Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
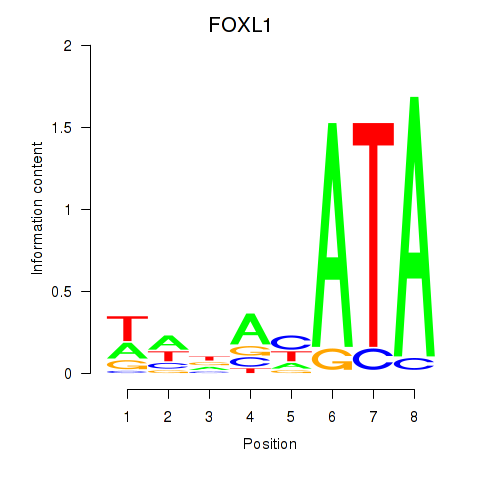
Results for FOXL1
Z-value: 1.71
Transcription factors associated with FOXL1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
FOXL1
|
ENSG00000176678.6 | FOXL1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| FOXL1 | hg38_v1_chr16_+_86578543_86578555 | 0.37 | 2.3e-08 | Click! |
Activity profile of FOXL1 motif
Sorted Z-values of FOXL1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of FOXL1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_90069662 | 52.21 |
ENST00000390271.2
|
IGKV6D-41
|
immunoglobulin kappa variable 6D-41 (non-functional) |
| chr2_-_89160329 | 47.50 |
ENST00000390256.2
|
IGKV6-21
|
immunoglobulin kappa variable 6-21 (non-functional) |
| chr12_+_69348372 | 46.02 |
ENST00000261267.7
ENST00000549690.1 ENST00000548839.1 |
LYZ
|
lysozyme |
| chr12_-_91180365 | 45.21 |
ENST00000547937.5
|
DCN
|
decorin |
| chr12_-_91179472 | 39.62 |
ENST00000550099.5
ENST00000546391.5 |
DCN
|
decorin |
| chr2_-_88992903 | 33.77 |
ENST00000495489.1
|
IGKV1-8
|
immunoglobulin kappa variable 1-8 |
| chr6_+_33080445 | 33.76 |
ENST00000428835.5
|
HLA-DPB1
|
major histocompatibility complex, class II, DP beta 1 |
| chr12_-_91179517 | 30.25 |
ENST00000551354.1
|
DCN
|
decorin |
| chr22_+_22900976 | 30.20 |
ENST00000390323.2
|
IGLC2
|
immunoglobulin lambda constant 2 |
| chr2_+_90021567 | 29.09 |
ENST00000436451.2
|
IGKV6D-21
|
immunoglobulin kappa variable 6D-21 (non-functional) |
| chr12_-_91153149 | 28.82 |
ENST00000550758.1
|
DCN
|
decorin |
| chr6_-_32589833 | 27.91 |
ENST00000360004.5
|
HLA-DRB1
|
major histocompatibility complex, class II, DR beta 1 |
| chr4_-_70666492 | 27.78 |
ENST00000254801.9
ENST00000391614.7 |
JCHAIN
|
joining chain of multimeric IgA and IgM |
| chr4_-_99352754 | 27.25 |
ENST00000639454.1
|
ADH1B
|
alcohol dehydrogenase 1B (class I), beta polypeptide |
| chr7_+_142791635 | 27.17 |
ENST00000633705.1
|
TRBC1
|
T cell receptor beta constant 1 |
| chr2_-_89320146 | 26.99 |
ENST00000498574.1
|
IGKV1-39
|
immunoglobulin kappa variable 1-39 |
| chr2_-_89222461 | 26.93 |
ENST00000482769.1
|
IGKV2-28
|
immunoglobulin kappa variable 2-28 |
| chr1_+_116754422 | 26.92 |
ENST00000369478.4
ENST00000369477.1 |
CD2
|
CD2 molecule |
| chr2_+_89936859 | 26.54 |
ENST00000474213.1
|
IGKV2D-30
|
immunoglobulin kappa variable 2D-30 |
| chr2_-_89100352 | 25.55 |
ENST00000479981.1
|
IGKV1-16
|
immunoglobulin kappa variable 1-16 |
| chr12_-_91179355 | 25.19 |
ENST00000550563.5
ENST00000546370.5 |
DCN
|
decorin |
| chr14_-_106470788 | 24.10 |
ENST00000434710.1
|
IGHV3-43
|
immunoglobulin heavy variable 3-43 |
| chr11_-_5227063 | 23.18 |
ENST00000335295.4
ENST00000485743.1 ENST00000647020.1 |
HBB
|
hemoglobin subunit beta |
| chr8_-_81483226 | 22.24 |
ENST00000256104.5
|
FABP4
|
fatty acid binding protein 4 |
| chr4_-_99321362 | 22.06 |
ENST00000625860.2
ENST00000305046.13 ENST00000506651.5 |
ADH1B
|
alcohol dehydrogenase 1B (class I), beta polypeptide |
| chr2_+_89862438 | 21.94 |
ENST00000448155.2
|
IGKV1D-39
|
immunoglobulin kappa variable 1D-39 |
| chr12_-_91178520 | 21.66 |
ENST00000425043.5
ENST00000420120.6 ENST00000441303.6 ENST00000456569.2 |
DCN
|
decorin |
| chr14_-_106025628 | 21.64 |
ENST00000631943.1
|
IGHV7-4-1
|
immunoglobulin heavy variable 7-4-1 |
| chr2_-_89245596 | 20.83 |
ENST00000468494.1
|
IGKV2-30
|
immunoglobulin kappa variable 2-30 |
| chr14_-_106675544 | 20.79 |
ENST00000390632.2
|
IGHV3-66
|
immunoglobulin heavy variable 3-66 |
| chr16_+_33802683 | 19.90 |
ENST00000570121.2
|
IGHV3OR16-12
|
immunoglobulin heavy variable 3/OR16-12 (non-functional) |
| chr2_-_88947820 | 19.79 |
ENST00000496168.1
|
IGKV1-5
|
immunoglobulin kappa variable 1-5 |
| chr4_+_70383123 | 19.74 |
ENST00000304915.8
|
SMR3B
|
submaxillary gland androgen regulated protein 3B |
| chr2_-_89213917 | 19.00 |
ENST00000498435.1
|
IGKV1-27
|
immunoglobulin kappa variable 1-27 |
| chr14_-_106593319 | 18.69 |
ENST00000390627.3
|
IGHV3-53
|
immunoglobulin heavy variable 3-53 |
| chr6_+_32741382 | 18.54 |
ENST00000374940.4
|
HLA-DQA2
|
major histocompatibility complex, class II, DQ alpha 2 |
| chr2_+_90209873 | 18.46 |
ENST00000468879.1
|
IGKV1D-43
|
immunoglobulin kappa variable 1D-43 |
| chr12_-_14961559 | 18.36 |
ENST00000228945.9
ENST00000541546.5 |
ARHGDIB
|
Rho GDP dissociation inhibitor beta |
| chr14_-_106185387 | 17.56 |
ENST00000390605.2
|
IGHV1-18
|
immunoglobulin heavy variable 1-18 |
| chr2_+_90154073 | 17.56 |
ENST00000611391.1
|
IGKV1D-13
|
immunoglobulin kappa variable 1D-13 |
| chr2_+_188974364 | 17.38 |
ENST00000304636.9
ENST00000317840.9 |
COL3A1
|
collagen type III alpha 1 chain |
| chr2_+_88885397 | 16.94 |
ENST00000390243.2
|
IGKV4-1
|
immunoglobulin kappa variable 4-1 |
| chr14_-_106538331 | 16.67 |
ENST00000390624.3
|
IGHV3-48
|
immunoglobulin heavy variable 3-48 |
| chr2_+_90159840 | 16.23 |
ENST00000377032.5
|
IGKV1D-12
|
immunoglobulin kappa variable 1D-12 |
| chr4_+_69995958 | 15.91 |
ENST00000381060.2
ENST00000246895.9 |
STATH
|
statherin |
| chr1_-_9943314 | 15.83 |
ENST00000377223.6
ENST00000377213.1 |
LZIC
|
leucine zipper and CTNNBIP1 domain containing |
| chr2_+_90220727 | 15.68 |
ENST00000471857.2
|
IGKV1D-8
|
immunoglobulin kappa variable 1D-8 |
| chr12_-_11310420 | 15.53 |
ENST00000621732.4
ENST00000445719.2 ENST00000279575.7 |
PRB4
|
proline rich protein BstNI subfamily 4 |
| chr2_-_89117844 | 15.29 |
ENST00000490686.1
|
IGKV1-17
|
immunoglobulin kappa variable 1-17 |
| chr3_+_189789672 | 15.06 |
ENST00000434928.5
|
TP63
|
tumor protein p63 |
| chr16_-_33845229 | 14.79 |
ENST00000569103.2
|
IGHV3OR16-17
|
immunoglobulin heavy variable 3/OR16-17 (non-functional) |
| chr1_+_103750406 | 14.75 |
ENST00000370079.3
|
AMY1C
|
amylase alpha 1C |
| chr22_-_17221841 | 14.69 |
ENST00000449907.7
ENST00000441548.1 ENST00000399839.5 |
ADA2
|
adenosine deaminase 2 |
| chrX_+_1591590 | 14.56 |
ENST00000313871.9
ENST00000381261.8 |
AKAP17A
|
A-kinase anchoring protein 17A |
| chr12_-_7503744 | 14.46 |
ENST00000396620.7
ENST00000432237.3 |
CD163
|
CD163 molecule |
| chr11_-_59866478 | 14.32 |
ENST00000257264.4
|
TCN1
|
transcobalamin 1 |
| chr2_-_89143133 | 14.15 |
ENST00000492167.1
|
IGKV3-20
|
immunoglobulin kappa variable 3-20 |
| chr14_-_106658251 | 14.12 |
ENST00000454421.2
|
IGHV3-64
|
immunoglobulin heavy variable 3-64 |
| chr14_-_106360320 | 14.11 |
ENST00000390615.2
|
IGHV3-33
|
immunoglobulin heavy variable 3-33 |
| chr9_-_92482350 | 13.98 |
ENST00000375543.2
|
ASPN
|
asporin |
| chr2_+_89884740 | 13.93 |
ENST00000509129.1
|
IGKV1D-37
|
immunoglobulin kappa variable 1D-37 (non-functional) |
| chr3_+_111292719 | 13.82 |
ENST00000460744.1
|
CD96
|
CD96 molecule |
| chr14_-_106269133 | 13.77 |
ENST00000390609.3
|
IGHV3-23
|
immunoglobulin heavy variable 3-23 |
| chr9_-_21368962 | 13.73 |
ENST00000610660.1
|
IFNA13
|
interferon alpha 13 |
| chr2_+_89913982 | 13.28 |
ENST00000390265.2
|
IGKV1D-33
|
immunoglobulin kappa variable 1D-33 |
| chr12_-_7503841 | 12.96 |
ENST00000359156.8
|
CD163
|
CD163 molecule |
| chr12_-_11395556 | 12.94 |
ENST00000565533.1
ENST00000389362.6 ENST00000546254.3 |
PRB2
PRB1
|
proline rich protein BstNI subfamily 2 proline rich protein BstNI subfamily 1 |
| chr3_-_195583931 | 12.91 |
ENST00000343267.8
ENST00000421243.5 ENST00000453131.1 |
APOD
|
apolipoprotein D |
| chr4_-_99352730 | 12.89 |
ENST00000510055.5
ENST00000515683.6 ENST00000511397.3 |
ADH1C
|
alcohol dehydrogenase 1C (class I), gamma polypeptide |
| chr1_-_35769958 | 12.87 |
ENST00000251195.9
ENST00000318121.8 |
CLSPN
|
claspin |
| chr4_-_185956652 | 12.84 |
ENST00000355634.9
|
SORBS2
|
sorbin and SH3 domain containing 2 |
| chr6_-_32941018 | 12.38 |
ENST00000418107.3
|
HLA-DMB
|
major histocompatibility complex, class II, DM beta |
| chrX_-_119943732 | 12.24 |
ENST00000371410.5
|
NKAP
|
NFKB activating protein |
| chr9_-_92482499 | 12.21 |
ENST00000375544.7
|
ASPN
|
asporin |
| chr10_-_67838019 | 12.00 |
ENST00000483798.6
|
DNAJC12
|
DnaJ heat shock protein family (Hsp40) member C12 |
| chr14_-_105940235 | 11.81 |
ENST00000390593.2
|
IGHV6-1
|
immunoglobulin heavy variable 6-1 |
| chr14_+_21990357 | 11.79 |
ENST00000390444.1
|
TRAV16
|
T cell receptor alpha variable 16 |
| chr2_+_90082635 | 11.70 |
ENST00000483379.1
|
IGKV1D-17
|
immunoglobulin kappa variable 1D-17 |
| chr12_-_10849464 | 11.61 |
ENST00000544994.5
ENST00000228811.8 ENST00000540107.2 |
PRR4
|
proline rich 4 |
| chr14_-_106062670 | 11.60 |
ENST00000390598.2
|
IGHV3-7
|
immunoglobulin heavy variable 3-7 |
| chr3_+_8501846 | 11.46 |
ENST00000454244.4
|
LMCD1
|
LIM and cysteine rich domains 1 |
| chr3_-_121660892 | 11.25 |
ENST00000428394.6
ENST00000314583.8 |
HCLS1
|
hematopoietic cell-specific Lyn substrate 1 |
| chr3_+_151814102 | 11.23 |
ENST00000232892.12
|
AADAC
|
arylacetamide deacetylase |
| chr4_+_40197023 | 10.85 |
ENST00000381799.10
|
RHOH
|
ras homolog family member H |
| chr4_-_83114715 | 10.84 |
ENST00000426923.2
ENST00000311507.9 ENST00000509973.5 |
PLAC8
|
placenta associated 8 |
| chr12_-_14951106 | 10.62 |
ENST00000541644.5
ENST00000545895.5 |
ARHGDIB
|
Rho GDP dissociation inhibitor beta |
| chr11_+_6863057 | 10.51 |
ENST00000641461.1
|
OR10A2
|
olfactory receptor family 10 subfamily A member 2 |
| chr19_+_9185594 | 10.44 |
ENST00000344248.4
|
OR7D2
|
olfactory receptor family 7 subfamily D member 2 |
| chr17_-_69150062 | 10.42 |
ENST00000522787.5
ENST00000521538.5 |
ABCA10
|
ATP binding cassette subfamily A member 10 |
| chr2_+_90234809 | 10.18 |
ENST00000443397.5
|
IGKV3D-7
|
immunoglobulin kappa variable 3D-7 |
| chr2_-_89268506 | 10.08 |
ENST00000473726.1
|
IGKV1-33
|
immunoglobulin kappa variable 1-33 |
| chr1_-_161307420 | 9.97 |
ENST00000491222.5
ENST00000672287.2 |
MPZ
|
myelin protein zero |
| chr11_-_5234475 | 9.80 |
ENST00000292901.7
ENST00000650601.1 ENST00000417377.1 |
HBD
|
hemoglobin subunit delta |
| chr5_-_140633690 | 9.69 |
ENST00000512545.1
ENST00000401743.6 |
CD14
|
CD14 molecule |
| chr1_+_196652022 | 9.69 |
ENST00000367429.9
ENST00000630130.2 ENST00000359637.2 |
CFH
|
complement factor H |
| chr2_+_89959979 | 9.68 |
ENST00000453166.2
|
IGKV2D-28
|
immunoglobulin kappa variable 2D-28 |
| chr18_+_44680093 | 9.58 |
ENST00000426838.8
ENST00000677068.1 |
SETBP1
|
SET binding protein 1 |
| chrY_+_2841594 | 9.49 |
ENST00000250784.13
|
RPS4Y1
|
ribosomal protein S4 Y-linked 1 |
| chr19_-_9219589 | 9.46 |
ENST00000641244.1
ENST00000641669.1 |
OR7D4
|
olfactory receptor family 7 subfamily D member 4 |
| chr2_+_90114838 | 9.44 |
ENST00000417279.3
|
IGKV3D-15
|
immunoglobulin kappa variable 3D-15 |
| chr6_-_33069242 | 9.42 |
ENST00000437811.1
|
HLA-DPA1
|
major histocompatibility complex, class II, DP alpha 1 |
| chr4_+_70195719 | 9.42 |
ENST00000683306.1
|
ODAM
|
odontogenic, ameloblast associated |
| chr14_-_105987068 | 9.31 |
ENST00000390594.3
|
IGHV1-2
|
immunoglobulin heavy variable 1-2 |
| chr4_+_87832917 | 9.26 |
ENST00000395102.8
ENST00000497649.6 ENST00000540395.1 ENST00000560249.5 ENST00000511670.5 ENST00000361056.3 |
MEPE
|
matrix extracellular phosphoglycoprotein |
| chr2_+_89947508 | 9.20 |
ENST00000491977.1
|
IGKV2D-29
|
immunoglobulin kappa variable 2D-29 |
| chr14_-_106622837 | 9.09 |
ENST00000390628.3
|
IGHV1-58
|
immunoglobulin heavy variable 1-58 |
| chr2_-_86105839 | 9.06 |
ENST00000263857.11
|
POLR1A
|
RNA polymerase I subunit A |
| chr17_+_56978111 | 8.94 |
ENST00000262288.8
ENST00000572710.5 ENST00000575395.5 ENST00000631024.1 |
SCPEP1
|
serine carboxypeptidase 1 |
| chr2_+_238138661 | 8.93 |
ENST00000409223.2
|
KLHL30
|
kelch like family member 30 |
| chr4_+_105710809 | 8.90 |
ENST00000360505.9
ENST00000510865.5 ENST00000509336.5 |
GSTCD
|
glutathione S-transferase C-terminal domain containing |
| chr11_-_85665077 | 8.86 |
ENST00000527447.2
|
CREBZF
|
CREB/ATF bZIP transcription factor |
| chr14_-_106811131 | 8.80 |
ENST00000424969.2
|
IGHV3-74
|
immunoglobulin heavy variable 3-74 |
| chr4_-_99435336 | 8.80 |
ENST00000437033.7
|
ADH7
|
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr2_+_113406368 | 8.70 |
ENST00000453673.3
|
IGKV1OR2-108
|
immunoglobulin kappa variable 1/OR2-108 (non-functional) |
| chr9_+_98943705 | 8.60 |
ENST00000610452.1
|
COL15A1
|
collagen type XV alpha 1 chain |
| chr14_-_106324743 | 8.57 |
ENST00000390612.3
|
IGHV4-28
|
immunoglobulin heavy variable 4-28 |
| chr8_-_85341705 | 8.57 |
ENST00000517618.5
|
CA1
|
carbonic anhydrase 1 |
| chr3_+_187024614 | 8.52 |
ENST00000416235.6
|
ST6GAL1
|
ST6 beta-galactoside alpha-2,6-sialyltransferase 1 |
| chr10_-_43574555 | 8.45 |
ENST00000374446.7
ENST00000535642.5 ENST00000426961.1 |
ZNF239
|
zinc finger protein 239 |
| chr1_+_207325629 | 8.43 |
ENST00000618707.2
|
CD55
|
CD55 molecule (Cromer blood group) |
| chr17_-_31314040 | 8.43 |
ENST00000330927.5
|
EVI2B
|
ecotropic viral integration site 2B |
| chr16_-_3372666 | 8.38 |
ENST00000399974.5
|
MTRNR2L4
|
MT-RNR2 like 4 |
| chr15_-_21718245 | 8.35 |
ENST00000630556.1
|
ENSG00000281179.1
|
novel gene identicle to IGHV1OR15-1 |
| chr14_-_106511856 | 8.26 |
ENST00000390622.2
|
IGHV1-46
|
immunoglobulin heavy variable 1-46 |
| chr15_-_19988117 | 7.96 |
ENST00000558565.2
|
IGHV3OR15-7
|
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr1_+_196819731 | 7.88 |
ENST00000320493.10
ENST00000367424.4 |
CFHR1
|
complement factor H related 1 |
| chr11_-_13496018 | 7.88 |
ENST00000529816.1
|
PTH
|
parathyroid hormone |
| chr14_-_106277039 | 7.85 |
ENST00000390610.2
|
IGHV1-24
|
immunoglobulin heavy variable 1-24 |
| chr7_+_80646436 | 7.71 |
ENST00000419819.2
|
CD36
|
CD36 molecule |
| chr13_-_37598750 | 7.70 |
ENST00000379743.8
ENST00000379742.4 ENST00000379749.8 ENST00000379747.9 ENST00000541179.5 ENST00000541481.5 |
POSTN
|
periostin |
| chr4_-_69961007 | 7.67 |
ENST00000353151.3
|
CSN2
|
casein beta |
| chr2_-_206086057 | 7.65 |
ENST00000403263.6
|
INO80D
|
INO80 complex subunit D |
| chr14_-_106737547 | 7.56 |
ENST00000632209.1
|
IGHV1-69-2
|
immunoglobulin heavy variable 1-69-2 |
| chr12_-_10884244 | 7.53 |
ENST00000543626.4
|
PRH1
|
proline rich protein HaeIII subfamily 1 |
| chr10_-_69409275 | 7.51 |
ENST00000373307.5
|
TACR2
|
tachykinin receptor 2 |
| chr1_+_207053229 | 7.50 |
ENST00000367080.8
ENST00000367079.3 |
PFKFB2
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 |
| chr7_-_38249572 | 7.47 |
ENST00000436911.6
|
TRGC2
|
T cell receptor gamma constant 2 |
| chr16_+_72056153 | 7.46 |
ENST00000576168.6
ENST00000567185.7 ENST00000567612.2 |
HP
|
haptoglobin |
| chr3_+_38496467 | 7.42 |
ENST00000453767.1
|
EXOG
|
exo/endonuclease G |
| chr19_+_35995176 | 7.41 |
ENST00000378887.4
|
SDHAF1
|
succinate dehydrogenase complex assembly factor 1 |
| chr14_-_106235582 | 7.35 |
ENST00000390607.2
|
IGHV3-21
|
immunoglobulin heavy variable 3-21 |
| chr6_-_27912396 | 7.31 |
ENST00000303324.4
|
OR2B2
|
olfactory receptor family 2 subfamily B member 2 |
| chr10_+_114239245 | 7.30 |
ENST00000392982.8
|
VWA2
|
von Willebrand factor A domain containing 2 |
| chr10_+_94762673 | 7.27 |
ENST00000480405.2
ENST00000371321.9 |
CYP2C19
|
cytochrome P450 family 2 subfamily C member 19 |
| chr1_-_207052980 | 7.27 |
ENST00000367084.1
|
YOD1
|
YOD1 deubiquitinase |
| chr14_-_106639589 | 7.19 |
ENST00000390630.3
|
IGHV4-61
|
immunoglobulin heavy variable 4-61 |
| chr6_+_131250375 | 7.16 |
ENST00000474850.2
|
AKAP7
|
A-kinase anchoring protein 7 |
| chr3_+_138347648 | 7.13 |
ENST00000614350.4
ENST00000289104.8 |
MRAS
|
muscle RAS oncogene homolog |
| chr2_-_162152404 | 7.13 |
ENST00000375497.3
|
GCG
|
glucagon |
| chr2_-_152098810 | 7.12 |
ENST00000636442.1
ENST00000638005.1 |
CACNB4
|
calcium voltage-gated channel auxiliary subunit beta 4 |
| chr9_+_137742957 | 7.10 |
ENST00000637318.1
ENST00000478940.1 |
EHMT1
|
euchromatic histone lysine methyltransferase 1 |
| chr4_+_67558719 | 7.02 |
ENST00000265404.7
ENST00000396225.1 |
STAP1
|
signal transducing adaptor family member 1 |
| chr8_+_24294044 | 6.93 |
ENST00000265769.9
|
ADAM28
|
ADAM metallopeptidase domain 28 |
| chr14_-_106389858 | 6.86 |
ENST00000390617.2
|
IGHV3-35
|
immunoglobulin heavy variable 3-35 (non-functional) |
| chr4_+_70360751 | 6.85 |
ENST00000226460.5
|
SMR3A
|
submaxillary gland androgen regulated protein 3A |
| chr15_-_21742799 | 6.83 |
ENST00000622410.2
|
ENSG00000278263.2
|
novel protein, identical to IGHV4-4 |
| chr14_-_106165730 | 6.81 |
ENST00000390604.2
|
IGHV3-16
|
immunoglobulin heavy variable 3-16 (non-functional) |
| chr6_-_169250825 | 6.76 |
ENST00000676869.1
ENST00000676760.1 |
THBS2
|
thrombospondin 2 |
| chr11_-_60183191 | 6.75 |
ENST00000412309.6
|
MS4A6A
|
membrane spanning 4-domains A6A |
| chr13_-_46438190 | 6.73 |
ENST00000409879.6
|
RUBCNL
|
rubicon like autophagy enhancer |
| chr6_-_116126120 | 6.70 |
ENST00000452729.1
ENST00000651968.1 ENST00000243222.8 |
COL10A1
|
collagen type X alpha 1 chain |
| chr22_+_22322452 | 6.67 |
ENST00000390290.3
|
IGLV1-51
|
immunoglobulin lambda variable 1-51 |
| chr12_+_71686473 | 6.61 |
ENST00000549735.5
|
TMEM19
|
transmembrane protein 19 |
| chr1_+_198638968 | 6.59 |
ENST00000348564.11
ENST00000530727.5 ENST00000442510.8 ENST00000645247.1 ENST00000367367.8 ENST00000367364.5 ENST00000413409.6 |
PTPRC
|
protein tyrosine phosphatase receptor type C |
| chrX_-_57910458 | 6.57 |
ENST00000358697.6
|
ZXDA
|
zinc finger X-linked duplicated A |
| chr16_-_1611985 | 6.54 |
ENST00000426508.7
|
IFT140
|
intraflagellar transport 140 |
| chr4_+_70430487 | 6.53 |
ENST00000413702.5
|
MUC7
|
mucin 7, secreted |
| chr6_-_169253835 | 6.52 |
ENST00000649844.1
ENST00000617924.6 |
THBS2
|
thrombospondin 2 |
| chr1_-_163202835 | 6.51 |
ENST00000527988.1
ENST00000531476.1 ENST00000530507.5 ENST00000313961.10 |
RGS5
|
regulator of G protein signaling 5 |
| chrX_+_37780049 | 6.51 |
ENST00000378588.5
|
CYBB
|
cytochrome b-245 beta chain |
| chr18_+_31447732 | 6.50 |
ENST00000257189.5
|
DSG3
|
desmoglein 3 |
| chr4_+_70397931 | 6.50 |
ENST00000399575.7
|
OPRPN
|
opiorphin prepropeptide |
| chr12_+_133181409 | 6.45 |
ENST00000416488.5
ENST00000228289.9 ENST00000541211.6 ENST00000536435.7 ENST00000500625.7 ENST00000539248.6 ENST00000542711.6 ENST00000536899.6 ENST00000542986.6 ENST00000611984.4 ENST00000541975.2 |
ZNF268
|
zinc finger protein 268 |
| chr22_+_22380766 | 6.45 |
ENST00000390297.3
|
IGLV1-44
|
immunoglobulin lambda variable 1-44 |
| chr5_-_39425187 | 6.39 |
ENST00000545653.5
|
DAB2
|
DAB adaptor protein 2 |
| chr9_-_92536031 | 6.38 |
ENST00000344604.9
ENST00000375540.5 |
ECM2
|
extracellular matrix protein 2 |
| chr10_+_122560679 | 6.35 |
ENST00000657942.1
|
DMBT1
|
deleted in malignant brain tumors 1 |
| chr5_-_170297746 | 6.33 |
ENST00000046794.10
|
LCP2
|
lymphocyte cytosolic protein 2 |
| chr1_+_13583762 | 6.26 |
ENST00000376057.8
ENST00000621990.5 ENST00000510906.5 |
PDPN
|
podoplanin |
| chr19_+_54769785 | 6.25 |
ENST00000336077.11
|
KIR2DL1
|
killer cell immunoglobulin like receptor, two Ig domains and long cytoplasmic tail 1 |
| chr4_-_99290975 | 6.21 |
ENST00000209668.3
|
ADH1A
|
alcohol dehydrogenase 1A (class I), alpha polypeptide |
| chr10_-_96271508 | 6.20 |
ENST00000427367.6
ENST00000413476.6 ENST00000371176.6 |
BLNK
|
B cell linker |
| chr5_-_79991237 | 6.17 |
ENST00000512560.5
ENST00000509852.6 ENST00000512528.3 ENST00000617335.4 |
MTX3
|
metaxin 3 |
| chr14_-_106088573 | 6.15 |
ENST00000632099.1
|
IGHV3-64D
|
immunoglobulin heavy variable 3-64D |
| chr19_-_54364863 | 6.14 |
ENST00000348231.8
|
LAIR1
|
leukocyte associated immunoglobulin like receptor 1 |
| chr7_-_150341615 | 6.09 |
ENST00000223271.8
ENST00000466675.5 ENST00000482669.1 ENST00000467793.5 |
RARRES2
|
retinoic acid receptor responder 2 |
| chr8_+_93754844 | 6.06 |
ENST00000684064.1
ENST00000498673.5 ENST00000518319.5 |
TMEM67
|
transmembrane protein 67 |
| chr3_+_119186716 | 6.05 |
ENST00000460625.1
|
UPK1B
|
uroplakin 1B |
| chr14_-_106557465 | 6.04 |
ENST00000390625.3
|
IGHV3-49
|
immunoglobulin heavy variable 3-49 |
| chr1_+_158931539 | 6.02 |
ENST00000368140.6
ENST00000368138.7 ENST00000392254.6 ENST00000392252.7 ENST00000368135.4 |
PYHIN1
|
pyrin and HIN domain family member 1 |
| chr8_-_85341659 | 6.01 |
ENST00000522389.5
|
CA1
|
carbonic anhydrase 1 |
| chr4_+_73404255 | 5.93 |
ENST00000621628.4
ENST00000621085.4 ENST00000415165.6 ENST00000295897.9 ENST00000503124.5 ENST00000509063.5 ENST00000401494.7 |
ALB
|
albumin |
| chr1_-_89126066 | 5.87 |
ENST00000370466.4
|
GBP2
|
guanylate binding protein 2 |
| chr8_+_93754879 | 5.86 |
ENST00000453906.6
ENST00000683362.1 ENST00000682036.1 ENST00000453321.8 ENST00000409623.8 ENST00000520680.2 ENST00000521517.6 ENST00000452276.6 |
TMEM67
|
transmembrane protein 67 |
| chr16_+_19285724 | 5.82 |
ENST00000636231.2
ENST00000493231.5 ENST00000465414.1 |
CLEC19A
|
C-type lectin domain containing 19A |
| chr6_-_131951364 | 5.82 |
ENST00000367976.4
|
CCN2
|
cellular communication network factor 2 |
| chr15_-_45378519 | 5.80 |
ENST00000558163.1
ENST00000396659.8 ENST00000675323.1 ENST00000558336.5 |
GATM
|
glycine amidinotransferase |
| chr5_+_141408000 | 5.78 |
ENST00000616430.1
|
PCDHGB6
|
protocadherin gamma subfamily B, 6 |
| chr19_+_36605292 | 5.77 |
ENST00000460670.5
ENST00000292928.7 ENST00000439428.5 |
ZNF382
|
zinc finger protein 382 |
| chr12_+_8843236 | 5.71 |
ENST00000541459.5
|
A2ML1
|
alpha-2-macroglobulin like 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 14.7 | 190.7 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 10.1 | 40.3 | GO:0002399 | MHC class II protein complex assembly(GO:0002399) |
| 9.7 | 29.0 | GO:0071461 | cellular response to redox state(GO:0071461) |
| 9.2 | 46.0 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 7.7 | 23.2 | GO:0030185 | nitric oxide transport(GO:0030185) |
| 7.0 | 84.2 | GO:0006069 | ethanol oxidation(GO:0006069) |
| 5.5 | 22.0 | GO:0007499 | ectoderm and mesoderm interaction(GO:0007499) |
| 5.2 | 10.4 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) |
| 4.4 | 26.2 | GO:0070171 | negative regulation of tooth mineralization(GO:0070171) |
| 4.3 | 12.9 | GO:2000097 | regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 4.1 | 608.2 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 4.0 | 16.1 | GO:0002370 | natural killer cell cytokine production(GO:0002370) regulation of natural killer cell cytokine production(GO:0002727) |
| 4.0 | 27.8 | GO:0060267 | positive regulation of respiratory burst(GO:0060267) |
| 3.8 | 26.9 | GO:0030885 | regulation of myeloid dendritic cell activation(GO:0030885) |
| 3.0 | 5.9 | GO:0019836 | hemolysis by symbiont of host erythrocytes(GO:0019836) hemolysis in other organism(GO:0044179) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 2.9 | 14.7 | GO:0046103 | adenosine catabolic process(GO:0006154) inosine biosynthetic process(GO:0046103) |
| 2.9 | 11.7 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 2.9 | 2.9 | GO:0046110 | xanthine metabolic process(GO:0046110) |
| 2.9 | 17.4 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 2.8 | 8.3 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 2.7 | 8.1 | GO:1990764 | regulation of myofibroblast contraction(GO:1904328) myofibroblast contraction(GO:1990764) |
| 2.6 | 5.2 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 2.6 | 7.7 | GO:1903487 | regulation of lactation(GO:1903487) |
| 2.5 | 7.5 | GO:2000296 | negative regulation of hydrogen peroxide catabolic process(GO:2000296) |
| 2.5 | 7.4 | GO:0034553 | respiratory chain complex II assembly(GO:0034552) mitochondrial respiratory chain complex II assembly(GO:0034553) mitochondrial respiratory chain complex II biogenesis(GO:0097032) |
| 2.4 | 7.3 | GO:1990168 | protein K33-linked deubiquitination(GO:1990168) |
| 2.3 | 7.0 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 2.2 | 6.6 | GO:0002378 | immunoglobulin biosynthetic process(GO:0002378) |
| 2.2 | 6.5 | GO:1904845 | response to L-glutamine(GO:1904844) cellular response to L-glutamine(GO:1904845) |
| 2.1 | 6.4 | GO:0045897 | positive regulation of transcription during mitosis(GO:0045897) |
| 2.1 | 6.4 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 2.1 | 10.6 | GO:0035106 | operant conditioning(GO:0035106) |
| 2.0 | 12.2 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 2.0 | 10.2 | GO:0071727 | toll-like receptor TLR1:TLR2 signaling pathway(GO:0038123) response to triacyl bacterial lipopeptide(GO:0071725) cellular response to triacyl bacterial lipopeptide(GO:0071727) |
| 2.0 | 6.1 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 2.0 | 15.9 | GO:0046541 | saliva secretion(GO:0046541) |
| 1.9 | 5.7 | GO:0071934 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 1.9 | 13.0 | GO:0060054 | positive regulation of epithelial cell proliferation involved in wound healing(GO:0060054) |
| 1.9 | 22.2 | GO:0071285 | cellular response to lithium ion(GO:0071285) |
| 1.8 | 3.6 | GO:0002581 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) |
| 1.8 | 19.6 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 1.7 | 5.1 | GO:0070429 | regulation of toll-like receptor 5 signaling pathway(GO:0034147) negative regulation of toll-like receptor 5 signaling pathway(GO:0034148) negative regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070429) tolerance induction to lipopolysaccharide(GO:0072573) |
| 1.7 | 5.1 | GO:0030573 | bile acid catabolic process(GO:0030573) |
| 1.7 | 23.7 | GO:0034196 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 1.7 | 10.0 | GO:1900004 | negative regulation of serine-type endopeptidase activity(GO:1900004) negative regulation of serine-type peptidase activity(GO:1902572) |
| 1.6 | 4.8 | GO:0060903 | positive regulation of meiosis I(GO:0060903) |
| 1.6 | 273.7 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 1.5 | 7.7 | GO:1904209 | regulation of chemokine (C-C motif) ligand 2 secretion(GO:1904207) positive regulation of chemokine (C-C motif) ligand 2 secretion(GO:1904209) |
| 1.5 | 1.5 | GO:0005989 | lactose metabolic process(GO:0005988) lactose biosynthetic process(GO:0005989) |
| 1.5 | 4.6 | GO:0001978 | regulation of systemic arterial blood pressure by carotid sinus baroreceptor feedback(GO:0001978) baroreceptor response to increased systemic arterial blood pressure(GO:0001983) |
| 1.4 | 5.7 | GO:1904749 | regulation of protein localization to nucleolus(GO:1904749) |
| 1.4 | 5.7 | GO:1902283 | negative regulation of primary amine oxidase activity(GO:1902283) |
| 1.4 | 4.2 | GO:0038178 | complement component C5a signaling pathway(GO:0038178) |
| 1.4 | 4.1 | GO:0014813 | skeletal muscle satellite cell commitment(GO:0014813) |
| 1.4 | 4.1 | GO:0002361 | CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0002361) |
| 1.4 | 1.4 | GO:0032661 | regulation of interleukin-18 production(GO:0032661) |
| 1.4 | 4.1 | GO:1990918 | meiotic DNA double-strand break processing(GO:0000706) double-strand break repair involved in meiotic recombination(GO:1990918) |
| 1.3 | 7.9 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 1.3 | 3.9 | GO:2001027 | negative regulation of endothelial cell chemotaxis(GO:2001027) |
| 1.3 | 5.1 | GO:0003366 | cell-matrix adhesion involved in ameboidal cell migration(GO:0003366) |
| 1.3 | 7.6 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 1.2 | 7.5 | GO:0033133 | positive regulation of glucokinase activity(GO:0033133) |
| 1.2 | 3.7 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 1.2 | 4.9 | GO:0046351 | sucrose biosynthetic process(GO:0005986) disaccharide biosynthetic process(GO:0046351) |
| 1.2 | 4.9 | GO:0002826 | negative regulation of T-helper 1 type immune response(GO:0002826) |
| 1.2 | 4.8 | GO:0050916 | sensory perception of sweet taste(GO:0050916) sensory perception of umami taste(GO:0050917) |
| 1.2 | 8.4 | GO:0016098 | monoterpenoid metabolic process(GO:0016098) |
| 1.2 | 14.3 | GO:0006824 | cobalt ion transport(GO:0006824) |
| 1.2 | 3.6 | GO:0019732 | antifungal humoral response(GO:0019732) |
| 1.1 | 2.3 | GO:0070309 | lens fiber cell morphogenesis(GO:0070309) |
| 1.1 | 3.3 | GO:1903348 | positive regulation of bicellular tight junction assembly(GO:1903348) |
| 1.1 | 5.6 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 1.1 | 5.5 | GO:1903936 | cellular response to sodium arsenite(GO:1903936) |
| 1.0 | 5.2 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 1.0 | 10.4 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 1.0 | 2.1 | GO:1904049 | negative regulation of spontaneous neurotransmitter secretion(GO:1904049) |
| 1.0 | 11.2 | GO:0010898 | positive regulation of triglyceride catabolic process(GO:0010898) |
| 1.0 | 7.0 | GO:0007161 | calcium-independent cell-matrix adhesion(GO:0007161) |
| 1.0 | 4.0 | GO:1902159 | regulation of cyclic nucleotide-gated ion channel activity(GO:1902159) |
| 1.0 | 3.0 | GO:0019878 | lysine biosynthetic process(GO:0009085) lysine biosynthetic process via aminoadipic acid(GO:0019878) L-lysine catabolic process to acetyl-CoA via saccharopine(GO:0033512) |
| 1.0 | 5.9 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 1.0 | 4.8 | GO:0032074 | negative regulation of nuclease activity(GO:0032074) |
| 1.0 | 3.8 | GO:0045356 | positive regulation of interferon-alpha biosynthetic process(GO:0045356) |
| 0.9 | 1.9 | GO:0097325 | melanocyte proliferation(GO:0097325) |
| 0.9 | 2.8 | GO:0061011 | hepatic duct development(GO:0061011) |
| 0.9 | 2.8 | GO:0002304 | gamma-delta intraepithelial T cell differentiation(GO:0002304) CD8-positive, gamma-delta intraepithelial T cell differentiation(GO:0002305) |
| 0.9 | 10.2 | GO:0002767 | immune response-inhibiting signal transduction(GO:0002765) immune response-inhibiting cell surface receptor signaling pathway(GO:0002767) |
| 0.9 | 3.7 | GO:1900228 | biofilm formation(GO:0042710) single-species biofilm formation(GO:0044010) single-species biofilm formation in or on host organism(GO:0044407) regulation of single-species biofilm formation(GO:1900190) negative regulation of single-species biofilm formation(GO:1900191) regulation of single-species biofilm formation in or on host organism(GO:1900228) negative regulation of single-species biofilm formation in or on host organism(GO:1900229) |
| 0.9 | 5.3 | GO:0009753 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 0.9 | 3.4 | GO:1902023 | L-arginine import(GO:0043091) arginine import(GO:0090467) L-arginine transport(GO:1902023) |
| 0.8 | 5.1 | GO:0019441 | tryptophan catabolic process to kynurenine(GO:0019441) |
| 0.8 | 14.7 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.8 | 7.3 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.8 | 3.2 | GO:0032848 | negative regulation of cellular pH reduction(GO:0032848) CD8-positive, alpha-beta T cell lineage commitment(GO:0043375) negative regulation of retinal cell programmed cell death(GO:0046671) |
| 0.8 | 4.7 | GO:0035723 | interleukin-15-mediated signaling pathway(GO:0035723) cellular response to interleukin-15(GO:0071350) |
| 0.8 | 8.4 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.8 | 5.3 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.8 | 15.8 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.7 | 2.9 | GO:0000451 | rRNA 2'-O-methylation(GO:0000451) |
| 0.7 | 6.5 | GO:0042904 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.7 | 2.9 | GO:0030806 | negative regulation of cyclic nucleotide catabolic process(GO:0030806) negative regulation of cAMP catabolic process(GO:0030821) |
| 0.7 | 10.8 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.7 | 48.0 | GO:0032729 | positive regulation of interferon-gamma production(GO:0032729) |
| 0.7 | 2.1 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.7 | 2.8 | GO:0031017 | response to chlorate(GO:0010157) exocrine pancreas development(GO:0031017) |
| 0.7 | 2.8 | GO:0003430 | growth plate cartilage chondrocyte growth(GO:0003430) growth plate cartilage chondrocyte development(GO:0003431) |
| 0.7 | 3.5 | GO:1904075 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.7 | 4.2 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.7 | 4.8 | GO:2000773 | negative regulation of cellular senescence(GO:2000773) |
| 0.7 | 3.5 | GO:0001555 | oocyte growth(GO:0001555) regulation of progesterone secretion(GO:2000870) |
| 0.7 | 2.1 | GO:0007341 | penetration of zona pellucida(GO:0007341) |
| 0.7 | 2.0 | GO:1902310 | positive regulation of peptidyl-serine dephosphorylation(GO:1902310) |
| 0.7 | 4.1 | GO:0003350 | pulmonary myocardium development(GO:0003350) |
| 0.7 | 3.4 | GO:0002725 | negative regulation of T cell cytokine production(GO:0002725) |
| 0.6 | 3.2 | GO:0032049 | cardiolipin biosynthetic process(GO:0032049) |
| 0.6 | 2.6 | GO:0002933 | lipid hydroxylation(GO:0002933) |
| 0.6 | 18.1 | GO:0098743 | cell aggregation(GO:0098743) |
| 0.6 | 0.6 | GO:0046668 | regulation of retinal cell programmed cell death(GO:0046668) |
| 0.6 | 2.4 | GO:0003330 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.6 | 3.0 | GO:0090309 | regulation of methylation-dependent chromatin silencing(GO:0090308) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.6 | 1.8 | GO:0033076 | isoquinoline alkaloid metabolic process(GO:0033076) serotonin biosynthetic process(GO:0042427) phytoalexin metabolic process(GO:0052314) |
| 0.6 | 58.8 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.6 | 3.5 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.6 | 7.5 | GO:1900017 | positive regulation of cytokine production involved in inflammatory response(GO:1900017) |
| 0.6 | 2.3 | GO:0006931 | substrate-dependent cell migration, cell attachment to substrate(GO:0006931) |
| 0.6 | 2.8 | GO:0046167 | glycerol-3-phosphate biosynthetic process(GO:0046167) |
| 0.6 | 2.2 | GO:0036369 | transcription factor catabolic process(GO:0036369) |
| 0.5 | 14.3 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.5 | 3.8 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.5 | 8.1 | GO:0030889 | negative regulation of B cell proliferation(GO:0030889) |
| 0.5 | 4.3 | GO:0070170 | regulation of tooth mineralization(GO:0070170) |
| 0.5 | 3.2 | GO:0019530 | taurine metabolic process(GO:0019530) |
| 0.5 | 11.0 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.5 | 3.1 | GO:1904306 | regulation of gastro-intestinal system smooth muscle contraction(GO:1904304) positive regulation of gastro-intestinal system smooth muscle contraction(GO:1904306) |
| 0.5 | 3.6 | GO:0043615 | astrocyte cell migration(GO:0043615) |
| 0.5 | 7.7 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.5 | 1.5 | GO:0015842 | aminergic neurotransmitter loading into synaptic vesicle(GO:0015842) |
| 0.5 | 5.5 | GO:0001865 | NK T cell differentiation(GO:0001865) |
| 0.5 | 3.0 | GO:0048050 | post-embryonic eye morphogenesis(GO:0048050) |
| 0.5 | 9.3 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.5 | 1.0 | GO:0036518 | chemorepulsion of dopaminergic neuron axon(GO:0036518) chemorepulsion of axon(GO:0061643) |
| 0.5 | 7.7 | GO:0006702 | androgen biosynthetic process(GO:0006702) |
| 0.5 | 3.3 | GO:0010819 | regulation of T cell chemotaxis(GO:0010819) |
| 0.5 | 3.3 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.5 | 8.0 | GO:0042573 | retinoic acid metabolic process(GO:0042573) |
| 0.5 | 1.4 | GO:0046603 | negative regulation of mitotic centrosome separation(GO:0046603) |
| 0.5 | 9.2 | GO:0097296 | activation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097296) |
| 0.5 | 12.8 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.4 | 2.7 | GO:2000483 | negative regulation of RNA polymerase II regulatory region sequence-specific DNA binding(GO:1903026) negative regulation of interleukin-8 secretion(GO:2000483) |
| 0.4 | 7.1 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.4 | 2.2 | GO:0018352 | protein-pyridoxal-5-phosphate linkage(GO:0018352) |
| 0.4 | 3.5 | GO:0015747 | urate transport(GO:0015747) |
| 0.4 | 4.7 | GO:0051791 | medium-chain fatty acid metabolic process(GO:0051791) |
| 0.4 | 0.4 | GO:0038162 | erythropoietin-mediated signaling pathway(GO:0038162) |
| 0.4 | 3.8 | GO:0006030 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.4 | 1.2 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.4 | 4.5 | GO:0010623 | programmed cell death involved in cell development(GO:0010623) |
| 0.4 | 2.8 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.4 | 4.3 | GO:0001675 | acrosome assembly(GO:0001675) |
| 0.4 | 4.2 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.4 | 3.0 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.4 | 23.4 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.4 | 2.6 | GO:0039663 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.4 | 12.2 | GO:0071425 | hematopoietic stem cell proliferation(GO:0071425) |
| 0.4 | 5.1 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.4 | 13.7 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.4 | 1.1 | GO:0006711 | estrogen catabolic process(GO:0006711) |
| 0.3 | 4.5 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.3 | 4.2 | GO:0034465 | response to carbon monoxide(GO:0034465) |
| 0.3 | 2.1 | GO:1904153 | negative regulation of protein exit from endoplasmic reticulum(GO:0070862) negative regulation of retrograde protein transport, ER to cytosol(GO:1904153) |
| 0.3 | 2.4 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 0.3 | 5.4 | GO:0031998 | regulation of fatty acid beta-oxidation(GO:0031998) |
| 0.3 | 1.0 | GO:0016256 | N-glycan processing to lysosome(GO:0016256) |
| 0.3 | 3.0 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.3 | 6.1 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.3 | 1.3 | GO:0042574 | retinal metabolic process(GO:0042574) |
| 0.3 | 1.3 | GO:0044026 | DNA hypermethylation(GO:0044026) |
| 0.3 | 9.1 | GO:0034035 | purine ribonucleoside bisphosphate metabolic process(GO:0034035) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) |
| 0.3 | 2.2 | GO:1902856 | negative regulation of nonmotile primary cilium assembly(GO:1902856) |
| 0.3 | 1.6 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.3 | 17.2 | GO:0045576 | mast cell activation(GO:0045576) |
| 0.3 | 1.5 | GO:0030070 | insulin processing(GO:0030070) |
| 0.3 | 1.8 | GO:0038016 | insulin receptor internalization(GO:0038016) |
| 0.3 | 14.6 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.3 | 2.3 | GO:0061365 | positive regulation of triglyceride lipase activity(GO:0061365) |
| 0.3 | 1.5 | GO:0035990 | tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) |
| 0.3 | 2.0 | GO:0010961 | cellular magnesium ion homeostasis(GO:0010961) |
| 0.3 | 3.1 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.3 | 2.5 | GO:0030207 | chondroitin sulfate catabolic process(GO:0030207) |
| 0.3 | 1.7 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.3 | 4.9 | GO:0010826 | negative regulation of centrosome duplication(GO:0010826) |
| 0.3 | 2.4 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.3 | 76.9 | GO:0002250 | adaptive immune response(GO:0002250) |
| 0.3 | 0.3 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 0.3 | 8.3 | GO:0022400 | regulation of rhodopsin mediated signaling pathway(GO:0022400) |
| 0.3 | 1.5 | GO:0071865 | cell proliferation in bone marrow(GO:0071838) regulation of cell proliferation in bone marrow(GO:0071863) positive regulation of cell proliferation in bone marrow(GO:0071864) regulation of apoptotic process in bone marrow(GO:0071865) negative regulation of apoptotic process in bone marrow(GO:0071866) |
| 0.2 | 0.5 | GO:1902080 | regulation of calcium ion import into sarcoplasmic reticulum(GO:1902080) negative regulation of calcium ion import into sarcoplasmic reticulum(GO:1902081) |
| 0.2 | 0.5 | GO:1902358 | sulfate transmembrane transport(GO:1902358) |
| 0.2 | 8.4 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.2 | 4.6 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.2 | 16.0 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.2 | 6.7 | GO:0007398 | ectoderm development(GO:0007398) |
| 0.2 | 3.1 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.2 | 4.0 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.2 | 0.9 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 0.2 | 1.4 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.2 | 1.1 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.2 | 2.7 | GO:0036500 | ATF6-mediated unfolded protein response(GO:0036500) |
| 0.2 | 1.8 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.2 | 0.7 | GO:1900738 | positive regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900738) |
| 0.2 | 0.7 | GO:2000661 | positive regulation of interleukin-1-mediated signaling pathway(GO:2000661) |
| 0.2 | 0.9 | GO:0060737 | prostate epithelial cord elongation(GO:0060523) prostate gland morphogenetic growth(GO:0060737) |
| 0.2 | 3.1 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.2 | 7.1 | GO:0050829 | defense response to Gram-negative bacterium(GO:0050829) |
| 0.2 | 1.8 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.2 | 0.6 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.2 | 5.1 | GO:0006825 | copper ion transport(GO:0006825) |
| 0.2 | 2.7 | GO:0035878 | nail development(GO:0035878) |
| 0.2 | 6.0 | GO:0044364 | killing of cells of other organism(GO:0031640) disruption of cells of other organism(GO:0044364) |
| 0.2 | 1.6 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.2 | 1.6 | GO:0010649 | regulation of cell communication by electrical coupling(GO:0010649) |
| 0.2 | 1.6 | GO:1901475 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.2 | 4.5 | GO:0046339 | diacylglycerol metabolic process(GO:0046339) |
| 0.2 | 10.4 | GO:0001895 | retina homeostasis(GO:0001895) |
| 0.2 | 0.4 | GO:0071316 | cellular response to nicotine(GO:0071316) positive regulation of miRNA metabolic process(GO:2000630) |
| 0.2 | 2.1 | GO:0034505 | tooth mineralization(GO:0034505) |
| 0.2 | 14.1 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.2 | 2.6 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.2 | 1.7 | GO:0010886 | positive regulation of cholesterol storage(GO:0010886) |
| 0.2 | 1.9 | GO:0098703 | calcium ion import across plasma membrane(GO:0098703) calcium ion import into cell(GO:1990035) |
| 0.2 | 1.9 | GO:0042737 | drug catabolic process(GO:0042737) |
| 0.2 | 13.8 | GO:0016525 | negative regulation of angiogenesis(GO:0016525) negative regulation of blood vessel morphogenesis(GO:2000181) |
| 0.2 | 0.7 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.2 | 1.4 | GO:0006682 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.2 | 2.3 | GO:0030210 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.2 | 3.5 | GO:0097151 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.2 | 1.1 | GO:0032754 | positive regulation of interleukin-5 production(GO:0032754) |
| 0.2 | 1.2 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.2 | 3.3 | GO:0030449 | regulation of complement activation(GO:0030449) |
| 0.2 | 0.5 | GO:0061026 | central nervous system morphogenesis(GO:0021551) cardiac muscle tissue regeneration(GO:0061026) |
| 0.2 | 2.9 | GO:0050860 | negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.2 | 6.1 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.2 | 0.3 | GO:0007500 | mesodermal cell fate determination(GO:0007500) |
| 0.2 | 2.3 | GO:0045793 | positive regulation of cell size(GO:0045793) |
| 0.1 | 2.2 | GO:1900116 | extracellular regulation of signal transduction(GO:1900115) extracellular negative regulation of signal transduction(GO:1900116) |
| 0.1 | 3.8 | GO:0035456 | response to interferon-beta(GO:0035456) |
| 0.1 | 0.4 | GO:1904354 | negative regulation of telomere capping(GO:1904354) |
| 0.1 | 2.2 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.1 | 2.6 | GO:0050651 | dermatan sulfate proteoglycan biosynthetic process(GO:0050651) |
| 0.1 | 1.7 | GO:0015871 | choline transport(GO:0015871) |
| 0.1 | 1.0 | GO:0046449 | creatinine metabolic process(GO:0046449) |
| 0.1 | 0.8 | GO:0045725 | positive regulation of glycogen biosynthetic process(GO:0045725) |
| 0.1 | 7.2 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.1 | 6.8 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.1 | 3.7 | GO:0010574 | regulation of vascular endothelial growth factor production(GO:0010574) |
| 0.1 | 7.4 | GO:0006959 | humoral immune response(GO:0006959) |
| 0.1 | 1.3 | GO:0034374 | low-density lipoprotein particle remodeling(GO:0034374) |
| 0.1 | 3.8 | GO:0032755 | positive regulation of interleukin-6 production(GO:0032755) |
| 0.1 | 0.4 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.1 | 0.5 | GO:0032733 | positive regulation of interleukin-10 production(GO:0032733) |
| 0.1 | 1.7 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.1 | 1.9 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.1 | 4.0 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.1 | 11.4 | GO:0070268 | cornification(GO:0070268) |
| 0.1 | 7.9 | GO:0030049 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.1 | 1.4 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 4.0 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.1 | 0.8 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.1 | 1.0 | GO:0043691 | reverse cholesterol transport(GO:0043691) |
| 0.1 | 0.5 | GO:0060574 | intestinal epithelial cell maturation(GO:0060574) |
| 0.1 | 1.1 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.1 | 0.2 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.1 | 2.9 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.1 | 1.2 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 0.1 | 1.0 | GO:0014010 | regulation of Schwann cell proliferation(GO:0010624) negative regulation of Schwann cell proliferation(GO:0010626) Schwann cell proliferation(GO:0014010) |
| 0.1 | 0.9 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.1 | 14.3 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 1.4 | GO:2001214 | positive regulation of vasculogenesis(GO:2001214) |
| 0.1 | 0.4 | GO:1903385 | regulation of homophilic cell adhesion(GO:1903385) |
| 0.1 | 1.2 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.1 | 1.7 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.1 | 2.4 | GO:0042102 | positive regulation of T cell proliferation(GO:0042102) |
| 0.1 | 0.4 | GO:0090118 | receptor-mediated endocytosis of low-density lipoprotein particle involved in cholesterol transport(GO:0090118) |
| 0.1 | 0.2 | GO:1905246 | regulation of choline O-acetyltransferase activity(GO:1902769) positive regulation of choline O-acetyltransferase activity(GO:1902771) negative regulation of tau-protein kinase activity(GO:1902948) positive regulation of early endosome to recycling endosome transport(GO:1902955) negative regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902960) regulation of neurofibrillary tangle assembly(GO:1902996) negative regulation of neurofibrillary tangle assembly(GO:1902997) negative regulation of aspartic-type peptidase activity(GO:1905246) |
| 0.1 | 0.4 | GO:0016127 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.1 | 1.1 | GO:0048011 | neurotrophin TRK receptor signaling pathway(GO:0048011) |
| 0.1 | 0.5 | GO:0042448 | progesterone metabolic process(GO:0042448) |
| 0.1 | 0.2 | GO:1904823 | pyrimidine nucleobase transport(GO:0015855) purine nucleoside transmembrane transport(GO:0015860) purine nucleobase transmembrane transport(GO:1904823) |
| 0.1 | 0.2 | GO:0035482 | gastric motility(GO:0035482) gastric emptying(GO:0035483) |
| 0.1 | 5.2 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.1 | 1.0 | GO:0043651 | linoleic acid metabolic process(GO:0043651) |
| 0.1 | 2.4 | GO:0035338 | long-chain fatty-acyl-CoA biosynthetic process(GO:0035338) |
| 0.1 | 0.7 | GO:0007250 | activation of NF-kappaB-inducing kinase activity(GO:0007250) |
| 0.1 | 3.2 | GO:0045739 | positive regulation of DNA repair(GO:0045739) |
| 0.1 | 1.0 | GO:0071353 | cellular response to interleukin-4(GO:0071353) |
| 0.1 | 4.3 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.1 | 0.8 | GO:0048240 | sperm capacitation(GO:0048240) |
| 0.0 | 2.3 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.0 | 0.6 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 0.0 | 0.2 | GO:0071839 | apoptotic process in bone marrow(GO:0071839) |
| 0.0 | 1.7 | GO:0046710 | GDP metabolic process(GO:0046710) |
| 0.0 | 4.9 | GO:0016575 | histone deacetylation(GO:0016575) |
| 0.0 | 2.7 | GO:0030282 | bone mineralization(GO:0030282) |
| 0.0 | 1.8 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.0 | 0.4 | GO:0060391 | positive regulation of SMAD protein import into nucleus(GO:0060391) |
| 0.0 | 0.1 | GO:0018874 | benzoate metabolic process(GO:0018874) butyrate metabolic process(GO:0019605) |
| 0.0 | 0.2 | GO:0006015 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 0.0 | 1.1 | GO:0016601 | Rac protein signal transduction(GO:0016601) |
| 0.0 | 0.6 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 0.6 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.0 | 0.8 | GO:0030220 | platelet formation(GO:0030220) |
| 0.0 | 3.0 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 0.9 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.0 | 1.1 | GO:0001938 | positive regulation of endothelial cell proliferation(GO:0001938) |
| 0.0 | 0.7 | GO:0014072 | response to isoquinoline alkaloid(GO:0014072) response to morphine(GO:0043278) |
| 0.0 | 0.5 | GO:0000729 | DNA double-strand break processing(GO:0000729) |
| 0.0 | 0.6 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.0 | 1.1 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.0 | 0.6 | GO:0002449 | lymphocyte mediated immunity(GO:0002449) |
| 0.0 | 2.6 | GO:0002576 | platelet degranulation(GO:0002576) |
| 0.0 | 0.3 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 0.2 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.0 | 0.5 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.0 | 1.1 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.0 | 0.1 | GO:0003010 | voluntary skeletal muscle contraction(GO:0003010) twitch skeletal muscle contraction(GO:0014721) fast-twitch skeletal muscle fiber contraction(GO:0031443) |
| 0.0 | 0.8 | GO:0060349 | bone morphogenesis(GO:0060349) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 14.7 | 190.7 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 14.0 | 41.9 | GO:0071756 | IgM immunoglobulin complex(GO:0071753) IgM immunoglobulin complex, circulating(GO:0071754) pentameric IgM immunoglobulin complex(GO:0071756) |
| 7.7 | 30.6 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 5.6 | 105.6 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 4.8 | 237.8 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 2.2 | 6.5 | GO:0097679 | other organism cytoplasm(GO:0097679) |
| 1.9 | 7.6 | GO:0030895 | apolipoprotein B mRNA editing enzyme complex(GO:0030895) |
| 1.6 | 6.4 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 1.5 | 7.4 | GO:0005826 | actomyosin contractile ring(GO:0005826) |
| 1.3 | 6.7 | GO:0005579 | membrane attack complex(GO:0005579) |
| 1.3 | 17.4 | GO:0098643 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 1.3 | 5.1 | GO:0034669 | integrin alpha4-beta7 complex(GO:0034669) |
| 1.3 | 6.3 | GO:0036398 | TCR signalosome(GO:0036398) |
| 1.2 | 4.8 | GO:0071546 | pi-body(GO:0071546) |
| 1.2 | 210.7 | GO:0072562 | blood microparticle(GO:0072562) |
| 1.2 | 3.5 | GO:0033150 | perinuclear theca(GO:0033011) cytoskeletal calyx(GO:0033150) |
| 1.1 | 9.7 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 1.1 | 20.4 | GO:0036038 | MKS complex(GO:0036038) |
| 1.1 | 10.5 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 1.0 | 3.0 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 0.9 | 6.5 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.9 | 4.7 | GO:0032807 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) DNA ligase IV complex(GO:0032807) |
| 0.8 | 2.4 | GO:0032437 | cuticular plate(GO:0032437) |
| 0.8 | 4.9 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.8 | 15.1 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.8 | 13.5 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.8 | 8.3 | GO:0098651 | basement membrane collagen trimer(GO:0098651) |
| 0.7 | 5.2 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.7 | 5.1 | GO:0030893 | meiotic cohesin complex(GO:0030893) |
| 0.7 | 7.2 | GO:0030478 | actin cap(GO:0030478) |
| 0.7 | 26.9 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.6 | 1.9 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.6 | 9.1 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.5 | 1.6 | GO:0000805 | X chromosome(GO:0000805) |
| 0.5 | 4.1 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.5 | 4.0 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.5 | 41.9 | GO:0035580 | specific granule lumen(GO:0035580) |
| 0.5 | 2.7 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.4 | 3.6 | GO:0097425 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.4 | 3.1 | GO:0043196 | varicosity(GO:0043196) |
| 0.4 | 3.9 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.4 | 3.5 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.4 | 6.5 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.4 | 14.3 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.4 | 3.0 | GO:0097209 | epidermal lamellar body(GO:0097209) |
| 0.4 | 4.2 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.4 | 2.1 | GO:0000836 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.4 | 2.1 | GO:1990761 | growth cone lamellipodium(GO:1990761) |
| 0.4 | 33.9 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.4 | 4.3 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.4 | 2.7 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.4 | 26.7 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.4 | 2.2 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.4 | 7.3 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.3 | 521.0 | GO:0005615 | extracellular space(GO:0005615) |
| 0.3 | 3.1 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.3 | 2.3 | GO:0060091 | kinocilium(GO:0060091) |
| 0.3 | 13.3 | GO:0001917 | photoreceptor inner segment(GO:0001917) |
| 0.3 | 8.1 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.3 | 4.2 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.3 | 0.8 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.3 | 1.4 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.3 | 3.3 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.3 | 49.6 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.3 | 6.1 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.3 | 10.4 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.2 | 3.2 | GO:0046930 | pore complex(GO:0046930) |
| 0.2 | 0.9 | GO:0042272 | nuclear RNA export factor complex(GO:0042272) |
| 0.2 | 3.5 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.2 | 4.6 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.2 | 3.0 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.2 | 2.0 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.2 | 1.6 | GO:0070449 | elongin complex(GO:0070449) |
| 0.2 | 2.3 | GO:0031462 | Cul2-RING ubiquitin ligase complex(GO:0031462) |
| 0.2 | 4.0 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.2 | 6.8 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.2 | 2.1 | GO:0042555 | MCM complex(GO:0042555) |
| 0.2 | 3.3 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.2 | 0.5 | GO:0035354 | Toll-like receptor 1-Toll-like receptor 2 protein complex(GO:0035354) |
| 0.1 | 0.4 | GO:0000333 | telomerase catalytic core complex(GO:0000333) |
| 0.1 | 1.7 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.1 | 11.6 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 0.5 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.1 | 5.5 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.1 | 4.2 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.1 | 1.5 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.1 | 0.5 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 0.1 | 9.6 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 6.0 | GO:0005844 | polysome(GO:0005844) |
| 0.1 | 1.1 | GO:0032797 | SMN complex(GO:0032797) |
| 0.1 | 0.7 | GO:0042611 | MHC protein complex(GO:0042611) |
| 0.1 | 0.5 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.1 | 2.7 | GO:0045095 | keratin filament(GO:0045095) |
| 0.1 | 10.5 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.1 | 2.9 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.1 | 0.7 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 2.2 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.1 | 1.1 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.1 | 0.5 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.1 | 1.6 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.0 | 2.2 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 3.8 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.0 | 0.8 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 2.4 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 0.3 | GO:0045179 | apical cortex(GO:0045179) |
| 0.0 | 0.3 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.0 | 0.2 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.0 | 0.3 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.0 | 1.3 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.3 | GO:0097228 | sperm principal piece(GO:0097228) |
| 0.0 | 0.7 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.1 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.5 | 84.2 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) |
| 7.7 | 46.0 | GO:0003796 | lysozyme activity(GO:0003796) |
| 6.1 | 18.2 | GO:0046848 | hydroxyapatite binding(GO:0046848) |
| 5.4 | 27.1 | GO:0019862 | IgA binding(GO:0019862) |
| 4.4 | 30.6 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 4.2 | 988.4 | GO:0003823 | antigen binding(GO:0003823) |
| 3.7 | 18.5 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 3.7 | 11.0 | GO:0035375 | zymogen binding(GO:0035375) |
| 3.6 | 29.0 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 3.5 | 10.6 | GO:0016497 | substance K receptor activity(GO:0016497) |
| 2.9 | 2.9 | GO:0004031 | aldehyde oxidase activity(GO:0004031) |
| 2.4 | 9.7 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 2.4 | 7.3 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 2.3 | 9.3 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 2.2 | 184.1 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 2.1 | 6.4 | GO:0070052 | collagen V binding(GO:0070052) |
| 2.1 | 6.3 | GO:0017129 | triglyceride binding(GO:0017129) |
| 1.9 | 5.7 | GO:0015234 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) |
| 1.8 | 12.9 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 1.7 | 8.5 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 1.7 | 22.0 | GO:0097371 | MDM2/MDM4 family protein binding(GO:0097371) |
| 1.6 | 13.2 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 1.6 | 14.7 | GO:0031685 | adenosine receptor binding(GO:0031685) |
| 1.6 | 4.9 | GO:0002113 | interleukin-33 binding(GO:0002113) |
| 1.6 | 1.6 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 1.6 | 16.1 | GO:0004064 | arylesterase activity(GO:0004064) |
| 1.6 | 17.4 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 1.6 | 4.7 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 1.5 | 4.5 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 1.5 | 4.4 | GO:0004556 | alpha-amylase activity(GO:0004556) |
| 1.4 | 7.2 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 1.4 | 5.7 | GO:0052595 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 1.4 | 14.3 | GO:0005549 | odorant binding(GO:0005549) |
| 1.4 | 7.0 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 1.4 | 4.2 | GO:0050528 | acyloxyacyl hydrolase activity(GO:0050528) |
| 1.4 | 5.5 | GO:0061752 | telomeric repeat-containing RNA binding(GO:0061752) |
| 1.4 | 4.2 | GO:0004878 | complement component C5a receptor activity(GO:0004878) |
| 1.4 | 4.1 | GO:0003916 | DNA topoisomerase activity(GO:0003916) |
| 1.3 | 11.4 | GO:0004875 | complement receptor activity(GO:0004875) |
| 1.2 | 7.5 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 1.2 | 3.7 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 1.2 | 14.7 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 1.2 | 4.9 | GO:0042132 | fructose 1,6-bisphosphate 1-phosphatase activity(GO:0042132) |
| 1.2 | 16.1 | GO:0031419 | cobalamin binding(GO:0031419) |
| 1.1 | 3.3 | GO:0016422 | mRNA (2'-O-methyladenosine-N6-)-methyltransferase activity(GO:0016422) |
| 1.1 | 3.2 | GO:0052816 | medium-chain acyl-CoA hydrolase activity(GO:0052815) long-chain acyl-CoA hydrolase activity(GO:0052816) |
| 1.0 | 4.2 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 1.0 | 2.9 | GO:0070039 | rRNA (guanosine-2'-O-)-methyltransferase activity(GO:0070039) |
| 0.9 | 4.7 | GO:0004771 | sterol esterase activity(GO:0004771) methylumbelliferyl-acetate deacetylase activity(GO:0047374) |
| 0.9 | 2.7 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.9 | 3.6 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.9 | 6.1 | GO:0050294 | steroid sulfotransferase activity(GO:0050294) |
| 0.8 | 2.4 | GO:0004833 | tryptophan 2,3-dioxygenase activity(GO:0004833) |
| 0.8 | 2.4 | GO:0052795 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.8 | 7.1 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.8 | 6.3 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.8 | 3.1 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.8 | 3.8 | GO:0004905 | type I interferon receptor activity(GO:0004905) |
| 0.8 | 2.3 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.7 | 2.2 | GO:0047751 | 3-oxo-5-alpha-steroid 4-dehydrogenase activity(GO:0003865) steroid dehydrogenase activity, acting on the CH-CH group of donors(GO:0033765) cholestenone 5-alpha-reductase activity(GO:0047751) |
| 0.7 | 3.0 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.7 | 3.7 | GO:0047086 | phenanthrene 9,10-monooxygenase activity(GO:0018636) ketosteroid monooxygenase activity(GO:0047086) trans-1,2-dihydrobenzene-1,2-diol dehydrogenase activity(GO:0047115) |
| 0.7 | 3.7 | GO:0030109 | HLA-B specific inhibitory MHC class I receptor activity(GO:0030109) |
| 0.7 | 13.8 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.7 | 2.6 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 0.7 | 13.7 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.6 | 5.2 | GO:0043394 | proteoglycan binding(GO:0043394) |
| 0.6 | 20.8 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.6 | 3.8 | GO:0003998 | acylphosphatase activity(GO:0003998) |
| 0.6 | 6.2 | GO:0032393 | MHC class I receptor activity(GO:0032393) |
| 0.6 | 5.6 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.6 | 14.6 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.6 | 3.6 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.6 | 28.4 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.6 | 5.2 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.6 | 8.1 | GO:0008517 | folic acid transporter activity(GO:0008517) |
| 0.6 | 2.3 | GO:0031716 | calcitonin receptor binding(GO:0031716) |
| 0.6 | 9.1 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.6 | 4.0 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.6 | 2.8 | GO:0004370 | glycerol kinase activity(GO:0004370) |
| 0.6 | 3.3 | GO:0038052 | RNA polymerase II transcription factor activity, estrogen-activated sequence-specific DNA binding(GO:0038052) |
| 0.6 | 4.5 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.6 | 1.7 | GO:0042806 | fucose binding(GO:0042806) |
| 0.5 | 10.4 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.5 | 1.6 | GO:0017153 | sodium:dicarboxylate symporter activity(GO:0017153) |
| 0.5 | 3.8 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.5 | 8.5 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.5 | 3.7 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.5 | 6.3 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.5 | 1.6 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.5 | 2.0 | GO:0030197 | extracellular matrix constituent, lubricant activity(GO:0030197) |
| 0.5 | 1.5 | GO:0033300 | dehydroascorbic acid transporter activity(GO:0033300) |
| 0.5 | 21.7 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.5 | 44.6 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.5 | 10.1 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.5 | 1.9 | GO:0002046 | opsin binding(GO:0002046) |
| 0.5 | 3.2 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.5 | 4.2 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.5 | 12.8 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.5 | 5.5 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.5 | 6.9 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.5 | 7.8 | GO:0019825 | oxygen binding(GO:0019825) |
| 0.5 | 5.0 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.5 | 2.3 | GO:0035673 | oligopeptide transmembrane transporter activity(GO:0035673) |
| 0.4 | 3.4 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.4 | 5.1 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.4 | 7.3 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.4 | 1.7 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.4 | 9.6 | GO:0031005 | filamin binding(GO:0031005) |
| 0.4 | 4.9 | GO:0005221 | intracellular cyclic nucleotide activated cation channel activity(GO:0005221) cyclic nucleotide-gated ion channel activity(GO:0043855) |
| 0.4 | 1.2 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.4 | 7.5 | GO:0008391 | arachidonic acid monooxygenase activity(GO:0008391) arachidonic acid epoxygenase activity(GO:0008392) |
| 0.4 | 1.6 | GO:0099580 | ion antiporter activity involved in regulation of postsynaptic membrane potential(GO:0099580) |
| 0.4 | 1.2 | GO:0004421 | hydroxymethylglutaryl-CoA synthase activity(GO:0004421) |
| 0.4 | 3.5 | GO:1901702 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.4 | 6.5 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.4 | 3.1 | GO:0016713 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced iron-sulfur protein as one donor, and incorporation of one atom of oxygen(GO:0016713) |
| 0.4 | 6.9 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.4 | 1.5 | GO:0015222 | serotonin transmembrane transporter activity(GO:0015222) |
| 0.4 | 3.0 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.4 | 8.3 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.4 | 3.8 | GO:0004568 | chitinase activity(GO:0004568) chitin binding(GO:0008061) |
| 0.4 | 2.2 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.4 | 4.4 | GO:0070330 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) aromatase activity(GO:0070330) |
| 0.4 | 5.1 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.4 | 4.3 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.4 | 4.3 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.4 | 1.8 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.4 | 2.1 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.3 | 2.1 | GO:0030345 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.3 | 2.0 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.3 | 2.3 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.3 | 6.7 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.3 | 2.7 | GO:0050664 | oxidoreductase activity, acting on NAD(P)H, oxygen as acceptor(GO:0050664) |
| 0.3 | 6.8 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.3 | 1.5 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.3 | 0.9 | GO:0003829 | beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0003829) |
| 0.3 | 7.2 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.3 | 8.6 | GO:0016646 | oxidoreductase activity, acting on the CH-NH group of donors, NAD or NADP as acceptor(GO:0016646) |
| 0.3 | 3.4 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.3 | 3.8 | GO:1902282 | voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1902282) |
| 0.3 | 1.6 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.3 | 5.6 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.3 | 2.5 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.3 | 1.0 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.2 | 6.6 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.2 | 0.7 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.2 | 3.8 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.2 | 4.8 | GO:0003708 | retinoic acid receptor activity(GO:0003708) |
| 0.2 | 29.1 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.2 | 7.2 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.2 | 1.1 | GO:0043141 | ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.2 | 3.4 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.2 | 2.7 | GO:0016645 | oxidoreductase activity, acting on the CH-NH group of donors(GO:0016645) |
| 0.2 | 6.9 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.2 | 5.1 | GO:0008409 | 5'-3' exonuclease activity(GO:0008409) |
| 0.2 | 2.2 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.2 | 3.7 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.2 | 1.7 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.2 | 0.6 | GO:0005260 | channel-conductance-controlling ATPase activity(GO:0005260) |
| 0.2 | 2.4 | GO:0097100 | supercoiled DNA binding(GO:0097100) |
| 0.2 | 15.6 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.2 | 2.2 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.2 | 1.4 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.2 | 1.2 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.2 | 0.6 | GO:0008511 | sodium:potassium:chloride symporter activity(GO:0008511) |
| 0.2 | 5.8 | GO:0016769 | transferase activity, transferring nitrogenous groups(GO:0016769) |
| 0.2 | 1.1 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.2 | 2.8 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.2 | 5.5 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.2 | 0.7 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 0.2 | 1.4 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.2 | 4.8 | GO:0031698 | beta-2 adrenergic receptor binding(GO:0031698) |
| 0.2 | 23.9 | GO:0008201 | heparin binding(GO:0008201) |
| 0.2 | 0.6 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 0.2 | 1.0 | GO:1904929 | coreceptor activity involved in Wnt signaling pathway, planar cell polarity pathway(GO:1904929) |
| 0.2 | 0.9 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.2 | 4.0 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.2 | 0.6 | GO:0004667 | prostaglandin-D synthase activity(GO:0004667) |
| 0.2 | 9.7 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.2 | 2.1 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.2 | 11.4 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.2 | 0.5 | GO:0035663 | Toll-like receptor 2 binding(GO:0035663) |
| 0.2 | 0.6 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.2 | 19.1 | GO:0005179 | hormone activity(GO:0005179) |
| 0.2 | 6.5 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 1.0 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.1 | 3.1 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.1 | 2.3 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 1.2 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 0.1 | 2.7 | GO:0070402 | NADPH binding(GO:0070402) |
| 0.1 | 1.2 | GO:0031545 | peptidyl-proline 4-dioxygenase activity(GO:0031545) |
| 0.1 | 0.9 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.1 | 0.9 | GO:0008271 | secondary active sulfate transmembrane transporter activity(GO:0008271) |
| 0.1 | 3.0 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 6.2 | GO:0032934 | sterol binding(GO:0032934) |
| 0.1 | 1.8 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 3.1 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 0.4 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.1 | 0.3 | GO:0036487 | nitric-oxide synthase inhibitor activity(GO:0036487) |
| 0.1 | 4.3 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.1 | 0.6 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.1 | 1.4 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.1 | 1.9 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.1 | 0.5 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.1 | 4.2 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 2.4 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.1 | 5.2 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.1 | 2.6 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 0.4 | GO:0019763 | immunoglobulin receptor activity(GO:0019763) |
| 0.1 | 10.1 | GO:0019210 | kinase inhibitor activity(GO:0019210) |
| 0.1 | 2.9 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.1 | 1.3 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.1 | 4.1 | GO:0004540 | ribonuclease activity(GO:0004540) |
| 0.1 | 3.3 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.1 | 0.9 | GO:0015037 | peptide disulfide oxidoreductase activity(GO:0015037) |
| 0.1 | 4.1 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.1 | 1.7 | GO:0031491 | nucleosome binding(GO:0031491) |
| 0.1 | 3.6 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.1 | 8.5 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.1 | 0.4 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
| 0.1 | 0.2 | GO:0015389 | pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.1 | 2.9 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.1 | 1.7 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.1 | 6.1 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.1 | 0.3 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.1 | 0.8 | GO:0015926 | glucosidase activity(GO:0015926) |
| 0.1 | 10.7 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 2.8 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.1 | 3.5 | GO:0004520 | endodeoxyribonuclease activity(GO:0004520) |
| 0.0 | 2.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 1.9 | GO:0017046 | peptide hormone binding(GO:0017046) |
| 0.0 | 3.5 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 0.8 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 1.6 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 3.9 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 0.5 | GO:0004716 | receptor signaling protein tyrosine kinase activity(GO:0004716) |
| 0.0 | 1.2 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 1.4 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.2 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 3.6 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 1.7 | GO:0046934 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) |
| 0.0 | 0.2 | GO:0004749 | ribose phosphate diphosphokinase activity(GO:0004749) |
| 0.0 | 1.0 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 1.8 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.7 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.0 | 9.0 | GO:0000987 | core promoter proximal region sequence-specific DNA binding(GO:0000987) |
| 0.0 | 1.6 | GO:0019838 | growth factor binding(GO:0019838) |
| 0.0 | 0.3 | GO:0008252 | nucleotidase activity(GO:0008252) |
| 0.0 | 0.2 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.0 | 0.1 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 1.0 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.0 | 1.2 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.0 | 0.1 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.0 | 0.0 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.1 | 209.7 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 1.1 | 8.4 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.8 | 3.0 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.7 | 31.8 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.6 | 36.7 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.6 | 3.4 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.5 | 9.9 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.5 | 9.4 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.4 | 61.1 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.4 | 3.6 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.4 | 10.2 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.3 | 2.1 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.3 | 1.6 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.3 | 7.7 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.3 | 8.2 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.3 | 4.9 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.3 | 9.3 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.3 | 58.2 | NABA CORE MATRISOME | Ensemble of genes encoding core extracellular matrix including ECM glycoproteins, collagens and proteoglycans |
| 0.3 | 20.0 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.2 | 5.1 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.2 | 3.8 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.2 | 16.7 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.2 | 18.1 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.2 | 8.1 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.2 | 6.8 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.2 | 1.2 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.2 | 2.7 | ST GAQ PATHWAY | G alpha q Pathway |
| 0.2 | 10.1 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.2 | 6.7 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.2 | 7.1 | PID TCR PATHWAY | TCR signaling in naïve CD4+ T cells |
| 0.2 | 7.8 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.2 | 2.8 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.2 | 1.7 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.2 | 7.9 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.2 | 2.2 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.2 | 7.9 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.2 | 3.2 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.2 | 4.4 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.1 | 39.8 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 7.0 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.1 | 23.7 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 4.5 | PID SHP2 PATHWAY | SHP2 signaling |
| 0.1 | 2.3 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.1 | 6.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 2.1 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 4.1 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.1 | 1.9 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
| 0.1 | 1.1 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 1.9 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 2.7 | PID MYC PATHWAY | C-MYC pathway |
| 0.1 | 1.5 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.1 | 2.9 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.1 | 2.8 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 0.8 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 1.5 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 1.2 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 3.2 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 1.0 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.8 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.0 | 2.3 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.9 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 12.8 | NABA MATRISOME | Ensemble of genes encoding extracellular matrix and extracellular matrix-associated proteins |
| 0.0 | 0.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 0.9 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 0.4 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.0 | 0.7 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 0.3 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.0 | 0.2 | PID BCR 5PATHWAY | BCR signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.4 | 84.2 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 7.1 | 190.7 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 4.5 | 94.3 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 1.6 | 6.6 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 1.5 | 18.6 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 1.3 | 60.9 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 1.0 | 16.1 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.7 | 16.8 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.7 | 5.1 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.7 | 15.2 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.6 | 9.0 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.6 | 55.6 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.6 | 8.6 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.6 | 9.6 | REACTOME TRAF6 MEDIATED INDUCTION OF TAK1 COMPLEX | Genes involved in TRAF6 mediated induction of TAK1 complex |
| 0.5 | 12.2 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.5 | 7.4 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.5 | 13.2 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.5 | 2.6 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 0.5 | 38.2 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.5 | 8.9 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.4 | 7.9 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.4 | 39.4 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.4 | 7.1 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.4 | 8.3 | REACTOME DEFENSINS | Genes involved in Defensins |
| 0.4 | 4.6 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.4 | 14.4 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.3 | 6.2 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.3 | 7.4 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.3 | 2.3 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.3 | 7.2 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.3 | 1.3 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS | Genes involved in Synthesis of bile acids and bile salts |
| 0.3 | 3.5 | REACTOME TCR SIGNALING | Genes involved in TCR signaling |
| 0.3 | 18.7 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.3 | 5.0 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.3 | 6.3 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.3 | 31.0 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.3 | 5.0 | REACTOME SIGNALING BY PDGF | Genes involved in Signaling by PDGF |
| 0.3 | 1.9 | REACTOME DOWNSTREAM SIGNALING EVENTS OF B CELL RECEPTOR BCR | Genes involved in Downstream Signaling Events Of B Cell Receptor (BCR) |
| 0.3 | 4.1 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.3 | 8.7 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.3 | 5.1 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.3 | 34.9 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.3 | 4.9 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.2 | 2.5 | REACTOME SIGNALING BY CONSTITUTIVELY ACTIVE EGFR | Genes involved in Signaling by constitutively active EGFR |
| 0.2 | 3.7 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.2 | 6.7 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.2 | 4.4 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.2 | 2.2 | REACTOME OPSINS | Genes involved in Opsins |
| 0.2 | 2.3 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.2 | 2.7 | REACTOME ACTIVATION OF CHAPERONES BY ATF6 ALPHA | Genes involved in Activation of Chaperones by ATF6-alpha |
| 0.2 | 3.4 | REACTOME TRAF3 DEPENDENT IRF ACTIVATION PATHWAY | Genes involved in TRAF3-dependent IRF activation pathway |
| 0.2 | 2.3 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.2 | 3.0 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.2 | 6.6 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.2 | 29.5 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.2 | 6.7 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.2 | 7.4 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 2.9 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.1 | 2.8 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.1 | 1.7 | REACTOME LIPOPROTEIN METABOLISM | Genes involved in Lipoprotein metabolism |
| 0.1 | 2.8 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 3.8 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 4.5 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 13.7 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.1 | 1.9 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.1 | 18.3 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.1 | 7.9 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 2.3 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.1 | 10.8 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.1 | 1.2 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.1 | 1.9 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.1 | 3.1 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.1 | 1.3 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.1 | 4.5 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 2.3 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.1 | 1.6 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 1.4 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.1 | 2.9 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.1 | 3.1 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.1 | 1.6 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.1 | 2.4 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.1 | 1.8 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.1 | 2.9 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.1 | 2.1 | REACTOME MRNA 3 END PROCESSING | Genes involved in mRNA 3'-end processing |
| 0.1 | 1.5 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.1 | 1.2 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.1 | 1.5 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 1.5 | REACTOME CELL CELL JUNCTION ORGANIZATION | Genes involved in Cell-cell junction organization |
| 0.1 | 6.5 | REACTOME MUSCLE CONTRACTION | Genes involved in Muscle contraction |
| 0.1 | 2.2 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.1 | 1.3 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 1.7 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.1 | 1.3 | REACTOME FGFR2C LIGAND BINDING AND ACTIVATION | Genes involved in FGFR2c ligand binding and activation |
| 0.1 | 2.2 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.0 | 5.0 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 4.3 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.9 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.7 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.0 | 2.3 | REACTOME MEIOSIS | Genes involved in Meiosis |
| 0.0 | 0.8 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 1.1 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.0 | 1.8 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 2.2 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.2 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 0.4 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 0.6 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |