Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
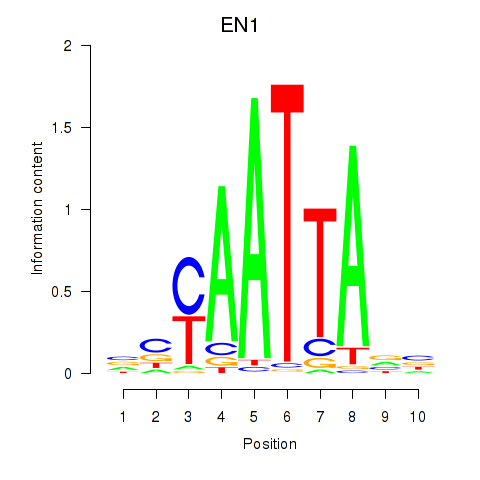
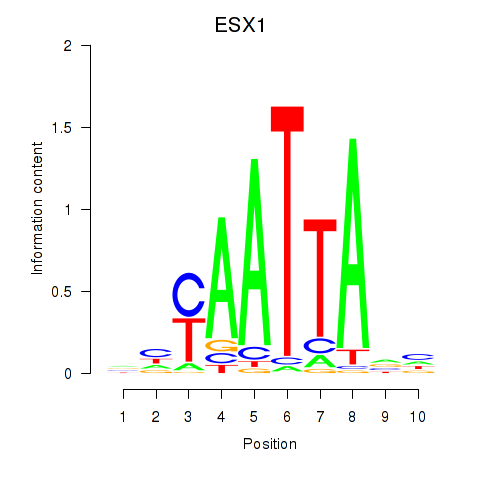
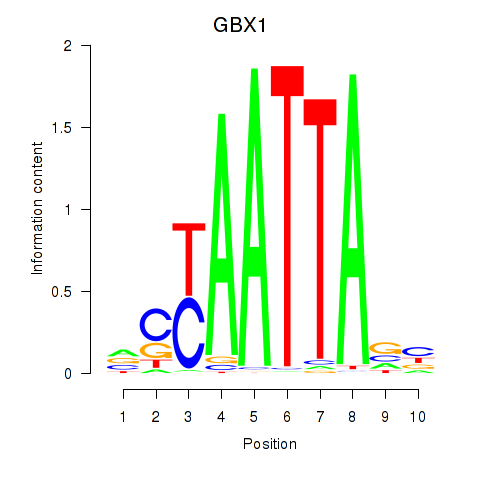
Results for EN1_ESX1_GBX1
Z-value: 1.11
Transcription factors associated with EN1_ESX1_GBX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
EN1
|
ENSG00000163064.7 | EN1 |
|
ESX1
|
ENSG00000123576.5 | ESX1 |
|
GBX1
|
ENSG00000164900.5 | GBX1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| GBX1 | hg38_v1_chr7_-_151167692_151167697 | 0.42 | 1.4e-10 | Click! |
| EN1 | hg38_v1_chr2_-_118847638_118847654 | 0.11 | 1.0e-01 | Click! |
Activity profile of EN1_ESX1_GBX1 motif
Sorted Z-values of EN1_ESX1_GBX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of EN1_ESX1_GBX1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.3 | 46.5 | GO:1901628 | positive regulation of postsynaptic membrane organization(GO:1901628) |
| 7.1 | 71.3 | GO:0036309 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 4.9 | 14.6 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 4.8 | 14.4 | GO:0044828 | negative regulation by host of viral genome replication(GO:0044828) |
| 4.5 | 13.6 | GO:0071139 | resolution of recombination intermediates(GO:0071139) resolution of mitotic recombination intermediates(GO:0071140) |
| 4.3 | 12.9 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 4.0 | 35.8 | GO:0061669 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 4.0 | 11.9 | GO:0002416 | IgG immunoglobulin transcytosis in epithelial cells mediated by FcRn immunoglobulin receptor(GO:0002416) |
| 3.7 | 11.1 | GO:0003218 | cardiac left ventricle formation(GO:0003218) |
| 3.6 | 25.0 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 3.3 | 9.8 | GO:1904908 | negative regulation of maintenance of sister chromatid cohesion(GO:0034092) negative regulation of maintenance of mitotic sister chromatid cohesion(GO:0034183) protein auto-ADP-ribosylation(GO:0070213) maintenance of mitotic sister chromatid cohesion, telomeric(GO:0099403) mitotic sister chromatid cohesion, telomeric(GO:0099404) regulation of telomeric DNA binding(GO:1904742) regulation of maintenance of mitotic sister chromatid cohesion, telomeric(GO:1904907) negative regulation of maintenance of mitotic sister chromatid cohesion, telomeric(GO:1904908) |
| 3.2 | 15.9 | GO:0060748 | tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
| 2.6 | 7.7 | GO:0019859 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 2.6 | 30.8 | GO:0051612 | negative regulation of neurotransmitter uptake(GO:0051581) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 2.5 | 9.8 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 2.4 | 7.1 | GO:1903989 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) negative regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002590) regulation of T cell antigen processing and presentation(GO:0002625) positive regulation of iron ion transport(GO:0034758) positive regulation of iron ion transmembrane transport(GO:0034761) regulation of iron ion import(GO:1900390) regulation of ferrous iron import into cell(GO:1903989) positive regulation of ferrous iron import into cell(GO:1903991) regulation of ferrous iron binding(GO:1904432) positive regulation of ferrous iron binding(GO:1904434) regulation of transferrin receptor binding(GO:1904435) positive regulation of transferrin receptor binding(GO:1904437) regulation of ferrous iron import across plasma membrane(GO:1904438) positive regulation of ferrous iron import across plasma membrane(GO:1904440) response to iron ion starvation(GO:1990641) |
| 2.2 | 13.4 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 2.1 | 6.4 | GO:1990451 | cellular stress response to acidic pH(GO:1990451) |
| 2.1 | 8.4 | GO:0051037 | regulation of transcription involved in meiotic cell cycle(GO:0051037) |
| 2.0 | 34.8 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 2.0 | 16.3 | GO:1900113 | negative regulation of histone H3-K9 trimethylation(GO:1900113) |
| 1.8 | 5.4 | GO:0098582 | innate vocalization behavior(GO:0098582) |
| 1.8 | 14.3 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 1.8 | 12.4 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 1.8 | 26.5 | GO:0097151 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 1.7 | 12.0 | GO:1902231 | positive regulation of keratinocyte apoptotic process(GO:1902174) positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902231) |
| 1.7 | 8.4 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 1.6 | 4.8 | GO:1990418 | response to insulin-like growth factor stimulus(GO:1990418) |
| 1.6 | 4.7 | GO:1900239 | phenotypic switching(GO:0036166) regulation of phenotypic switching(GO:1900239) |
| 1.6 | 32.6 | GO:0001829 | trophectodermal cell differentiation(GO:0001829) |
| 1.5 | 6.1 | GO:0051490 | negative regulation of filopodium assembly(GO:0051490) |
| 1.5 | 40.0 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 1.5 | 4.4 | GO:0070409 | carbamoyl phosphate metabolic process(GO:0070408) carbamoyl phosphate biosynthetic process(GO:0070409) cellular response to oleic acid(GO:0071400) response to ammonia(GO:1903717) cellular response to ammonia(GO:1903718) |
| 1.5 | 4.4 | GO:0072144 | positive regulation of vascular wound healing(GO:0035470) mesangial cell development(GO:0072143) glomerular mesangial cell development(GO:0072144) |
| 1.4 | 4.2 | GO:0019836 | hemolysis by symbiont of host erythrocytes(GO:0019836) hemolysis in other organism(GO:0044179) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 1.4 | 19.0 | GO:0048245 | eosinophil chemotaxis(GO:0048245) |
| 1.3 | 3.9 | GO:0015847 | putrescine transport(GO:0015847) |
| 1.3 | 2.5 | GO:2000296 | regulation of hydrogen peroxide catabolic process(GO:2000295) negative regulation of hydrogen peroxide catabolic process(GO:2000296) |
| 1.2 | 6.2 | GO:0045872 | positive regulation of rhodopsin gene expression(GO:0045872) |
| 1.2 | 8.5 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 1.2 | 6.9 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 1.1 | 5.6 | GO:0051902 | negative regulation of mitochondrial depolarization(GO:0051902) |
| 1.1 | 2.2 | GO:0070309 | lens fiber cell morphogenesis(GO:0070309) |
| 1.1 | 4.3 | GO:0031443 | fast-twitch skeletal muscle fiber contraction(GO:0031443) |
| 1.1 | 13.0 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 1.1 | 7.6 | GO:0048241 | epinephrine transport(GO:0048241) |
| 1.1 | 1.1 | GO:0086092 | regulation of the force of heart contraction by cardiac conduction(GO:0086092) |
| 1.0 | 2.1 | GO:2000170 | positive regulation of peptidyl-cysteine S-nitrosylation(GO:2000170) |
| 1.0 | 11.1 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 1.0 | 7.9 | GO:0035878 | nail development(GO:0035878) |
| 1.0 | 8.9 | GO:0016198 | axon choice point recognition(GO:0016198) |
| 1.0 | 1.0 | GO:1900109 | regulation of histone H3-K9 dimethylation(GO:1900109) |
| 1.0 | 83.4 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.9 | 6.6 | GO:0001550 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) negative regulation of chondrocyte proliferation(GO:1902731) |
| 0.9 | 2.8 | GO:0071934 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 0.9 | 1.9 | GO:2001180 | negative regulation of interleukin-10 secretion(GO:2001180) |
| 0.9 | 12.7 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) |
| 0.9 | 5.4 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.9 | 4.5 | GO:0093001 | glycolysis from storage polysaccharide through glucose-1-phosphate(GO:0093001) |
| 0.9 | 20.5 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.9 | 3.5 | GO:1990737 | response to manganese-induced endoplasmic reticulum stress(GO:1990737) |
| 0.9 | 4.4 | GO:0014809 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 0.9 | 5.2 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.9 | 17.2 | GO:0097186 | amelogenesis(GO:0097186) |
| 0.9 | 8.6 | GO:0006707 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.9 | 2.6 | GO:1903892 | negative regulation of ATF6-mediated unfolded protein response(GO:1903892) |
| 0.8 | 2.5 | GO:0002326 | B cell lineage commitment(GO:0002326) |
| 0.8 | 5.7 | GO:1902514 | regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1902514) positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.8 | 5.5 | GO:0007000 | nucleolus organization(GO:0007000) |
| 0.8 | 15.6 | GO:0009756 | carbohydrate mediated signaling(GO:0009756) |
| 0.7 | 3.7 | GO:0070778 | L-aspartate transport(GO:0070778) L-aspartate transmembrane transport(GO:0089712) |
| 0.7 | 2.1 | GO:0033024 | mast cell homeostasis(GO:0033023) mast cell apoptotic process(GO:0033024) regulation of mast cell apoptotic process(GO:0033025) mast cell proliferation(GO:0070662) |
| 0.7 | 3.4 | GO:0060729 | intestinal epithelial structure maintenance(GO:0060729) |
| 0.7 | 2.7 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.6 | 1.9 | GO:0021538 | epithalamus development(GO:0021538) habenula development(GO:0021986) |
| 0.6 | 1.9 | GO:0003219 | cardiac right ventricle formation(GO:0003219) |
| 0.6 | 1.9 | GO:0044771 | meiotic cell cycle phase transition(GO:0044771) regulation of meiotic cell cycle phase transition(GO:1901993) negative regulation of meiotic cell cycle phase transition(GO:1901994) |
| 0.6 | 8.1 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.6 | 2.5 | GO:0090290 | hormone-mediated apoptotic signaling pathway(GO:0008628) positive regulation of osteoclast proliferation(GO:0090290) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.6 | 1.9 | GO:0010968 | regulation of microtubule nucleation(GO:0010968) |
| 0.6 | 1.2 | GO:0060369 | positive regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060369) |
| 0.6 | 9.8 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.6 | 1.8 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.6 | 2.9 | GO:0010746 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) |
| 0.6 | 16.4 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.6 | 54.0 | GO:1902600 | hydrogen ion transmembrane transport(GO:1902600) |
| 0.6 | 4.0 | GO:0086036 | regulation of cardiac muscle cell membrane potential(GO:0086036) |
| 0.6 | 1.7 | GO:0061074 | neuroblast differentiation(GO:0014016) regulation of neural retina development(GO:0061074) regulation of retina development in camera-type eye(GO:1902866) |
| 0.6 | 3.3 | GO:0006001 | fructose catabolic process(GO:0006001) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) |
| 0.6 | 1.7 | GO:2000053 | regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000053) |
| 0.6 | 1.7 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) sarcomerogenesis(GO:0048769) |
| 0.6 | 2.2 | GO:0003331 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.5 | 0.5 | GO:1903625 | negative regulation of DNA catabolic process(GO:1903625) |
| 0.5 | 1.6 | GO:0015808 | L-alanine transport(GO:0015808) |
| 0.5 | 4.3 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.5 | 4.2 | GO:0050882 | voluntary musculoskeletal movement(GO:0050882) |
| 0.5 | 3.1 | GO:0050955 | thermoception(GO:0050955) |
| 0.5 | 7.3 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.5 | 1.5 | GO:0015860 | purine nucleoside transmembrane transport(GO:0015860) |
| 0.5 | 28.5 | GO:0006458 | 'de novo' protein folding(GO:0006458) |
| 0.5 | 14.9 | GO:0045494 | photoreceptor cell maintenance(GO:0045494) |
| 0.5 | 5.2 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.5 | 17.9 | GO:0035066 | positive regulation of histone acetylation(GO:0035066) |
| 0.5 | 1.9 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.5 | 1.4 | GO:0072366 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) regulation of cellular ketone metabolic process by positive regulation of transcription from RNA polymerase II promoter(GO:0072366) |
| 0.5 | 4.2 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.4 | 5.8 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 0.4 | 1.8 | GO:0051140 | positive regulation of activation of Janus kinase activity(GO:0010536) regulation of NK T cell proliferation(GO:0051140) positive regulation of NK T cell proliferation(GO:0051142) |
| 0.4 | 2.2 | GO:0051573 | negative regulation of histone H3-K9 methylation(GO:0051573) |
| 0.4 | 2.2 | GO:0030070 | insulin processing(GO:0030070) |
| 0.4 | 3.4 | GO:1901529 | positive regulation of anion channel activity(GO:1901529) |
| 0.4 | 8.5 | GO:0006228 | UTP biosynthetic process(GO:0006228) |
| 0.4 | 9.7 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.4 | 3.8 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.4 | 2.1 | GO:0031022 | nuclear migration along microfilament(GO:0031022) |
| 0.4 | 2.5 | GO:0071395 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 0.4 | 2.8 | GO:0072513 | positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.4 | 2.3 | GO:0006710 | androgen catabolic process(GO:0006710) |
| 0.4 | 7.3 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.4 | 4.2 | GO:0021942 | radial glia guided migration of Purkinje cell(GO:0021942) |
| 0.4 | 5.0 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.4 | 13.4 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.4 | 2.2 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.4 | 4.1 | GO:0051791 | medium-chain fatty acid metabolic process(GO:0051791) |
| 0.4 | 2.8 | GO:0097039 | protein linear polyubiquitination(GO:0097039) |
| 0.3 | 4.5 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.3 | 1.7 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.3 | 2.3 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.3 | 4.9 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.3 | 4.5 | GO:0032196 | transposition(GO:0032196) |
| 0.3 | 0.9 | GO:0097272 | ammonia homeostasis(GO:0097272) |
| 0.3 | 3.4 | GO:0051344 | negative regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051344) |
| 0.3 | 0.9 | GO:0035425 | autocrine signaling(GO:0035425) |
| 0.3 | 1.7 | GO:0035063 | nuclear speck organization(GO:0035063) |
| 0.3 | 2.9 | GO:2000773 | negative regulation of cellular senescence(GO:2000773) |
| 0.3 | 2.0 | GO:0060087 | relaxation of vascular smooth muscle(GO:0060087) |
| 0.3 | 1.1 | GO:0021815 | cell motility involved in cerebral cortex radial glia guided migration(GO:0021814) modulation of microtubule cytoskeleton involved in cerebral cortex radial glia guided migration(GO:0021815) |
| 0.3 | 1.3 | GO:0019276 | UDP-N-acetylgalactosamine metabolic process(GO:0019276) |
| 0.3 | 2.4 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.3 | 0.8 | GO:0070375 | ERK5 cascade(GO:0070375) |
| 0.3 | 2.1 | GO:0042905 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.3 | 5.4 | GO:0060122 | inner ear receptor stereocilium organization(GO:0060122) |
| 0.3 | 1.3 | GO:0021559 | trigeminal nerve development(GO:0021559) |
| 0.2 | 6.0 | GO:1904380 | endoplasmic reticulum mannose trimming(GO:1904380) |
| 0.2 | 2.4 | GO:0045657 | positive regulation of monocyte differentiation(GO:0045657) |
| 0.2 | 4.6 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.2 | 0.7 | GO:0055020 | apoptotic process involved in endocardial cushion morphogenesis(GO:0003277) intermediate mesoderm development(GO:0048389) intermediate mesoderm morphogenesis(GO:0048390) intermediate mesoderm formation(GO:0048391) intermediate mesodermal cell differentiation(GO:0048392) regulation of cardiac muscle fiber development(GO:0055018) positive regulation of cardiac muscle fiber development(GO:0055020) bud dilation involved in lung branching(GO:0060503) regulation of transcription from RNA polymerase II promoter involved in kidney development(GO:0060994) BMP signaling pathway involved in ureter morphogenesis(GO:0061149) renal system segmentation(GO:0061150) BMP signaling pathway involved in renal system segmentation(GO:0061151) pulmonary artery endothelial tube morphogenesis(GO:0061155) regulation of transcription from RNA polymerase II promoter involved in mesonephros development(GO:0061216) pattern specification involved in mesonephros development(GO:0061227) BMP signaling pathway involved in nephric duct formation(GO:0071893) negative regulation of branch elongation involved in ureteric bud branching(GO:0072096) negative regulation of branch elongation involved in ureteric bud branching by BMP signaling pathway(GO:0072097) anterior/posterior pattern specification involved in kidney development(GO:0072098) anterior/posterior pattern specification involved in ureteric bud development(GO:0072099) specification of ureteric bud anterior/posterior symmetry(GO:0072100) specification of ureteric bud anterior/posterior symmetry by BMP signaling pathway(GO:0072101) ureter urothelium development(GO:0072190) ureter epithelial cell differentiation(GO:0072192) ureter morphogenesis(GO:0072197) negative regulation of mesenchymal cell proliferation involved in ureter development(GO:0072200) positive regulation of cell proliferation involved in outflow tract morphogenesis(GO:1901964) cardiac jelly development(GO:1905072) regulation of metanephric S-shaped body morphogenesis(GO:2000004) negative regulation of metanephric S-shaped body morphogenesis(GO:2000005) regulation of metanephric comma-shaped body morphogenesis(GO:2000006) negative regulation of metanephric comma-shaped body morphogenesis(GO:2000007) negative regulation of cell proliferation involved in heart morphogenesis(GO:2000137) |
| 0.2 | 13.8 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.2 | 6.3 | GO:0034204 | lipid translocation(GO:0034204) phospholipid translocation(GO:0045332) |
| 0.2 | 4.8 | GO:0006704 | glucocorticoid biosynthetic process(GO:0006704) |
| 0.2 | 0.7 | GO:0018874 | benzoate metabolic process(GO:0018874) butyrate metabolic process(GO:0019605) |
| 0.2 | 7.4 | GO:0071711 | basement membrane organization(GO:0071711) |
| 0.2 | 8.1 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.2 | 4.8 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.2 | 0.2 | GO:2000538 | regulation of B cell chemotaxis(GO:2000537) positive regulation of B cell chemotaxis(GO:2000538) |
| 0.2 | 30.1 | GO:0006757 | glycolytic process(GO:0006096) ATP generation from ADP(GO:0006757) |
| 0.2 | 1.5 | GO:0009048 | dosage compensation(GO:0007549) dosage compensation by inactivation of X chromosome(GO:0009048) |
| 0.2 | 6.2 | GO:0009083 | branched-chain amino acid catabolic process(GO:0009083) |
| 0.2 | 4.7 | GO:1902475 | L-alpha-amino acid transmembrane transport(GO:1902475) |
| 0.2 | 31.7 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.2 | 3.1 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.2 | 2.0 | GO:0006069 | ethanol oxidation(GO:0006069) |
| 0.2 | 1.7 | GO:0051901 | positive regulation of mitochondrial depolarization(GO:0051901) |
| 0.2 | 4.4 | GO:0007530 | sex determination(GO:0007530) |
| 0.2 | 3.1 | GO:0006706 | steroid catabolic process(GO:0006706) |
| 0.2 | 0.4 | GO:0042320 | regulation of circadian sleep/wake cycle, REM sleep(GO:0042320) |
| 0.2 | 4.6 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.2 | 2.0 | GO:0009642 | response to light intensity(GO:0009642) |
| 0.2 | 1.6 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.2 | 2.3 | GO:0031987 | locomotion involved in locomotory behavior(GO:0031987) |
| 0.2 | 0.9 | GO:0061051 | positive regulation of cell growth involved in cardiac muscle cell development(GO:0061051) |
| 0.2 | 2.5 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.2 | 0.5 | GO:1902951 | negative regulation of dendritic spine maintenance(GO:1902951) |
| 0.2 | 1.7 | GO:0007638 | mechanosensory behavior(GO:0007638) |
| 0.2 | 0.9 | GO:0015842 | aminergic neurotransmitter loading into synaptic vesicle(GO:0015842) |
| 0.1 | 0.4 | GO:0071481 | cellular response to X-ray(GO:0071481) |
| 0.1 | 1.0 | GO:0010898 | positive regulation of triglyceride catabolic process(GO:0010898) |
| 0.1 | 2.1 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.1 | 7.9 | GO:0007368 | determination of left/right symmetry(GO:0007368) |
| 0.1 | 2.5 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.1 | 6.8 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.1 | 1.4 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.1 | 0.6 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.1 | 1.6 | GO:0046149 | porphyrin-containing compound catabolic process(GO:0006787) tetrapyrrole catabolic process(GO:0033015) heme catabolic process(GO:0042167) pigment catabolic process(GO:0046149) |
| 0.1 | 2.2 | GO:0003298 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.1 | 2.5 | GO:0060397 | JAK-STAT cascade involved in growth hormone signaling pathway(GO:0060397) |
| 0.1 | 5.4 | GO:0070306 | lens fiber cell differentiation(GO:0070306) |
| 0.1 | 1.4 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.1 | 3.5 | GO:0015813 | L-glutamate transport(GO:0015813) |
| 0.1 | 2.8 | GO:0014046 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.1 | 1.3 | GO:0042789 | mRNA transcription from RNA polymerase II promoter(GO:0042789) |
| 0.1 | 2.1 | GO:0048730 | epidermis morphogenesis(GO:0048730) |
| 0.1 | 2.8 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.1 | 4.0 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.1 | 0.4 | GO:0003199 | endocardial cushion to mesenchymal transition involved in heart valve formation(GO:0003199) positive regulation of blood vessel remodeling(GO:2000504) |
| 0.1 | 2.7 | GO:0000132 | establishment of mitotic spindle orientation(GO:0000132) |
| 0.1 | 14.3 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.1 | 2.9 | GO:2000398 | regulation of T cell differentiation in thymus(GO:0033081) regulation of thymocyte aggregation(GO:2000398) |
| 0.1 | 1.8 | GO:0035090 | maintenance of apical/basal cell polarity(GO:0035090) maintenance of epithelial cell apical/basal polarity(GO:0045199) |
| 0.1 | 27.2 | GO:0060271 | cilium morphogenesis(GO:0060271) |
| 0.1 | 3.3 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.1 | 1.4 | GO:0034391 | smooth muscle cell apoptotic process(GO:0034390) regulation of smooth muscle cell apoptotic process(GO:0034391) |
| 0.1 | 0.4 | GO:0071418 | cellular response to amine stimulus(GO:0071418) |
| 0.1 | 0.8 | GO:0030450 | regulation of complement activation, classical pathway(GO:0030450) negative regulation of complement activation, classical pathway(GO:0045959) regulation of opsonization(GO:1903027) |
| 0.1 | 3.2 | GO:0007194 | negative regulation of adenylate cyclase activity(GO:0007194) |
| 0.1 | 1.1 | GO:2000169 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) regulation of peptidyl-cysteine S-nitrosylation(GO:2000169) |
| 0.1 | 0.4 | GO:1902725 | regulation of skeletal muscle tissue growth(GO:0048631) negative regulation of satellite cell differentiation(GO:1902725) |
| 0.1 | 2.8 | GO:0007616 | long-term memory(GO:0007616) |
| 0.1 | 7.7 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.1 | 0.3 | GO:0021965 | spinal cord ventral commissure morphogenesis(GO:0021965) |
| 0.1 | 0.7 | GO:0009957 | epidermal cell fate specification(GO:0009957) |
| 0.1 | 6.4 | GO:0035690 | cellular response to drug(GO:0035690) |
| 0.1 | 2.1 | GO:0002792 | negative regulation of peptide secretion(GO:0002792) negative regulation of insulin secretion(GO:0046676) negative regulation of peptide hormone secretion(GO:0090278) |
| 0.1 | 0.7 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 0.1 | 7.6 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.1 | 5.7 | GO:0051703 | social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.1 | 69.7 | GO:0007417 | central nervous system development(GO:0007417) |
| 0.1 | 3.8 | GO:0086010 | membrane depolarization during action potential(GO:0086010) |
| 0.1 | 2.3 | GO:0060765 | regulation of androgen receptor signaling pathway(GO:0060765) |
| 0.1 | 1.0 | GO:0032411 | positive regulation of transporter activity(GO:0032411) |
| 0.1 | 1.3 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.1 | 0.5 | GO:0045078 | positive regulation of interferon-gamma biosynthetic process(GO:0045078) |
| 0.1 | 3.4 | GO:0035196 | production of miRNAs involved in gene silencing by miRNA(GO:0035196) |
| 0.1 | 27.0 | GO:0051056 | regulation of small GTPase mediated signal transduction(GO:0051056) |
| 0.1 | 1.1 | GO:0035563 | positive regulation of chromatin binding(GO:0035563) |
| 0.1 | 1.0 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 2.4 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.1 | 0.1 | GO:0044339 | canonical Wnt signaling pathway involved in osteoblast differentiation(GO:0044339) |
| 0.1 | 3.8 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.1 | 7.3 | GO:0048024 | regulation of mRNA splicing, via spliceosome(GO:0048024) |
| 0.1 | 4.6 | GO:0032981 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.1 | 1.8 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.1 | 0.2 | GO:0043017 | negative regulation of T cell tolerance induction(GO:0002665) negative regulation of T cell anergy(GO:0002668) negative regulation of lymphocyte anergy(GO:0002912) regulation of lymphotoxin A production(GO:0032681) positive regulation of lymphotoxin A production(GO:0032761) regulation of lymphotoxin A biosynthetic process(GO:0043016) positive regulation of lymphotoxin A biosynthetic process(GO:0043017) |
| 0.1 | 4.0 | GO:0031214 | biomineral tissue development(GO:0031214) |
| 0.1 | 4.7 | GO:0036498 | IRE1-mediated unfolded protein response(GO:0036498) |
| 0.1 | 0.7 | GO:0061436 | regulation of water loss via skin(GO:0033561) establishment of skin barrier(GO:0061436) |
| 0.1 | 2.4 | GO:0075733 | intracellular transport of virus(GO:0075733) |
| 0.1 | 1.4 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.1 | 0.2 | GO:1904381 | Golgi apparatus mannose trimming(GO:1904381) |
| 0.1 | 5.1 | GO:0097194 | execution phase of apoptosis(GO:0097194) |
| 0.1 | 2.0 | GO:0050918 | positive chemotaxis(GO:0050918) |
| 0.1 | 5.6 | GO:0006937 | regulation of muscle contraction(GO:0006937) |
| 0.1 | 2.4 | GO:0042246 | tissue regeneration(GO:0042246) |
| 0.1 | 1.3 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 0.6 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.0 | 2.4 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.0 | 2.0 | GO:1904837 | beta-catenin-TCF complex assembly(GO:1904837) |
| 0.0 | 0.6 | GO:0031573 | intra-S DNA damage checkpoint(GO:0031573) |
| 0.0 | 6.3 | GO:0007411 | axon guidance(GO:0007411) |
| 0.0 | 0.4 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.0 | 1.6 | GO:0072164 | ureteric bud development(GO:0001657) mesonephros development(GO:0001823) mesonephric epithelium development(GO:0072163) mesonephric tubule development(GO:0072164) |
| 0.0 | 1.0 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.0 | 0.6 | GO:0052695 | cellular glucuronidation(GO:0052695) |
| 0.0 | 2.0 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.0 | 0.6 | GO:0042761 | fatty acid elongation(GO:0030497) very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.0 | 2.4 | GO:0032418 | lysosome localization(GO:0032418) |
| 0.0 | 0.5 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) |
| 0.0 | 2.7 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.0 | 1.3 | GO:0080171 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.0 | 1.0 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.0 | 3.9 | GO:0002576 | platelet degranulation(GO:0002576) |
| 0.0 | 0.1 | GO:0042635 | positive regulation of hair cycle(GO:0042635) positive regulation of hair follicle development(GO:0051798) |
| 0.0 | 1.4 | GO:0042771 | intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:0042771) |
| 0.0 | 4.3 | GO:0071805 | cellular potassium ion transport(GO:0071804) potassium ion transmembrane transport(GO:0071805) |
| 0.0 | 0.5 | GO:0014823 | response to activity(GO:0014823) |
| 0.0 | 1.5 | GO:2000117 | negative regulation of cysteine-type endopeptidase activity(GO:2000117) |
| 0.0 | 0.7 | GO:0030819 | positive regulation of cAMP biosynthetic process(GO:0030819) |
| 0.0 | 0.3 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 0.2 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 1.8 | GO:0007548 | sex differentiation(GO:0007548) |
| 0.0 | 1.2 | GO:0008277 | regulation of G-protein coupled receptor protein signaling pathway(GO:0008277) |
| 0.0 | 2.1 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 1.0 | GO:0050796 | regulation of insulin secretion(GO:0050796) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.8 | 31.1 | GO:0034365 | discoidal high-density lipoprotein particle(GO:0034365) |
| 5.1 | 35.6 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 3.6 | 14.4 | GO:0002945 | cyclin K-CDK13 complex(GO:0002945) |
| 3.1 | 12.4 | GO:0071546 | pi-body(GO:0071546) |
| 3.0 | 48.4 | GO:0045277 | respiratory chain complex IV(GO:0045277) |
| 2.6 | 41.5 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 2.2 | 13.5 | GO:0033063 | DNA recombinase mediator complex(GO:0033061) Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 2.0 | 5.9 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 1.7 | 13.4 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 1.6 | 6.2 | GO:0030895 | apolipoprotein B mRNA editing enzyme complex(GO:0030895) |
| 1.5 | 8.9 | GO:0032584 | growth cone membrane(GO:0032584) |
| 1.4 | 17.3 | GO:0016013 | syntrophin complex(GO:0016013) |
| 1.4 | 67.8 | GO:0031430 | M band(GO:0031430) |
| 1.3 | 26.5 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 1.3 | 20.4 | GO:0030914 | STAGA complex(GO:0030914) |
| 1.2 | 6.0 | GO:0000835 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 1.2 | 7.1 | GO:1990357 | terminal web(GO:1990357) |
| 1.1 | 62.3 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 1.0 | 22.7 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.9 | 19.0 | GO:0031045 | dense core granule(GO:0031045) |
| 0.9 | 4.4 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.8 | 12.5 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.8 | 14.7 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.8 | 5.4 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.8 | 10.0 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.7 | 4.5 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.7 | 16.3 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.7 | 6.8 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.7 | 25.4 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.7 | 8.5 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.6 | 3.2 | GO:0036398 | TCR signalosome(GO:0036398) |
| 0.6 | 8.9 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.6 | 2.5 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.6 | 4.3 | GO:0043034 | costamere(GO:0043034) |
| 0.6 | 7.1 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.6 | 1.7 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 0.6 | 3.4 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.6 | 6.6 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.5 | 6.4 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.5 | 3.6 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.5 | 2.5 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.5 | 4.9 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.5 | 2.4 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 0.5 | 129.9 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.5 | 2.8 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.4 | 3.1 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.4 | 3.1 | GO:0060091 | kinocilium(GO:0060091) |
| 0.4 | 4.0 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.4 | 5.6 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.4 | 3.4 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.4 | 3.3 | GO:0005593 | FACIT collagen trimer(GO:0005593) |
| 0.4 | 4.5 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.4 | 7.9 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.4 | 14.7 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.4 | 6.9 | GO:0036038 | MKS complex(GO:0036038) |
| 0.4 | 4.2 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.4 | 9.0 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.3 | 5.5 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.3 | 1.4 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.3 | 4.6 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.3 | 7.2 | GO:0031672 | A band(GO:0031672) |
| 0.3 | 29.5 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.3 | 2.0 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.3 | 5.7 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.3 | 2.4 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.3 | 2.0 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.2 | 5.1 | GO:0097342 | ripoptosome(GO:0097342) |
| 0.2 | 2.1 | GO:0034993 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.2 | 2.4 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.2 | 1.1 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.2 | 1.5 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.2 | 12.1 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.2 | 7.1 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.2 | 62.4 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.2 | 1.8 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.2 | 5.4 | GO:0032420 | stereocilium(GO:0032420) |
| 0.2 | 3.8 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.2 | 0.9 | GO:0070081 | clathrin-sculpted monoamine transport vesicle(GO:0070081) clathrin-sculpted monoamine transport vesicle membrane(GO:0070083) |
| 0.1 | 6.2 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 8.6 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 4.7 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.1 | 10.6 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.1 | 1.8 | GO:0031906 | late endosome lumen(GO:0031906) |
| 0.1 | 15.2 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.1 | 3.4 | GO:0001917 | photoreceptor inner segment(GO:0001917) |
| 0.1 | 18.3 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.1 | 13.8 | GO:0043204 | perikaryon(GO:0043204) |
| 0.1 | 2.4 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.1 | 3.0 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 4.4 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 4.2 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 5.9 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.1 | 1.5 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 0.9 | GO:0044298 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.1 | 2.5 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.1 | 6.0 | GO:0030017 | sarcomere(GO:0030017) |
| 0.1 | 6.6 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 2.5 | GO:0031904 | endosome lumen(GO:0031904) |
| 0.1 | 1.9 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 0.9 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.1 | 2.2 | GO:0097060 | synaptic membrane(GO:0097060) |
| 0.1 | 9.3 | GO:0034399 | nuclear periphery(GO:0034399) |
| 0.0 | 0.9 | GO:0090533 | cation-transporting ATPase complex(GO:0090533) |
| 0.0 | 0.4 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.0 | 80.1 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 1.3 | GO:0098858 | actin-based cell projection(GO:0098858) |
| 0.0 | 0.8 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 2.6 | GO:0043235 | receptor complex(GO:0043235) |
| 0.0 | 9.1 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 2.1 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 13.8 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 2.3 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 7.6 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 5.6 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 1.9 | GO:0008023 | transcription elongation factor complex(GO:0008023) |
| 0.0 | 0.6 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 1.2 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 16.3 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 2.6 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.0 | 1.4 | GO:0055029 | DNA-directed RNA polymerase complex(GO:0000428) RNA polymerase complex(GO:0030880) nuclear DNA-directed RNA polymerase complex(GO:0055029) |
| 0.0 | 0.4 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.5 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 0.3 | GO:0005905 | clathrin-coated pit(GO:0005905) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.8 | 31.1 | GO:0070326 | very-low-density lipoprotein particle receptor binding(GO:0070326) |
| 7.1 | 35.6 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 5.3 | 31.6 | GO:0000285 | 1-phosphatidylinositol-3-phosphate 5-kinase activity(GO:0000285) |
| 3.9 | 15.4 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 3.7 | 18.7 | GO:0047273 | galactosylgalactosylglucosylceramide beta-D-acetylgalactosaminyltransferase activity(GO:0047273) |
| 3.0 | 11.9 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 2.9 | 17.3 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 2.4 | 7.3 | GO:0016838 | carbon-oxygen lyase activity, acting on phosphates(GO:0016838) |
| 2.1 | 6.4 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 2.1 | 8.4 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 2.0 | 12.0 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 1.9 | 13.6 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 1.9 | 7.7 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 1.9 | 7.6 | GO:0005277 | acetylcholine transmembrane transporter activity(GO:0005277) secondary active organic cation transmembrane transporter activity(GO:0008513) acetate ester transmembrane transporter activity(GO:1901375) |
| 1.7 | 50.1 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 1.6 | 49.1 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 1.6 | 9.8 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 1.6 | 4.8 | GO:0003845 | 11-beta-hydroxysteroid dehydrogenase [NAD(P)] activity(GO:0003845) |
| 1.6 | 18.8 | GO:0008430 | selenium binding(GO:0008430) |
| 1.5 | 4.6 | GO:0090555 | phosphatidylethanolamine-translocating ATPase activity(GO:0090555) |
| 1.4 | 1.4 | GO:0001034 | RNA polymerase III transcription factor activity, sequence-specific DNA binding(GO:0001034) |
| 1.3 | 3.9 | GO:0015203 | polyamine transmembrane transporter activity(GO:0015203) putrescine transmembrane transporter activity(GO:0015489) |
| 1.3 | 85.7 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 1.3 | 5.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 1.2 | 11.1 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 1.1 | 70.5 | GO:0030507 | spectrin binding(GO:0030507) |
| 1.1 | 3.4 | GO:0017129 | triglyceride binding(GO:0017129) |
| 1.1 | 3.3 | GO:0061609 | fructose-1-phosphate aldolase activity(GO:0061609) |
| 1.1 | 31.0 | GO:0019865 | immunoglobulin binding(GO:0019865) |
| 1.1 | 4.4 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 1.0 | 8.9 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.9 | 5.7 | GO:0005223 | intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.9 | 2.8 | GO:0015234 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) |
| 0.9 | 3.8 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.9 | 2.8 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.9 | 4.5 | GO:0004803 | transposase activity(GO:0004803) |
| 0.9 | 2.7 | GO:0008267 | poly-glutamine tract binding(GO:0008267) |
| 0.9 | 16.3 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.8 | 5.1 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.8 | 5.0 | GO:0042285 | UDP-xylosyltransferase activity(GO:0035252) xylosyltransferase activity(GO:0042285) |
| 0.8 | 2.5 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 0.8 | 4.1 | GO:0047374 | sterol esterase activity(GO:0004771) methylumbelliferyl-acetate deacetylase activity(GO:0047374) |
| 0.8 | 12.0 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.8 | 3.1 | GO:0004773 | steryl-sulfatase activity(GO:0004773) |
| 0.7 | 13.4 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.7 | 3.7 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.7 | 7.9 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.7 | 2.1 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.7 | 4.2 | GO:0017098 | sulfonylurea receptor binding(GO:0017098) |
| 0.7 | 4.7 | GO:0000182 | rDNA binding(GO:0000182) |
| 0.7 | 14.4 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.6 | 2.5 | GO:0005148 | prolactin receptor binding(GO:0005148) |
| 0.6 | 13.3 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.6 | 144.7 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.6 | 5.9 | GO:0004064 | arylesterase activity(GO:0004064) |
| 0.6 | 7.9 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.6 | 6.2 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.6 | 4.5 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.6 | 6.6 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.5 | 8.7 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.5 | 4.2 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.5 | 4.6 | GO:1904315 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.5 | 4.0 | GO:0030345 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.5 | 5.5 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.5 | 16.4 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.5 | 1.9 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.5 | 6.4 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.5 | 17.4 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.5 | 10.5 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.5 | 10.4 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.4 | 1.3 | GO:0035248 | alpha-1,4-N-acetylgalactosaminyltransferase activity(GO:0035248) |
| 0.4 | 12.9 | GO:0071949 | FAD binding(GO:0071949) |
| 0.4 | 4.4 | GO:0047035 | testosterone dehydrogenase (NAD+) activity(GO:0047035) |
| 0.4 | 4.3 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.4 | 2.1 | GO:0031711 | tripeptidyl-peptidase activity(GO:0008240) peptidyl-dipeptidase activity(GO:0008241) bradykinin receptor binding(GO:0031711) metallodipeptidase activity(GO:0070573) |
| 0.4 | 1.6 | GO:0015180 | L-alanine transmembrane transporter activity(GO:0015180) alanine transmembrane transporter activity(GO:0022858) |
| 0.4 | 2.4 | GO:0047696 | beta-adrenergic receptor kinase activity(GO:0047696) |
| 0.4 | 3.2 | GO:0030275 | LRR domain binding(GO:0030275) |
| 0.4 | 2.0 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.4 | 7.7 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.4 | 1.5 | GO:0005415 | nucleoside:sodium symporter activity(GO:0005415) |
| 0.4 | 3.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.4 | 2.5 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.3 | 5.5 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.3 | 1.0 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.3 | 4.7 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.3 | 4.0 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.3 | 3.9 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.3 | 3.4 | GO:0046625 | sphingolipid binding(GO:0046625) |
| 0.3 | 53.9 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.3 | 4.0 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.3 | 2.4 | GO:0043813 | phosphatidylinositol-3,5-bisphosphate 5-phosphatase activity(GO:0043813) |
| 0.3 | 21.0 | GO:0048365 | Rac GTPase binding(GO:0048365) |
| 0.3 | 0.9 | GO:0015222 | serotonin transmembrane transporter activity(GO:0015222) |
| 0.3 | 1.8 | GO:0005143 | interleukin-12 receptor binding(GO:0005143) |
| 0.3 | 8.5 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.3 | 0.9 | GO:0050815 | phosphoserine binding(GO:0050815) |
| 0.3 | 1.7 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.3 | 25.3 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.3 | 5.8 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.3 | 2.4 | GO:0001056 | RNA polymerase III activity(GO:0001056) |
| 0.3 | 1.1 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.3 | 6.3 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.3 | 5.2 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.3 | 17.7 | GO:0070888 | E-box binding(GO:0070888) |
| 0.3 | 20.0 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.3 | 9.5 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.3 | 2.0 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) |
| 0.2 | 1.7 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.2 | 6.8 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.2 | 1.4 | GO:0048101 | calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.2 | 6.9 | GO:0038187 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.2 | 1.6 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.2 | 20.4 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.2 | 1.1 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.2 | 24.3 | GO:0003774 | motor activity(GO:0003774) |
| 0.2 | 4.9 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.2 | 14.4 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.2 | 3.3 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.2 | 0.6 | GO:0004466 | long-chain-acyl-CoA dehydrogenase activity(GO:0004466) |
| 0.2 | 3.5 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.2 | 0.5 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 0.2 | 6.3 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.2 | 0.9 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.2 | 4.7 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.2 | 0.7 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 0.2 | 1.0 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.2 | 6.6 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.2 | 2.2 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.2 | 0.8 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.2 | 3.8 | GO:1905030 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.2 | 1.8 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.2 | 4.4 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.2 | 1.0 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.2 | 4.0 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.2 | 0.9 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.2 | 3.2 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.2 | 0.8 | GO:0023024 | MHC class I protein complex binding(GO:0023024) |
| 0.1 | 1.9 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.1 | 4.7 | GO:0015175 | neutral amino acid transmembrane transporter activity(GO:0015175) |
| 0.1 | 4.4 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.1 | 1.7 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.1 | 3.5 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.1 | 2.5 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.1 | 5.6 | GO:0004532 | exoribonuclease activity(GO:0004532) |
| 0.1 | 7.3 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 43.9 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.1 | 4.8 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.1 | 36.2 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.1 | 4.4 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 1.0 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.1 | 4.9 | GO:0008536 | nuclear localization sequence binding(GO:0008139) Ran GTPase binding(GO:0008536) |
| 0.1 | 11.5 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 2.4 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.1 | 0.5 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 2.2 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.1 | 3.2 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 2.8 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.1 | 3.1 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 0.4 | GO:0004852 | uroporphyrinogen-III synthase activity(GO:0004852) |
| 0.1 | 0.6 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) |
| 0.1 | 0.3 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.1 | 0.4 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.1 | 0.7 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.1 | 1.7 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 0.5 | GO:0019958 | C-X-C chemokine binding(GO:0019958) |
| 0.1 | 1.8 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.1 | 1.3 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 2.3 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 16.7 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) ubiquitin-like protein ligase activity(GO:0061659) |
| 0.1 | 0.7 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.1 | 2.7 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 1.9 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 0.6 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.1 | 0.8 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.1 | 1.0 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.1 | 3.8 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 6.5 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 3.4 | GO:0002039 | p53 binding(GO:0002039) |
| 0.0 | 10.8 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 4.2 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.9 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 1.2 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 3.9 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 3.0 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 3.0 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.0 | 3.4 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 1.4 | GO:0004527 | exonuclease activity(GO:0004527) |
| 0.0 | 5.7 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 1.3 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 0.2 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.0 | 0.8 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.0 | 2.0 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 1.9 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.0 | 2.6 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 6.7 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 0.5 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) lysophosphatidic acid acyltransferase activity(GO:0042171) lysophospholipid acyltransferase activity(GO:0071617) |
| 0.0 | 11.5 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 0.1 | GO:0048185 | activin binding(GO:0048185) |
| 0.0 | 0.4 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 1.0 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 0.9 | GO:0004540 | ribonuclease activity(GO:0004540) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 15.9 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.5 | 14.0 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.3 | 12.0 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.3 | 16.5 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.3 | 12.8 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.3 | 6.4 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.3 | 15.3 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.3 | 5.2 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.2 | 2.9 | SIG IL4RECEPTOR IN B LYPHOCYTES | Genes related to IL4 rceptor signaling in B lymphocytes |
| 0.2 | 4.5 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.2 | 13.1 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.2 | 8.9 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.2 | 4.2 | SIG CHEMOTAXIS | Genes related to chemotaxis |
| 0.2 | 2.8 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.2 | 3.0 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.2 | 13.2 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.2 | 5.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 5.6 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.1 | 7.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 8.2 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 6.4 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.1 | 3.2 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.1 | 8.8 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
| 0.1 | 24.2 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 1.5 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.1 | 3.4 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 2.9 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.1 | 4.0 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 7.9 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.1 | 2.0 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.1 | 1.7 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 9.2 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.1 | 1.0 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.1 | 2.5 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.1 | 1.7 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 2.1 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 1.0 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 2.4 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 3.0 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 0.8 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 1.4 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 1.7 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.0 | 2.4 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 1.8 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 1.0 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 1.7 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 0.4 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 1.9 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 3.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.4 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 0.2 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 0.3 | PID EPHB FWD PATHWAY | EPHB forward signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 29.5 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 1.7 | 36.7 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 1.2 | 26.5 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.8 | 11.0 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.7 | 8.6 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.6 | 13.8 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.5 | 9.7 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.5 | 13.4 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.5 | 5.6 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.5 | 11.1 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.4 | 7.6 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.4 | 4.5 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.4 | 16.3 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.4 | 5.9 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.3 | 2.4 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.3 | 8.1 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.3 | 10.7 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.3 | 5.5 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.3 | 3.1 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.3 | 5.1 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.3 | 7.7 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.3 | 2.9 | REACTOME OPSINS | Genes involved in Opsins |
| 0.2 | 3.7 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.2 | 14.6 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.2 | 17.8 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.2 | 2.0 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.2 | 3.3 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.2 | 5.5 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.2 | 2.4 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.2 | 15.3 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.2 | 7.1 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.2 | 2.1 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.2 | 5.5 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.2 | 5.8 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.2 | 3.1 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.2 | 3.0 | REACTOME INITIAL TRIGGERING OF COMPLEMENT | Genes involved in Initial triggering of complement |
| 0.2 | 2.9 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.2 | 1.4 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.1 | 5.5 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.1 | 5.7 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.1 | 2.8 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.1 | 6.4 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 2.1 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.1 | 1.7 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.1 | 11.1 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 3.1 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.1 | 2.2 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 1.8 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.1 | 3.6 | REACTOME INWARDLY RECTIFYING K CHANNELS | Genes involved in Inwardly rectifying K+ channels |
| 0.1 | 1.0 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.1 | 5.2 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 3.1 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.1 | 1.2 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.1 | 8.5 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 3.3 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.1 | 2.2 | REACTOME PERK REGULATED GENE EXPRESSION | Genes involved in PERK regulated gene expression |
| 0.1 | 2.0 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.1 | 4.2 | REACTOME POST NMDA RECEPTOR ACTIVATION EVENTS | Genes involved in Post NMDA receptor activation events |
| 0.1 | 11.9 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.1 | 1.7 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.1 | 0.9 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 2.1 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 1.5 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.0 | 0.5 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.6 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.0 | 1.2 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 3.6 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.3 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 0.8 | REACTOME NGF SIGNALLING VIA TRKA FROM THE PLASMA MEMBRANE | Genes involved in NGF signalling via TRKA from the plasma membrane |
| 0.0 | 1.2 | REACTOME CELL CELL JUNCTION ORGANIZATION | Genes involved in Cell-cell junction organization |
| 0.0 | 1.9 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 3.1 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 1.1 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 1.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 1.9 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.4 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.0 | 0.2 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.0 | 0.6 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.0 | 1.9 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 1.6 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.9 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 1.5 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.4 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 0.3 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 0.9 | REACTOME INTEGRATION OF ENERGY METABOLISM | Genes involved in Integration of energy metabolism |