Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
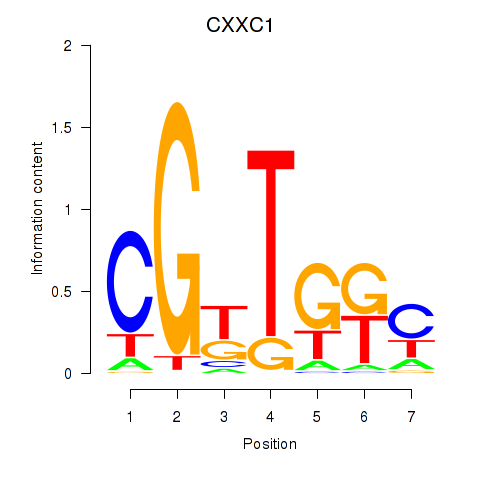
Results for CXXC1
Z-value: 1.87
Transcription factors associated with CXXC1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
CXXC1
|
ENSG00000154832.16 | CXXC1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| CXXC1 | hg38_v1_chr18_-_50287570_50287709 | 0.22 | 8.4e-04 | Click! |
Activity profile of CXXC1 motif
Sorted Z-values of CXXC1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of CXXC1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 14.6 | 43.8 | GO:0060623 | regulation of chromosome condensation(GO:0060623) cellular response to iron(III) ion(GO:0071283) |
| 12.8 | 38.4 | GO:0044878 | mitotic cytokinesis checkpoint(GO:0044878) |
| 11.2 | 33.5 | GO:0030327 | prenylated protein catabolic process(GO:0030327) |
| 10.9 | 43.5 | GO:0000961 | negative regulation of mitochondrial RNA catabolic process(GO:0000961) |
| 10.4 | 31.1 | GO:1990022 | RNA polymerase II complex import to nucleus(GO:0044376) RNA polymerase III complex localization to nucleus(GO:1990022) |
| 10.0 | 39.9 | GO:1902544 | regulation of DNA N-glycosylase activity(GO:1902544) |
| 9.4 | 37.8 | GO:0009224 | CMP salvage(GO:0006238) CMP biosynthetic process(GO:0009224) CMP metabolic process(GO:0046035) |
| 9.3 | 112.0 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 8.3 | 33.4 | GO:1903609 | negative regulation of peptidyl-tyrosine autophosphorylation(GO:1900085) negative regulation of inward rectifier potassium channel activity(GO:1903609) |
| 8.3 | 49.7 | GO:0006235 | dTTP biosynthetic process(GO:0006235) pyrimidine deoxyribonucleoside triphosphate biosynthetic process(GO:0009212) |
| 7.6 | 22.8 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 7.4 | 22.2 | GO:1903093 | regulation of protein K48-linked deubiquitination(GO:1903093) negative regulation of protein K48-linked deubiquitination(GO:1903094) negative regulation of ubiquitin-specific protease activity(GO:2000157) |
| 7.1 | 21.4 | GO:0005999 | xylulose biosynthetic process(GO:0005999) |
| 7.0 | 7.0 | GO:0043504 | mitochondrial DNA repair(GO:0043504) |
| 6.8 | 34.2 | GO:1902626 | assembly of large subunit precursor of preribosome(GO:1902626) |
| 6.6 | 13.3 | GO:0046075 | dTTP metabolic process(GO:0046075) |
| 6.1 | 24.4 | GO:0072683 | T cell extravasation(GO:0072683) |
| 6.0 | 42.0 | GO:0046940 | nucleoside monophosphate phosphorylation(GO:0046940) |
| 6.0 | 17.9 | GO:0046292 | formaldehyde metabolic process(GO:0046292) |
| 5.5 | 38.3 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 5.1 | 15.4 | GO:1902990 | mitotic telomere maintenance via semi-conservative replication(GO:1902990) |
| 5.1 | 40.9 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 5.1 | 15.3 | GO:0043465 | fermentation(GO:0006113) regulation of fermentation(GO:0043465) |
| 5.0 | 15.0 | GO:0046452 | dihydrofolate metabolic process(GO:0046452) |
| 4.8 | 14.5 | GO:0006097 | glyoxylate cycle(GO:0006097) |
| 4.8 | 28.7 | GO:0007079 | mitotic chromosome movement towards spindle pole(GO:0007079) |
| 4.7 | 18.9 | GO:1901873 | regulation of post-translational protein modification(GO:1901873) |
| 4.7 | 14.1 | GO:0006781 | succinyl-CoA pathway(GO:0006781) |
| 4.7 | 4.7 | GO:0072434 | signal transduction involved in G2 DNA damage checkpoint(GO:0072425) signal transduction involved in mitotic G2 DNA damage checkpoint(GO:0072434) |
| 4.6 | 18.6 | GO:0019418 | sulfide oxidation(GO:0019418) sulfide oxidation, using sulfide:quinone oxidoreductase(GO:0070221) |
| 4.6 | 13.9 | GO:0034239 | macrophage fusion(GO:0034238) regulation of macrophage fusion(GO:0034239) positive regulation of macrophage fusion(GO:0034241) |
| 4.4 | 8.8 | GO:1903719 | regulation of I-kappaB phosphorylation(GO:1903719) positive regulation of I-kappaB phosphorylation(GO:1903721) |
| 4.4 | 26.1 | GO:0016321 | female meiosis chromosome segregation(GO:0016321) |
| 4.2 | 12.7 | GO:2000048 | negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 4.2 | 8.4 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 4.2 | 29.2 | GO:0046985 | histone H4-R3 methylation(GO:0043985) negative regulation of megakaryocyte differentiation(GO:0045653) positive regulation of hemoglobin biosynthetic process(GO:0046985) |
| 4.1 | 49.6 | GO:0010917 | negative regulation of mitochondrial membrane potential(GO:0010917) |
| 4.1 | 16.3 | GO:0060715 | syncytiotrophoblast cell differentiation involved in labyrinthine layer development(GO:0060715) |
| 4.0 | 8.0 | GO:0000454 | snoRNA guided rRNA pseudouridine synthesis(GO:0000454) |
| 4.0 | 15.9 | GO:0055014 | atrial cardiac muscle cell differentiation(GO:0055011) atrial cardiac muscle cell development(GO:0055014) |
| 3.9 | 15.4 | GO:0009438 | methylglyoxal metabolic process(GO:0009438) |
| 3.8 | 19.0 | GO:0090069 | regulation of ribosome biogenesis(GO:0090069) |
| 3.6 | 21.6 | GO:0006729 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 3.6 | 10.7 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 3.5 | 14.2 | GO:0090094 | metanephric cap development(GO:0072185) metanephric cap morphogenesis(GO:0072186) metanephric cap mesenchymal cell proliferation involved in metanephros development(GO:0090094) regulation of metanephric cap mesenchymal cell proliferation(GO:0090095) positive regulation of metanephric cap mesenchymal cell proliferation(GO:0090096) |
| 3.5 | 10.4 | GO:0036245 | cellular response to menadione(GO:0036245) |
| 3.4 | 20.4 | GO:0006021 | inositol biosynthetic process(GO:0006021) |
| 3.4 | 10.1 | GO:0061011 | hepatic duct development(GO:0061011) |
| 3.3 | 6.7 | GO:0005997 | xylulose metabolic process(GO:0005997) |
| 3.3 | 10.0 | GO:0090149 | synaptic vesicle recycling via endosome(GO:0036466) mitochondrial membrane fission(GO:0090149) |
| 3.3 | 19.7 | GO:0032383 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 3.2 | 22.7 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 3.2 | 16.2 | GO:0006231 | dTMP biosynthetic process(GO:0006231) dTMP metabolic process(GO:0046073) |
| 3.2 | 63.7 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 3.1 | 18.7 | GO:0002176 | male germ cell proliferation(GO:0002176) germ cell proliferation(GO:0036093) |
| 3.1 | 9.3 | GO:1903644 | regulation of chaperone-mediated protein folding(GO:1903644) |
| 3.1 | 3.1 | GO:1903630 | regulation of aminoacyl-tRNA ligase activity(GO:1903630) |
| 3.1 | 18.6 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 3.1 | 18.5 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 3.0 | 12.2 | GO:0007144 | female meiosis I(GO:0007144) |
| 3.0 | 21.2 | GO:0045047 | protein targeting to ER(GO:0045047) |
| 3.0 | 12.1 | GO:1902775 | mitochondrial ribosome assembly(GO:0061668) mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 3.0 | 12.1 | GO:1904154 | positive regulation of retrograde protein transport, ER to cytosol(GO:1904154) |
| 3.0 | 9.0 | GO:0046504 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) ether biosynthetic process(GO:1901503) |
| 3.0 | 14.9 | GO:0031120 | snRNA pseudouridine synthesis(GO:0031120) |
| 3.0 | 17.8 | GO:0042780 | tRNA 3'-end processing(GO:0042780) |
| 3.0 | 17.8 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 2.9 | 8.7 | GO:0007057 | spindle assembly involved in female meiosis I(GO:0007057) positive regulation of oocyte maturation(GO:1900195) |
| 2.8 | 14.1 | GO:0046654 | tetrahydrofolate biosynthetic process(GO:0046654) |
| 2.7 | 13.7 | GO:1904925 | positive regulation of macromitophagy(GO:1901526) positive regulation of mitophagy in response to mitochondrial depolarization(GO:1904925) |
| 2.7 | 21.6 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 2.7 | 8.1 | GO:0034402 | mRNA export from nucleus in response to heat stress(GO:0031990) recruitment of 3'-end processing factors to RNA polymerase II holoenzyme complex(GO:0034402) |
| 2.7 | 26.7 | GO:0009146 | purine nucleoside triphosphate catabolic process(GO:0009146) |
| 2.7 | 21.4 | GO:1901856 | negative regulation of cellular respiration(GO:1901856) |
| 2.7 | 26.7 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 2.7 | 202.0 | GO:0070125 | mitochondrial translational elongation(GO:0070125) |
| 2.6 | 7.7 | GO:2000616 | negative regulation of histone H3-K9 acetylation(GO:2000616) |
| 2.6 | 10.2 | GO:0018106 | peptidyl-histidine phosphorylation(GO:0018106) |
| 2.5 | 38.2 | GO:0045324 | late endosome to vacuole transport(GO:0045324) |
| 2.5 | 7.6 | GO:0072019 | proximal convoluted tubule development(GO:0072019) metanephric proximal convoluted tubule development(GO:0072229) |
| 2.5 | 7.6 | GO:0030920 | N-terminal peptidyl-serine acetylation(GO:0017198) N-terminal peptidyl-glutamic acid acetylation(GO:0018002) peptidyl-serine acetylation(GO:0030920) |
| 2.5 | 7.5 | GO:1905000 | regulation of membrane repolarization during atrial cardiac muscle cell action potential(GO:1905000) |
| 2.5 | 42.1 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 2.5 | 81.3 | GO:0007094 | mitotic spindle assembly checkpoint(GO:0007094) spindle assembly checkpoint(GO:0071173) |
| 2.5 | 22.1 | GO:0060613 | fat pad development(GO:0060613) |
| 2.5 | 29.4 | GO:0071550 | death-inducing signaling complex assembly(GO:0071550) |
| 2.4 | 19.5 | GO:2000816 | negative regulation of mitotic sister chromatid separation(GO:2000816) |
| 2.4 | 7.3 | GO:1904732 | regulation of electron carrier activity(GO:1904732) |
| 2.4 | 9.7 | GO:0035606 | peptidyl-cysteine S-trans-nitrosylation(GO:0035606) |
| 2.4 | 11.9 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 2.4 | 23.6 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 2.3 | 30.3 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 2.3 | 16.2 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 2.3 | 11.3 | GO:1902527 | positive regulation of protein monoubiquitination(GO:1902527) |
| 2.2 | 9.0 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 2.2 | 17.8 | GO:0097411 | hypoxia-inducible factor-1alpha signaling pathway(GO:0097411) |
| 2.2 | 57.8 | GO:0097341 | inhibition of cysteine-type endopeptidase activity(GO:0097340) zymogen inhibition(GO:0097341) |
| 2.2 | 13.3 | GO:0015866 | ADP transport(GO:0015866) |
| 2.1 | 12.6 | GO:0035549 | positive regulation of interferon-beta secretion(GO:0035549) |
| 2.1 | 10.5 | GO:0039019 | posttranslational protein targeting to membrane(GO:0006620) pronephric nephron development(GO:0039019) |
| 2.1 | 18.8 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 2.1 | 8.3 | GO:1904116 | response to vasopressin(GO:1904116) cellular response to vasopressin(GO:1904117) |
| 2.1 | 20.5 | GO:0070601 | centromeric sister chromatid cohesion(GO:0070601) |
| 2.0 | 6.0 | GO:0048250 | mitochondrial iron ion transport(GO:0048250) |
| 2.0 | 7.9 | GO:0051182 | coenzyme transport(GO:0051182) |
| 2.0 | 9.9 | GO:0001315 | age-dependent response to oxidative stress(GO:0001306) age-dependent response to reactive oxygen species(GO:0001315) regulation of systemic arterial blood pressure by acetylcholine(GO:0003068) vasodilation by acetylcholine involved in regulation of systemic arterial blood pressure(GO:0003069) regulation of systemic arterial blood pressure by neurotransmitter(GO:0003070) age-dependent general metabolic decline(GO:0007571) |
| 1.9 | 7.6 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 1.9 | 9.4 | GO:0044791 | modulation by host of viral release from host cell(GO:0044789) positive regulation by host of viral release from host cell(GO:0044791) |
| 1.9 | 22.6 | GO:0070986 | left/right axis specification(GO:0070986) |
| 1.9 | 11.2 | GO:1904885 | beta-catenin destruction complex assembly(GO:1904885) |
| 1.9 | 9.3 | GO:0033383 | geranyl diphosphate metabolic process(GO:0033383) geranyl diphosphate biosynthetic process(GO:0033384) farnesyl diphosphate biosynthetic process(GO:0045337) |
| 1.8 | 14.3 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 1.8 | 7.1 | GO:0097327 | response to antineoplastic agent(GO:0097327) |
| 1.8 | 17.5 | GO:1901409 | positive regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901409) |
| 1.7 | 15.4 | GO:0006528 | asparagine metabolic process(GO:0006528) |
| 1.7 | 6.8 | GO:0032262 | pyrimidine ribonucleotide salvage(GO:0010138) pyrimidine nucleotide salvage(GO:0032262) UMP salvage(GO:0044206) |
| 1.7 | 10.1 | GO:0060356 | leucine import(GO:0060356) |
| 1.7 | 8.4 | GO:0070197 | meiotic telomere tethering at nuclear periphery(GO:0044821) meiotic attachment of telomere to nuclear envelope(GO:0070197) chromosome attachment to the nuclear envelope(GO:0097240) |
| 1.7 | 10.1 | GO:0006196 | AMP catabolic process(GO:0006196) adenosine biosynthetic process(GO:0046086) |
| 1.7 | 8.3 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 1.6 | 52.7 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 1.6 | 9.8 | GO:0019060 | intracellular transport of viral protein in host cell(GO:0019060) symbiont intracellular protein transport in host(GO:0030581) intracellular protein transport in other organism involved in symbiotic interaction(GO:0051708) |
| 1.6 | 8.2 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 1.6 | 11.3 | GO:0032218 | riboflavin transport(GO:0032218) |
| 1.6 | 4.8 | GO:0042357 | thiamine diphosphate metabolic process(GO:0042357) |
| 1.6 | 22.0 | GO:0000920 | cell separation after cytokinesis(GO:0000920) |
| 1.6 | 6.3 | GO:0030961 | peptidyl-arginine hydroxylation(GO:0030961) |
| 1.6 | 7.8 | GO:0046391 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 1.5 | 20.1 | GO:1900625 | regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 1.5 | 13.9 | GO:0075525 | viral translational termination-reinitiation(GO:0075525) |
| 1.5 | 4.6 | GO:0042939 | glutathione transport(GO:0034635) tripeptide transport(GO:0042939) |
| 1.5 | 18.1 | GO:0070141 | response to UV-A(GO:0070141) |
| 1.5 | 3.0 | GO:0071603 | endothelial cell-cell adhesion(GO:0071603) |
| 1.5 | 19.3 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 1.5 | 20.6 | GO:1900364 | negative regulation of mRNA polyadenylation(GO:1900364) |
| 1.5 | 4.4 | GO:0045554 | TRAIL biosynthetic process(GO:0045553) regulation of TRAIL biosynthetic process(GO:0045554) positive regulation of TRAIL biosynthetic process(GO:0045556) |
| 1.4 | 39.7 | GO:0008334 | histone mRNA metabolic process(GO:0008334) |
| 1.4 | 6.8 | GO:0015862 | uridine transport(GO:0015862) |
| 1.4 | 2.7 | GO:0019249 | lactate biosynthetic process(GO:0019249) |
| 1.4 | 16.3 | GO:0010965 | regulation of mitotic sister chromatid separation(GO:0010965) mitotic sister chromatid separation(GO:0051306) |
| 1.3 | 4.0 | GO:0033313 | meiotic cell cycle checkpoint(GO:0033313) |
| 1.3 | 4.0 | GO:0036493 | positive regulation of translation in response to endoplasmic reticulum stress(GO:0036493) |
| 1.3 | 21.3 | GO:0032968 | positive regulation of transcription elongation from RNA polymerase II promoter(GO:0032968) |
| 1.3 | 10.6 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 1.3 | 21.1 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 1.3 | 5.2 | GO:0050916 | sensory perception of sweet taste(GO:0050916) sensory perception of umami taste(GO:0050917) |
| 1.3 | 15.5 | GO:0000491 | small nucleolar ribonucleoprotein complex assembly(GO:0000491) box C/D snoRNP assembly(GO:0000492) |
| 1.3 | 9.0 | GO:0090666 | scaRNA localization to Cajal body(GO:0090666) |
| 1.3 | 9.0 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 1.3 | 11.4 | GO:0032057 | negative regulation of translational initiation in response to stress(GO:0032057) |
| 1.3 | 12.5 | GO:0002934 | desmosome organization(GO:0002934) |
| 1.2 | 32.3 | GO:0032201 | telomere maintenance via semi-conservative replication(GO:0032201) |
| 1.2 | 3.7 | GO:0035544 | negative regulation of SNARE complex assembly(GO:0035544) |
| 1.2 | 4.9 | GO:1903758 | regulation of transcription from RNA polymerase II promoter by histone modification(GO:1903756) negative regulation of transcription from RNA polymerase II promoter by histone modification(GO:1903758) |
| 1.2 | 27.9 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 1.2 | 2.3 | GO:0052501 | induction of programmed cell death(GO:0012502) positive regulation of apoptotic process in other organism(GO:0044533) positive regulation by symbiont of host programmed cell death(GO:0052042) positive regulation by organism of programmed cell death in other organism involved in symbiotic interaction(GO:0052330) positive regulation by organism of apoptotic process in other organism involved in symbiotic interaction(GO:0052501) |
| 1.1 | 14.7 | GO:0006085 | acetyl-CoA biosynthetic process(GO:0006085) |
| 1.1 | 102.4 | GO:0051439 | negative regulation of ubiquitin-protein ligase activity involved in mitotic cell cycle(GO:0051436) regulation of ubiquitin-protein ligase activity involved in mitotic cell cycle(GO:0051439) |
| 1.1 | 7.7 | GO:0000389 | mRNA 3'-splice site recognition(GO:0000389) |
| 1.1 | 4.4 | GO:0048388 | endosomal lumen acidification(GO:0048388) |
| 1.1 | 4.3 | GO:0036518 | chemorepulsion of dopaminergic neuron axon(GO:0036518) chemorepulsion of axon(GO:0061643) |
| 1.1 | 10.7 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 1.1 | 6.3 | GO:0002943 | tRNA dihydrouridine synthesis(GO:0002943) |
| 1.0 | 10.4 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 1.0 | 3.1 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 1.0 | 33.9 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 1.0 | 14.2 | GO:0048262 | determination of dorsal/ventral asymmetry(GO:0048262) |
| 1.0 | 130.5 | GO:0006613 | cotranslational protein targeting to membrane(GO:0006613) |
| 1.0 | 27.9 | GO:0002183 | cytoplasmic translational initiation(GO:0002183) |
| 1.0 | 2.9 | GO:0006117 | acetaldehyde metabolic process(GO:0006117) |
| 1.0 | 9.7 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 1.0 | 5.8 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 1.0 | 10.7 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 1.0 | 2.9 | GO:2000767 | positive regulation of cytoplasmic translation(GO:2000767) |
| 0.9 | 2.8 | GO:2000850 | negative regulation of steroid hormone secretion(GO:2000832) negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 0.9 | 5.7 | GO:0097384 | cellular lipid biosynthetic process(GO:0097384) |
| 0.9 | 34.4 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.9 | 5.5 | GO:1990564 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.9 | 5.5 | GO:0060368 | regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060368) |
| 0.9 | 23.0 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.9 | 3.6 | GO:1903755 | regulation of SUMO transferase activity(GO:1903182) positive regulation of SUMO transferase activity(GO:1903755) |
| 0.9 | 0.9 | GO:0021539 | subthalamus development(GO:0021539) |
| 0.9 | 4.5 | GO:1904796 | synaptic vesicle targeting(GO:0016080) regulation of core promoter binding(GO:1904796) |
| 0.9 | 9.6 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 0.9 | 5.1 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.9 | 13.7 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.9 | 12.0 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.8 | 5.1 | GO:0008582 | regulation of synaptic growth at neuromuscular junction(GO:0008582) |
| 0.8 | 5.0 | GO:0090206 | negative regulation of cholesterol biosynthetic process(GO:0045541) negative regulation of cholesterol metabolic process(GO:0090206) |
| 0.8 | 2.5 | GO:1903978 | regulation of microglial cell activation(GO:1903978) positive regulation of lysosomal protein catabolic process(GO:1905167) |
| 0.8 | 6.7 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.8 | 7.5 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 0.8 | 7.5 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.8 | 3.3 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.8 | 5.8 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.8 | 4.1 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.8 | 5.7 | GO:2000074 | regulation of type B pancreatic cell development(GO:2000074) |
| 0.8 | 3.2 | GO:0060743 | epithelial cell maturation involved in prostate gland development(GO:0060743) |
| 0.8 | 23.8 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.8 | 6.2 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.8 | 4.6 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.8 | 6.9 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.8 | 6.1 | GO:0009169 | ribonucleoside monophosphate catabolic process(GO:0009158) purine ribonucleoside monophosphate catabolic process(GO:0009169) |
| 0.7 | 3.0 | GO:1903553 | positive regulation of extracellular exosome assembly(GO:1903553) |
| 0.7 | 21.9 | GO:0031639 | plasminogen activation(GO:0031639) |
| 0.7 | 11.6 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.7 | 8.7 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.7 | 4.3 | GO:0050747 | positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.7 | 2.1 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.7 | 35.9 | GO:0000281 | mitotic cytokinesis(GO:0000281) |
| 0.7 | 9.1 | GO:2000622 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.7 | 6.2 | GO:0061000 | negative regulation of dendritic spine development(GO:0061000) |
| 0.7 | 46.6 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.7 | 15.0 | GO:1904406 | negative regulation of nitric oxide biosynthetic process(GO:0045019) negative regulation of nitric oxide metabolic process(GO:1904406) |
| 0.7 | 10.9 | GO:0031268 | pseudopodium organization(GO:0031268) |
| 0.7 | 4.7 | GO:0032328 | alanine transport(GO:0032328) |
| 0.7 | 19.3 | GO:0000715 | nucleotide-excision repair, DNA damage recognition(GO:0000715) |
| 0.7 | 4.6 | GO:1990253 | cellular response to leucine starvation(GO:1990253) |
| 0.7 | 5.9 | GO:0042074 | cell migration involved in gastrulation(GO:0042074) |
| 0.7 | 12.4 | GO:0043981 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.6 | 6.4 | GO:0015781 | pyrimidine nucleotide-sugar transport(GO:0015781) |
| 0.6 | 3.8 | GO:0035986 | senescence-associated heterochromatin focus assembly(GO:0035986) |
| 0.6 | 3.8 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.6 | 3.8 | GO:0045006 | DNA deamination(GO:0045006) DNA cytosine deamination(GO:0070383) |
| 0.6 | 3.8 | GO:0090270 | fibroblast growth factor production(GO:0090269) regulation of fibroblast growth factor production(GO:0090270) |
| 0.6 | 17.0 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.6 | 19.9 | GO:0032435 | negative regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032435) |
| 0.6 | 23.6 | GO:0071431 | tRNA export from nucleus(GO:0006409) tRNA transport(GO:0051031) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.6 | 1.2 | GO:0040031 | snRNA modification(GO:0040031) |
| 0.6 | 3.5 | GO:0098838 | methotrexate transport(GO:0051958) reduced folate transmembrane transport(GO:0098838) |
| 0.6 | 17.4 | GO:0030207 | chondroitin sulfate catabolic process(GO:0030207) |
| 0.6 | 8.7 | GO:0034356 | NAD biosynthesis via nicotinamide riboside salvage pathway(GO:0034356) |
| 0.6 | 2.3 | GO:0097676 | cell migration involved in vasculogenesis(GO:0035441) histone H3-K36 dimethylation(GO:0097676) |
| 0.6 | 2.3 | GO:0060335 | positive regulation of response to interferon-gamma(GO:0060332) positive regulation of interferon-gamma-mediated signaling pathway(GO:0060335) |
| 0.6 | 3.9 | GO:0030421 | defecation(GO:0030421) |
| 0.6 | 14.0 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.6 | 28.9 | GO:0050434 | positive regulation of viral transcription(GO:0050434) |
| 0.6 | 2.8 | GO:0042986 | positive regulation of amyloid precursor protein biosynthetic process(GO:0042986) |
| 0.6 | 24.9 | GO:0043330 | response to exogenous dsRNA(GO:0043330) |
| 0.6 | 6.1 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.5 | 13.6 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.5 | 15.2 | GO:0050855 | regulation of B cell receptor signaling pathway(GO:0050855) |
| 0.5 | 2.2 | GO:0006868 | glutamine transport(GO:0006868) glutamine secretion(GO:0010585) L-glutamine import(GO:0036229) L-glutamine import into cell(GO:1903803) |
| 0.5 | 21.6 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.5 | 10.7 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.5 | 2.1 | GO:0051873 | killing by host of symbiont cells(GO:0051873) |
| 0.5 | 11.6 | GO:0060397 | JAK-STAT cascade involved in growth hormone signaling pathway(GO:0060397) |
| 0.5 | 1.6 | GO:2000520 | regulation of immunological synapse formation(GO:2000520) negative regulation of immunological synapse formation(GO:2000521) |
| 0.5 | 2.1 | GO:1900041 | negative regulation of interleukin-2 secretion(GO:1900041) |
| 0.5 | 11.7 | GO:0000732 | strand displacement(GO:0000732) |
| 0.5 | 2.0 | GO:1905205 | positive regulation of connective tissue replacement(GO:1905205) |
| 0.5 | 4.5 | GO:0031584 | activation of phospholipase D activity(GO:0031584) |
| 0.5 | 6.5 | GO:1904355 | positive regulation of telomere capping(GO:1904355) |
| 0.5 | 9.9 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.5 | 22.6 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.5 | 6.5 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.5 | 44.4 | GO:0001909 | leukocyte mediated cytotoxicity(GO:0001909) |
| 0.4 | 1.8 | GO:0032913 | B cell receptor transport within lipid bilayer(GO:0032595) B cell receptor transport into membrane raft(GO:0032597) protein transport out of membrane raft(GO:0032599) chemokine receptor transport out of membrane raft(GO:0032600) negative regulation of transforming growth factor beta3 production(GO:0032913) chemokine receptor transport within lipid bilayer(GO:0033606) |
| 0.4 | 3.1 | GO:1904526 | regulation of microtubule binding(GO:1904526) |
| 0.4 | 1.7 | GO:0090472 | viral protein processing(GO:0019082) regulation of nerve growth factor production(GO:0032903) negative regulation of nerve growth factor production(GO:0032904) dibasic protein processing(GO:0090472) |
| 0.4 | 9.6 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.4 | 21.3 | GO:0009301 | snRNA transcription(GO:0009301) snRNA transcription from RNA polymerase II promoter(GO:0042795) |
| 0.4 | 3.8 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.4 | 30.2 | GO:0032981 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.4 | 7.2 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.4 | 5.9 | GO:2000659 | regulation of interleukin-1-mediated signaling pathway(GO:2000659) |
| 0.4 | 12.1 | GO:0072663 | protein targeting to peroxisome(GO:0006625) protein localization to peroxisome(GO:0072662) establishment of protein localization to peroxisome(GO:0072663) |
| 0.4 | 15.7 | GO:0015949 | nucleobase-containing small molecule interconversion(GO:0015949) |
| 0.4 | 6.2 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.4 | 8.1 | GO:0032516 | positive regulation of phosphoprotein phosphatase activity(GO:0032516) |
| 0.4 | 7.7 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.4 | 1.2 | GO:0097527 | necroptotic signaling pathway(GO:0097527) |
| 0.4 | 1.5 | GO:0031064 | negative regulation of histone deacetylation(GO:0031064) negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.4 | 3.7 | GO:0046325 | negative regulation of glucose import(GO:0046325) |
| 0.4 | 76.4 | GO:0000375 | RNA splicing, via transesterification reactions(GO:0000375) |
| 0.4 | 2.2 | GO:0061101 | neuroendocrine cell differentiation(GO:0061101) |
| 0.3 | 2.8 | GO:0034626 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.3 | 19.1 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.3 | 10.8 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.3 | 4.2 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.3 | 8.9 | GO:0042347 | negative regulation of NF-kappaB import into nucleus(GO:0042347) |
| 0.3 | 1.8 | GO:0006548 | histamine metabolic process(GO:0001692) histidine catabolic process(GO:0006548) imidazole-containing compound catabolic process(GO:0052805) |
| 0.3 | 6.1 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 0.3 | 1.8 | GO:0015680 | intracellular copper ion transport(GO:0015680) |
| 0.3 | 6.2 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.3 | 16.9 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.3 | 1.7 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.3 | 2.6 | GO:1903830 | magnesium ion transmembrane transport(GO:1903830) |
| 0.3 | 1.1 | GO:0017062 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) |
| 0.3 | 1.4 | GO:0033058 | directional locomotion(GO:0033058) |
| 0.3 | 10.1 | GO:0035455 | response to interferon-alpha(GO:0035455) |
| 0.3 | 1.9 | GO:0071692 | sequestering of TGFbeta in extracellular matrix(GO:0035583) protein localization to extracellular region(GO:0071692) maintenance of protein location in extracellular region(GO:0071694) |
| 0.3 | 10.6 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.3 | 10.2 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.3 | 3.5 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.3 | 2.1 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.3 | 0.5 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.3 | 0.8 | GO:0006982 | response to lipid hydroperoxide(GO:0006982) |
| 0.3 | 1.6 | GO:0098795 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing by siRNA(GO:0090625) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.3 | 3.2 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.3 | 24.7 | GO:0000910 | cytokinesis(GO:0000910) |
| 0.3 | 12.2 | GO:0043486 | histone exchange(GO:0043486) |
| 0.3 | 3.1 | GO:0090286 | cytoskeletal anchoring at nuclear membrane(GO:0090286) |
| 0.3 | 3.8 | GO:1904886 | beta-catenin destruction complex disassembly(GO:1904886) |
| 0.2 | 9.1 | GO:0006656 | phosphatidylcholine biosynthetic process(GO:0006656) |
| 0.2 | 2.6 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.2 | 6.8 | GO:0009060 | aerobic respiration(GO:0009060) |
| 0.2 | 2.3 | GO:2000543 | positive regulation of gastrulation(GO:2000543) |
| 0.2 | 3.0 | GO:0010452 | histone H3-K36 methylation(GO:0010452) |
| 0.2 | 1.6 | GO:1905123 | regulation of glucosylceramidase activity(GO:1905123) |
| 0.2 | 0.7 | GO:0071934 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 0.2 | 3.6 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.2 | 0.7 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.2 | 0.7 | GO:0070650 | endoplasmic reticulum polarization(GO:0061163) actin filament bundle retrograde transport(GO:0061573) actin filament bundle distribution(GO:0070650) |
| 0.2 | 4.4 | GO:0016578 | histone deubiquitination(GO:0016578) |
| 0.2 | 2.4 | GO:0017085 | response to insecticide(GO:0017085) |
| 0.2 | 3.5 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.2 | 1.1 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.2 | 0.9 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.2 | 6.4 | GO:0043537 | negative regulation of blood vessel endothelial cell migration(GO:0043537) |
| 0.2 | 2.1 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.2 | 1.7 | GO:0045647 | negative regulation of erythrocyte differentiation(GO:0045647) |
| 0.2 | 4.3 | GO:0036295 | cellular response to increased oxygen levels(GO:0036295) |
| 0.2 | 3.3 | GO:0007084 | mitotic nuclear envelope reassembly(GO:0007084) |
| 0.2 | 0.4 | GO:0046222 | mycotoxin metabolic process(GO:0043385) aflatoxin metabolic process(GO:0046222) organic heteropentacyclic compound metabolic process(GO:1901376) |
| 0.2 | 1.4 | GO:0071557 | histone H3-K27 demethylation(GO:0071557) |
| 0.2 | 4.8 | GO:0060716 | labyrinthine layer blood vessel development(GO:0060716) |
| 0.2 | 3.0 | GO:0034067 | protein localization to Golgi apparatus(GO:0034067) |
| 0.2 | 15.5 | GO:0033275 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.2 | 13.5 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.2 | 1.0 | GO:2001137 | positive regulation of endocytic recycling(GO:2001137) |
| 0.2 | 1.2 | GO:0033152 | immunoglobulin V(D)J recombination(GO:0033152) |
| 0.2 | 1.2 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.2 | 1.2 | GO:0051832 | negative regulation of cardiac muscle adaptation(GO:0010616) evasion or tolerance of host defenses by virus(GO:0019049) avoidance of host defenses(GO:0044413) evasion or tolerance of host defenses(GO:0044415) avoidance of defenses of other organism involved in symbiotic interaction(GO:0051832) evasion or tolerance of defenses of other organism involved in symbiotic interaction(GO:0051834) response to defenses of other organism involved in symbiotic interaction(GO:0052173) response to host defenses(GO:0052200) negative regulation of lung blood pressure(GO:0061767) response to host(GO:0075136) negative regulation of cardiac muscle hypertrophy in response to stress(GO:1903243) |
| 0.2 | 4.4 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.2 | 5.2 | GO:0021801 | cerebral cortex radial glia guided migration(GO:0021801) telencephalon glial cell migration(GO:0022030) |
| 0.2 | 11.7 | GO:0006901 | vesicle coating(GO:0006901) vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.2 | 0.7 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.2 | 1.1 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.2 | 0.7 | GO:0035562 | negative regulation of chromatin binding(GO:0035562) |
| 0.2 | 9.8 | GO:0045071 | negative regulation of viral genome replication(GO:0045071) |
| 0.2 | 8.7 | GO:0034383 | low-density lipoprotein particle clearance(GO:0034383) |
| 0.2 | 6.8 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.2 | 9.2 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.2 | 4.0 | GO:0097502 | mannosylation(GO:0097502) |
| 0.2 | 0.8 | GO:1902231 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902231) |
| 0.2 | 2.4 | GO:0097150 | neuronal stem cell population maintenance(GO:0097150) |
| 0.2 | 5.5 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.2 | 6.4 | GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process(GO:0030433) |
| 0.2 | 4.4 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.2 | 2.1 | GO:0021535 | cell migration in hindbrain(GO:0021535) |
| 0.2 | 2.0 | GO:0090005 | negative regulation of establishment of protein localization to plasma membrane(GO:0090005) |
| 0.1 | 10.3 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0006919) |
| 0.1 | 0.3 | GO:0061010 | external genitalia morphogenesis(GO:0035261) gall bladder development(GO:0061010) |
| 0.1 | 10.2 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.1 | 3.1 | GO:0032967 | positive regulation of collagen biosynthetic process(GO:0032967) |
| 0.1 | 1.3 | GO:0097688 | AMPA glutamate receptor clustering(GO:0097113) glutamate receptor clustering(GO:0097688) |
| 0.1 | 1.1 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.1 | 7.2 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.1 | 3.4 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.1 | 3.2 | GO:0035162 | embryonic hemopoiesis(GO:0035162) |
| 0.1 | 0.4 | GO:0014707 | branchiomeric skeletal muscle development(GO:0014707) |
| 0.1 | 3.2 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.1 | 1.8 | GO:0032463 | negative regulation of protein homooligomerization(GO:0032463) |
| 0.1 | 3.7 | GO:0042573 | retinoic acid metabolic process(GO:0042573) |
| 0.1 | 1.0 | GO:0043984 | histone H4-K16 acetylation(GO:0043984) |
| 0.1 | 2.8 | GO:0032098 | regulation of appetite(GO:0032098) |
| 0.1 | 0.5 | GO:0033089 | positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 0.1 | 5.4 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 5.6 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.1 | 1.5 | GO:0036344 | platelet morphogenesis(GO:0036344) |
| 0.1 | 0.6 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.1 | 2.8 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.1 | 0.6 | GO:0098704 | fructose transport(GO:0015755) fructose import(GO:0032445) carbohydrate import into cell(GO:0097319) carbohydrate import across plasma membrane(GO:0098704) fructose import across plasma membrane(GO:1990539) |
| 0.1 | 0.3 | GO:1901899 | positive regulation of relaxation of muscle(GO:1901079) positive regulation of relaxation of cardiac muscle(GO:1901899) regulation of calcium ion import into sarcoplasmic reticulum(GO:1902080) negative regulation of calcium ion import into sarcoplasmic reticulum(GO:1902081) |
| 0.1 | 0.8 | GO:0015986 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.1 | 3.4 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.1 | 7.8 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.1 | 1.3 | GO:0045898 | regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) |
| 0.1 | 1.3 | GO:0040032 | parathyroid hormone secretion(GO:0035898) post-embryonic body morphogenesis(GO:0040032) regulation of parathyroid hormone secretion(GO:2000828) |
| 0.1 | 3.8 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.1 | 1.8 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.1 | 4.3 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.1 | 0.8 | GO:0016233 | telomere capping(GO:0016233) |
| 0.1 | 3.2 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.1 | 0.8 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.1 | 1.7 | GO:0006688 | glycosphingolipid biosynthetic process(GO:0006688) |
| 0.1 | 0.5 | GO:0033631 | cell-cell adhesion mediated by integrin(GO:0033631) |
| 0.1 | 2.4 | GO:0007032 | endosome organization(GO:0007032) |
| 0.1 | 2.7 | GO:0045103 | intermediate filament-based process(GO:0045103) |
| 0.1 | 1.4 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 1.5 | GO:0006851 | mitochondrial calcium ion transport(GO:0006851) |
| 0.1 | 4.5 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.1 | 1.2 | GO:0006308 | DNA catabolic process(GO:0006308) |
| 0.1 | 2.0 | GO:0090502 | RNA phosphodiester bond hydrolysis, endonucleolytic(GO:0090502) |
| 0.0 | 0.5 | GO:0046599 | regulation of centriole replication(GO:0046599) |
| 0.0 | 1.9 | GO:0006735 | NADH regeneration(GO:0006735) canonical glycolysis(GO:0061621) glucose catabolic process to pyruvate(GO:0061718) |
| 0.0 | 3.0 | GO:0030148 | sphingolipid biosynthetic process(GO:0030148) |
| 0.0 | 0.2 | GO:0070072 | proton-transporting V-type ATPase complex assembly(GO:0070070) vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.0 | 0.1 | GO:0050720 | interleukin-1 biosynthetic process(GO:0042222) interleukin-1 beta biosynthetic process(GO:0050720) |
| 0.0 | 0.7 | GO:0019370 | leukotriene biosynthetic process(GO:0019370) |
| 0.0 | 0.7 | GO:0045652 | regulation of megakaryocyte differentiation(GO:0045652) |
| 0.0 | 0.5 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |
| 0.0 | 0.1 | GO:0035247 | peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) peptidyl-arginine omega-N-methylation(GO:0035247) |
| 0.0 | 0.2 | GO:1904424 | regulation of GTP binding(GO:1904424) |
| 0.0 | 1.2 | GO:0030042 | actin filament depolymerization(GO:0030042) |
| 0.0 | 1.5 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) |
| 0.0 | 1.4 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 0.2 | GO:0070120 | ciliary neurotrophic factor-mediated signaling pathway(GO:0070120) |
| 0.0 | 0.3 | GO:1900027 | regulation of ruffle assembly(GO:1900027) |
| 0.0 | 0.0 | GO:0033499 | galactose catabolic process via UDP-galactose(GO:0033499) |
| 0.0 | 0.1 | GO:0098914 | membrane repolarization during atrial cardiac muscle cell action potential(GO:0098914) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 11.8 | 35.3 | GO:1990298 | mitotic checkpoint complex(GO:0033597) bub1-bub3 complex(GO:1990298) |
| 10.9 | 43.8 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 7.7 | 23.2 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 6.2 | 24.9 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 6.0 | 18.0 | GO:0034455 | t-UTP complex(GO:0034455) |
| 5.8 | 46.4 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 5.7 | 40.2 | GO:0005683 | U7 snRNP(GO:0005683) |
| 5.3 | 21.0 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 5.2 | 47.1 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 5.0 | 35.0 | GO:0031415 | NatA complex(GO:0031415) |
| 5.0 | 15.0 | GO:0034272 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 4.8 | 133.7 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 4.7 | 32.8 | GO:0032021 | NELF complex(GO:0032021) |
| 4.6 | 13.7 | GO:0036284 | tubulobulbar complex(GO:0036284) |
| 4.4 | 43.8 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 4.1 | 16.3 | GO:0005715 | late recombination nodule(GO:0005715) |
| 4.0 | 40.3 | GO:0034709 | methylosome(GO:0034709) |
| 3.7 | 14.7 | GO:0071942 | XPC complex(GO:0071942) |
| 3.7 | 18.3 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 3.6 | 14.3 | GO:0000811 | GINS complex(GO:0000811) |
| 3.6 | 32.1 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 3.5 | 17.7 | GO:0033503 | HULC complex(GO:0033503) |
| 3.5 | 10.6 | GO:1990723 | cytoplasmic periphery of the nuclear pore complex(GO:1990723) |
| 3.4 | 10.3 | GO:0009328 | phenylalanine-tRNA ligase complex(GO:0009328) |
| 3.4 | 23.5 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 3.4 | 6.7 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 3.3 | 23.0 | GO:0072589 | box H/ACA snoRNP complex(GO:0031429) box H/ACA RNP complex(GO:0072588) box H/ACA scaRNP complex(GO:0072589) box H/ACA telomerase RNP complex(GO:0090661) |
| 3.3 | 13.1 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 3.1 | 9.4 | GO:0034991 | nuclear meiotic cohesin complex(GO:0034991) |
| 3.1 | 15.4 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 3.0 | 27.0 | GO:0044754 | autolysosome(GO:0044754) |
| 3.0 | 9.0 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 3.0 | 35.4 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 2.9 | 49.9 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 2.8 | 31.3 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 2.8 | 14.2 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 2.7 | 16.2 | GO:0002178 | palmitoyltransferase complex(GO:0002178) |
| 2.4 | 14.6 | GO:0042721 | mitochondrial inner membrane protein insertion complex(GO:0042721) |
| 2.4 | 28.8 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 2.4 | 21.2 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 2.2 | 17.8 | GO:0000796 | condensin complex(GO:0000796) |
| 2.2 | 13.1 | GO:0031371 | ubiquitin conjugating enzyme complex(GO:0031371) |
| 2.2 | 35.0 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 2.1 | 18.9 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 2.1 | 6.2 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 2.1 | 28.9 | GO:0005686 | U2 snRNP(GO:0005686) |
| 2.1 | 16.5 | GO:0005688 | U6 snRNP(GO:0005688) |
| 2.0 | 12.1 | GO:0071256 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 2.0 | 39.3 | GO:0016580 | Sin3 complex(GO:0016580) |
| 1.9 | 17.5 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 1.9 | 153.1 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 1.8 | 20.1 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 1.7 | 5.1 | GO:0033185 | dolichol-phosphate-mannose synthase complex(GO:0033185) |
| 1.7 | 6.8 | GO:0071986 | Ragulator complex(GO:0071986) |
| 1.7 | 18.6 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 1.7 | 23.2 | GO:0019774 | proteasome core complex, beta-subunit complex(GO:0019774) |
| 1.6 | 17.8 | GO:0070449 | elongin complex(GO:0070449) |
| 1.6 | 8.0 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 1.6 | 17.3 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 1.6 | 9.4 | GO:0097422 | tubular endosome(GO:0097422) |
| 1.6 | 14.1 | GO:0042382 | paraspeckles(GO:0042382) |
| 1.6 | 4.7 | GO:0070939 | Dsl1p complex(GO:0070939) |
| 1.5 | 3.1 | GO:0031414 | N-terminal protein acetyltransferase complex(GO:0031414) |
| 1.5 | 18.5 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 1.5 | 7.5 | GO:0031523 | Myb complex(GO:0031523) |
| 1.5 | 17.8 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 1.4 | 5.8 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 1.4 | 11.4 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 1.4 | 18.4 | GO:0045275 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 1.4 | 25.2 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 1.3 | 52.0 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 1.3 | 140.2 | GO:0015934 | large ribosomal subunit(GO:0015934) |
| 1.3 | 9.0 | GO:0098560 | cytoplasmic side of late endosome membrane(GO:0098560) |
| 1.3 | 8.9 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 1.3 | 15.2 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 1.3 | 18.9 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 1.2 | 7.4 | GO:0061617 | MICOS complex(GO:0061617) |
| 1.2 | 3.7 | GO:0043259 | laminin-1 complex(GO:0005606) laminin-10 complex(GO:0043259) |
| 1.2 | 9.7 | GO:0097452 | GAIT complex(GO:0097452) |
| 1.2 | 7.2 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 1.2 | 25.0 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 1.2 | 10.7 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 1.2 | 3.5 | GO:0005674 | transcription factor TFIIF complex(GO:0005674) |
| 1.1 | 37.3 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 1.1 | 15.5 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 1.1 | 29.7 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 1.1 | 5.4 | GO:0016589 | NURF complex(GO:0016589) |
| 1.0 | 4.2 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 1.0 | 12.5 | GO:0030008 | TRAPP complex(GO:0030008) |
| 1.0 | 4.1 | GO:1990876 | cytoplasmic side of nuclear pore(GO:1990876) |
| 1.0 | 26.1 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 1.0 | 2.0 | GO:0097135 | cyclin E2-CDK2 complex(GO:0097135) |
| 1.0 | 2.9 | GO:0035525 | NF-kappaB p50/p65 complex(GO:0035525) |
| 1.0 | 4.9 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.9 | 8.5 | GO:0044194 | cytolytic granule(GO:0044194) |
| 0.9 | 3.6 | GO:1990356 | sumoylated E2 ligase complex(GO:1990356) |
| 0.9 | 12.2 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.9 | 4.3 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.8 | 25.2 | GO:0022624 | proteasome regulatory particle(GO:0005838) proteasome accessory complex(GO:0022624) |
| 0.8 | 4.0 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.8 | 1.6 | GO:0071817 | MMXD complex(GO:0071817) |
| 0.8 | 3.2 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 0.8 | 11.0 | GO:0090543 | Flemming body(GO:0090543) |
| 0.8 | 44.0 | GO:0015030 | Cajal body(GO:0015030) |
| 0.7 | 9.7 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.7 | 3.0 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.7 | 8.1 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.7 | 62.3 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.7 | 13.1 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.7 | 11.3 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.7 | 7.1 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.7 | 86.6 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.7 | 9.0 | GO:0002199 | zona pellucida receptor complex(GO:0002199) |
| 0.7 | 2.1 | GO:0044393 | microspike(GO:0044393) |
| 0.7 | 4.1 | GO:0009368 | endopeptidase Clp complex(GO:0009368) |
| 0.7 | 7.5 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.7 | 15.1 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.7 | 7.8 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.6 | 7.7 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.6 | 20.8 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.6 | 29.6 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.6 | 5.5 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.6 | 13.5 | GO:0030137 | COPI-coated vesicle(GO:0030137) |
| 0.6 | 3.7 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.6 | 2.4 | GO:0031264 | death-inducing signaling complex(GO:0031264) |
| 0.6 | 12.4 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.6 | 6.4 | GO:0005845 | mRNA cap binding complex(GO:0005845) |
| 0.6 | 2.9 | GO:0005761 | organellar ribosome(GO:0000313) mitochondrial ribosome(GO:0005761) |
| 0.6 | 60.0 | GO:0000784 | nuclear chromosome, telomeric region(GO:0000784) |
| 0.5 | 3.8 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.5 | 10.4 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.5 | 4.8 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.5 | 3.1 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.5 | 17.1 | GO:0005844 | polysome(GO:0005844) |
| 0.5 | 1.0 | GO:0072487 | MSL complex(GO:0072487) |
| 0.5 | 1.5 | GO:0030312 | external encapsulating structure(GO:0030312) |
| 0.5 | 4.3 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.5 | 26.0 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.5 | 33.6 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.5 | 9.1 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.4 | 6.7 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.4 | 1.8 | GO:0071920 | cleavage body(GO:0071920) |
| 0.4 | 3.1 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.4 | 9.6 | GO:0031304 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.4 | 10.4 | GO:0031904 | endosome lumen(GO:0031904) |
| 0.4 | 5.1 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.4 | 1.7 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.4 | 2.4 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.4 | 27.0 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.4 | 4.4 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.4 | 13.1 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.4 | 4.3 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.4 | 17.4 | GO:0045095 | keratin filament(GO:0045095) |
| 0.4 | 1.2 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.4 | 5.7 | GO:0000178 | exosome (RNase complex)(GO:0000178) |
| 0.4 | 3.0 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.4 | 5.2 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.4 | 51.6 | GO:0030496 | midbody(GO:0030496) |
| 0.4 | 5.6 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.4 | 7.4 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.3 | 22.0 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.3 | 15.9 | GO:0031430 | M band(GO:0031430) |
| 0.3 | 2.0 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.3 | 8.0 | GO:0030057 | desmosome(GO:0030057) |
| 0.3 | 7.6 | GO:0000776 | kinetochore(GO:0000776) |
| 0.3 | 36.7 | GO:0000922 | spindle pole(GO:0000922) |
| 0.3 | 7.6 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.3 | 1.3 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.3 | 22.0 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.3 | 16.3 | GO:0043034 | costamere(GO:0043034) |
| 0.3 | 113.9 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.3 | 8.3 | GO:0032420 | stereocilium(GO:0032420) |
| 0.3 | 2.9 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.3 | 41.6 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.3 | 3.8 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.3 | 27.4 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.3 | 5.3 | GO:0035580 | specific granule lumen(GO:0035580) |
| 0.3 | 27.4 | GO:0016605 | PML body(GO:0016605) |
| 0.3 | 2.3 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.3 | 2.6 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.3 | 7.6 | GO:0000502 | proteasome complex(GO:0000502) |
| 0.3 | 2.5 | GO:0097443 | sorting endosome(GO:0097443) |
| 0.3 | 8.6 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.3 | 7.8 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.3 | 38.4 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.3 | 36.1 | GO:0005903 | brush border(GO:0005903) |
| 0.2 | 1.0 | GO:0020018 | ciliary pocket(GO:0020016) ciliary pocket membrane(GO:0020018) |
| 0.2 | 2.6 | GO:0032039 | integrator complex(GO:0032039) |
| 0.2 | 9.2 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.2 | 8.7 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.2 | 24.0 | GO:0101002 | ficolin-1-rich granule(GO:0101002) ficolin-1-rich granule lumen(GO:1904813) |
| 0.2 | 15.1 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.2 | 4.0 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.2 | 2.0 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.2 | 0.9 | GO:0043541 | UDP-N-acetylglucosamine transferase complex(GO:0043541) |
| 0.2 | 2.4 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.2 | 5.6 | GO:0035267 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.2 | 9.2 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.2 | 23.3 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.2 | 2.1 | GO:0044815 | nucleosome(GO:0000786) DNA packaging complex(GO:0044815) |
| 0.2 | 0.4 | GO:0033565 | ESCRT-0 complex(GO:0033565) |
| 0.2 | 0.7 | GO:0034274 | Atg12-Atg5-Atg16 complex(GO:0034274) |
| 0.2 | 1.2 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.1 | 1.4 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.1 | 0.8 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 0.1 | 2.0 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.1 | 1.1 | GO:0032806 | holo TFIIH complex(GO:0005675) carboxy-terminal domain protein kinase complex(GO:0032806) |
| 0.1 | 10.5 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
| 0.1 | 4.6 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 36.7 | GO:0098857 | membrane raft(GO:0045121) membrane microdomain(GO:0098857) |
| 0.1 | 4.7 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.1 | 2.6 | GO:0005605 | basal lamina(GO:0005605) |
| 0.1 | 2.4 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.1 | 0.3 | GO:0032116 | SMC loading complex(GO:0032116) |
| 0.1 | 6.0 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 1.9 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 3.0 | GO:0045178 | basal part of cell(GO:0045178) |
| 0.1 | 3.8 | GO:0032432 | actin filament bundle(GO:0032432) |
| 0.1 | 4.5 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 1.5 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.1 | 8.3 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 1.9 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 1.5 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.1 | 2.1 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 2.6 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.0 | 0.2 | GO:0097059 | CNTFR-CLCF1 complex(GO:0097059) |
| 0.0 | 0.2 | GO:0043291 | RAVE complex(GO:0043291) |
| 0.0 | 2.3 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.0 | 0.7 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.2 | GO:0000322 | storage vacuole(GO:0000322) |
| 0.0 | 0.8 | GO:0042627 | chylomicron(GO:0042627) |
| 0.0 | 0.7 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 21.7 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 1.9 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.0 | 5.0 | GO:0016324 | apical plasma membrane(GO:0016324) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 28.0 | 112.0 | GO:0044378 | non-sequence-specific DNA binding, bending(GO:0044378) |
| 13.3 | 39.9 | GO:0043739 | G/U mismatch-specific uracil-DNA glycosylase activity(GO:0043739) |
| 10.4 | 31.2 | GO:0035529 | NADH pyrophosphatase activity(GO:0035529) |
| 10.4 | 10.4 | GO:0051717 | inositol-1,3,4,5-tetrakisphosphate 3-phosphatase activity(GO:0051717) |
| 8.8 | 26.5 | GO:0034416 | bisphosphoglycerate phosphatase activity(GO:0034416) |
| 8.4 | 42.0 | GO:0050220 | prostaglandin-E synthase activity(GO:0050220) |
| 8.3 | 33.4 | GO:0070320 | inward rectifier potassium channel inhibitor activity(GO:0070320) |
| 8.0 | 32.1 | GO:0004775 | succinate-CoA ligase (ADP-forming) activity(GO:0004775) |
| 7.3 | 43.8 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 7.0 | 48.9 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 7.0 | 20.9 | GO:0004618 | phosphoglycerate kinase activity(GO:0004618) |
| 6.6 | 19.7 | GO:0070538 | oleic acid binding(GO:0070538) |
| 5.9 | 17.7 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 5.8 | 57.8 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 5.4 | 21.6 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 5.4 | 16.2 | GO:0004132 | dCMP deaminase activity(GO:0004132) |
| 5.2 | 47.1 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 5.1 | 15.4 | GO:0030337 | DNA polymerase processivity factor activity(GO:0030337) |
| 5.0 | 14.9 | GO:0000252 | C-3 sterol dehydrogenase (C-4 sterol decarboxylase) activity(GO:0000252) sterol-4-alpha-carboxylate 3-dehydrogenase (decarboxylating) activity(GO:0047012) |
| 4.9 | 29.2 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 4.9 | 29.2 | GO:0044020 | histone methyltransferase activity (H4-R3 specific)(GO:0044020) |
| 4.8 | 14.5 | GO:0004450 | isocitrate dehydrogenase (NADP+) activity(GO:0004450) |
| 4.8 | 19.3 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 4.5 | 18.2 | GO:0016532 | superoxide dismutase copper chaperone activity(GO:0016532) |
| 4.5 | 27.0 | GO:0017108 | 5'-flap endonuclease activity(GO:0017108) |
| 4.5 | 22.3 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 4.3 | 21.6 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 4.3 | 21.4 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 4.0 | 23.9 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 3.9 | 30.9 | GO:0050145 | nucleoside phosphate kinase activity(GO:0050145) |
| 3.8 | 49.9 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 3.7 | 33.0 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 3.6 | 24.9 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 3.5 | 21.2 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 3.5 | 10.4 | GO:0005046 | KDEL sequence binding(GO:0005046) |
| 3.3 | 26.6 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 3.3 | 23.0 | GO:0034513 | box H/ACA snoRNA binding(GO:0034513) |
| 3.2 | 9.7 | GO:0030156 | benzodiazepine receptor binding(GO:0030156) |
| 3.2 | 9.7 | GO:0035605 | peptidyl-cysteine S-nitrosylase activity(GO:0035605) |
| 3.2 | 38.3 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) N6-methyladenosine-containing RNA binding(GO:1990247) |
| 3.2 | 9.5 | GO:0070259 | tyrosyl-DNA phosphodiesterase activity(GO:0070259) |
| 3.1 | 25.2 | GO:0052833 | inositol monophosphate 1-phosphatase activity(GO:0008934) inositol monophosphate 3-phosphatase activity(GO:0052832) inositol monophosphate 4-phosphatase activity(GO:0052833) inositol monophosphate phosphatase activity(GO:0052834) |
| 3.1 | 9.3 | GO:0010698 | acetyltransferase activator activity(GO:0010698) |
| 2.9 | 14.7 | GO:0032795 | heterotrimeric G-protein binding(GO:0032795) |
| 2.9 | 8.6 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 2.7 | 24.4 | GO:0089720 | caspase binding(GO:0089720) |
| 2.7 | 8.1 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 2.6 | 7.7 | GO:0047374 | methylumbelliferyl-acetate deacetylase activity(GO:0047374) |
| 2.6 | 2.6 | GO:0008160 | protein tyrosine phosphatase activator activity(GO:0008160) |
| 2.6 | 40.9 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 2.5 | 35.3 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 2.4 | 9.7 | GO:0033677 | DNA/RNA helicase activity(GO:0033677) |
| 2.4 | 11.9 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 2.3 | 18.5 | GO:0034604 | pyruvate dehydrogenase activity(GO:0004738) pyruvate dehydrogenase [NAD(P)+] activity(GO:0034603) pyruvate dehydrogenase (NAD+) activity(GO:0034604) |
| 2.1 | 14.7 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 2.1 | 18.8 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 2.0 | 12.0 | GO:0036033 | mediator complex binding(GO:0036033) |
| 2.0 | 37.5 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 1.9 | 7.8 | GO:0002060 | purine nucleobase binding(GO:0002060) |
| 1.9 | 13.3 | GO:0005347 | adenine nucleotide transmembrane transporter activity(GO:0000295) purine ribonucleotide transmembrane transporter activity(GO:0005346) ATP transmembrane transporter activity(GO:0005347) purine nucleotide transmembrane transporter activity(GO:0015216) ADP transmembrane transporter activity(GO:0015217) |
| 1.9 | 7.6 | GO:0003985 | acetyl-CoA C-acetyltransferase activity(GO:0003985) |
| 1.9 | 5.6 | GO:0016531 | copper chaperone activity(GO:0016531) |
| 1.9 | 14.9 | GO:0016721 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 1.9 | 9.3 | GO:0004337 | dimethylallyltranstransferase activity(GO:0004161) geranyltranstransferase activity(GO:0004337) |
| 1.9 | 52.0 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 1.8 | 5.5 | GO:0071566 | UFM1 activating enzyme activity(GO:0071566) |
| 1.8 | 7.3 | GO:0043515 | kinetochore binding(GO:0043515) |
| 1.8 | 18.2 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 1.8 | 52.3 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 1.8 | 16.2 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 1.7 | 8.7 | GO:0004771 | sterol esterase activity(GO:0004771) |
| 1.7 | 5.1 | GO:0004582 | dolichyl-phosphate beta-D-mannosyltransferase activity(GO:0004582) |
| 1.7 | 8.5 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 1.6 | 11.3 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 1.6 | 14.2 | GO:0008865 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 1.6 | 9.5 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 1.6 | 1.6 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 1.6 | 12.4 | GO:0046972 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
| 1.5 | 6.1 | GO:0043262 | adenosine-diphosphatase activity(GO:0043262) |
| 1.5 | 7.6 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 1.5 | 7.5 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 1.4 | 5.8 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 1.4 | 4.3 | GO:0004608 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 1.4 | 28.6 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 1.4 | 6.9 | GO:0004905 | type I interferon receptor activity(GO:0004905) |
| 1.4 | 23.2 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 1.4 | 16.3 | GO:0032137 | guanine/thymine mispair binding(GO:0032137) |
| 1.4 | 10.8 | GO:0015288 | porin activity(GO:0015288) |
| 1.3 | 13.3 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 1.3 | 18.4 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 1.3 | 11.8 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 1.3 | 5.2 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 1.3 | 10.4 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 1.3 | 7.7 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 1.3 | 6.4 | GO:0004376 | glycolipid mannosyltransferase activity(GO:0004376) |
| 1.3 | 3.8 | GO:0052794 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 1.3 | 16.5 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 1.3 | 47.6 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 1.2 | 16.7 | GO:0031386 | protein tag(GO:0031386) |
| 1.2 | 7.1 | GO:0015166 | polyol transmembrane transporter activity(GO:0015166) |
| 1.2 | 5.9 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 1.2 | 33.5 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 1.1 | 20.6 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 1.1 | 24.7 | GO:0008494 | translation activator activity(GO:0008494) |
| 1.1 | 4.4 | GO:0045134 | uridine-diphosphatase activity(GO:0045134) |
| 1.1 | 13.3 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 1.1 | 32.0 | GO:0019843 | rRNA binding(GO:0019843) |
| 1.1 | 4.4 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 1.1 | 15.3 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 1.1 | 45.4 | GO:0031369 | translation initiation factor binding(GO:0031369) |
| 1.1 | 6.3 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 1.1 | 20.0 | GO:0000339 | RNA cap binding(GO:0000339) |
| 1.0 | 9.4 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 1.0 | 6.2 | GO:0043008 | ATP-dependent protein binding(GO:0043008) |
| 1.0 | 8.1 | GO:0015235 | cobalamin transporter activity(GO:0015235) |
| 1.0 | 7.7 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 1.0 | 14.3 | GO:0015215 | nucleotide transmembrane transporter activity(GO:0015215) |
| 0.9 | 5.7 | GO:0004310 | farnesyl-diphosphate farnesyltransferase activity(GO:0004310) squalene synthase activity(GO:0051996) |
| 0.9 | 3.8 | GO:0008469 | histone-arginine N-methyltransferase activity(GO:0008469) protein-arginine omega-N asymmetric methyltransferase activity(GO:0035242) |
| 0.9 | 19.7 | GO:0043495 | protein anchor(GO:0043495) |
| 0.9 | 193.9 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.9 | 10.2 | GO:0004673 | protein histidine kinase activity(GO:0004673) |
| 0.9 | 2.8 | GO:0034736 | sterol O-acyltransferase activity(GO:0004772) cholesterol O-acyltransferase activity(GO:0034736) |
| 0.9 | 24.8 | GO:0070717 | poly-purine tract binding(GO:0070717) |
| 0.9 | 3.6 | GO:0061656 | SUMO conjugating enzyme activity(GO:0061656) |
| 0.9 | 14.2 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.9 | 23.0 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.9 | 7.0 | GO:0003910 | DNA ligase (ATP) activity(GO:0003910) |
| 0.9 | 11.1 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.8 | 13.2 | GO:0031404 | chloride ion binding(GO:0031404) |
| 0.8 | 17.9 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.8 | 3.2 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.8 | 10.1 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.8 | 41.2 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.8 | 21.4 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.8 | 27.1 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.8 | 6.0 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.7 | 5.9 | GO:0036435 | K48-linked polyubiquitin binding(GO:0036435) |
| 0.7 | 4.4 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.7 | 10.1 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.7 | 12.9 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.7 | 5.6 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.7 | 4.9 | GO:0030267 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.7 | 4.9 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.7 | 20.7 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.7 | 36.8 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.7 | 4.8 | GO:0016778 | diphosphotransferase activity(GO:0016778) |
| 0.7 | 32.5 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.7 | 20.2 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.7 | 15.5 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.7 | 4.0 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.7 | 30.2 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.7 | 16.6 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.7 | 8.5 | GO:0070567 | cytidylyltransferase activity(GO:0070567) |
| 0.6 | 7.1 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.6 | 71.6 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.6 | 3.2 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.6 | 20.7 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.6 | 2.9 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.6 | 3.5 | GO:0008518 | reduced folate carrier activity(GO:0008518) methotrexate transporter activity(GO:0015350) |
| 0.6 | 8.7 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.6 | 27.5 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.6 | 4.5 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.6 | 15.1 | GO:0043021 | ribonucleoprotein complex binding(GO:0043021) |
| 0.5 | 1.6 | GO:0050509 | N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase activity(GO:0050509) |
| 0.5 | 6.0 | GO:0005381 | iron ion transmembrane transporter activity(GO:0005381) |
| 0.5 | 8.2 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.5 | 36.9 | GO:0008235 | metalloexopeptidase activity(GO:0008235) |
| 0.5 | 4.3 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 0.5 | 2.1 | GO:0008177 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) |
| 0.5 | 32.9 | GO:0050681 | androgen receptor binding(GO:0050681) |
| 0.5 | 6.8 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.5 | 10.0 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.5 | 11.4 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.5 | 15.7 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.5 | 9.6 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.5 | 2.5 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.5 | 5.4 | GO:0035591 | signaling adaptor activity(GO:0035591) |
| 0.5 | 3.9 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.5 | 5.4 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.5 | 2.4 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.5 | 3.3 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.5 | 5.1 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.5 | 24.6 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.5 | 2.3 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.5 | 21.2 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.4 | 7.0 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.4 | 9.2 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.4 | 3.5 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.4 | 4.3 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.4 | 5.8 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.4 | 6.7 | GO:0016655 | oxidoreductase activity, acting on NAD(P)H, quinone or similar compound as acceptor(GO:0016655) |
| 0.4 | 6.1 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.4 | 3.7 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.4 | 13.0 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.4 | 2.0 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.4 | 2.4 | GO:0070513 | death domain binding(GO:0070513) |
| 0.4 | 3.2 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.4 | 27.9 | GO:0070888 | E-box binding(GO:0070888) |
| 0.4 | 7.5 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.4 | 6.3 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.4 | 2.7 | GO:0004457 | lactate dehydrogenase activity(GO:0004457) L-lactate dehydrogenase activity(GO:0004459) |
| 0.4 | 1.9 | GO:0050436 | microfibril binding(GO:0050436) |
| 0.4 | 5.8 | GO:0016004 | phospholipase activator activity(GO:0016004) |
| 0.4 | 10.7 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.4 | 1.5 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.4 | 6.5 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.3 | 2.8 | GO:0102336 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.3 | 18.7 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.3 | 4.4 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.3 | 13.2 | GO:0043236 | laminin binding(GO:0043236) |
| 0.3 | 10.7 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.3 | 8.6 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.3 | 0.7 | GO:0016972 | flavin-linked sulfhydryl oxidase activity(GO:0016971) thiol oxidase activity(GO:0016972) |
| 0.3 | 7.9 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.3 | 30.8 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.3 | 19.5 | GO:0009055 | electron carrier activity(GO:0009055) |
| 0.3 | 6.1 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.3 | 3.9 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.3 | 4.4 | GO:0009881 | photoreceptor activity(GO:0009881) |
| 0.3 | 1.2 | GO:0005148 | prolactin receptor binding(GO:0005148) |
| 0.3 | 4.3 | GO:0016421 | CoA carboxylase activity(GO:0016421) |
| 0.3 | 10.9 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.3 | 12.1 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.3 | 1.4 | GO:0004140 | dephospho-CoA kinase activity(GO:0004140) |
| 0.3 | 4.9 | GO:0016876 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.3 | 6.5 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.3 | 2.3 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.3 | 25.6 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.3 | 2.3 | GO:0030346 | protein phosphatase 2B binding(GO:0030346) |
| 0.3 | 6.9 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.3 | 3.3 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.2 | 30.0 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.2 | 5.5 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.2 | 11.0 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.2 | 0.7 | GO:0015234 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) |
| 0.2 | 1.1 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 0.2 | 2.2 | GO:0019211 | phosphatase activator activity(GO:0019211) |
| 0.2 | 3.8 | GO:0016814 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amidines(GO:0016814) |
| 0.2 | 33.0 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.2 | 4.5 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.2 | 1.7 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.2 | 6.5 | GO:0008135 | translation factor activity, RNA binding(GO:0008135) |
| 0.2 | 1.3 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.2 | 5.9 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.2 | 1.4 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.2 | 10.5 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.2 | 9.2 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.2 | 15.5 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.2 | 1.7 | GO:0003831 | beta-N-acetylglucosaminylglycopeptide beta-1,4-galactosyltransferase activity(GO:0003831) |
| 0.2 | 1.9 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.2 | 2.6 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.2 | 8.9 | GO:0004386 | helicase activity(GO:0004386) |
| 0.2 | 5.1 | GO:0042974 | retinoic acid receptor binding(GO:0042974) |
| 0.2 | 3.4 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.2 | 217.8 | GO:0003723 | RNA binding(GO:0003723) |
| 0.2 | 4.3 | GO:0046961 | ATPase activity, coupled to transmembrane movement of ions, rotational mechanism(GO:0044769) proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.2 | 2.3 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.2 | 1.8 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.2 | 5.5 | GO:0008200 | ion channel inhibitor activity(GO:0008200) channel inhibitor activity(GO:0016248) |
| 0.1 | 0.7 | GO:0019776 | Atg8 ligase activity(GO:0019776) |
| 0.1 | 1.4 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.1 | 3.5 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.1 | 4.5 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 22.6 | GO:0061659 | ubiquitin-like protein ligase activity(GO:0061659) |
| 0.1 | 0.5 | GO:0004906 | interferon-gamma receptor activity(GO:0004906) |
| 0.1 | 0.5 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.1 | 5.4 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.1 | 3.8 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.1 | 1.2 | GO:0017081 | chloride channel regulator activity(GO:0017081) |
| 0.1 | 7.5 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.1 | 0.9 | GO:0003857 | 3-hydroxyacyl-CoA dehydrogenase activity(GO:0003857) |
| 0.1 | 0.6 | GO:0005353 | fructose transmembrane transporter activity(GO:0005353) |
| 0.1 | 2.1 | GO:0005537 | mannose binding(GO:0005537) |
| 0.1 | 0.6 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 0.1 | 2.8 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 0.8 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.1 | 0.3 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.1 | 0.1 | GO:0004904 | interferon receptor activity(GO:0004904) |
| 0.1 | 0.8 | GO:0008428 | ribonuclease inhibitor activity(GO:0008428) |
| 0.1 | 1.9 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.1 | 1.4 | GO:0048531 | beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.1 | 1.7 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 2.1 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.1 | 3.7 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.0 | 2.4 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.5 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 5.7 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 1.5 | GO:0004536 | deoxyribonuclease activity(GO:0004536) |
| 0.0 | 11.0 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.0 | 0.9 | GO:0051721 | protein phosphatase 2A binding(GO:0051721) |
| 0.0 | 1.5 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.0 | 0.1 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 0.0 | 0.7 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.2 | GO:0004897 | ciliary neurotrophic factor receptor activity(GO:0004897) |
| 0.0 | 0.6 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.0 | 0.1 | GO:0032453 | histone demethylase activity (H3-K4 specific)(GO:0032453) |
| 0.0 | 0.7 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 150.2 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 1.1 | 44.1 | PID MYC PATHWAY | C-MYC pathway |
| 1.1 | 53.5 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 1.0 | 46.7 | PID AURORA A PATHWAY | Aurora A signaling |
| 1.0 | 10.7 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 1.0 | 43.3 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.8 | 57.8 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.8 | 2.3 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.8 | 25.8 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.7 | 20.1 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.7 | 14.2 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.7 | 56.7 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.6 | 7.1 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.6 | 70.6 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.6 | 105.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.6 | 18.3 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.6 | 11.2 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.5 | 8.8 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.5 | 15.9 | PID ATR PATHWAY | ATR signaling pathway |
| 0.5 | 35.9 | PID E2F PATHWAY | E2F transcription factor network |
| 0.5 | 45.9 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.5 | 7.0 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.5 | 7.4 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.4 | 34.2 | PID P73PATHWAY | p73 transcription factor network |
| 0.4 | 11.1 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.4 | 16.7 | PID FOXO PATHWAY | FoxO family signaling |
| 0.4 | 16.7 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.4 | 5.1 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.4 | 6.5 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.4 | 12.0 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.3 | 19.2 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.3 | 14.1 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.3 | 13.0 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.3 | 7.8 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.3 | 4.2 | PID ATM PATHWAY | ATM pathway |
| 0.2 | 2.8 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.2 | 4.9 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.2 | 2.5 | PID EPO PATHWAY | EPO signaling pathway |
| 0.2 | 4.1 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.2 | 10.9 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.2 | 3.7 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.2 | 10.5 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.2 | 1.3 | PID CXCR4 PATHWAY | CXCR4-mediated signaling events |
| 0.2 | 4.9 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.2 | 12.5 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.2 | 9.6 | PID TCR PATHWAY | TCR signaling in naïve CD4+ T cells |
| 0.2 | 8.1 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.2 | 11.4 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.2 | 7.6 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.1 | 3.4 | PID GMCSF PATHWAY | GMCSF-mediated signaling events |
| 0.1 | 4.7 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.1 | 2.7 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
| 0.1 | 5.4 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 3.9 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.1 | 1.3 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 3.3 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.1 | 4.5 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.1 | 2.4 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 2.0 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 2.6 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.1 | 2.2 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.1 | 1.5 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.1 | 1.2 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 3.0 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.1 | 3.2 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.1 | 1.9 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.1 | 0.7 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 1.6 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 1.7 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 0.6 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 0.7 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.8 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 1.1 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.7 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.5 | PID SHP2 PATHWAY | SHP2 signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.4 | 111.6 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 7.1 | 35.3 | REACTOME APC C CDC20 MEDIATED DEGRADATION OF MITOTIC PROTEINS | Genes involved in APC/C:Cdc20 mediated degradation of mitotic proteins |
| 4.4 | 65.5 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 4.0 | 52.4 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 2.7 | 40.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 2.6 | 39.4 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 2.2 | 33.6 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 2.2 | 17.9 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 2.1 | 210.1 | REACTOME APC C CDH1 MEDIATED DEGRADATION OF CDC20 AND OTHER APC C CDH1 TARGETED PROTEINS IN LATE MITOSIS EARLY G1 | Genes involved in APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 |
| 2.1 | 81.4 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 2.1 | 22.9 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 2.1 | 39.3 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 1.6 | 26.9 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 1.6 | 90.7 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 1.5 | 13.4 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 1.5 | 29.5 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 1.4 | 27.4 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 1.4 | 42.1 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 1.3 | 71.1 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 1.3 | 19.7 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |
| 1.3 | 34.3 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 1.2 | 22.4 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 1.2 | 92.6 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 1.2 | 18.5 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 1.1 | 18.0 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |
| 1.1 | 65.7 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 1.1 | 123.6 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 1.0 | 52.0 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 1.0 | 6.1 | REACTOME INFLUENZA VIRAL RNA TRANSCRIPTION AND REPLICATION | Genes involved in Influenza Viral RNA Transcription and Replication |
| 1.0 | 27.2 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 1.0 | 20.1 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 1.0 | 33.1 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 1.0 | 15.0 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 1.0 | 33.4 | REACTOME ENOS ACTIVATION AND REGULATION | Genes involved in eNOS activation and regulation |
| 1.0 | 71.2 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 1.0 | 6.7 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.9 | 79.7 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.9 | 15.0 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.9 | 29.3 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.9 | 13.6 | REACTOME REGULATED PROTEOLYSIS OF P75NTR | Genes involved in Regulated proteolysis of p75NTR |
| 0.9 | 2.7 | REACTOME SPRY REGULATION OF FGF SIGNALING | Genes involved in Spry regulation of FGF signaling |
| 0.9 | 10.7 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.9 | 14.8 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.9 | 10.4 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.8 | 19.3 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.8 | 32.5 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.8 | 13.7 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.8 | 8.5 | REACTOME TRANSCRIPTIONAL ACTIVITY OF SMAD2 SMAD3 SMAD4 HETEROTRIMER | Genes involved in Transcriptional activity of SMAD2/SMAD3:SMAD4 heterotrimer |
| 0.8 | 16.2 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.7 | 7.3 | REACTOME ACTIVATION OF ATR IN RESPONSE TO REPLICATION STRESS | Genes involved in Activation of ATR in response to replication stress |
| 0.7 | 24.3 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.7 | 2.8 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.7 | 23.3 | REACTOME S PHASE | Genes involved in S Phase |
| 0.7 | 7.3 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.7 | 10.4 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.6 | 9.0 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.6 | 5.7 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.6 | 17.1 | REACTOME TRANSLATION | Genes involved in Translation |
| 0.6 | 16.3 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.6 | 7.5 | REACTOME ARMS MEDIATED ACTIVATION | Genes involved in ARMS-mediated activation |
| 0.6 | 14.5 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.5 | 25.6 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.5 | 13.5 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.5 | 42.3 | REACTOME APOPTOTIC EXECUTION PHASE | Genes involved in Apoptotic execution phase |
| 0.5 | 27.9 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.5 | 23.0 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.5 | 15.6 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
| 0.5 | 47.9 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.4 | 9.0 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.4 | 12.3 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.4 | 9.2 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.4 | 10.8 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.4 | 18.2 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.4 | 24.7 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.4 | 28.4 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.4 | 32.6 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.4 | 8.0 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.4 | 12.4 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.3 | 8.0 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.3 | 7.2 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.3 | 10.5 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.3 | 46.6 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.3 | 13.3 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.3 | 11.5 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.3 | 5.7 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.3 | 5.1 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.3 | 7.6 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.2 | 3.9 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.2 | 6.2 | REACTOME INTEGRIN ALPHAIIB BETA3 SIGNALING | Genes involved in Integrin alphaIIb beta3 signaling |
| 0.2 | 3.4 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.2 | 3.2 | REACTOME NONSENSE MEDIATED DECAY ENHANCED BY THE EXON JUNCTION COMPLEX | Genes involved in Nonsense Mediated Decay Enhanced by the Exon Junction Complex |
| 0.2 | 3.4 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.2 | 5.7 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.2 | 3.8 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.2 | 4.4 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.2 | 7.7 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.2 | 4.0 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.2 | 3.6 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 3.9 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.1 | 2.3 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.1 | 5.8 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 1.5 | REACTOME OPSINS | Genes involved in Opsins |
| 0.1 | 0.9 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.1 | 18.2 | REACTOME INTEGRATION OF ENERGY METABOLISM | Genes involved in Integration of energy metabolism |
| 0.1 | 3.3 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.1 | 1.6 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.1 | 6.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 2.0 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.1 | 4.7 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.1 | 1.7 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.1 | 1.1 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.1 | 1.7 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 1.1 | REACTOME DEADENYLATION DEPENDENT MRNA DECAY | Genes involved in Deadenylation-dependent mRNA decay |
| 0.1 | 0.8 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 0.5 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.0 | 1.1 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.0 | 0.7 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.2 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.4 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |