNA binding, signal transduction associated 1
|
|
chr2_+_71148936
|
1.273
|
|
NAGK
|
N-acetylglucosamine kinase
|
|
chr2_+_100545827
|
1.272
|
NM_024065
|
PDCL3
|
phosducin-like 3
|
|
chr6_-_150011551
|
1.263
|
|
KATNA1
|
katanin p60 (ATPase containing) subunit A 1
|
|
chr19_+_10932792
|
1.260
|
NM_001128849
|
SMARCA4
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4
|
|
chr1_-_1699723
|
1.259
|
NM_023018
|
NADK
|
NAD kinase
|
|
chr2_-_69518108
|
1.256
|
|
NFU1
|
NFU1 iron-sulfur cluster scaffold homolog (S. cerevisiae)
|
|
chr22_+_19686434
|
1.255
|
|
FLJ39582
|
hypothetical LOC439931
|
|
chr8_-_22054939
|
1.254
|
NM_025232
|
REEP4
|
receptor accessory protein 4
|
|
chr6_+_44463418
|
1.251
|
|
CDC5L
|
CDC5 cell division cycle 5-like (S. pombe)
|
|
chr16_+_88280594
|
1.248
|
|
CDK10
|
cyclin-dependent kinase 10
|
|
chr5_+_271441
|
1.239
|
|
SDHA
|
succinate dehydrogenase complex, subunit A, flavoprotein (Fp)
|
|
chr16_+_88280569
|
1.230
|
NM_001098533
NM_001160367
NM_052987
NM_052988
|
CDK10
|
cyclin-dependent kinase 10
|
|
chr22_-_19543012
|
1.228
|
|
PI4KA
|
phosphatidylinositol 4-kinase, catalytic, alpha
|
|
chr3_+_49701965
|
1.227
|
|
RNF123
|
ring finger protein 123
|
|
chr1_-_40903897
|
1.226
|
NM_014747
|
RIMS3
|
regulating synaptic membrane exocytosis 3
|
|
chr6_-_24775106
|
1.224
|
|
TDP2
|
tyrosyl-DNA phosphodiesterase 2
|
|
chr19_-_47619992
|
1.221
|
|
LIPE
|
lipase, hormone-sensitive
|
|
chr20_-_35013589
|
1.216
|
|
SAMHD1
|
SAM domain and HD domain 1
|
|
chr10_-_73281047
|
1.215
|
|
PSAP
|
prosaposin
|
|
chr7_+_44802798
|
1.213
|
|
PPIA
|
peptidylprolyl isomerase A (cyclophilin A)
|
|
chr1_-_40903700
|
1.205
|
|
RIMS3
|
regulating synaptic membrane exocytosis 3
|
|
chr10_-_73281056
|
1.196
|
NM_001042465
NM_001042466
NM_002778
|
PSAP
|
prosaposin
|
|
chr1_-_229181203
|
1.190
|
|
TTC13
|
tetratricopeptide repeat domain 13
|
|
chr7_+_149651202
|
1.188
|
NM_001142928
NM_023942
|
LRRC61
|
leucine rich repeat containing 61
|
|
chr3_+_196870122
|
1.187
|
|
SDHAP2
|
succinate dehydrogenase complex, subunit A, flavoprotein pseudogene 2
|
|
chr16_-_82736186
|
1.184
|
NM_001146051
NM_031463
|
HSDL1
|
hydroxysteroid dehydrogenase like 1
|
|
chr21_+_44377974
|
1.184
|
|
C21orf33
|
chromosome 21 open reading frame 33
|
|
chr3_+_49701978
|
1.182
|
NM_022064
|
RNF123
|
ring finger protein 123
|
|
chr22_-_19543099
|
1.177
|
|
PI4KA
|
phosphatidylinositol 4-kinase, catalytic, alpha
|
|
chr5_-_1852902
|
1.175
|
|
MRPL36
|
mitochondrial ribosomal protein L36
|
|
chr6_+_44463265
|
1.174
|
NM_001253
|
CDC5L
|
CDC5 cell division cycle 5-like (S. pombe)
|
|
chr1_+_231152942
|
1.166
|
NM_032324
|
NTPCR
|
nucleoside-triphosphatase, cancer-related
|
|
chr16_+_28472726
|
1.163
|
NM_138414
|
CCDC101
|
coiled-coil domain containing 101
|
|
chr12_+_123023521
|
1.160
|
NM_152437
|
ZNF664
|
zinc finger protein 664
|
|
chr2_-_69518051
|
1.160
|
|
NFU1
|
NFU1 iron-sulfur cluster scaffold homolog (S. cerevisiae)
|
|
chr20_-_35013600
|
1.157
|
NM_015474
|
SAMHD1
|
SAM domain and HD domain 1
|
|
chr8_+_21833162
|
1.149
|
|
XPO7
|
exportin 7
|
|
chr3_-_198839026
|
1.149
|
|
LOC220729
|
succinate dehydrogenase complex, subunit A, flavoprotein pseudogene
|
|
chr1_+_210275752
|
1.147
|
|
DTL
|
denticleless homolog (Drosophila)
|
|
chr14_-_22127902
|
1.137
|
|
DAD1
|
defender against cell death 1
|
|
chr16_+_23754997
|
1.132
|
|
PRKCB
|
protein kinase C, beta
|
|
chr16_+_29734785
|
1.129
|
|
C16orf53
|
chromosome 16 open reading frame 53
|
|
chr5_-_271235
|
1.128
|
NM_145265
|
CCDC127
|
coiled-coil domain containing 127
|
|
chr7_+_138675644
|
1.126
|
|
LUC7L2
|
LUC7-like 2 (S. cerevisiae)
|
|
chr3_+_25806552
|
1.124
|
NM_001145391
NM_017897
|
OXSM
|
3-oxoacyl-ACP synthase, mitochondrial
|
|
chr16_-_4106186
|
1.117
|
NM_001116
|
ADCY9
|
adenylate cyclase 9
|
|
chr10_-_16899342
|
1.114
|
|
RSU1
|
Ras suppressor protein 1
|
|
chr2_+_197212522
|
1.110
|
NM_001080539
|
CCDC150
|
coiled-coil domain containing 150
|
|
chr7_-_149651594
|
1.107
|
NM_001164458
|
ACTR3C
|
ARP3 actin-related protein 3 homolog C (yeast)
|
|
chr7_-_86526880
|
1.101
|
NM_001142749
|
KIAA1324L
|
KIAA1324-like
|
|
chr6_-_150011542
|
1.099
|
|
KATNA1
|
katanin p60 (ATPase containing) subunit A 1
|
|
chr1_-_218329773
|
1.096
|
NM_006085
|
BPNT1
|
3'(2'), 5'-bisphosphate nucleotidase 1
|
|
chr12_-_40824699
|
1.090
|
NM_001099650
NM_173601
|
GXYLT1
|
glucoside xylosyltransferase 1
|
|
chr7_+_149651481
|
1.085
|
|
LRRC61
|
leucine rich repeat containing 61
|
|
chr16_+_82736365
|
1.083
|
NM_178452
|
LRRC50
|
leucine rich repeat containing 50
|
|
chr10_-_73281011
|
1.082
|
|
PSAP
|
prosaposin
|
|
chr16_+_88566485
|
1.081
|
|
AFG3L1P
|
AFG3 ATPase family gene 3-like 1 (S. cerevisiae), pseudogene
|
|
chr1_-_115102109
|
1.077
|
|
CSDE1
|
cold shock domain containing E1, RNA-binding
|
|
chr16_-_88566370
|
1.071
|
NM_145039
|
CENPBD1
|
CENPB DNA-binding domains containing 1
|
|
chr7_-_139523187
|
1.066
|
NM_030647
|
JHDM1D
|
jumonji C domain containing histone demethylase 1 homolog D (S. cerevisiae)
|
|
chr17_+_55325268
|
1.061
|
|
RPS6KB1
|
ribosomal protein S6 kinase, 70kDa, polypeptide 1
|
|
chr1_-_229181219
|
1.061
|
|
TTC13
|
tetratricopeptide repeat domain 13
|
|
chr22_+_19543248
|
1.057
|
NM_004782
|
SNAP29
|
synaptosomal-associated protein, 29kDa
|
|
chr1_-_115102106
|
1.047
|
|
CSDE1
|
cold shock domain containing E1, RNA-binding
|
|
chr16_+_29734990
|
1.047
|
NM_024516
|
C16orf53
|
chromosome 16 open reading frame 53
|
|
chr9_+_130491291
|
1.045
|
NM_003011
|
SET
|
SET nuclear oncogene
|