Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
Results for HMX3
Z-value: 0.86
Transcription factors associated with HMX3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HMX3
|
ENSG00000188620.11 | HMX3 |
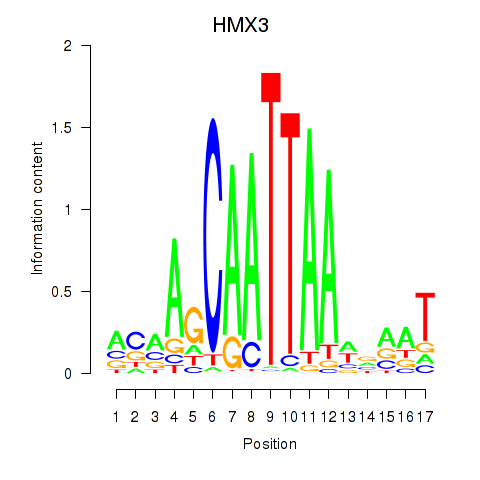
Activity profile of HMX3 motif
Sorted Z-values of HMX3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HMX3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.5 | 22.4 | GO:0006982 | response to lipid hydroperoxide(GO:0006982) |
| 6.6 | 19.9 | GO:0001869 | regulation of complement activation, lectin pathway(GO:0001868) negative regulation of complement activation, lectin pathway(GO:0001869) |
| 2.7 | 35.6 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 2.5 | 7.4 | GO:1901074 | cellular response to interleukin-13(GO:0035963) regulation of engulfment of apoptotic cell(GO:1901074) lipoxin biosynthetic process(GO:2001301) lipoxin A4 metabolic process(GO:2001302) lipoxin A4 biosynthetic process(GO:2001303) |
| 2.5 | 14.8 | GO:1903282 | negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) regulation of peroxidase activity(GO:2000468) positive regulation of peroxidase activity(GO:2000470) |
| 2.3 | 9.0 | GO:0050904 | diapedesis(GO:0050904) |
| 2.2 | 6.7 | GO:1903717 | carbamoyl phosphate metabolic process(GO:0070408) carbamoyl phosphate biosynthetic process(GO:0070409) cellular response to oleic acid(GO:0071400) response to ammonia(GO:1903717) cellular response to ammonia(GO:1903718) |
| 2.2 | 6.6 | GO:1900920 | regulation of amino acid uptake involved in synaptic transmission(GO:0051941) regulation of glutamate uptake involved in transmission of nerve impulse(GO:0051946) regulation of L-glutamate import(GO:1900920) |
| 2.2 | 6.5 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 2.1 | 8.3 | GO:0006258 | UDP-glucose catabolic process(GO:0006258) positive regulation of adrenergic receptor signaling pathway(GO:0071879) negative regulation of type B pancreatic cell development(GO:2000077) |
| 1.9 | 5.6 | GO:0036292 | DNA rewinding(GO:0036292) |
| 1.8 | 5.4 | GO:0006210 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 1.7 | 5.2 | GO:0010025 | wax biosynthetic process(GO:0010025) wax metabolic process(GO:0010166) |
| 1.5 | 4.5 | GO:0044179 | hemolysis by symbiont of host erythrocytes(GO:0019836) hemolysis in other organism(GO:0044179) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 1.5 | 4.4 | GO:1903487 | regulation of lactation(GO:1903487) |
| 1.4 | 5.5 | GO:0035790 | platelet-derived growth factor receptor-alpha signaling pathway(GO:0035790) |
| 1.3 | 4.0 | GO:0033082 | regulation of extrathymic T cell differentiation(GO:0033082) sebum secreting cell proliferation(GO:1990654) |
| 1.3 | 4.0 | GO:0050653 | chondroitin sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0050653) |
| 1.3 | 5.3 | GO:0071918 | urea transmembrane transport(GO:0071918) |
| 1.3 | 6.4 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 1.2 | 8.5 | GO:1990253 | cellular response to leucine starvation(GO:1990253) |
| 1.2 | 4.9 | GO:0072660 | maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 1.2 | 3.5 | GO:0002416 | IgG immunoglobulin transcytosis in epithelial cells mediated by FcRn immunoglobulin receptor(GO:0002416) |
| 1.1 | 4.6 | GO:0000023 | maltose metabolic process(GO:0000023) |
| 1.1 | 3.4 | GO:1902498 | regulation of protein autoubiquitination(GO:1902498) |
| 1.1 | 4.4 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 1.1 | 3.3 | GO:0030573 | bile acid catabolic process(GO:0030573) |
| 1.1 | 6.4 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 1.1 | 13.7 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 1.1 | 13.7 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 1.0 | 3.0 | GO:2000439 | positive regulation of monocyte extravasation(GO:2000439) |
| 1.0 | 2.9 | GO:0001300 | chronological cell aging(GO:0001300) |
| 1.0 | 4.9 | GO:1903385 | regulation of homophilic cell adhesion(GO:1903385) |
| 1.0 | 2.9 | GO:0030187 | melatonin metabolic process(GO:0030186) melatonin biosynthetic process(GO:0030187) |
| 0.9 | 6.6 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 0.9 | 10.4 | GO:0006030 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.9 | 6.4 | GO:0015705 | iodide transport(GO:0015705) |
| 0.9 | 3.6 | GO:0071395 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 0.9 | 4.4 | GO:0014738 | regulation of muscle hyperplasia(GO:0014738) |
| 0.9 | 2.6 | GO:0015847 | putrescine transport(GO:0015847) positive regulation of polyamine transmembrane transport(GO:1902269) |
| 0.9 | 9.4 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.8 | 2.5 | GO:0071109 | superior temporal gyrus development(GO:0071109) |
| 0.8 | 6.8 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 0.8 | 4.9 | GO:0070829 | response to vitamin B2(GO:0033274) heterochromatin maintenance(GO:0070829) |
| 0.8 | 7.9 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.8 | 2.3 | GO:0010040 | response to iron(II) ion(GO:0010040) |
| 0.8 | 5.3 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.8 | 6.8 | GO:0042904 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.8 | 2.3 | GO:0072233 | thick ascending limb development(GO:0072023) metanephric thick ascending limb development(GO:0072233) |
| 0.7 | 4.4 | GO:0060084 | synaptic transmission involved in micturition(GO:0060084) |
| 0.7 | 6.5 | GO:0070315 | G1 to G0 transition involved in cell differentiation(GO:0070315) |
| 0.7 | 3.6 | GO:0009233 | menaquinone metabolic process(GO:0009233) |
| 0.7 | 2.9 | GO:0044026 | DNA hypermethylation(GO:0044026) |
| 0.7 | 7.2 | GO:0006537 | glutamate biosynthetic process(GO:0006537) gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 0.7 | 2.2 | GO:0060988 | lipid tube assembly(GO:0060988) |
| 0.7 | 4.3 | GO:0061624 | fructose catabolic process(GO:0006001) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) |
| 0.7 | 7.0 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.7 | 8.9 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.7 | 2.0 | GO:1901097 | negative regulation of autophagosome maturation(GO:1901097) |
| 0.7 | 6.1 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.7 | 4.7 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.7 | 2.6 | GO:1904381 | Golgi apparatus mannose trimming(GO:1904381) |
| 0.6 | 4.5 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.6 | 3.2 | GO:1904073 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.6 | 3.2 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.6 | 3.8 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 0.6 | 9.4 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 0.6 | 3.0 | GO:0093001 | glycolysis from storage polysaccharide through glucose-1-phosphate(GO:0093001) |
| 0.6 | 3.0 | GO:0060748 | tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
| 0.6 | 1.8 | GO:0002503 | peptide antigen assembly with MHC class II protein complex(GO:0002503) |
| 0.6 | 3.5 | GO:0010044 | response to aluminum ion(GO:0010044) positive regulation of catecholamine metabolic process(GO:0045915) positive regulation of dopamine metabolic process(GO:0045964) |
| 0.6 | 9.3 | GO:0060907 | positive regulation of macrophage cytokine production(GO:0060907) |
| 0.6 | 1.7 | GO:0045356 | positive regulation of interferon-alpha biosynthetic process(GO:0045356) |
| 0.6 | 1.7 | GO:1900673 | phenylpropanoid catabolic process(GO:0046271) olefin metabolic process(GO:1900673) |
| 0.6 | 1.7 | GO:0060369 | positive regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060369) |
| 0.6 | 7.9 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) |
| 0.5 | 4.4 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.5 | 1.6 | GO:0003099 | vasodilation by norepinephrine-epinephrine involved in regulation of systemic arterial blood pressure(GO:0002025) positive regulation of the force of heart contraction by chemical signal(GO:0003099) |
| 0.5 | 3.2 | GO:0061517 | microglial cell activation involved in immune response(GO:0002282) macrophage proliferation(GO:0061517) microglial cell proliferation(GO:0061518) |
| 0.5 | 5.8 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.5 | 8.2 | GO:0097151 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.5 | 1.0 | GO:0010840 | regulation of circadian sleep/wake cycle, wakefulness(GO:0010840) circadian sleep/wake cycle, wakefulness(GO:0042746) |
| 0.5 | 3.5 | GO:0072733 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 0.5 | 3.0 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.5 | 1.5 | GO:0061588 | calcium activated phospholipid scrambling(GO:0061588) calcium activated phosphatidylcholine scrambling(GO:0061590) calcium activated galactosylceramide scrambling(GO:0061591) |
| 0.5 | 1.4 | GO:0071461 | cellular response to redox state(GO:0071461) |
| 0.5 | 5.8 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 0.5 | 0.5 | GO:1904339 | negative regulation of dopaminergic neuron differentiation(GO:1904339) |
| 0.5 | 1.0 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 0.5 | 2.8 | GO:0007341 | penetration of zona pellucida(GO:0007341) |
| 0.5 | 4.2 | GO:0042670 | retinal cone cell differentiation(GO:0042670) retinal cone cell development(GO:0046549) |
| 0.5 | 2.7 | GO:0046092 | deoxycytidine metabolic process(GO:0046092) |
| 0.5 | 3.6 | GO:0048630 | skeletal muscle tissue growth(GO:0048630) |
| 0.4 | 1.3 | GO:0018963 | dibenzo-p-dioxin metabolic process(GO:0018894) phthalate metabolic process(GO:0018963) cellular response to luteinizing hormone stimulus(GO:0071373) |
| 0.4 | 2.1 | GO:1901909 | diadenosine polyphosphate catabolic process(GO:0015961) diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 0.4 | 1.7 | GO:1900158 | positive regulation of osteoclast proliferation(GO:0090290) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.4 | 1.3 | GO:2000342 | negative regulation of chemokine (C-X-C motif) ligand 2 production(GO:2000342) |
| 0.4 | 5.7 | GO:0048245 | eosinophil chemotaxis(GO:0048245) |
| 0.4 | 3.2 | GO:0016139 | glycoside catabolic process(GO:0016139) |
| 0.4 | 4.4 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.4 | 2.0 | GO:0015811 | L-cystine transport(GO:0015811) |
| 0.4 | 2.3 | GO:0050955 | thermoception(GO:0050955) |
| 0.4 | 1.9 | GO:0046208 | spermine catabolic process(GO:0046208) |
| 0.4 | 1.2 | GO:0032661 | regulation of interleukin-18 production(GO:0032661) |
| 0.4 | 3.4 | GO:1903142 | positive regulation of endothelial cell development(GO:1901552) positive regulation of establishment of endothelial barrier(GO:1903142) |
| 0.4 | 1.9 | GO:0038163 | thrombopoietin-mediated signaling pathway(GO:0038163) |
| 0.4 | 13.3 | GO:0072600 | establishment of protein localization to Golgi(GO:0072600) |
| 0.4 | 5.1 | GO:1990440 | ATF6-mediated unfolded protein response(GO:0036500) positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.4 | 1.1 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.4 | 10.2 | GO:1904659 | hexose transmembrane transport(GO:0035428) glucose transmembrane transport(GO:1904659) |
| 0.4 | 1.4 | GO:0046601 | positive regulation of centriole replication(GO:0046601) |
| 0.4 | 2.5 | GO:1902514 | regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1902514) positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.3 | 1.4 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.3 | 3.4 | GO:0042908 | xenobiotic transport(GO:0042908) |
| 0.3 | 3.7 | GO:0006069 | ethanol oxidation(GO:0006069) |
| 0.3 | 4.7 | GO:0015074 | DNA integration(GO:0015074) |
| 0.3 | 6.6 | GO:0021513 | spinal cord dorsal/ventral patterning(GO:0021513) |
| 0.3 | 3.3 | GO:1902857 | positive regulation of nonmotile primary cilium assembly(GO:1902857) |
| 0.3 | 2.3 | GO:2000698 | positive regulation of epithelial cell differentiation involved in kidney development(GO:2000698) positive regulation of nephron tubule epithelial cell differentiation(GO:2000768) |
| 0.3 | 0.9 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 0.3 | 0.9 | GO:0097272 | ammonia homeostasis(GO:0097272) |
| 0.3 | 2.5 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.3 | 2.5 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.3 | 4.0 | GO:0045046 | protein import into peroxisome membrane(GO:0045046) |
| 0.3 | 1.5 | GO:0010748 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) plasma membrane long-chain fatty acid transport(GO:0015911) |
| 0.3 | 0.9 | GO:0010046 | response to mycotoxin(GO:0010046) |
| 0.3 | 0.6 | GO:1990418 | response to insulin-like growth factor stimulus(GO:1990418) |
| 0.3 | 5.1 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.3 | 2.7 | GO:0086024 | adrenergic receptor signaling pathway involved in positive regulation of heart rate(GO:0086024) |
| 0.3 | 4.4 | GO:0072540 | T-helper 17 cell lineage commitment(GO:0072540) |
| 0.3 | 5.7 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.3 | 4.0 | GO:0090026 | positive regulation of monocyte chemotaxis(GO:0090026) |
| 0.3 | 1.7 | GO:0033029 | regulation of neutrophil apoptotic process(GO:0033029) |
| 0.3 | 2.2 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.3 | 8.3 | GO:0050427 | purine ribonucleoside bisphosphate metabolic process(GO:0034035) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) |
| 0.3 | 1.3 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.3 | 2.7 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.3 | 14.0 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.3 | 1.3 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.3 | 0.5 | GO:0014016 | neuroblast differentiation(GO:0014016) |
| 0.3 | 2.6 | GO:0032914 | positive regulation of transforming growth factor beta1 production(GO:0032914) |
| 0.3 | 1.0 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.3 | 4.3 | GO:0019934 | cGMP-mediated signaling(GO:0019934) |
| 0.3 | 5.8 | GO:0071294 | cellular response to zinc ion(GO:0071294) |
| 0.3 | 0.5 | GO:1990167 | protein K27-linked deubiquitination(GO:1990167) |
| 0.2 | 2.5 | GO:0048102 | autophagic cell death(GO:0048102) |
| 0.2 | 2.7 | GO:0034219 | carbohydrate transmembrane transport(GO:0034219) |
| 0.2 | 2.0 | GO:0035234 | ectopic germ cell programmed cell death(GO:0035234) |
| 0.2 | 1.2 | GO:0010966 | regulation of phosphate transport(GO:0010966) |
| 0.2 | 0.7 | GO:1904350 | regulation of protein catabolic process in the vacuole(GO:1904350) positive regulation of protein catabolic process in the vacuole(GO:1904352) |
| 0.2 | 1.4 | GO:0070305 | response to cGMP(GO:0070305) |
| 0.2 | 1.6 | GO:1990416 | cellular response to brain-derived neurotrophic factor stimulus(GO:1990416) |
| 0.2 | 2.8 | GO:0071285 | cellular response to lithium ion(GO:0071285) |
| 0.2 | 3.2 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.2 | 7.7 | GO:0008209 | androgen metabolic process(GO:0008209) |
| 0.2 | 2.7 | GO:0001514 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.2 | 0.7 | GO:0072709 | cellular response to sorbitol(GO:0072709) |
| 0.2 | 5.3 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.2 | 2.7 | GO:0071318 | cellular response to ATP(GO:0071318) |
| 0.2 | 1.8 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.2 | 3.9 | GO:1901663 | ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.2 | 0.9 | GO:0006663 | platelet activating factor biosynthetic process(GO:0006663) |
| 0.2 | 2.1 | GO:0048263 | determination of dorsal identity(GO:0048263) |
| 0.2 | 0.8 | GO:0060133 | somatotropin secreting cell development(GO:0060133) |
| 0.2 | 0.6 | GO:1903059 | regulation of protein lipidation(GO:1903059) positive regulation of protein lipidation(GO:1903061) |
| 0.2 | 2.1 | GO:1902525 | regulation of protein monoubiquitination(GO:1902525) |
| 0.2 | 2.7 | GO:1902018 | negative regulation of cilium assembly(GO:1902018) |
| 0.2 | 0.8 | GO:0010836 | negative regulation of protein ADP-ribosylation(GO:0010836) |
| 0.2 | 0.6 | GO:0015798 | myo-inositol transport(GO:0015798) |
| 0.2 | 8.1 | GO:0070884 | regulation of calcineurin-NFAT signaling cascade(GO:0070884) |
| 0.2 | 2.1 | GO:0034312 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) sphingoid biosynthetic process(GO:0046520) |
| 0.2 | 1.3 | GO:0070862 | negative regulation of protein exit from endoplasmic reticulum(GO:0070862) negative regulation of retrograde protein transport, ER to cytosol(GO:1904153) |
| 0.2 | 4.6 | GO:0006907 | pinocytosis(GO:0006907) |
| 0.2 | 0.2 | GO:2001183 | negative regulation of interleukin-12 secretion(GO:2001183) |
| 0.2 | 2.3 | GO:0098719 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.2 | 3.2 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.2 | 0.9 | GO:0010519 | negative regulation of phospholipase activity(GO:0010519) |
| 0.2 | 1.1 | GO:0070172 | positive regulation of tooth mineralization(GO:0070172) |
| 0.2 | 1.6 | GO:0006682 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.2 | 2.2 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.2 | 3.0 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.2 | 1.6 | GO:0034723 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.2 | 2.2 | GO:0014809 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 0.2 | 4.5 | GO:1901687 | glutathione derivative metabolic process(GO:1901685) glutathione derivative biosynthetic process(GO:1901687) |
| 0.2 | 6.9 | GO:0007618 | mating(GO:0007618) |
| 0.2 | 1.0 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.2 | 1.3 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.2 | 4.9 | GO:0007398 | ectoderm development(GO:0007398) |
| 0.2 | 5.8 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.2 | 1.1 | GO:0072539 | T-helper 17 cell differentiation(GO:0072539) |
| 0.2 | 1.3 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.2 | 2.4 | GO:1900452 | regulation of long term synaptic depression(GO:1900452) |
| 0.2 | 0.8 | GO:0046167 | glycerol-3-phosphate biosynthetic process(GO:0046167) |
| 0.2 | 3.6 | GO:0045332 | lipid translocation(GO:0034204) phospholipid translocation(GO:0045332) |
| 0.1 | 2.8 | GO:0090022 | regulation of neutrophil chemotaxis(GO:0090022) |
| 0.1 | 1.6 | GO:0002775 | antimicrobial peptide production(GO:0002775) antibacterial peptide production(GO:0002778) |
| 0.1 | 2.6 | GO:0090036 | regulation of protein kinase C signaling(GO:0090036) |
| 0.1 | 1.6 | GO:0048007 | antigen processing and presentation of lipid antigen via MHC class Ib(GO:0048003) antigen processing and presentation, exogenous lipid antigen via MHC class Ib(GO:0048007) |
| 0.1 | 4.3 | GO:0042462 | eye photoreceptor cell development(GO:0042462) |
| 0.1 | 11.1 | GO:0007032 | endosome organization(GO:0007032) |
| 0.1 | 2.2 | GO:0010826 | negative regulation of centrosome duplication(GO:0010826) |
| 0.1 | 22.0 | GO:0046718 | viral entry into host cell(GO:0046718) |
| 0.1 | 5.2 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 13.6 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.1 | 2.7 | GO:0006271 | DNA strand elongation involved in DNA replication(GO:0006271) |
| 0.1 | 4.9 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.1 | 4.5 | GO:0000002 | mitochondrial genome maintenance(GO:0000002) |
| 0.1 | 21.5 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.1 | 0.5 | GO:0045829 | negative regulation of isotype switching(GO:0045829) |
| 0.1 | 4.9 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.1 | 0.9 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 0.1 | 3.6 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.1 | 0.9 | GO:0050882 | voluntary musculoskeletal movement(GO:0050882) |
| 0.1 | 4.3 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.1 | 1.2 | GO:0010756 | positive regulation of plasminogen activation(GO:0010756) positive regulation of extracellular matrix disassembly(GO:0090091) |
| 0.1 | 4.4 | GO:0007194 | negative regulation of adenylate cyclase activity(GO:0007194) |
| 0.1 | 1.4 | GO:0042355 | fucose catabolic process(GO:0019317) L-fucose metabolic process(GO:0042354) L-fucose catabolic process(GO:0042355) |
| 0.1 | 7.9 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.1 | 2.8 | GO:0035640 | exploration behavior(GO:0035640) |
| 0.1 | 1.9 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.1 | 4.7 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.1 | 13.8 | GO:0007608 | sensory perception of smell(GO:0007608) |
| 0.1 | 2.2 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.1 | 2.4 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.1 | 6.1 | GO:0019933 | cAMP-mediated signaling(GO:0019933) |
| 0.1 | 0.4 | GO:0009698 | phenylpropanoid metabolic process(GO:0009698) |
| 0.1 | 0.4 | GO:2000504 | positive regulation of blood vessel remodeling(GO:2000504) |
| 0.1 | 0.3 | GO:0097032 | respiratory chain complex II assembly(GO:0034552) mitochondrial respiratory chain complex II assembly(GO:0034553) mitochondrial respiratory chain complex II biogenesis(GO:0097032) |
| 0.1 | 4.1 | GO:0002755 | MyD88-dependent toll-like receptor signaling pathway(GO:0002755) |
| 0.1 | 1.3 | GO:0044804 | nucleophagy(GO:0044804) |
| 0.1 | 7.0 | GO:0030101 | natural killer cell activation(GO:0030101) |
| 0.1 | 1.8 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.1 | 1.7 | GO:0034505 | tooth mineralization(GO:0034505) |
| 0.1 | 5.8 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.1 | 3.3 | GO:0043304 | regulation of mast cell activation involved in immune response(GO:0033006) regulation of mast cell degranulation(GO:0043304) |
| 0.1 | 0.8 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.1 | 2.6 | GO:0097503 | sialylation(GO:0097503) |
| 0.1 | 1.7 | GO:0008535 | respiratory chain complex IV assembly(GO:0008535) |
| 0.1 | 3.3 | GO:0033687 | osteoblast proliferation(GO:0033687) |
| 0.1 | 0.4 | GO:0035992 | tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) |
| 0.1 | 1.4 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.1 | 1.2 | GO:1903608 | protein localization to cytoplasmic stress granule(GO:1903608) |
| 0.1 | 2.4 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.1 | 0.4 | GO:0031119 | tRNA pseudouridine synthesis(GO:0031119) |
| 0.1 | 0.9 | GO:0001573 | ganglioside metabolic process(GO:0001573) |
| 0.1 | 0.2 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) |
| 0.1 | 2.8 | GO:0051145 | smooth muscle cell differentiation(GO:0051145) |
| 0.1 | 0.2 | GO:2000705 | dense core granule biogenesis(GO:0061110) amniotic stem cell differentiation(GO:0097086) regulation of dense core granule biogenesis(GO:2000705) negative regulation of dense core granule biogenesis(GO:2000706) negative regulation of mesenchymal stem cell differentiation(GO:2000740) regulation of amniotic stem cell differentiation(GO:2000797) negative regulation of amniotic stem cell differentiation(GO:2000798) |
| 0.1 | 3.5 | GO:0051693 | actin filament capping(GO:0051693) |
| 0.1 | 2.7 | GO:0001895 | retina homeostasis(GO:0001895) |
| 0.1 | 0.6 | GO:0046940 | nucleoside monophosphate phosphorylation(GO:0046940) |
| 0.1 | 1.0 | GO:0090344 | negative regulation of cell aging(GO:0090344) |
| 0.1 | 0.7 | GO:0034067 | protein localization to Golgi apparatus(GO:0034067) |
| 0.1 | 6.5 | GO:0008585 | female gonad development(GO:0008585) |
| 0.1 | 1.1 | GO:0048016 | inositol phosphate-mediated signaling(GO:0048016) |
| 0.1 | 7.8 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.1 | 1.0 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) arachidonic acid secretion(GO:0050482) arachidonate transport(GO:1903963) |
| 0.1 | 1.9 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.1 | 1.8 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.1 | 1.7 | GO:0045022 | early endosome to late endosome transport(GO:0045022) |
| 0.1 | 3.3 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.1 | 1.4 | GO:0031640 | killing of cells of other organism(GO:0031640) disruption of cells of other organism(GO:0044364) |
| 0.1 | 0.1 | GO:0005989 | lactose metabolic process(GO:0005988) lactose biosynthetic process(GO:0005989) |
| 0.1 | 0.2 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 0.1 | 2.3 | GO:0060997 | dendritic spine morphogenesis(GO:0060997) |
| 0.1 | 1.1 | GO:0014046 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.1 | 0.9 | GO:0006590 | thyroid hormone generation(GO:0006590) |
| 0.1 | 0.7 | GO:1901750 | leukotriene D4 metabolic process(GO:1901748) leukotriene D4 biosynthetic process(GO:1901750) |
| 0.1 | 1.1 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.1 | 1.3 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.1 | 4.0 | GO:0034644 | cellular response to UV(GO:0034644) |
| 0.0 | 1.7 | GO:0032760 | positive regulation of tumor necrosis factor production(GO:0032760) |
| 0.0 | 2.9 | GO:0009411 | response to UV(GO:0009411) |
| 0.0 | 16.9 | GO:0051056 | regulation of small GTPase mediated signal transduction(GO:0051056) |
| 0.0 | 1.4 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 1.8 | GO:0046835 | carbohydrate phosphorylation(GO:0046835) |
| 0.0 | 1.2 | GO:0050982 | detection of mechanical stimulus(GO:0050982) |
| 0.0 | 0.8 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 2.7 | GO:0006687 | glycosphingolipid metabolic process(GO:0006687) |
| 0.0 | 0.5 | GO:0070935 | 3'-UTR-mediated mRNA stabilization(GO:0070935) |
| 0.0 | 3.2 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.0 | 3.3 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 0.3 | GO:0071801 | regulation of podosome assembly(GO:0071801) positive regulation of podosome assembly(GO:0071803) |
| 0.0 | 0.3 | GO:1902236 | negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway(GO:1902236) |
| 0.0 | 2.3 | GO:0016079 | synaptic vesicle exocytosis(GO:0016079) |
| 0.0 | 0.9 | GO:0032418 | lysosome localization(GO:0032418) |
| 0.0 | 0.8 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.0 | 1.2 | GO:0070206 | protein trimerization(GO:0070206) |
| 0.0 | 0.4 | GO:0030208 | chondroitin sulfate catabolic process(GO:0030207) dermatan sulfate biosynthetic process(GO:0030208) |
| 0.0 | 0.2 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.0 | 0.9 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation(GO:0010862) |
| 0.0 | 1.3 | GO:0007585 | respiratory gaseous exchange(GO:0007585) |
| 0.0 | 1.6 | GO:0021549 | cerebellum development(GO:0021549) |
| 0.0 | 0.6 | GO:0046850 | regulation of bone remodeling(GO:0046850) |
| 0.0 | 3.2 | GO:0006909 | phagocytosis(GO:0006909) |
| 0.0 | 2.2 | GO:0060337 | type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 0.0 | 3.2 | GO:0043484 | regulation of RNA splicing(GO:0043484) |
| 0.0 | 3.6 | GO:0007411 | axon guidance(GO:0007411) |
| 0.0 | 0.4 | GO:0008089 | anterograde axonal transport(GO:0008089) |
| 0.0 | 2.1 | GO:0008217 | regulation of blood pressure(GO:0008217) |
| 0.0 | 0.7 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 1.3 | GO:0043473 | pigmentation(GO:0043473) |
| 0.0 | 0.1 | GO:2000809 | synaptic vesicle clustering(GO:0097091) regulation of synaptic vesicle clustering(GO:2000807) positive regulation of synaptic vesicle clustering(GO:2000809) |
| 0.0 | 0.7 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 0.1 | GO:0042953 | lipoprotein transport(GO:0042953) lipoprotein localization(GO:0044872) |
| 0.0 | 1.6 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.0 | 0.5 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 0.3 | GO:0042359 | vitamin D metabolic process(GO:0042359) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 35.6 | GO:0098647 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 2.1 | 6.4 | GO:0009328 | phenylalanine-tRNA ligase complex(GO:0009328) |
| 1.6 | 4.8 | GO:0043159 | acrosomal matrix(GO:0043159) |
| 1.6 | 4.7 | GO:0090571 | RNA polymerase II transcription repressor complex(GO:0090571) |
| 1.0 | 2.9 | GO:0072563 | endothelial microparticle(GO:0072563) |
| 0.9 | 16.5 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.8 | 6.4 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.7 | 3.0 | GO:0005965 | protein farnesyltransferase complex(GO:0005965) |
| 0.7 | 27.1 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.7 | 2.2 | GO:0060987 | lipid tube(GO:0060987) |
| 0.7 | 4.3 | GO:0034448 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.7 | 8.3 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.7 | 1.4 | GO:0033011 | perinuclear theca(GO:0033011) cytoskeletal calyx(GO:0033150) |
| 0.6 | 9.6 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.6 | 3.5 | GO:0032437 | cuticular plate(GO:0032437) |
| 0.6 | 2.8 | GO:0032807 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) DNA ligase IV complex(GO:0032807) |
| 0.5 | 3.0 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.5 | 2.0 | GO:0000801 | central element(GO:0000801) |
| 0.5 | 6.6 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.5 | 1.4 | GO:0098536 | deuterosome(GO:0098536) |
| 0.5 | 1.8 | GO:0034678 | integrin alpha8-beta1 complex(GO:0034678) |
| 0.5 | 3.2 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.4 | 8.9 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.4 | 6.6 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.4 | 3.1 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.4 | 8.3 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.4 | 4.3 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.4 | 3.8 | GO:0031414 | N-terminal protein acetyltransferase complex(GO:0031414) |
| 0.4 | 2.5 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.4 | 2.0 | GO:0036398 | TCR signalosome(GO:0036398) |
| 0.4 | 2.8 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 0.4 | 2.0 | GO:0072558 | NLRP1 inflammasome complex(GO:0072558) |
| 0.4 | 1.6 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.4 | 3.1 | GO:0097425 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.4 | 3.4 | GO:0032010 | phagolysosome(GO:0032010) |
| 0.4 | 6.8 | GO:0042627 | chylomicron(GO:0042627) |
| 0.4 | 2.6 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.4 | 6.6 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.3 | 2.1 | GO:0070554 | synaptobrevin 2-SNAP-25-syntaxin-3-complexin complex(GO:0070554) |
| 0.3 | 5.0 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.3 | 9.2 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.3 | 34.4 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 0.3 | 9.2 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.3 | 27.0 | GO:0005902 | microvillus(GO:0005902) |
| 0.3 | 5.8 | GO:0031045 | dense core granule(GO:0031045) |
| 0.3 | 4.9 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.3 | 1.3 | GO:0000836 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.2 | 5.7 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.2 | 2.9 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.2 | 3.6 | GO:0097433 | dense body(GO:0097433) |
| 0.2 | 2.5 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.2 | 5.0 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.2 | 0.7 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 0.2 | 6.9 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.2 | 5.8 | GO:0032420 | stereocilium(GO:0032420) |
| 0.2 | 9.6 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.2 | 2.5 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.2 | 2.2 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.2 | 1.2 | GO:0097165 | nuclear stress granule(GO:0097165) |
| 0.2 | 3.8 | GO:0036020 | endolysosome membrane(GO:0036020) |
| 0.2 | 6.2 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 8.2 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 1.6 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.1 | 4.3 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.1 | 2.2 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.1 | 0.9 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.1 | 2.5 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.1 | 2.2 | GO:0036038 | MKS complex(GO:0036038) |
| 0.1 | 2.1 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.1 | 5.8 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.1 | 1.3 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.1 | 8.5 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.1 | 8.2 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.1 | 3.2 | GO:0032982 | myosin filament(GO:0032982) |
| 0.1 | 0.5 | GO:0005889 | hydrogen:potassium-exchanging ATPase complex(GO:0005889) |
| 0.1 | 6.1 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 2.2 | GO:0031082 | BLOC complex(GO:0031082) |
| 0.1 | 3.8 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.1 | 16.1 | GO:0043204 | perikaryon(GO:0043204) |
| 0.1 | 10.2 | GO:0005811 | lipid particle(GO:0005811) |
| 0.1 | 1.2 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.1 | 3.5 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 10.8 | GO:0030665 | clathrin-coated vesicle membrane(GO:0030665) |
| 0.1 | 4.9 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 4.2 | GO:0005844 | polysome(GO:0005844) |
| 0.1 | 1.8 | GO:0032809 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.1 | 1.4 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.1 | 1.3 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 25.9 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 4.3 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 1.6 | GO:0071011 | precatalytic spliceosome(GO:0071011) |
| 0.1 | 14.2 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.1 | 9.7 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.1 | 2.0 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 1.1 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.1 | 0.5 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.1 | 1.2 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.1 | 0.7 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 0.1 | 3.8 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.1 | 3.5 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 68.2 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.2 | GO:0097123 | cyclin A1-CDK2 complex(GO:0097123) |
| 0.0 | 9.6 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 9.0 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.0 | 4.1 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.0 | 1.3 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 2.3 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 3.7 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 1.8 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 1.7 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 1.8 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 6.2 | GO:0030139 | endocytic vesicle(GO:0030139) |
| 0.0 | 52.3 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 0.2 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 76.6 | GO:0016021 | integral component of membrane(GO:0016021) |
| 0.0 | 0.5 | GO:0031011 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 0.7 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.0 | 0.1 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.6 | 10.7 | GO:0019959 | interleukin-8 binding(GO:0019959) |
| 2.5 | 7.4 | GO:0047977 | hepoxilin-epoxide hydrolase activity(GO:0047977) |
| 2.5 | 14.8 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 2.4 | 9.5 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 2.3 | 7.0 | GO:0036134 | thromboxane-A synthase activity(GO:0004796) 12-hydroxyheptadecatrienoic acid synthase activity(GO:0036134) |
| 2.3 | 13.9 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 2.1 | 25.1 | GO:0008430 | selenium binding(GO:0008430) |
| 1.9 | 7.5 | GO:0004982 | N-formyl peptide receptor activity(GO:0004982) scavenger receptor binding(GO:0005124) |
| 1.9 | 5.6 | GO:0008969 | phosphohistidine phosphatase activity(GO:0008969) |
| 1.7 | 5.2 | GO:0080019 | fatty-acyl-CoA reductase (alcohol-forming) activity(GO:0080019) |
| 1.7 | 6.7 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 1.6 | 4.9 | GO:0004577 | N-acetylglucosaminyldiphosphodolichol N-acetylglucosaminyltransferase activity(GO:0004577) |
| 1.5 | 4.6 | GO:0004339 | glucan 1,4-alpha-glucosidase activity(GO:0004339) |
| 1.4 | 4.3 | GO:0061609 | fructose-1-phosphate aldolase activity(GO:0061609) |
| 1.3 | 5.3 | GO:0015204 | urea transmembrane transporter activity(GO:0015204) |
| 1.3 | 6.4 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 1.2 | 11.6 | GO:0004064 | arylesterase activity(GO:0004064) |
| 1.1 | 4.4 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 1.1 | 4.4 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 1.1 | 3.3 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 1.1 | 4.3 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 1.0 | 2.0 | GO:0047115 | phenanthrene 9,10-monooxygenase activity(GO:0018636) ketosteroid monooxygenase activity(GO:0047086) trans-1,2-dihydrobenzene-1,2-diol dehydrogenase activity(GO:0047115) |
| 1.0 | 7.0 | GO:0015198 | oligopeptide transporter activity(GO:0015198) |
| 0.9 | 10.4 | GO:0004568 | chitinase activity(GO:0004568) chitin binding(GO:0008061) |
| 0.9 | 2.8 | GO:0004040 | amidase activity(GO:0004040) fucose binding(GO:0042806) |
| 0.9 | 4.7 | GO:0004803 | transposase activity(GO:0004803) |
| 0.9 | 8.2 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.9 | 4.6 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.9 | 2.7 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.9 | 2.6 | GO:0052798 | beta-galactoside alpha-2,3-sialyltransferase activity(GO:0052798) |
| 0.9 | 2.6 | GO:0015489 | polyamine transmembrane transporter activity(GO:0015203) putrescine transmembrane transporter activity(GO:0015489) |
| 0.9 | 3.5 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.8 | 4.9 | GO:0004489 | methylenetetrahydrofolate reductase (NAD(P)H) activity(GO:0004489) |
| 0.8 | 4.0 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.8 | 4.0 | GO:0047238 | glucuronosyl-N-acetylgalactosaminyl-proteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047238) |
| 0.8 | 3.1 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.8 | 6.1 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.7 | 3.0 | GO:0004660 | protein farnesyltransferase activity(GO:0004660) |
| 0.7 | 10.2 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.7 | 5.0 | GO:0050294 | steroid sulfotransferase activity(GO:0050294) |
| 0.7 | 2.0 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 0.7 | 5.9 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.6 | 3.2 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.6 | 54.6 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.6 | 1.9 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.5 | 1.6 | GO:0004939 | beta-adrenergic receptor activity(GO:0004939) |
| 0.5 | 9.3 | GO:0031432 | titin binding(GO:0031432) |
| 0.5 | 1.6 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 0.5 | 3.7 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.5 | 1.6 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 0.5 | 4.2 | GO:0044547 | DNA topoisomerase binding(GO:0044547) |
| 0.5 | 0.5 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 0.5 | 7.2 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.5 | 6.6 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.5 | 5.6 | GO:0036310 | annealing helicase activity(GO:0036310) |
| 0.5 | 14.7 | GO:0071949 | FAD binding(GO:0071949) |
| 0.5 | 2.5 | GO:0005298 | proline:sodium symporter activity(GO:0005298) |
| 0.5 | 6.8 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.5 | 1.9 | GO:0052901 | polyamine oxidase activity(GO:0046592) spermine:oxygen oxidoreductase (spermidine-forming) activity(GO:0052901) |
| 0.5 | 8.2 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.5 | 3.8 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 0.5 | 1.4 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 0.5 | 1.9 | GO:0016426 | tRNA (adenine) methyltransferase activity(GO:0016426) |
| 0.5 | 3.7 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) |
| 0.4 | 8.0 | GO:0016405 | CoA-ligase activity(GO:0016405) |
| 0.4 | 6.5 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.4 | 2.1 | GO:0052848 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) inositol diphosphate tetrakisphosphate diphosphatase activity(GO:0052840) inositol bisdiphosphate tetrakisphosphate diphosphatase activity(GO:0052841) inositol diphosphate pentakisphosphate diphosphatase activity(GO:0052842) inositol-1-diphosphate-2,3,4,5,6-pentakisphosphate diphosphatase activity(GO:0052843) inositol-3-diphosphate-1,2,4,5,6-pentakisphosphate diphosphatase activity(GO:0052844) inositol-5-diphosphate-1,2,3,4,6-pentakisphosphate diphosphatase activity(GO:0052845) inositol-1,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 1-diphosphatase activity(GO:0052846) inositol-1,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 5-diphosphatase activity(GO:0052847) inositol-3,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 5-diphosphatase activity(GO:0052848) |
| 0.4 | 3.4 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.4 | 4.2 | GO:0005549 | odorant binding(GO:0005549) |
| 0.4 | 2.9 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.4 | 3.3 | GO:0035374 | chondroitin sulfate binding(GO:0035374) |
| 0.4 | 2.4 | GO:0016807 | cysteine-type carboxypeptidase activity(GO:0016807) cysteine-type exopeptidase activity(GO:0070004) |
| 0.4 | 2.0 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.4 | 2.7 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) thymidine kinase activity(GO:0004797) |
| 0.4 | 6.5 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.4 | 8.7 | GO:0008409 | 5'-3' exonuclease activity(GO:0008409) |
| 0.4 | 3.0 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.4 | 1.9 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.4 | 1.1 | GO:0046848 | hydroxyapatite binding(GO:0046848) |
| 0.4 | 10.7 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.4 | 2.9 | GO:0005534 | galactose binding(GO:0005534) |
| 0.4 | 9.8 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.4 | 6.1 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.4 | 2.1 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.4 | 2.1 | GO:0005223 | intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.4 | 3.5 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.3 | 1.4 | GO:0047708 | biotinidase activity(GO:0047708) |
| 0.3 | 3.1 | GO:0003696 | satellite DNA binding(GO:0003696) |
| 0.3 | 1.0 | GO:0008267 | poly-glutamine tract binding(GO:0008267) |
| 0.3 | 2.6 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.3 | 11.1 | GO:1905030 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.3 | 4.5 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.3 | 4.7 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.3 | 1.5 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) xenobiotic transporter activity(GO:0042910) |
| 0.3 | 3.6 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.3 | 0.9 | GO:0002113 | interleukin-33 binding(GO:0002113) |
| 0.3 | 10.1 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.3 | 4.5 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.3 | 8.3 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.3 | 1.7 | GO:0030021 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.3 | 2.2 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.3 | 4.9 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.3 | 4.1 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.3 | 3.8 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.3 | 8.5 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.3 | 2.6 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.3 | 5.5 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.3 | 5.1 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.3 | 2.0 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.2 | 5.4 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.2 | 3.2 | GO:0019864 | IgG binding(GO:0019864) |
| 0.2 | 2.2 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.2 | 1.7 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.2 | 2.9 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.2 | 1.2 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.2 | 0.7 | GO:0097258 | tocopherol omega-hydroxylase activity(GO:0052870) alpha-tocopherol omega-hydroxylase activity(GO:0052871) 20-hydroxy-leukotriene B4 omega oxidase activity(GO:0097258) 20-aldehyde-leukotriene B4 20-monooxygenase activity(GO:0097259) |
| 0.2 | 4.6 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.2 | 3.0 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.2 | 2.3 | GO:0015385 | sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.2 | 2.5 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.2 | 2.7 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.2 | 0.7 | GO:0050528 | acyloxyacyl hydrolase activity(GO:0050528) |
| 0.2 | 24.1 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.2 | 3.4 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.2 | 0.8 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.2 | 3.7 | GO:0070513 | death domain binding(GO:0070513) |
| 0.2 | 4.8 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.2 | 1.6 | GO:0050694 | galactosylceramide sulfotransferase activity(GO:0001733) galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.2 | 2.6 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.2 | 0.8 | GO:0051734 | ATP-dependent polydeoxyribonucleotide 5'-hydroxyl-kinase activity(GO:0046404) polydeoxyribonucleotide kinase activity(GO:0051733) ATP-dependent polynucleotide kinase activity(GO:0051734) |
| 0.2 | 1.8 | GO:0004340 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.2 | 1.3 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.2 | 0.9 | GO:0004447 | iodide peroxidase activity(GO:0004447) |
| 0.2 | 0.2 | GO:0070336 | flap-structured DNA binding(GO:0070336) |
| 0.2 | 0.7 | GO:0001602 | pancreatic polypeptide receptor activity(GO:0001602) |
| 0.2 | 1.8 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.2 | 1.4 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 0.2 | 3.4 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
| 0.2 | 1.6 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.2 | 1.6 | GO:0030884 | lipid antigen binding(GO:0030882) endogenous lipid antigen binding(GO:0030883) exogenous lipid antigen binding(GO:0030884) |
| 0.2 | 1.9 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.2 | 1.9 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.2 | 3.8 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.2 | 1.0 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.2 | 8.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.2 | 3.5 | GO:0008175 | tRNA methyltransferase activity(GO:0008175) |
| 0.2 | 5.7 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.2 | 1.0 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.2 | 0.5 | GO:0030290 | sphingolipid activator protein activity(GO:0030290) beta-N-acetylgalactosaminidase activity(GO:0032428) |
| 0.2 | 0.8 | GO:0004370 | glycerol kinase activity(GO:0004370) |
| 0.1 | 2.5 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.1 | 3.2 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.1 | 1.3 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.1 | 1.4 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.1 | 2.6 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.1 | 3.3 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.1 | 3.9 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 0.6 | GO:0046899 | nucleoside triphosphate adenylate kinase activity(GO:0046899) |
| 0.1 | 11.1 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 2.6 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 2.6 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.1 | 2.4 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.1 | 0.9 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.1 | 9.0 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.1 | 6.6 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.1 | 2.2 | GO:0031005 | filamin binding(GO:0031005) |
| 0.1 | 3.5 | GO:0008188 | neuropeptide receptor activity(GO:0008188) |
| 0.1 | 2.2 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.1 | 2.7 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.1 | 0.9 | GO:0016724 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.1 | 29.6 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 1.0 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 2.2 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.1 | 0.3 | GO:1902271 | D3 vitamins binding(GO:1902271) |
| 0.1 | 1.3 | GO:0017127 | cholesterol transporter activity(GO:0017127) |
| 0.1 | 17.5 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.1 | 2.8 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.1 | 3.1 | GO:0047555 | 3',5'-cyclic-GMP phosphodiesterase activity(GO:0047555) |
| 0.1 | 0.3 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 0.1 | 0.7 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.1 | 9.5 | GO:0101005 | thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.1 | 1.0 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.1 | 3.1 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.1 | 4.7 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 9.6 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.1 | 11.9 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.1 | 0.8 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.1 | 0.9 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.1 | 1.7 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.1 | 8.1 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.1 | 5.6 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.1 | 6.4 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 2.7 | GO:1901476 | carbohydrate transmembrane transporter activity(GO:0015144) carbohydrate transporter activity(GO:1901476) |
| 0.1 | 0.1 | GO:0042954 | lipoprotein transporter activity(GO:0042954) |
| 0.1 | 5.8 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.1 | 1.1 | GO:0008252 | nucleotidase activity(GO:0008252) |
| 0.1 | 5.6 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.1 | 1.9 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 1.6 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.1 | 2.1 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.1 | 1.4 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.1 | 2.6 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.1 | 3.1 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 3.2 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.1 | 5.3 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.1 | 1.3 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 0.7 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.1 | 1.1 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 2.4 | GO:0042805 | actinin binding(GO:0042805) |
| 0.1 | 3.9 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.1 | 5.6 | GO:0008201 | heparin binding(GO:0008201) |
| 0.1 | 0.8 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 7.8 | GO:0016810 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds(GO:0016810) |
| 0.0 | 2.5 | GO:0050681 | androgen receptor binding(GO:0050681) |
| 0.0 | 0.5 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.0 | 1.0 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.6 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.0 | 3.2 | GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds(GO:0004553) |
| 0.0 | 1.3 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.0 | 1.4 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.0 | 28.5 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 1.0 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.0 | 1.3 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 1.8 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 0.3 | GO:0030020 | extracellular matrix structural constituent conferring tensile strength(GO:0030020) |
| 0.0 | 2.4 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.0 | 1.2 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.0 | 0.4 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.0 | 1.6 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.0 | 0.1 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.0 | 1.1 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 3.2 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 0.3 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.0 | 3.2 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 0.8 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.3 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.0 | 0.9 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.0 | 1.5 | GO:0015297 | antiporter activity(GO:0015297) |
| 0.0 | 3.0 | GO:0005057 | receptor signaling protein activity(GO:0005057) |
| 0.0 | 0.2 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.0 | 0.8 | GO:0008237 | metallopeptidase activity(GO:0008237) |
| 0.0 | 0.1 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.0 | 0.1 | GO:0070697 | activin receptor binding(GO:0070697) type II activin receptor binding(GO:0070699) |
| 0.0 | 0.9 | GO:0043130 | ubiquitin binding(GO:0043130) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 32.8 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.3 | 16.5 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.3 | 18.3 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.2 | 3.6 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.2 | 51.4 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.2 | 1.3 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.2 | 8.4 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.2 | 3.6 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.2 | 9.1 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.2 | 5.9 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.2 | 4.6 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.2 | 3.3 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 3.0 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.1 | 4.6 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.1 | 8.2 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 10.6 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 1.7 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.1 | 4.1 | PID S1P S1P3 PATHWAY | S1P3 pathway |
| 0.1 | 3.5 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 3.9 | PID SHP2 PATHWAY | SHP2 signaling |
| 0.1 | 2.0 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.1 | 2.8 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.1 | 6.1 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 1.2 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.1 | 0.9 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.1 | 1.5 | SIG IL4RECEPTOR IN B LYPHOCYTES | Genes related to IL4 rceptor signaling in B lymphocytes |
| 0.1 | 5.8 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.1 | 13.9 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.1 | 4.9 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.1 | 4.7 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 1.0 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.1 | 2.9 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.1 | 2.1 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.1 | 3.5 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.1 | 4.6 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.1 | 2.5 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 4.7 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.1 | 2.4 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 1.3 | PID IL23 PATHWAY | IL23-mediated signaling events |
| 0.1 | 0.8 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.1 | 4.1 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.1 | 2.6 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 1.3 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.1 | 1.3 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.1 | 2.1 | PID FGF PATHWAY | FGF signaling pathway |
| 0.1 | 4.2 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 2.1 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 3.0 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 2.1 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 1.8 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.0 | 1.0 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 6.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.4 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 1.5 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 0.1 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 36.0 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.9 | 20.8 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.7 | 11.6 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.7 | 6.4 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.7 | 13.9 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.6 | 4.4 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.6 | 14.1 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.6 | 12.8 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.6 | 8.5 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.6 | 11.8 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.5 | 11.5 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.5 | 10.2 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.5 | 22.0 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.5 | 19.4 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.4 | 11.2 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.4 | 7.2 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.4 | 9.2 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.4 | 7.9 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.4 | 5.9 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.4 | 8.2 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.4 | 4.1 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.4 | 6.2 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.3 | 3.7 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.3 | 3.1 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.3 | 4.0 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.3 | 2.8 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.3 | 6.7 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.3 | 10.7 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.2 | 2.7 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.2 | 12.4 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.2 | 18.9 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.2 | 4.1 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.2 | 5.2 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.2 | 7.7 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.2 | 2.5 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.2 | 6.9 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.2 | 2.2 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.2 | 2.0 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.2 | 1.4 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.2 | 1.7 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.2 | 3.8 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.2 | 3.6 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.2 | 13.2 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.2 | 6.0 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.2 | 4.6 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.2 | 4.9 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.1 | 9.2 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 2.7 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.1 | 10.2 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 12.7 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.1 | 2.6 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 1.5 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.1 | 4.4 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.1 | 2.1 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 0.9 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.1 | 5.2 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.1 | 1.1 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.1 | 1.8 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.1 | 2.1 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.1 | 1.3 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.1 | 5.1 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.1 | 2.0 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.1 | 6.1 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 4.9 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 2.2 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.1 | 2.2 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.1 | 3.1 | REACTOME PLATELET AGGREGATION PLUG FORMATION | Genes involved in Platelet Aggregation (Plug Formation) |
| 0.1 | 4.6 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 3.2 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 4.3 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.1 | 1.6 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 2.0 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.1 | 2.5 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.1 | 1.8 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.1 | 2.1 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 1.8 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.1 | 0.7 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 2.7 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.1 | 1.1 | REACTOME GABA B RECEPTOR ACTIVATION | Genes involved in GABA B receptor activation |
| 0.1 | 8.3 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.1 | 0.5 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.0 | 1.1 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 3.0 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 3.7 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.0 | 0.9 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.0 | 1.8 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 3.0 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 1.3 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.0 | 0.7 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 0.0 | 0.3 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.0 | 0.8 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.6 | REACTOME SIGNALING BY SCF KIT | Genes involved in Signaling by SCF-KIT |
| 0.0 | 2.9 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.5 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.0 | 1.4 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 1.8 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 2.2 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 1.2 | REACTOME SIGNALING BY PDGF | Genes involved in Signaling by PDGF |
| 0.0 | 1.3 | REACTOME NGF SIGNALLING VIA TRKA FROM THE PLASMA MEMBRANE | Genes involved in NGF signalling via TRKA from the plasma membrane |
| 0.0 | 0.1 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.0 | 0.1 | REACTOME MEIOSIS | Genes involved in Meiosis |