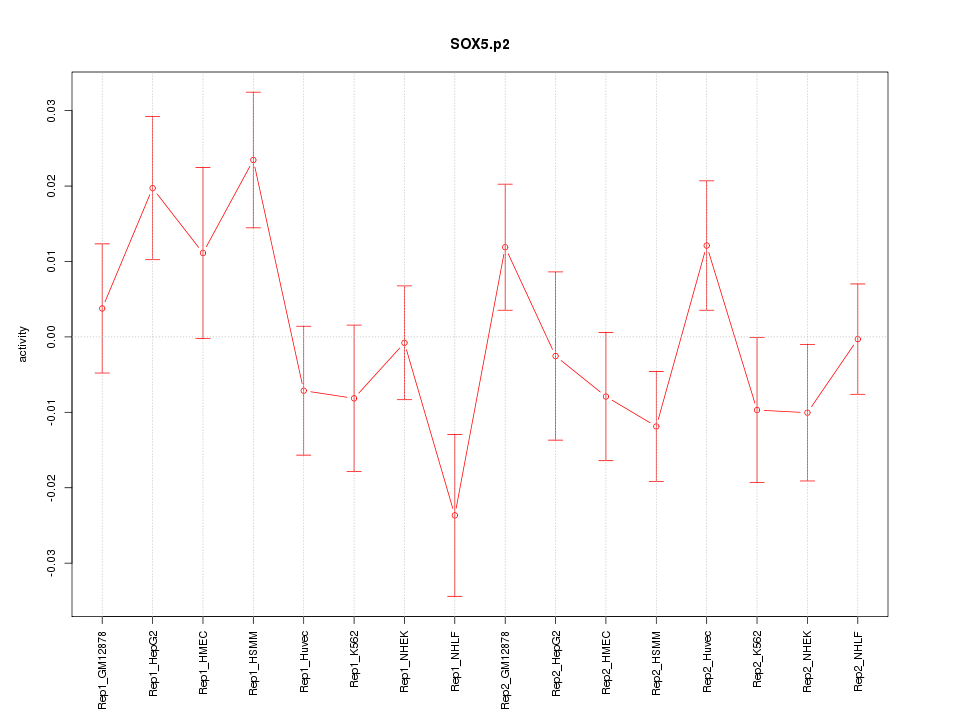
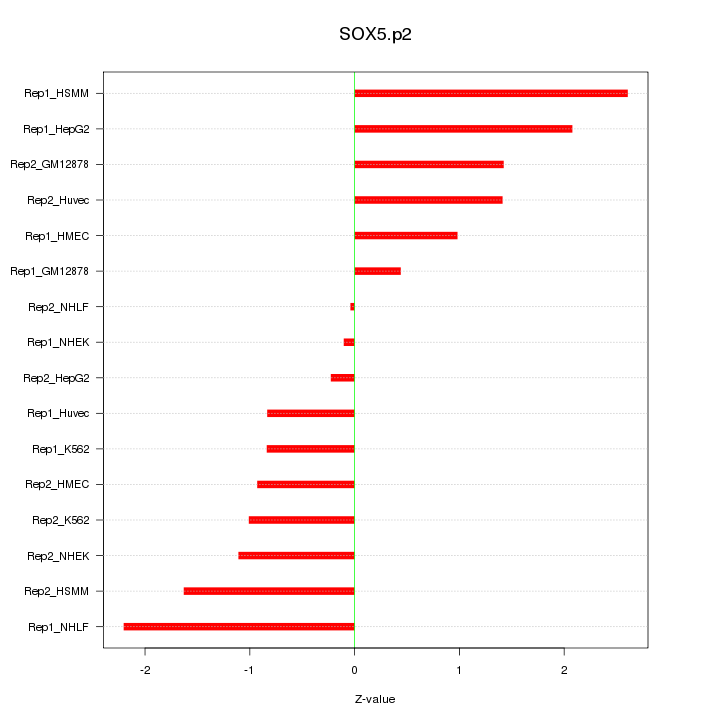
|
chr4_+_110968594
|
2.850
|
NM_006583
|
RRH
|
retinal pigment epithelium-derived rhodopsin homolog
|
|
chr11_-_16380967
|
2.254
|
NM_001145819
NM_017508
|
SOX6
|
SRY (sex determining region Y)-box 6
|
|
chr3_-_150422474
|
1.990
|
NM_000096
|
CP
|
ceruloplasmin (ferroxidase)
|
|
chr17_-_1195096
|
1.852
|
|
PAFAH1B1
|
platelet-activating factor acetylhydrolase 1b, regulatory subunit 1 (45kDa)
|
|
chr3_+_88270951
|
1.739
|
NM_018293
|
ZNF654
|
zinc finger protein 654
|
|
chr14_-_22974707
|
1.670
|
NM_000257
|
MYH7
|
myosin, heavy chain 7, cardiac muscle, beta
|
|
chr7_-_11838260
|
1.416
|
NM_015204
|
THSD7A
|
thrombospondin, type I, domain containing 7A
|
|
chr4_-_105635499
|
1.385
|
|
CXXC4
|
CXXC finger protein 4
|
|
chr1_+_154855709
|
1.278
|
NM_021817
|
HAPLN2
|
hyaluronan and proteoglycan link protein 2
|
|
chr3_-_150533948
|
1.276
|
NM_001184723
NM_138786
|
TM4SF18
|
transmembrane 4 L six family member 18
|
|
chr14_-_106184988
|
1.274
|
|
IGHA1
IGHG1
|
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant gamma 1 (G1m marker)
|
|
chr12_-_10453409
|
1.241
|
NM_013431
|
KLRC4-KLRK1
KLRC4
|
KLRC4-KLRK1 read-through transcript
killer cell lectin-like receptor subfamily C, member 4
|
|
chr11_-_55460451
|
1.216
|
NM_006637
|
OR5I1
|
olfactory receptor, family 5, subfamily I, member 1
|
|
chr16_+_55373241
|
1.163
|
|
NUP93
|
nucleoporin 93kDa
|
|
chr10_-_32707549
|
1.159
|
|
EPC1
|
enhancer of polycomb homolog 1 (Drosophila)
|
|
chr4_-_101658094
|
1.108
|
NM_001159694
NM_016242
|
EMCN
|
endomucin
|
|
chr1_+_50344550
|
1.098
|
NM_001144775
|
ELAVL4
|
ELAV (embryonic lethal, abnormal vision, Drosophila)-like 4 (Hu antigen D)
|
|
chr6_-_135312899
|
1.083
|
NM_001193480
NM_022568
NM_170771
|
ALDH8A1
|
aldehyde dehydrogenase 8 family, member A1
|
|
chr7_-_26198673
|
1.051
|
|
HNRNPA2B1
|
heterogeneous nuclear ribonucleoprotein A2/B1
|
|
chr2_-_215953818
|
1.004
|
|
FN1
|
fibronectin 1
|
|
chr2_+_228386800
|
0.997
|
NM_001130046
NM_004591
|
CCL20
|
chemokine (C-C motif) ligand 20
|
|
chr1_-_150564302
|
0.988
|
NM_002016
|
FLG
|
filaggrin
|
|
chr10_-_62002672
|
0.958
|
|
ANK3
|
ankyrin 3, node of Ranvier (ankyrin G)
|
|
chr4_+_113437947
|
0.957
|
NM_001102406
NM_025144
|
ALPK1
|
alpha-kinase 1
|
|
chr1_-_51660328
|
0.918
|
NM_001159969
|
EPS15
|
epidermal growth factor receptor pathway substrate 15
|
|
chr4_+_53975597
|
0.893
|
|
FIP1L1
|
FIP1 like 1 (S. cerevisiae)
|
|
chr2_+_169631777
|
0.869
|
NM_199204
|
DHRS9
|
dehydrogenase/reductase (SDR family) member 9
|
|
chr2_+_143351536
|
0.863
|
NM_001032998
NM_003937
|
KYNU
|
kynureninase
|
|
chr21_-_36870736
|
0.836
|
NM_001146077
|
CLDN14
|
claudin 14
|
|
chr3_-_69212192
|
0.802
|
NM_003968
NM_198195
|
UBA3
|
ubiquitin-like modifier activating enzyme 3
|
|
chr2_-_174475528
|
0.795
|
|
|
|
|
chr1_-_230717850
|
0.776
|
NM_020808
|
SIPA1L2
|
signal-induced proliferation-associated 1 like 2
|
|
chr5_-_111340421
|
0.770
|
NM_001142474
NM_001142475
|
C5orf13
|
chromosome 5 open reading frame 13
|
|
chr2_-_21087435
|
0.768
|
|
APOB
|
apolipoprotein B (including Ag(x) antigen)
|
|
chr4_-_170314382
|
0.762
|
|
SH3RF1
|
SH3 domain containing ring finger 1
|
|
chr3_+_173954940
|
0.762
|
NM_018098
|
ECT2
|
epithelial cell transforming sequence 2 oncogene
|
|
chr3_-_48729593
|
0.737
|
NM_001005909
NM_001005910
NM_001005911
NM_001146178
NM_001146179
NM_001190316
NM_001190317
NM_016291
|
IP6K2
|
inositol hexakisphosphate kinase 2
|
|
chr2_+_211166323
|
0.729
|
NM_001122634
|
CPS1
|
carbamoyl-phosphate synthase 1, mitochondrial
|
|
chr3_+_173043832
|
0.728
|
NM_001164436
|
TMEM212
|
transmembrane protein 212
|
|
chr14_-_93493519
|
0.724
|
NM_016150
|
ASB2
|
ankyrin repeat and SOCS box containing 2
|
|
chr5_+_140227951
|
0.721
|
NM_018902
NM_031861
|
PCDHA11
|
protocadherin alpha 11
|
|
chr3_-_121065112
|
0.718
|
|
|
|
|
chr1_-_49014993
|
0.713
|
NM_024603
|
BEND5
|
BEN domain containing 5
|
|
chr2_+_167468220
|
0.712
|
NM_001079810
NM_152381
|
XIRP2
|
xin actin-binding repeat containing 2
|
|
chr15_+_47502666
|
0.711
|
NM_002009
|
FGF7
|
fibroblast growth factor 7
|
|
chr1_+_116727514
|
0.711
|
NM_001160234
|
ATP1A1
|
ATPase, Na+/K+ transporting, alpha 1 polypeptide
|
|
chr3_+_190831909
|
0.708
|
NM_001114978
NM_001114979
NM_003722
|
TP63
|
tumor protein p63
|
|
chr17_+_3047559
|
0.705
|
NM_012352
|
OR1A2
|
olfactory receptor, family 1, subfamily A, member 2
|
|
chr20_-_7869068
|
0.703
|
NM_017545
|
HAO1
|
hydroxyacid oxidase (glycolate oxidase) 1
|
|
chr2_-_56266408
|
0.694
|
|
LOC100129434
|
hypothetical LOC100129434
|
|
chr2_+_166137091
|
0.692
|
NM_024969
|
CSRNP3
|
cysteine-serine-rich nuclear protein 3
|
|
chr10_-_56230953
|
0.684
|
NM_001142763
NM_001142764
NM_001142765
NM_001142766
NM_001142767
NM_001142768
NM_001142769
NM_001142770
NM_001142771
NM_001142772
NM_001142773
NM_033056
|
PCDH15
|
protocadherin-related 15
|
|
chr6_+_29902734
|
0.680
|
NM_002127
|
HLA-G
|
major histocompatibility complex, class I, G
|
|
chr12_-_10893318
|
0.679
|
NM_007244
|
PRR4
|
proline rich 4 (lacrimal)
|
|
chr15_-_39195621
|
0.668
|
NM_017553
|
INO80
|
INO80 homolog (S. cerevisiae)
|
|
chr12_-_11354620
|
0.654
|
NM_002723
|
PRB4
|
proline-rich protein BstNI subfamily 4
|
|
chr9_-_4142182
|
0.652
|
NM_152629
|
GLIS3
|
GLIS family zinc finger 3
|
|
chr3_+_44665220
|
0.646
|
NM_003420
|
ZNF35
|
zinc finger protein 35
|
|
chr12_-_101115677
|
0.639
|
NM_002674
|
PMCH
|
pro-melanin-concentrating hormone
|
|
chr2_-_225073357
|
0.636
|
|
CUL3
|
cullin 3
|
|
chr8_-_59575249
|
0.635
|
NM_000780
|
CYP7A1
|
cytochrome P450, family 7, subfamily A, polypeptide 1
|
|
chr4_+_71418886
|
0.627
|
NM_212557
|
AMTN
|
amelotin
|
|
chr17_-_53847937
|
0.619
|
|
RNF43
|
ring finger protein 43
|
|
chr4_-_64957595
|
0.606
|
NM_001010874
|
TECRL
|
trans-2,3-enoyl-CoA reductase-like
|
|
chr6_-_128263868
|
0.605
|
NM_001010923
NM_001164685
|
THEMIS
|
thymocyte selection associated
|
|
chr1_-_152431175
|
0.599
|
NM_152263
|
TPM3
|
tropomyosin 3
|
|
chr9_-_28709302
|
0.596
|
NM_152570
|
LINGO2
|
leucine rich repeat and Ig domain containing 2
|
|
chr8_+_85258007
|
0.589
|
NM_001100392
NM_001100393
|
RALYL
|
RALY RNA binding protein-like
|
|
chr4_-_68766392
|
0.589
|
NM_001129907
|
TMPRSS11BNL
|
TMPRSS11B N terminal-like
|
|
chr9_+_109131814
|
0.586
|
|
RAD23B
|
RAD23 homolog B (S. cerevisiae)
|
|
chr17_-_46479127
|
0.585
|
|
SPAG9
|
sperm associated antigen 9
|
|
chr5_+_140573692
|
0.581
|
NM_018933
|
PCDHB13
|
protocadherin beta 13
|
|
chr10_+_94584449
|
0.580
|
NM_001013848
|
EXOC6
|
exocyst complex component 6
|
|
chr8_-_38324809
|
0.579
|
|
WHSC1L1
|
Wolf-Hirschhorn syndrome candidate 1-like 1
|
|
chr3_-_101077605
|
0.578
|
NM_014890
|
FILIP1L
|
filamin A interacting protein 1-like
|
|
chr20_-_45418818
|
0.573
|
NM_012408
NM_183047
NM_183048
|
ZMYND8
|
zinc finger, MYND-type containing 8
|
|
chr7_-_84589182
|
0.571
|
NM_152754
|
SEMA3D
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3D
|
|
chr16_-_20323533
|
0.569
|
NM_174924
|
PDILT
|
protein disulfide isomerase-like, testis expressed
|
|
chr4_+_71096830
|
0.567
|
NM_017855
|
ODAM
|
odontogenic, ameloblast asssociated
|
|
chr7_+_50314801
|
0.566
|
NM_006060
|
IKZF1
|
IKAROS family zinc finger 1 (Ikaros)
|
|
chr12_-_11399776
|
0.564
|
NM_005039
NM_199353
NM_199354
|
PRB1
|
proline-rich protein BstNI subfamily 1
|
|
chr3_-_56673113
|
0.564
|
NM_015224
|
C3orf63
|
chromosome 3 open reading frame 63
|
|
chr18_-_55136673
|
0.562
|
NM_181654
|
CPLX4
|
complexin 4
|
|
chr15_+_22751162
|
0.561
|
NM_003097
NM_022804
NM_005678
|
SNRPN
SNURF
|
small nuclear ribonucleoprotein polypeptide N
SNRPN upstream reading frame
|
|
chr2_+_114935623
|
0.550
|
NM_001178036
|
DPP10
|
dipeptidyl-peptidase 10 (non-functional)
|
|
chr17_-_36451189
|
0.538
|
NM_030967
|
KRTAP1-3
KRTAP1-1
|
keratin associated protein 1-3
keratin associated protein 1-1
|
|
chr2_+_143603439
|
0.535
|
|
ARHGAP15
|
Rho GTPase activating protein 15
|
|
chr3_+_142587973
|
0.533
|
|
|
|
|
chr3_+_85858321
|
0.528
|
NM_153184
|
CADM2
|
cell adhesion molecule 2
|
|
chr14_+_19318240
|
0.526
|
NM_001005500
|
OR4M1
|
olfactory receptor, family 4, subfamily M, member 1
|
|
chr2_+_191009265
|
0.525
|
|
MFSD6
|
major facilitator superfamily domain containing 6
|
|
chr6_-_48186868
|
0.522
|
|
C6orf138
|
chromosome 6 open reading frame 138
|
|
chr3_-_129777618
|
0.522
|
NM_007354
|
C3orf27
|
chromosome 3 open reading frame 27
|
|
chr11_-_121492010
|
0.522
|
NM_001001786
|
BLID
|
BH3-like motif containing, cell death inducer
|
|
chr20_-_14266200
|
0.518
|
NM_013281
NM_198391
|
FLRT3
|
fibronectin leucine rich transmembrane protein 3
|
|
chr12_+_54172413
|
0.517
|
NM_001005519
|
OR6C68
|
olfactory receptor, family 6, subfamily C, member 68
|
|
chr14_+_19414230
|
0.511
|
NM_001005501
|
OR4K2
|
olfactory receptor, family 4, subfamily K, member 2
|
|
chr17_+_73511843
|
0.510
|
NM_001142640
NM_018996
|
TNRC6C
|
trinucleotide repeat containing 6C
|
|
chr2_-_162717159
|
0.509
|
NM_002054
|
GCG
|
glucagon
|
|
chr12_+_111980044
|
0.509
|
NM_004416
|
DTX1
|
deltex homolog 1 (Drosophila)
|
|
chr10_-_74863303
|
0.505
|
NM_001024593
|
ZMYND17
|
zinc finger, MYND-type containing 17
|
|
chr5_-_14706486
|
0.503
|
|
|
|
|
chr11_-_123414917
|
0.503
|
NM_001004463
|
OR10G7
|
olfactory receptor, family 10, subfamily G, member 7
|
|
chr5_-_64813459
|
0.503
|
NM_197941
|
ADAMTS6
|
ADAM metallopeptidase with thrombospondin type 1 motif, 6
|
|
chr6_-_76838944
|
0.502
|
NM_001563
|
IMPG1
|
interphotoreceptor matrix proteoglycan 1
|
|
chr6_-_111995213
|
0.501
|
NM_001164283
|
TRAF3IP2
|
TRAF3 interacting protein 2
|
|
chr2_+_143603368
|
0.500
|
NM_018460
|
ARHGAP15
|
Rho GTPase activating protein 15
|
|
chr7_+_106897737
|
0.495
|
NM_005295
|
GPR22
|
G protein-coupled receptor 22
|
|
chr14_-_19843969
|
0.494
|
NM_138376
|
TTC5
|
tetratricopeptide repeat domain 5
|
|
chr10_+_96433240
|
0.494
|
NM_000772
NM_001128925
|
CYP2C18
|
cytochrome P450, family 2, subfamily C, polypeptide 18
|
|
chr14_-_94669547
|
0.492
|
NM_001195573
|
DICER1
|
dicer 1, ribonuclease type III
|
|
chr12_+_8499788
|
0.487
|
NM_001007033
|
CLEC6A
|
C-type lectin domain family 6, member A
|
|
chr3_+_82117978
|
0.485
|
|
|
|
|
chr1_-_7998265
|
0.482
|
|
ERRFI1
|
ERBB receptor feedback inhibitor 1
|
|
chr9_-_4656665
|
0.482
|
NM_001039395
|
C9orf68
|
chromosome 9 open reading frame 68
|
|
chr14_-_106202262
|
0.479
|
|
IGLJ3
IGHA1
|
immunoglobulin lambda joining 3
immunoglobulin heavy constant alpha 1
|
|
chr11_-_92570626
|
0.477
|
|
|
|
|
chr1_-_74971679
|
0.476
|
NM_001130042
NM_001130043
NM_001134759
NM_001889
|
CRYZ
|
crystallin, zeta (quinone reductase)
|
|
chr1_+_148603208
|
0.476
|
|
RPRD2
|
regulation of nuclear pre-mRNA domain containing 2
|
|
chr2_-_97775300
|
0.475
|
|
TMEM131
|
transmembrane protein 131
|
|
chr12_-_11066436
|
0.474
|
NM_176888
|
TAS2R19
|
taste receptor, type 2, member 19
|
|
chr6_-_24491498
|
0.474
|
NM_001195610
|
DCDC2
|
doublecortin domain containing 2
|
|
chr17_+_31415752
|
0.469
|
NM_002988
|
CCL18
|
chemokine (C-C motif) ligand 18 (pulmonary and activation-regulated)
|
|
chr4_+_1375339
|
0.464
|
NM_175918
|
CRIPAK
|
cysteine-rich PAK1 inhibitor
|
|
chr7_-_22518200
|
0.463
|
|
EEF1A1
|
eukaryotic translation elongation factor 1 alpha 1
|
|
chr5_-_13997588
|
0.460
|
NM_001369
|
DNAH5
|
dynein, axonemal, heavy chain 5
|
|
chr13_+_23742846
|
0.455
|
|
SPATA13
|
spermatogenesis associated 13
|
|
chrX_-_50573759
|
0.451
|
NM_020717
|
SHROOM4
|
shroom family member 4
|
|
chr5_-_75505874
|
0.449
|
|
RAP1BL
|
RAP1B, member of RAS oncogene family pseudogene
|
|
chr11_+_4564596
|
0.447
|
NM_001005170
|
OR52I2
|
olfactory receptor, family 52, subfamily I, member 2
|
|
chr6_+_100161370
|
0.447
|
NM_021620
|
PRDM13
|
PR domain containing 13
|
|
chr6_+_114285209
|
0.445
|
NM_002356
|
MARCKS
|
myristoylated alanine-rich protein kinase C substrate
|
|
chr3_+_46591048
|
0.444
|
NM_001174136
|
TDGF1
|
teratocarcinoma-derived growth factor 1
|
|
chr1_-_195302973
|
0.442
|
NM_001994
|
F13B
|
coagulation factor XIII, B polypeptide
|
|
chr8_-_82769761
|
0.437
|
NM_001010893
|
SLC10A5
|
solute carrier family 10 (sodium/bile acid cotransporter family), member 5
|
|
chr13_-_32678142
|
0.435
|
NM_052851
|
STARD13
|
StAR-related lipid transfer (START) domain containing 13
|
|
chr6_-_116488458
|
0.432
|
NM_002031
|
FRK
|
fyn-related kinase
|
|
chr5_+_140835788
|
0.430
|
|
PCDHGC3
|
protocadherin gamma subfamily C, 3
|
|
chr14_+_38653177
|
0.429
|
NM_001009182
NM_001009183
NM_003616
|
SIP1
|
survival of motor neuron protein interacting protein 1
|
|
chr9_+_22103542
|
0.428
|
|
CDKN2B-AS1
|
CDKN2B antisense RNA 1 (non-protein coding)
|
|
chr8_-_72437020
|
0.427
|
NM_000503
|
EYA1
|
eyes absent homolog 1 (Drosophila)
|
|
chr6_-_128883125
|
0.427
|
NM_002844
NM_001135648
|
PTPRK
|
protein tyrosine phosphatase, receptor type, K
|
|
chr11_+_5862076
|
0.425
|
NM_001005165
|
OR52E4
|
olfactory receptor, family 52, subfamily E, member 4
|
|
chr15_+_71953020
|
0.425
|
NM_153356
|
TBC1D21
|
TBC1 domain family, member 21
|
|
chr7_+_54577512
|
0.424
|
NM_182546
|
VSTM2A
|
V-set and transmembrane domain containing 2A
|
|
chr14_-_19843918
|
0.424
|
|
TTC5
|
tetratricopeptide repeat domain 5
|
|
chr14_+_38772870
|
0.423
|
NM_054024
|
MIA2
|
melanoma inhibitory activity 2
|
|
chr11_+_54891935
|
0.423
|
NM_001005275
|
OR4A15
|
olfactory receptor, family 4, subfamily A, member 15
|
|
chr2_-_165768798
|
0.422
|
NM_001081676
NM_001081677
NM_006922
|
SCN3A
|
sodium channel, voltage-gated, type III, alpha subunit
|
|
chr8_+_24207524
|
0.422
|
NM_014265
NM_021777
|
ADAM28
|
ADAM metallopeptidase domain 28
|
|
chr3_+_99699137
|
0.417
|
NM_001004737
|
OR5K2
|
olfactory receptor, family 5, subfamily K, member 2
|
|
chr13_+_107753764
|
0.417
|
|
|
|
|
chr17_+_58625086
|
0.417
|
|
TANC2
|
tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 2
|
|
chr6_-_130578110
|
0.417
|
NM_152552
|
SAMD3
|
sterile alpha motif domain containing 3
|
|
chr11_+_9552809
|
0.415
|
NM_001143976
|
WEE1
|
WEE1 homolog (S. pombe)
|
|
chr2_-_128138435
|
0.414
|
NM_017980
|
LIMS2
|
LIM and senescent cell antigen-like domains 2
|
|
chr13_+_77213295
|
0.414
|
NM_144595
|
SLAIN1
|
SLAIN motif family, member 1
|
|
chr21_-_30583702
|
0.413
|
NM_001128598
|
KRTAP25-1
|
keratin associated protein 25-1
|
|
chr16_+_68284252
|
0.413
|
|
NFAT5
|
nuclear factor of activated T-cells 5, tonicity-responsive
|
|
chr7_+_156128431
|
0.412
|
NM_001184997
|
RNF32
|
ring finger protein 32
|
|
chr11_-_123130435
|
0.411
|
NM_001005188
|
OR6X1
|
olfactory receptor, family 6, subfamily X, member 1
|
|
chr8_+_24297742
|
0.409
|
NM_001145271
NM_001145272
NM_014479
|
ADAMDEC1
|
ADAM-like, decysin 1
|
|
chr8_+_67201684
|
0.406
|
NM_033058
NM_184085
NM_184086
NM_184087
|
TRIM55
|
tripartite motif containing 55
|
|
chr2_-_179380301
|
0.405
|
NM_003319
NM_133378
NM_133379
NM_133432
NM_133437
|
TTN
|
titin
|
|
chr15_-_39195701
|
0.404
|
|
INO80
|
INO80 homolog (S. cerevisiae)
|
|
chr9_+_115265827
|
0.403
|
NM_017790
|
RGS3
|
regulator of G-protein signaling 3
|
|
chr19_+_12810258
|
0.403
|
NM_014975
|
MAST1
|
microtubule associated serine/threonine kinase 1
|
|
chr2_-_180319013
|
0.402
|
NM_001113397
|
ZNF385B
|
zinc finger protein 385B
|
|
chr7_-_105009160
|
0.401
|
|
EFCAB10
|
EF-hand calcium binding domain 10
|
|
chr5_+_140835727
|
0.400
|
NM_002588
NM_032402
NM_032403
|
PCDHGC3
|
protocadherin gamma subfamily C, 3
|
|
chr1_+_84402962
|
0.399
|
|
PRKACB
|
protein kinase, cAMP-dependent, catalytic, beta
|
|
chrX_+_15435351
|
0.399
|
NM_001721
|
BMX
|
BMX non-receptor tyrosine kinase
|
|
chr10_-_98326435
|
0.397
|
|
TM9SF3
|
transmembrane 9 superfamily member 3
|
|
chr2_-_236837726
|
0.396
|
NM_212556
|
ASB18
|
ankyrin repeat and SOCS box containing 18
|
|
chr3_+_99484421
|
0.394
|
NM_001005482
|
OR5H2
|
olfactory receptor, family 5, subfamily H, member 2
|
|
chr2_-_178461710
|
0.394
|
NM_001077196
|
PDE11A
|
phosphodiesterase 11A
|
|
chr1_+_176577228
|
0.391
|
NM_004841
|
RASAL2
|
RAS protein activator like 2
|
|
chr11_-_35398077
|
0.391
|
NM_001195728
|
SLC1A2
|
solute carrier family 1 (glial high affinity glutamate transporter), member 2
|
|
chr4_+_124537334
|
0.390
|
NM_199327
|
SPRY1
|
sprouty homolog 1, antagonist of FGF signaling (Drosophila)
|
|
chr2_+_108637698
|
0.388
|
NM_001193485
|
LIMS1
|
LIM and senescent cell antigen-like domains 1
|
|
chr11_+_5925152
|
0.388
|
NM_001003443
|
OR56A3
|
olfactory receptor, family 56, subfamily A, member 3
|
|
chr2_-_180135559
|
0.386
|
NM_001113398
|
ZNF385B
|
zinc finger protein 385B
|
|
chr19_-_6230797
|
0.386
|
|
|
|
|
chr3_-_147805715
|
0.386
|
|
PLSCR5
|
phospholipid scramblase family, member 5
|
|
chr9_+_134884627
|
0.385
|
|
EEF1A1
|
eukaryotic translation elongation factor 1 alpha 1
|
|
chr2_+_108230082
|
0.384
|
NM_001008743
|
SULT1C3
|
sulfotransferase family, cytosolic, 1C, member 3
|
|
chr2_+_211129567
|
0.382
|
NM_001875
|
CPS1
|
carbamoyl-phosphate synthase 1, mitochondrial
|
|
chrX_-_116991728
|
0.382
|
NM_033495
NM_001168300
|
KLHL13
|
kelch-like 13 (Drosophila)
|
|
chr3_-_113701073
|
0.379
|
NM_001085357
NM_181780
|
BTLA
|
B and T lymphocyte associated
|
|
chr18_-_49944986
|
0.379
|
|
MBD2
|
methyl-CpG binding domain protein 2
|
|
chr3_+_186529376
|
0.377
|
|
MAP3K13
|
mitogen-activated protein kinase kinase kinase 13
|
|
chr14_-_87859284
|
0.375
|
NM_138317
|
KCNK10
|
potassium channel, subfamily K, member 10
|
|
chr10_-_98311813
|
0.373
|
|
TM9SF3
|
transmembrane 9 superfamily member 3
|
|
chr6_-_52818851
|
0.373
|
NM_153699
|
GSTA5
|
glutathione S-transferase alpha 5
|
|
chr12_-_368517
|
0.372
|
|
KDM5A
|
lysine (K)-specific demethylase 5A
|
|
chr16_+_48857962
|
0.372
|
|
ADCY7
|
adenylate cyclase 7
|
|
chr4_-_68432293
|
0.370
|
NM_004262
|
TMPRSS11D
|
transmembrane protease, serine 11D
|
|
chr5_-_38631263
|
0.370
|
NM_002310
|
LIFR
|
leukemia inhibitory factor receptor alpha
|
|
chr11_-_51269023
|
0.368
|
NM_001005272
|
OR4A5
|
olfactory receptor, family 4, subfamily A, member 5
|
|
chr5_-_147266257
|
0.367
|
NM_206966
|
C5orf46
|
chromosome 5 open reading frame 46
|