Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
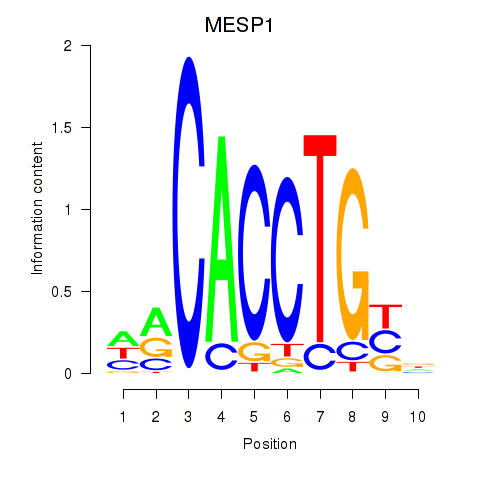
Results for MESP1
Z-value: 1.72
Transcription factors associated with MESP1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
MESP1
|
ENSG00000166823.5 | MESP1 |
Activity profile of MESP1 motif
Sorted Z-values of MESP1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of MESP1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 28.5 | 114.2 | GO:1903609 | negative regulation of peptidyl-tyrosine autophosphorylation(GO:1900085) negative regulation of inward rectifier potassium channel activity(GO:1903609) |
| 12.5 | 37.5 | GO:0061163 | endoplasmic reticulum polarization(GO:0061163) actin filament bundle retrograde transport(GO:0061573) actin filament bundle distribution(GO:0070650) |
| 12.3 | 74.0 | GO:0007296 | vitellogenesis(GO:0007296) |
| 9.0 | 45.1 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 8.9 | 26.6 | GO:1902269 | positive regulation of polyamine transmembrane transport(GO:1902269) |
| 8.1 | 24.3 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 7.9 | 23.7 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 7.7 | 38.7 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 6.4 | 38.4 | GO:0051563 | smooth endoplasmic reticulum calcium ion homeostasis(GO:0051563) |
| 6.1 | 18.4 | GO:0021592 | fourth ventricle development(GO:0021592) third ventricle development(GO:0021678) |
| 4.8 | 43.6 | GO:0010994 | regulation of ubiquitin homeostasis(GO:0010993) free ubiquitin chain polymerization(GO:0010994) positive regulation of exit from mitosis(GO:0031536) |
| 4.8 | 62.8 | GO:1900028 | negative regulation of ruffle assembly(GO:1900028) |
| 4.8 | 62.7 | GO:1900623 | regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 4.5 | 17.9 | GO:0031179 | peptide amidation(GO:0001519) protein amidation(GO:0018032) peptide modification(GO:0031179) |
| 4.3 | 17.3 | GO:0070295 | renal water absorption(GO:0070295) |
| 4.3 | 12.9 | GO:0055014 | atrial cardiac muscle cell differentiation(GO:0055011) atrial cardiac muscle cell development(GO:0055014) |
| 4.0 | 16.1 | GO:0035750 | protein localization to myelin sheath abaxonal region(GO:0035750) |
| 3.9 | 11.7 | GO:0036333 | hepatocyte homeostasis(GO:0036333) response to tetrachloromethane(GO:1904772) |
| 3.9 | 50.3 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 3.8 | 11.4 | GO:0090155 | negative regulation of sphingolipid biosynthetic process(GO:0090155) negative regulation of ceramide biosynthetic process(GO:1900060) |
| 3.6 | 46.6 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 3.5 | 24.6 | GO:0048050 | post-embryonic eye morphogenesis(GO:0048050) |
| 3.5 | 20.7 | GO:0048549 | positive regulation of pinocytosis(GO:0048549) |
| 3.4 | 20.6 | GO:0003164 | His-Purkinje system development(GO:0003164) |
| 3.3 | 9.9 | GO:1901503 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) ether biosynthetic process(GO:1901503) |
| 3.2 | 9.6 | GO:0060739 | mesenchymal-epithelial cell signaling involved in prostate gland development(GO:0060739) |
| 3.2 | 9.5 | GO:1904693 | midbrain morphogenesis(GO:1904693) |
| 3.1 | 12.5 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 3.0 | 21.2 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 3.0 | 14.9 | GO:0009240 | isopentenyl diphosphate biosynthetic process(GO:0009240) isopentenyl diphosphate metabolic process(GO:0046490) |
| 2.8 | 8.5 | GO:0043311 | positive regulation of eosinophil degranulation(GO:0043311) positive regulation of eosinophil activation(GO:1902568) |
| 2.8 | 8.5 | GO:0018160 | peptidyl-pyrromethane cofactor linkage(GO:0018160) |
| 2.8 | 8.4 | GO:1903490 | regulation of cytokinetic process(GO:0032954) regulation of mitotic cytokinetic process(GO:1903436) positive regulation of mitotic cytokinetic process(GO:1903438) positive regulation of mitotic cytokinesis(GO:1903490) |
| 2.7 | 74.8 | GO:0034063 | stress granule assembly(GO:0034063) |
| 2.6 | 28.1 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 2.5 | 7.5 | GO:1901388 | regulation of transforming growth factor beta activation(GO:1901388) negative regulation of transforming growth factor beta activation(GO:1901389) |
| 2.5 | 7.5 | GO:0036111 | very long-chain fatty-acyl-CoA metabolic process(GO:0036111) |
| 2.5 | 17.4 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 2.5 | 12.4 | GO:1902361 | mitochondrial pyruvate transport(GO:0006850) mitochondrial pyruvate transmembrane transport(GO:1902361) |
| 2.4 | 7.3 | GO:2000360 | negative regulation of binding of sperm to zona pellucida(GO:2000360) |
| 2.4 | 7.1 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 2.4 | 49.4 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 2.3 | 23.4 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 2.3 | 18.4 | GO:2000483 | negative regulation of interleukin-8 secretion(GO:2000483) |
| 2.3 | 15.9 | GO:0015866 | ADP transport(GO:0015866) |
| 2.2 | 10.9 | GO:0042374 | phylloquinone metabolic process(GO:0042374) phylloquinone catabolic process(GO:0042376) quinone catabolic process(GO:1901662) |
| 2.2 | 24.0 | GO:0006552 | leucine catabolic process(GO:0006552) |
| 2.1 | 17.1 | GO:1902961 | positive regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902961) positive regulation of aspartic-type peptidase activity(GO:1905247) |
| 2.1 | 25.4 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 2.1 | 12.7 | GO:0048254 | snoRNA localization(GO:0048254) |
| 2.1 | 14.6 | GO:0006990 | positive regulation of transcription from RNA polymerase II promoter involved in unfolded protein response(GO:0006990) |
| 2.1 | 2.1 | GO:0060920 | cardiac pacemaker cell differentiation(GO:0060920) cardiac pacemaker cell development(GO:0060926) |
| 2.1 | 10.4 | GO:0030242 | pexophagy(GO:0030242) |
| 2.0 | 18.1 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 2.0 | 10.0 | GO:0002439 | chronic inflammatory response to antigenic stimulus(GO:0002439) |
| 2.0 | 5.9 | GO:0015746 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 1.9 | 9.7 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 1.9 | 15.1 | GO:0046836 | glycolipid transport(GO:0046836) |
| 1.9 | 16.9 | GO:0006167 | AMP biosynthetic process(GO:0006167) |
| 1.9 | 7.5 | GO:0033606 | B cell receptor transport within lipid bilayer(GO:0032595) B cell receptor transport into membrane raft(GO:0032597) protein transport out of membrane raft(GO:0032599) chemokine receptor transport out of membrane raft(GO:0032600) negative regulation of transforming growth factor beta3 production(GO:0032913) chemokine receptor transport within lipid bilayer(GO:0033606) |
| 1.9 | 37.5 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 1.8 | 16.5 | GO:0006930 | substrate-dependent cell migration, cell extension(GO:0006930) |
| 1.8 | 23.7 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 1.8 | 16.0 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 1.8 | 8.8 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 1.7 | 10.4 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 1.7 | 10.2 | GO:1903862 | positive regulation of oxidative phosphorylation(GO:1903862) |
| 1.7 | 126.1 | GO:0032981 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 1.7 | 8.3 | GO:0043126 | regulation of 1-phosphatidylinositol 4-kinase activity(GO:0043126) positive regulation of 1-phosphatidylinositol 4-kinase activity(GO:0043128) |
| 1.7 | 8.3 | GO:0010286 | heat acclimation(GO:0010286) cellular heat acclimation(GO:0070370) |
| 1.6 | 18.1 | GO:0007144 | female meiosis I(GO:0007144) |
| 1.6 | 8.2 | GO:0015862 | uridine transport(GO:0015862) |
| 1.6 | 47.3 | GO:0031116 | positive regulation of microtubule polymerization(GO:0031116) |
| 1.6 | 13.0 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 1.6 | 67.9 | GO:0097284 | hepatocyte apoptotic process(GO:0097284) |
| 1.6 | 4.7 | GO:1903826 | arginine transmembrane transport(GO:1903826) |
| 1.6 | 26.9 | GO:0003334 | keratinocyte development(GO:0003334) |
| 1.6 | 7.9 | GO:0006933 | negative regulation of cell adhesion involved in substrate-bound cell migration(GO:0006933) |
| 1.5 | 9.3 | GO:0006050 | mannosamine metabolic process(GO:0006050) N-acetylmannosamine metabolic process(GO:0006051) |
| 1.5 | 7.5 | GO:0030309 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 1.5 | 4.5 | GO:0016340 | calcium-dependent cell-matrix adhesion(GO:0016340) regulation of embryonic cell shape(GO:0016476) |
| 1.5 | 21.0 | GO:0043562 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 1.5 | 16.5 | GO:0048194 | Golgi vesicle budding(GO:0048194) |
| 1.5 | 42.9 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 1.5 | 7.3 | GO:0042695 | thelarche(GO:0042695) mammary gland branching involved in thelarche(GO:0060744) |
| 1.5 | 5.8 | GO:0072396 | response to cell cycle checkpoint signaling(GO:0072396) response to DNA integrity checkpoint signaling(GO:0072402) response to DNA damage checkpoint signaling(GO:0072423) |
| 1.4 | 28.9 | GO:1904816 | positive regulation of protein localization to chromosome, telomeric region(GO:1904816) |
| 1.4 | 7.1 | GO:1901350 | cell-cell signaling involved in cell-cell junction organization(GO:1901350) |
| 1.4 | 9.9 | GO:0035879 | plasma membrane lactate transport(GO:0035879) |
| 1.4 | 5.6 | GO:0043314 | negative regulation of neutrophil degranulation(GO:0043314) |
| 1.4 | 5.6 | GO:0051611 | negative regulation of neurotransmitter uptake(GO:0051581) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 1.4 | 15.2 | GO:0032914 | positive regulation of transforming growth factor beta1 production(GO:0032914) |
| 1.4 | 55.6 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 1.4 | 1.4 | GO:0060467 | negative regulation of fertilization(GO:0060467) |
| 1.3 | 20.0 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 1.3 | 12.0 | GO:1903003 | positive regulation of protein deubiquitination(GO:1903003) |
| 1.3 | 50.4 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 1.3 | 2.6 | GO:0010728 | regulation of hydrogen peroxide biosynthetic process(GO:0010728) |
| 1.3 | 13.0 | GO:0070863 | positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 1.3 | 6.4 | GO:0003056 | regulation of vascular smooth muscle contraction(GO:0003056) |
| 1.3 | 6.4 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 1.3 | 3.8 | GO:1903926 | signal transduction involved in intra-S DNA damage checkpoint(GO:0072428) response to bisphenol A(GO:1903925) cellular response to bisphenol A(GO:1903926) |
| 1.3 | 12.7 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 1.3 | 11.3 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 1.2 | 7.4 | GO:0014835 | myoblast differentiation involved in skeletal muscle regeneration(GO:0014835) positive regulation of interleukin-12 secretion(GO:2001184) |
| 1.2 | 6.1 | GO:0006041 | glucosamine metabolic process(GO:0006041) |
| 1.2 | 6.1 | GO:0018197 | peptidyl-aspartic acid modification(GO:0018197) |
| 1.2 | 9.4 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 1.2 | 4.7 | GO:0043387 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) regulation of glutathione biosynthetic process(GO:1903786) positive regulation of glutathione biosynthetic process(GO:1903788) |
| 1.2 | 7.0 | GO:1901164 | negative regulation of trophoblast cell migration(GO:1901164) |
| 1.1 | 10.2 | GO:1902416 | positive regulation of mRNA binding(GO:1902416) |
| 1.1 | 14.5 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 1.1 | 8.6 | GO:0009597 | detection of virus(GO:0009597) |
| 1.1 | 5.4 | GO:1902513 | regulation of organelle transport along microtubule(GO:1902513) |
| 1.1 | 5.3 | GO:2000542 | negative regulation of gastrulation(GO:2000542) |
| 1.1 | 3.2 | GO:0038195 | urokinase plasminogen activator signaling pathway(GO:0038195) |
| 1.0 | 7.3 | GO:1990592 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 1.0 | 6.2 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 1.0 | 12.3 | GO:0010668 | ectodermal cell differentiation(GO:0010668) |
| 1.0 | 15.3 | GO:1903760 | regulation of voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1903760) |
| 1.0 | 42.9 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 1.0 | 4.9 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 1.0 | 12.6 | GO:0090084 | negative regulation of inclusion body assembly(GO:0090084) |
| 0.9 | 6.5 | GO:0046726 | positive regulation by virus of viral protein levels in host cell(GO:0046726) |
| 0.9 | 6.5 | GO:0042421 | norepinephrine biosynthetic process(GO:0042421) |
| 0.9 | 19.4 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.9 | 13.7 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.9 | 5.4 | GO:1902231 | positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage(GO:1902231) |
| 0.9 | 5.3 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.9 | 22.0 | GO:2000637 | positive regulation of posttranscriptional gene silencing(GO:0060148) positive regulation of gene silencing by miRNA(GO:2000637) |
| 0.9 | 22.6 | GO:0045948 | positive regulation of translational initiation(GO:0045948) |
| 0.9 | 39.9 | GO:0048255 | mRNA stabilization(GO:0048255) |
| 0.9 | 2.6 | GO:0070902 | mitochondrial tRNA pseudouridine synthesis(GO:0070902) |
| 0.9 | 2.6 | GO:0010693 | negative regulation of alkaline phosphatase activity(GO:0010693) uterine wall breakdown(GO:0042704) |
| 0.8 | 3.4 | GO:0010637 | negative regulation of mitochondrial fusion(GO:0010637) |
| 0.8 | 0.8 | GO:0098917 | retrograde trans-synaptic signaling(GO:0098917) |
| 0.8 | 1.7 | GO:0009912 | auditory receptor cell fate commitment(GO:0009912) inner ear receptor cell fate commitment(GO:0060120) |
| 0.8 | 2.5 | GO:0002752 | cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 0.8 | 3.2 | GO:0034627 | nicotinamide nucleotide biosynthetic process from aspartate(GO:0019355) 'de novo' NAD biosynthetic process(GO:0034627) 'de novo' NAD biosynthetic process from aspartate(GO:0034628) |
| 0.8 | 3.2 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.8 | 3.2 | GO:0072683 | T cell extravasation(GO:0072683) |
| 0.8 | 8.7 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.8 | 7.9 | GO:1902669 | positive regulation of axon guidance(GO:1902669) |
| 0.8 | 6.2 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.8 | 3.1 | GO:1990637 | response to prolactin(GO:1990637) |
| 0.8 | 7.7 | GO:0045716 | positive regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045716) |
| 0.8 | 84.0 | GO:0061418 | regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061418) |
| 0.8 | 3.1 | GO:1904685 | positive regulation of metalloendopeptidase activity(GO:1904685) |
| 0.8 | 11.3 | GO:0034975 | protein folding in endoplasmic reticulum(GO:0034975) |
| 0.7 | 9.7 | GO:0034134 | toll-like receptor 2 signaling pathway(GO:0034134) |
| 0.7 | 4.4 | GO:0060574 | intestinal epithelial cell maturation(GO:0060574) |
| 0.7 | 8.0 | GO:0070234 | positive regulation of T cell apoptotic process(GO:0070234) |
| 0.7 | 8.7 | GO:0048711 | positive regulation of astrocyte differentiation(GO:0048711) |
| 0.7 | 7.1 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.7 | 4.1 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 0.7 | 4.0 | GO:1903243 | negative regulation of cardiac muscle adaptation(GO:0010616) negative regulation of lung blood pressure(GO:0061767) negative regulation of cardiac muscle hypertrophy in response to stress(GO:1903243) |
| 0.7 | 4.0 | GO:0089700 | protein kinase D signaling(GO:0089700) positive regulation of histone deacetylase activity(GO:1901727) |
| 0.7 | 2.7 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 0.7 | 13.9 | GO:0097062 | dendritic spine maintenance(GO:0097062) |
| 0.6 | 7.8 | GO:0030050 | vesicle transport along actin filament(GO:0030050) |
| 0.6 | 4.4 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.6 | 3.8 | GO:0002191 | cap-dependent translational initiation(GO:0002191) |
| 0.6 | 2.5 | GO:0003025 | regulation of systemic arterial blood pressure by baroreceptor feedback(GO:0003025) |
| 0.6 | 1.9 | GO:0034499 | late endosome to Golgi transport(GO:0034499) |
| 0.6 | 13.0 | GO:0009162 | deoxyribonucleoside monophosphate metabolic process(GO:0009162) |
| 0.6 | 13.6 | GO:0043457 | regulation of cellular respiration(GO:0043457) |
| 0.6 | 5.5 | GO:0060125 | negative regulation of growth hormone secretion(GO:0060125) |
| 0.6 | 10.4 | GO:0042473 | outer ear morphogenesis(GO:0042473) |
| 0.6 | 2.4 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.6 | 1.2 | GO:0006272 | leading strand elongation(GO:0006272) |
| 0.6 | 14.9 | GO:0042753 | positive regulation of circadian rhythm(GO:0042753) |
| 0.6 | 2.4 | GO:0035407 | histone H3-T11 phosphorylation(GO:0035407) |
| 0.6 | 1.8 | GO:0071109 | superior temporal gyrus development(GO:0071109) |
| 0.6 | 4.1 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.6 | 5.3 | GO:0098909 | regulation of cardiac muscle cell action potential involved in regulation of contraction(GO:0098909) |
| 0.6 | 5.8 | GO:0031953 | negative regulation of protein autophosphorylation(GO:0031953) |
| 0.6 | 19.7 | GO:0035066 | positive regulation of histone acetylation(GO:0035066) |
| 0.6 | 6.3 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.6 | 2.8 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.6 | 13.3 | GO:0042347 | negative regulation of NF-kappaB import into nucleus(GO:0042347) |
| 0.6 | 0.6 | GO:0060129 | thyroid-stimulating hormone-secreting cell differentiation(GO:0060129) |
| 0.6 | 1.7 | GO:0045062 | extrathymic T cell selection(GO:0045062) |
| 0.5 | 6.0 | GO:0070424 | regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070424) |
| 0.5 | 1.6 | GO:0003099 | positive regulation of the force of heart contraction by chemical signal(GO:0003099) |
| 0.5 | 8.7 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) |
| 0.5 | 3.8 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.5 | 4.3 | GO:1990416 | cellular response to brain-derived neurotrophic factor stimulus(GO:1990416) |
| 0.5 | 2.2 | GO:0018199 | peptidyl-glutamine modification(GO:0018199) |
| 0.5 | 8.3 | GO:0090190 | positive regulation of branching involved in ureteric bud morphogenesis(GO:0090190) |
| 0.5 | 1.0 | GO:0010986 | positive regulation of lipoprotein particle clearance(GO:0010986) |
| 0.5 | 2.0 | GO:0015868 | purine nucleotide transport(GO:0015865) purine ribonucleotide transport(GO:0015868) |
| 0.5 | 7.6 | GO:0038166 | angiotensin-activated signaling pathway(GO:0038166) |
| 0.5 | 2.5 | GO:1903553 | positive regulation of extracellular exosome assembly(GO:1903553) |
| 0.5 | 3.5 | GO:0003402 | planar cell polarity pathway involved in axis elongation(GO:0003402) |
| 0.5 | 6.5 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.5 | 14.2 | GO:0048745 | smooth muscle tissue development(GO:0048745) |
| 0.5 | 6.7 | GO:0035826 | rubidium ion transport(GO:0035826) regulation of rubidium ion transport(GO:2000680) |
| 0.5 | 13.5 | GO:0006625 | protein targeting to peroxisome(GO:0006625) protein localization to peroxisome(GO:0072662) establishment of protein localization to peroxisome(GO:0072663) |
| 0.5 | 12.4 | GO:0006379 | mRNA cleavage(GO:0006379) |
| 0.4 | 1.8 | GO:0033690 | positive regulation of osteoblast proliferation(GO:0033690) retinal blood vessel morphogenesis(GO:0061304) |
| 0.4 | 1.8 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.4 | 10.7 | GO:0010800 | positive regulation of peptidyl-threonine phosphorylation(GO:0010800) |
| 0.4 | 6.2 | GO:0030497 | fatty acid elongation(GO:0030497) very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.4 | 0.4 | GO:1901420 | negative regulation of response to alcohol(GO:1901420) |
| 0.4 | 4.8 | GO:0007379 | segment specification(GO:0007379) |
| 0.4 | 16.9 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.4 | 9.1 | GO:0046040 | IMP metabolic process(GO:0046040) |
| 0.4 | 15.3 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.4 | 11.9 | GO:0046475 | glycerophospholipid catabolic process(GO:0046475) |
| 0.4 | 13.2 | GO:0008334 | histone mRNA metabolic process(GO:0008334) |
| 0.4 | 2.4 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.4 | 2.8 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 0.4 | 2.0 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.4 | 8.4 | GO:0032486 | Rap protein signal transduction(GO:0032486) |
| 0.4 | 1.6 | GO:0003409 | optic cup structural organization(GO:0003409) |
| 0.4 | 11.0 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.4 | 10.6 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.4 | 3.8 | GO:0060056 | mammary gland involution(GO:0060056) |
| 0.4 | 3.4 | GO:0006335 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.4 | 2.6 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.4 | 2.5 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.4 | 3.2 | GO:0007042 | lysosomal lumen acidification(GO:0007042) |
| 0.3 | 1.4 | GO:0032848 | negative regulation of cellular pH reduction(GO:0032848) CD8-positive, alpha-beta T cell lineage commitment(GO:0043375) negative regulation of retinal cell programmed cell death(GO:0046671) |
| 0.3 | 3.1 | GO:0035745 | T-helper 2 cell cytokine production(GO:0035745) |
| 0.3 | 1.7 | GO:1902202 | regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 0.3 | 6.1 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.3 | 3.0 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.3 | 3.0 | GO:0032286 | central nervous system myelin maintenance(GO:0032286) |
| 0.3 | 2.3 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) |
| 0.3 | 8.6 | GO:0005980 | glycogen catabolic process(GO:0005980) |
| 0.3 | 9.2 | GO:0001913 | T cell mediated cytotoxicity(GO:0001913) |
| 0.3 | 11.1 | GO:0070265 | necrotic cell death(GO:0070265) |
| 0.3 | 5.8 | GO:0006002 | fructose 6-phosphate metabolic process(GO:0006002) |
| 0.3 | 15.7 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.3 | 11.5 | GO:0070306 | lens fiber cell differentiation(GO:0070306) |
| 0.3 | 3.5 | GO:0034244 | negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.3 | 6.8 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.3 | 16.5 | GO:0033683 | nucleotide-excision repair, DNA incision(GO:0033683) |
| 0.3 | 19.7 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.3 | 7.8 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.3 | 3.7 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.3 | 1.4 | GO:0061743 | motor learning(GO:0061743) |
| 0.3 | 2.0 | GO:1902031 | regulation of NADP metabolic process(GO:1902031) |
| 0.3 | 8.8 | GO:0045745 | positive regulation of G-protein coupled receptor protein signaling pathway(GO:0045745) |
| 0.3 | 4.0 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.3 | 1.0 | GO:0003285 | septum secundum development(GO:0003285) atrial septum secundum morphogenesis(GO:0003290) |
| 0.3 | 6.8 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.3 | 25.2 | GO:0006413 | translational initiation(GO:0006413) |
| 0.3 | 0.5 | GO:0060313 | negative regulation of blood vessel remodeling(GO:0060313) |
| 0.3 | 0.5 | GO:0045631 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 0.2 | 3.2 | GO:2000052 | positive regulation of non-canonical Wnt signaling pathway(GO:2000052) |
| 0.2 | 5.4 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.2 | 6.1 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.2 | 22.1 | GO:0006415 | translational termination(GO:0006415) |
| 0.2 | 4.4 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.2 | 3.4 | GO:0060575 | intestinal epithelial cell differentiation(GO:0060575) |
| 0.2 | 6.0 | GO:1904376 | negative regulation of protein localization to plasma membrane(GO:1903077) negative regulation of protein localization to cell periphery(GO:1904376) |
| 0.2 | 0.6 | GO:2000504 | endocardial cushion to mesenchymal transition involved in heart valve formation(GO:0003199) positive regulation of blood vessel remodeling(GO:2000504) |
| 0.2 | 4.5 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) |
| 0.2 | 0.6 | GO:0045974 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.2 | 1.0 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.2 | 4.6 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.2 | 1.1 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.2 | 3.2 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.2 | 2.2 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.2 | 73.2 | GO:0043687 | post-translational protein modification(GO:0043687) |
| 0.2 | 1.3 | GO:0098908 | regulation of neuronal action potential(GO:0098908) |
| 0.2 | 0.7 | GO:0098957 | anterograde axonal transport of mitochondrion(GO:0098957) |
| 0.2 | 3.9 | GO:0007143 | female meiotic division(GO:0007143) |
| 0.2 | 1.9 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.2 | 1.5 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.2 | 0.7 | GO:0035811 | negative regulation of urine volume(GO:0035811) |
| 0.2 | 5.0 | GO:0006370 | 7-methylguanosine mRNA capping(GO:0006370) |
| 0.2 | 10.8 | GO:0071349 | interleukin-12-mediated signaling pathway(GO:0035722) response to interleukin-12(GO:0070671) cellular response to interleukin-12(GO:0071349) |
| 0.2 | 11.4 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.2 | 1.3 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.2 | 2.3 | GO:0060445 | branching involved in salivary gland morphogenesis(GO:0060445) |
| 0.1 | 6.0 | GO:0060071 | Wnt signaling pathway, planar cell polarity pathway(GO:0060071) |
| 0.1 | 7.8 | GO:0072583 | clathrin-mediated endocytosis(GO:0072583) |
| 0.1 | 0.6 | GO:0007028 | cytoplasm organization(GO:0007028) |
| 0.1 | 1.4 | GO:0036309 | T-tubule organization(GO:0033292) protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.1 | 6.7 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.1 | 3.9 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.1 | 0.8 | GO:0070257 | positive regulation of mucus secretion(GO:0070257) |
| 0.1 | 3.0 | GO:0006582 | melanin metabolic process(GO:0006582) melanin biosynthetic process(GO:0042438) |
| 0.1 | 1.0 | GO:0032310 | prostaglandin secretion(GO:0032310) |
| 0.1 | 2.0 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.1 | 0.3 | GO:0050928 | negative regulation of positive chemotaxis(GO:0050928) |
| 0.1 | 2.0 | GO:0001829 | trophectodermal cell differentiation(GO:0001829) |
| 0.1 | 1.6 | GO:0019852 | L-ascorbic acid metabolic process(GO:0019852) |
| 0.1 | 3.6 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.1 | 9.1 | GO:0007368 | determination of left/right symmetry(GO:0007368) |
| 0.1 | 3.3 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.1 | 17.2 | GO:0006613 | cotranslational protein targeting to membrane(GO:0006613) |
| 0.1 | 34.5 | GO:0016050 | vesicle organization(GO:0016050) |
| 0.1 | 2.2 | GO:0099514 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.1 | 7.1 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.1 | 8.9 | GO:0006493 | protein O-linked glycosylation(GO:0006493) |
| 0.1 | 0.6 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.1 | 0.5 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.1 | 2.4 | GO:1901998 | toxin transport(GO:1901998) |
| 0.1 | 2.1 | GO:0032469 | endoplasmic reticulum calcium ion homeostasis(GO:0032469) |
| 0.1 | 1.3 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.1 | 2.6 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.1 | 1.0 | GO:0032801 | receptor catabolic process(GO:0032801) |
| 0.1 | 0.8 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.1 | 0.7 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.1 | 4.7 | GO:0021766 | hippocampus development(GO:0021766) |
| 0.1 | 0.1 | GO:0003221 | right ventricular cardiac muscle tissue morphogenesis(GO:0003221) |
| 0.1 | 0.2 | GO:0006117 | acetaldehyde metabolic process(GO:0006117) |
| 0.1 | 2.9 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 0.6 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.1 | 8.4 | GO:0006818 | hydrogen transport(GO:0006818) |
| 0.1 | 14.5 | GO:0016042 | lipid catabolic process(GO:0016042) |
| 0.1 | 0.3 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.0 | 1.6 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.0 | 0.1 | GO:1904350 | regulation of protein catabolic process in the vacuole(GO:1904350) positive regulation of protein catabolic process in the vacuole(GO:1904352) |
| 0.0 | 1.1 | GO:0006517 | protein deglycosylation(GO:0006517) |
| 0.0 | 1.5 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.2 | GO:0021894 | cerebral cortex GABAergic interneuron development(GO:0021894) |
| 0.0 | 1.2 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.0 | 2.0 | GO:0032508 | DNA duplex unwinding(GO:0032508) |
| 0.0 | 0.3 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.0 | 0.0 | GO:0032425 | regulation of mismatch repair(GO:0032423) positive regulation of mismatch repair(GO:0032425) |
| 0.0 | 0.3 | GO:0019720 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) molybdopterin cofactor biosynthetic process(GO:0032324) molybdopterin cofactor metabolic process(GO:0043545) prosthetic group metabolic process(GO:0051189) |
| 0.0 | 0.7 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 2.3 | GO:0031424 | keratinization(GO:0031424) |
| 0.0 | 0.6 | GO:0006505 | GPI anchor metabolic process(GO:0006505) GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 0.9 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.0 | 2.9 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.0 | 2.5 | GO:0002576 | platelet degranulation(GO:0002576) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.1 | 50.3 | GO:0097149 | centralspindlin complex(GO:0097149) |
| 6.6 | 112.6 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 6.4 | 70.3 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 5.3 | 15.8 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 5.0 | 15.1 | GO:0034515 | proteasome storage granule(GO:0034515) |
| 4.4 | 17.6 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 4.0 | 16.0 | GO:0034667 | integrin alpha3-beta1 complex(GO:0034667) |
| 3.8 | 34.1 | GO:1990761 | growth cone lamellipodium(GO:1990761) |
| 3.5 | 17.6 | GO:0097513 | myosin II filament(GO:0097513) |
| 3.3 | 16.3 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 3.2 | 9.7 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 3.1 | 31.1 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 3.1 | 37.0 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 3.0 | 24.0 | GO:1905202 | 3-methylcrotonyl-CoA carboxylase complex, mitochondrial(GO:0002169) methylcrotonoyl-CoA carboxylase complex(GO:1905202) |
| 2.9 | 11.4 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 2.6 | 76.0 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 2.6 | 13.0 | GO:0045272 | plasma membrane respiratory chain complex I(GO:0045272) |
| 2.6 | 7.8 | GO:0035370 | UBC13-UEV1A complex(GO:0035370) |
| 2.5 | 10.2 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 2.5 | 7.5 | GO:1990913 | sperm head plasma membrane(GO:1990913) ooplasm(GO:1990917) |
| 2.4 | 7.3 | GO:0002081 | outer acrosomal membrane(GO:0002081) |
| 2.3 | 9.4 | GO:1990246 | uniplex complex(GO:1990246) |
| 2.2 | 100.5 | GO:0045095 | keratin filament(GO:0045095) |
| 2.2 | 8.8 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 2.1 | 14.7 | GO:0098560 | cytoplasmic side of late endosome membrane(GO:0098560) |
| 2.0 | 26.6 | GO:0005915 | zonula adherens(GO:0005915) |
| 2.0 | 50.9 | GO:0044295 | axonal growth cone(GO:0044295) |
| 1.9 | 24.8 | GO:0035749 | myelin sheath adaxonal region(GO:0035749) |
| 1.9 | 13.2 | GO:0034715 | U7 snRNP(GO:0005683) pICln-Sm protein complex(GO:0034715) |
| 1.9 | 11.2 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 1.9 | 22.2 | GO:0031209 | SCAR complex(GO:0031209) |
| 1.8 | 23.7 | GO:0042555 | MCM complex(GO:0042555) |
| 1.8 | 5.3 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 1.7 | 10.4 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 1.7 | 3.4 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 1.6 | 6.6 | GO:0044307 | dendritic branch(GO:0044307) |
| 1.6 | 12.7 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 1.5 | 6.1 | GO:0031166 | integral component of vacuolar membrane(GO:0031166) |
| 1.4 | 5.6 | GO:0071986 | Ragulator complex(GO:0071986) |
| 1.3 | 3.9 | GO:1990393 | 3M complex(GO:1990393) |
| 1.3 | 9.1 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 1.3 | 11.3 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 1.2 | 54.1 | GO:0032420 | stereocilium(GO:0032420) |
| 1.2 | 20.7 | GO:0033643 | host cell part(GO:0033643) |
| 1.1 | 18.3 | GO:0030663 | COPI-coated vesicle membrane(GO:0030663) |
| 1.1 | 13.1 | GO:0030008 | TRAPP complex(GO:0030008) |
| 1.1 | 6.5 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 1.1 | 6.4 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 1.1 | 13.9 | GO:0002199 | zona pellucida receptor complex(GO:0002199) |
| 1.0 | 21.9 | GO:0030056 | hemidesmosome(GO:0030056) |
| 1.0 | 24.8 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 1.0 | 15.3 | GO:0043219 | lateral loop(GO:0043219) |
| 1.0 | 8.0 | GO:0030891 | VCB complex(GO:0030891) |
| 1.0 | 6.8 | GO:0005818 | astral microtubule(GO:0000235) aster(GO:0005818) |
| 1.0 | 12.4 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.9 | 14.1 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.9 | 54.4 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.9 | 20.5 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.9 | 4.4 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 0.8 | 2.3 | GO:0043260 | laminin-10 complex(GO:0043259) laminin-11 complex(GO:0043260) |
| 0.8 | 47.3 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.8 | 7.7 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.8 | 2.3 | GO:0016442 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.7 | 10.4 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.7 | 2.8 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.7 | 3.5 | GO:0032044 | DSIF complex(GO:0032044) |
| 0.7 | 67.5 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.7 | 14.9 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.7 | 10.4 | GO:0070578 | RISC-loading complex(GO:0070578) |
| 0.6 | 2.5 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.6 | 5.5 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.6 | 6.1 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.6 | 3.1 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.6 | 8.4 | GO:0090543 | Flemming body(GO:0090543) |
| 0.6 | 11.8 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.6 | 57.1 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.6 | 16.4 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.6 | 5.3 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.6 | 15.7 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.6 | 4.0 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.6 | 10.2 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.6 | 6.2 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.6 | 6.7 | GO:0097487 | multivesicular body, internal vesicle(GO:0097487) |
| 0.5 | 15.5 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.5 | 36.1 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.5 | 4.4 | GO:0097443 | sorting endosome(GO:0097443) |
| 0.5 | 28.6 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.5 | 5.8 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.5 | 4.7 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.5 | 12.8 | GO:0030057 | desmosome(GO:0030057) |
| 0.5 | 69.7 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.4 | 4.0 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.4 | 1.8 | GO:1990851 | Wnt-Frizzled-LRP5/6 complex(GO:1990851) |
| 0.4 | 24.2 | GO:0002102 | podosome(GO:0002102) |
| 0.4 | 3.5 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.4 | 12.4 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.4 | 14.8 | GO:0000795 | synaptonemal complex(GO:0000795) |
| 0.4 | 7.1 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.4 | 6.2 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.4 | 19.0 | GO:0090544 | BAF-type complex(GO:0090544) |
| 0.4 | 6.1 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.4 | 4.4 | GO:0032039 | integrator complex(GO:0032039) |
| 0.4 | 6.2 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.4 | 6.4 | GO:0000145 | exocyst(GO:0000145) |
| 0.4 | 20.7 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.4 | 4.5 | GO:0070938 | contractile ring(GO:0070938) |
| 0.4 | 3.0 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.4 | 12.0 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.3 | 6.3 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.3 | 11.8 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.3 | 30.7 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.3 | 2.7 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.3 | 3.2 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.3 | 6.2 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.3 | 4.5 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.2 | 3.0 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.2 | 15.4 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.2 | 7.1 | GO:0005921 | gap junction(GO:0005921) |
| 0.2 | 1.2 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.2 | 10.2 | GO:0000313 | organellar ribosome(GO:0000313) mitochondrial ribosome(GO:0005761) |
| 0.2 | 21.3 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.2 | 1.3 | GO:0009368 | endopeptidase Clp complex(GO:0009368) |
| 0.2 | 17.3 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.2 | 3.8 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.2 | 8.7 | GO:0043034 | costamere(GO:0043034) |
| 0.2 | 6.5 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.2 | 2.6 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.2 | 2.1 | GO:0005664 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.2 | 0.6 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.2 | 2.2 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.2 | 2.9 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.2 | 1.3 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.2 | 7.2 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.2 | 7.1 | GO:0005746 | mitochondrial respiratory chain(GO:0005746) |
| 0.2 | 13.7 | GO:0101003 | ficolin-1-rich granule membrane(GO:0101003) |
| 0.2 | 25.5 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.2 | 31.5 | GO:0101002 | ficolin-1-rich granule(GO:0101002) ficolin-1-rich granule lumen(GO:1904813) |
| 0.2 | 88.0 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.2 | 9.8 | GO:0000502 | proteasome complex(GO:0000502) |
| 0.2 | 9.8 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.1 | 0.8 | GO:0043196 | varicosity(GO:0043196) |
| 0.1 | 8.2 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.1 | 4.9 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 38.4 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.1 | 8.8 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.1 | 13.3 | GO:0043204 | perikaryon(GO:0043204) |
| 0.1 | 17.5 | GO:0005819 | spindle(GO:0005819) |
| 0.1 | 2.1 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.1 | 29.0 | GO:0019866 | organelle inner membrane(GO:0019866) |
| 0.1 | 30.5 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 6.8 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 2.9 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.1 | 1.8 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.1 | 32.6 | GO:0005874 | microtubule(GO:0005874) |
| 0.1 | 6.3 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.1 | 0.6 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.1 | 1.3 | GO:0042827 | platelet dense granule(GO:0042827) |
| 0.1 | 0.2 | GO:0071159 | NF-kappaB p50/p65 complex(GO:0035525) NF-kappaB complex(GO:0071159) |
| 0.1 | 1.4 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 0.1 | 9.6 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 0.8 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.1 | 2.5 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 31.7 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 1.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 3.6 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 0.1 | GO:0032449 | CBM complex(GO:0032449) |
| 0.0 | 0.9 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 76.6 | GO:0070062 | extracellular exosome(GO:0070062) |
| 0.0 | 1.4 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 28.5 | 114.2 | GO:0070320 | inward rectifier potassium channel inhibitor activity(GO:0070320) |
| 15.5 | 46.6 | GO:0003974 | UDP-N-acetylglucosamine 4-epimerase activity(GO:0003974) UDP-glucose 4-epimerase activity(GO:0003978) |
| 9.5 | 47.5 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 6.0 | 18.1 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 5.5 | 38.7 | GO:0016403 | dimethylargininase activity(GO:0016403) |
| 5.5 | 16.5 | GO:0070336 | flap-structured DNA binding(GO:0070336) |
| 5.3 | 15.8 | GO:0005046 | KDEL sequence binding(GO:0005046) |
| 4.5 | 17.9 | GO:0004504 | peptidylglycine monooxygenase activity(GO:0004504) peptidylamidoglycolate lyase activity(GO:0004598) |
| 4.3 | 12.9 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 4.2 | 42.4 | GO:0051425 | PTB domain binding(GO:0051425) |
| 3.8 | 26.6 | GO:0042978 | ornithine decarboxylase activator activity(GO:0042978) |
| 3.8 | 15.1 | GO:0017089 | glycolipid transporter activity(GO:0017089) |
| 3.7 | 14.8 | GO:0008192 | RNA guanylyltransferase activity(GO:0008192) |
| 3.7 | 11.1 | GO:0004766 | spermidine synthase activity(GO:0004766) |
| 3.6 | 28.5 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 3.5 | 17.3 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 3.4 | 16.9 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 3.3 | 6.5 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |
| 3.2 | 12.8 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 3.2 | 15.8 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) |
| 3.0 | 9.1 | GO:0050146 | nucleoside phosphotransferase activity(GO:0050146) |
| 3.0 | 24.0 | GO:0004485 | methylcrotonoyl-CoA carboxylase activity(GO:0004485) |
| 3.0 | 14.8 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 2.9 | 17.6 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 2.9 | 20.0 | GO:0004459 | lactate dehydrogenase activity(GO:0004457) L-lactate dehydrogenase activity(GO:0004459) |
| 2.8 | 8.5 | GO:0004418 | hydroxymethylbilane synthase activity(GO:0004418) |
| 2.7 | 8.0 | GO:0048030 | disaccharide binding(GO:0048030) |
| 2.5 | 7.5 | GO:0002135 | CTP binding(GO:0002135) |
| 2.5 | 7.5 | GO:0044594 | 3alpha,7alpha,12alpha-trihydroxy-5beta-cholest-24-enoyl-CoA hydratase activity(GO:0033989) 17-beta-hydroxysteroid dehydrogenase (NAD+) activity(GO:0044594) |
| 2.4 | 9.4 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 2.4 | 18.8 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 2.3 | 9.3 | GO:0045127 | N-acetylglucosamine kinase activity(GO:0045127) |
| 2.3 | 15.9 | GO:0015216 | adenine nucleotide transmembrane transporter activity(GO:0000295) purine ribonucleotide transmembrane transporter activity(GO:0005346) ATP transmembrane transporter activity(GO:0005347) purine nucleotide transmembrane transporter activity(GO:0015216) ADP transmembrane transporter activity(GO:0015217) |
| 2.3 | 11.3 | GO:0038051 | glucocorticoid receptor activity(GO:0004883) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 2.2 | 9.0 | GO:0004466 | long-chain-acyl-CoA dehydrogenase activity(GO:0004466) |
| 2.2 | 17.9 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 2.2 | 11.0 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) |
| 2.2 | 8.7 | GO:0004853 | uroporphyrinogen decarboxylase activity(GO:0004853) |
| 2.1 | 61.2 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 2.1 | 6.2 | GO:0052796 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 2.0 | 16.4 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 2.0 | 5.9 | GO:0015137 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 1.9 | 27.1 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 1.9 | 84.6 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 1.8 | 40.6 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 1.8 | 39.7 | GO:0008494 | translation activator activity(GO:0008494) |
| 1.8 | 5.3 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 1.7 | 59.7 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 1.6 | 4.9 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 1.6 | 9.7 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 1.6 | 6.4 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 1.6 | 4.7 | GO:0015181 | arginine transmembrane transporter activity(GO:0015181) |
| 1.6 | 12.4 | GO:0050833 | pyruvate transmembrane transporter activity(GO:0050833) |
| 1.5 | 4.6 | GO:0008466 | glycogenin glucosyltransferase activity(GO:0008466) |
| 1.5 | 7.5 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 1.5 | 10.4 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 1.5 | 4.4 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 1.5 | 10.2 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 1.4 | 8.7 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 1.4 | 17.2 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 1.4 | 25.4 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 1.4 | 26.0 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 1.3 | 14.8 | GO:0038132 | neuregulin binding(GO:0038132) |
| 1.3 | 37.0 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 1.3 | 15.8 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 1.3 | 11.8 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 1.3 | 6.5 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 1.3 | 57.0 | GO:0003785 | actin monomer binding(GO:0003785) |
| 1.3 | 26.6 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 1.3 | 26.3 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 1.2 | 14.9 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 1.2 | 4.9 | GO:1990948 | ligase inhibitor activity(GO:0055104) ubiquitin ligase inhibitor activity(GO:1990948) |
| 1.2 | 6.1 | GO:1903135 | cupric ion binding(GO:1903135) |
| 1.1 | 12.4 | GO:0001056 | RNA polymerase III activity(GO:0001056) |
| 1.1 | 22.3 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 1.1 | 54.4 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 1.1 | 6.6 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 1.1 | 4.4 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 1.1 | 9.9 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 1.1 | 62.3 | GO:0019894 | kinesin binding(GO:0019894) |
| 1.1 | 9.6 | GO:0045545 | syndecan binding(GO:0045545) |
| 1.1 | 3.2 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 1.0 | 10.4 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 1.0 | 18.4 | GO:0004859 | phospholipase inhibitor activity(GO:0004859) |
| 1.0 | 6.1 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 1.0 | 4.0 | GO:0016404 | 15-hydroxyprostaglandin dehydrogenase (NAD+) activity(GO:0016404) |
| 1.0 | 8.0 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 1.0 | 3.0 | GO:0005010 | insulin-like growth factor-activated receptor activity(GO:0005010) |
| 1.0 | 12.8 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 1.0 | 125.2 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 1.0 | 5.8 | GO:0048256 | flap endonuclease activity(GO:0048256) |
| 1.0 | 17.4 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 1.0 | 8.7 | GO:0032190 | acrosin binding(GO:0032190) |
| 1.0 | 4.8 | GO:0004471 | malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) oxaloacetate decarboxylase activity(GO:0008948) |
| 0.9 | 8.3 | GO:1990459 | transferrin receptor binding(GO:1990459) |
| 0.9 | 10.7 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.9 | 6.2 | GO:0071535 | RING-like zinc finger domain binding(GO:0071535) |
| 0.9 | 4.4 | GO:0019912 | cyclin-dependent protein kinase activating kinase activity(GO:0019912) |
| 0.9 | 10.6 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.9 | 24.4 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.9 | 6.0 | GO:0099602 | acetylcholine receptor regulator activity(GO:0030548) neurotransmitter receptor regulator activity(GO:0099602) |
| 0.9 | 2.6 | GO:0004730 | pseudouridylate synthase activity(GO:0004730) |
| 0.9 | 2.6 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.9 | 15.3 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.8 | 21.0 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.8 | 6.5 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.8 | 3.2 | GO:0004515 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.8 | 12.8 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.8 | 9.4 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.8 | 2.3 | GO:0070984 | SET domain binding(GO:0070984) |
| 0.8 | 16.9 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.8 | 30.3 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.8 | 8.3 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.8 | 15.8 | GO:0070878 | primary miRNA binding(GO:0070878) |
| 0.7 | 2.2 | GO:0016295 | [acyl-carrier-protein] S-malonyltransferase activity(GO:0004314) 3-oxoacyl-[acyl-carrier-protein] synthase activity(GO:0004315) oleoyl-[acyl-carrier-protein] hydrolase activity(GO:0004320) myristoyl-[acyl-carrier-protein] hydrolase activity(GO:0016295) palmitoyl-[acyl-carrier-protein] hydrolase activity(GO:0016296) acyl-[acyl-carrier-protein] hydrolase activity(GO:0016297) S-malonyltransferase activity(GO:0016419) malonyltransferase activity(GO:0016420) phosphopantetheine binding(GO:0031177) |
| 0.7 | 4.4 | GO:0038085 | vascular endothelial growth factor binding(GO:0038085) |
| 0.7 | 55.9 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.7 | 12.8 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.7 | 2.8 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.7 | 10.3 | GO:0001094 | TFIID-class transcription factor binding(GO:0001094) |
| 0.7 | 2.7 | GO:0016524 | latrotoxin receptor activity(GO:0016524) |
| 0.7 | 7.4 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) NFAT protein binding(GO:0051525) |
| 0.7 | 8.5 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.6 | 15.5 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.6 | 45.1 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.6 | 8.2 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.6 | 2.5 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.6 | 4.3 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.6 | 5.5 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.6 | 25.7 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.6 | 1.8 | GO:1904928 | coreceptor activity involved in canonical Wnt signaling pathway(GO:1904928) |
| 0.6 | 1.8 | GO:0005280 | hydrogen:amino acid symporter activity(GO:0005280) |
| 0.6 | 7.7 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.6 | 2.9 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.6 | 15.0 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.6 | 6.1 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.6 | 9.4 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.5 | 9.2 | GO:0031404 | chloride ion binding(GO:0031404) |
| 0.5 | 16.8 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.5 | 6.4 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.5 | 2.6 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.5 | 11.5 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.5 | 26.6 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.5 | 2.0 | GO:1990189 | peptide-serine-N-acetyltransferase activity(GO:1990189) |
| 0.5 | 9.1 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.5 | 19.7 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.5 | 4.0 | GO:0090599 | alpha-glucosidase activity(GO:0090599) |
| 0.5 | 10.9 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.5 | 8.2 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.5 | 7.9 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.4 | 12.0 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.4 | 23.2 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.4 | 1.3 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.4 | 1.2 | GO:0004782 | sulfinoalanine decarboxylase activity(GO:0004782) |
| 0.4 | 14.2 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.4 | 26.1 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.4 | 6.5 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.4 | 1.8 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 0.4 | 4.0 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.4 | 4.6 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.3 | 7.6 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.3 | 6.7 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.3 | 13.7 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.3 | 13.0 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.3 | 10.2 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.3 | 15.2 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.3 | 11.3 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.3 | 14.7 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.3 | 4.0 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.3 | 3.0 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.3 | 1.6 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.3 | 5.4 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.3 | 1.0 | GO:0042799 | histone methyltransferase activity (H4-K20 specific)(GO:0042799) |
| 0.3 | 6.5 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.3 | 1.0 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.2 | 13.8 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.2 | 6.8 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.2 | 2.1 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 0.2 | 3.8 | GO:0001637 | G-protein coupled chemoattractant receptor activity(GO:0001637) chemokine receptor activity(GO:0004950) |
| 0.2 | 1.4 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 0.2 | 101.0 | GO:0005525 | GTP binding(GO:0005525) |
| 0.2 | 1.4 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.2 | 23.6 | GO:0005518 | collagen binding(GO:0005518) |
| 0.2 | 1.6 | GO:0015038 | glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.2 | 3.5 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.2 | 12.8 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.2 | 5.5 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.2 | 3.8 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.2 | 5.9 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.2 | 6.8 | GO:0019956 | chemokine binding(GO:0019956) |
| 0.2 | 0.6 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.2 | 5.5 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.2 | 5.4 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.2 | 1.0 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.2 | 1.0 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.2 | 3.5 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.2 | 6.7 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 22.3 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.1 | 0.7 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.1 | 4.9 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 2.8 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.1 | 1.6 | GO:0015250 | water channel activity(GO:0015250) |
| 0.1 | 14.9 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.1 | 6.4 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.1 | 4.3 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.1 | 12.5 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.1 | 1.7 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.1 | 6.9 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 1.1 | GO:0097493 | structural molecule activity conferring elasticity(GO:0097493) |
| 0.1 | 0.8 | GO:0031386 | protein tag(GO:0031386) |
| 0.1 | 0.8 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 0.3 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.1 | 0.3 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.1 | 10.5 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 1.9 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 1.2 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.1 | 0.9 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 1.6 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.1 | 1.5 | GO:0005328 | neurotransmitter:sodium symporter activity(GO:0005328) |
| 0.1 | 1.1 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.1 | 2.6 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.1 | 1.8 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 1.2 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 2.1 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.1 | 1.2 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.1 | 4.6 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 1.2 | GO:0016876 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.1 | 3.4 | GO:0008276 | protein methyltransferase activity(GO:0008276) |
| 0.0 | 8.0 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.3 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.0 | 0.5 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.0 | 0.7 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 0.3 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.0 | 0.8 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.0 | 0.7 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.8 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 0.6 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 0.3 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.0 | 3.1 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 0.1 | GO:0004727 | prenylated protein tyrosine phosphatase activity(GO:0004727) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 110.2 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 1.6 | 97.6 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 1.6 | 60.7 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 1.3 | 55.8 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 1.3 | 85.9 | PID PLK1 PATHWAY | PLK1 signaling events |
| 1.2 | 13.6 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 1.1 | 43.3 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 1.1 | 43.0 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 1.0 | 68.7 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.8 | 53.2 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.8 | 11.8 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.8 | 7.7 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.8 | 74.0 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.7 | 43.0 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.7 | 13.9 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.6 | 21.6 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.6 | 19.4 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.6 | 13.3 | PID ATR PATHWAY | ATR signaling pathway |
| 0.6 | 17.4 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.5 | 14.1 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.5 | 26.6 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.5 | 33.3 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.4 | 33.9 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.4 | 5.5 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.4 | 14.2 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.4 | 50.4 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.4 | 4.1 | PID MYC PATHWAY | C-MYC pathway |
| 0.4 | 9.7 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.4 | 3.2 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.3 | 9.3 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.3 | 9.1 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.3 | 10.5 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.2 | 10.4 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.2 | 1.7 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.2 | 2.6 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.2 | 6.7 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.2 | 4.8 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.2 | 5.5 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.2 | 2.4 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.2 | 9.2 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.2 | 6.1 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.2 | 10.4 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.2 | 10.0 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.2 | 4.8 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.2 | 31.5 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.2 | 6.3 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.2 | 4.5 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.2 | 3.8 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 3.9 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.1 | 2.6 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 6.2 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.1 | 7.2 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.1 | 2.3 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.1 | 1.5 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.1 | 4.1 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.1 | 3.3 | PID NFAT 3PATHWAY | Role of Calcineurin-dependent NFAT signaling in lymphocytes |
| 0.1 | 2.2 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 4.5 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 6.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.1 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 0.7 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.2 | 108.3 | REACTOME PHOSPHORYLATION OF THE APC C | Genes involved in Phosphorylation of the APC/C |
| 3.7 | 125.8 | REACTOME ENOS ACTIVATION AND REGULATION | Genes involved in eNOS activation and regulation |
| 3.7 | 87.7 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 2.3 | 41.6 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 2.0 | 33.8 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 1.6 | 23.7 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 1.5 | 81.5 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 1.5 | 49.0 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 1.5 | 36.5 | REACTOME P130CAS LINKAGE TO MAPK SIGNALING FOR INTEGRINS | Genes involved in p130Cas linkage to MAPK signaling for integrins |
| 1.3 | 62.5 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 1.2 | 42.4 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 1.2 | 48.8 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 1.2 | 43.4 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 1.2 | 85.6 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 1.1 | 16.9 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 1.1 | 9.7 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 1.0 | 16.5 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.9 | 13.2 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.9 | 37.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.9 | 13.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.9 | 56.1 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.8 | 24.0 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.8 | 67.4 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.7 | 24.3 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.7 | 53.9 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.6 | 21.4 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.6 | 14.0 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.6 | 1.3 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.6 | 9.2 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.6 | 12.4 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.6 | 17.2 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.6 | 62.1 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.5 | 13.1 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.5 | 3.8 | REACTOME P53 INDEPENDENT G1 S DNA DAMAGE CHECKPOINT | Genes involved in p53-Independent G1/S DNA damage checkpoint |
| 0.5 | 6.4 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.5 | 11.4 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.5 | 8.6 | REACTOME SCFSKP2 MEDIATED DEGRADATION OF P27 P21 | Genes involved in SCF(Skp2)-mediated degradation of p27/p21 |
| 0.5 | 37.1 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.5 | 13.5 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.5 | 23.8 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.5 | 11.2 | REACTOME RAS ACTIVATION UOPN CA2 INFUX THROUGH NMDA RECEPTOR | Genes involved in Ras activation uopn Ca2+ infux through NMDA receptor |
| 0.5 | 6.0 | REACTOME SIGNALING BY NOTCH3 | Genes involved in Signaling by NOTCH3 |
| 0.5 | 6.4 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.4 | 8.5 | REACTOME ABORTIVE ELONGATION OF HIV1 TRANSCRIPT IN THE ABSENCE OF TAT | Genes involved in Abortive elongation of HIV-1 transcript in the absence of Tat |
| 0.4 | 8.6 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.4 | 8.9 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.4 | 7.5 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.4 | 14.3 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.4 | 8.1 | REACTOME PYRUVATE METABOLISM | Genes involved in Pyruvate metabolism |
| 0.4 | 4.6 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.4 | 8.3 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.4 | 6.3 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.4 | 22.8 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.3 | 6.1 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.3 | 10.3 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.3 | 8.2 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.3 | 6.2 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.3 | 8.9 | REACTOME SHC1 EVENTS IN ERBB4 SIGNALING | Genes involved in SHC1 events in ERBB4 signaling |
| 0.3 | 7.4 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.3 | 2.4 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.3 | 3.2 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.3 | 4.4 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.2 | 3.7 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.2 | 5.9 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.2 | 27.5 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.2 | 17.6 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.2 | 13.8 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.2 | 5.5 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.2 | 3.8 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.2 | 3.6 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.2 | 6.6 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
| 0.2 | 6.2 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.2 | 6.2 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.2 | 9.2 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.2 | 4.0 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.2 | 4.0 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.2 | 2.4 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.2 | 7.2 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.2 | 8.8 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 2.8 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.1 | 14.4 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 2.3 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.1 | 3.0 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.1 | 17.4 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.1 | 4.0 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.1 | 1.6 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.1 | 2.5 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 3.2 | REACTOME TRANSFERRIN ENDOCYTOSIS AND RECYCLING | Genes involved in Transferrin endocytosis and recycling |
| 0.1 | 14.2 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.1 | 0.6 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.1 | 3.0 | REACTOME CLASS I MHC MEDIATED ANTIGEN PROCESSING PRESENTATION | Genes involved in Class I MHC mediated antigen processing & presentation |
| 0.1 | 1.6 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.1 | 0.2 | REACTOME REGULATED PROTEOLYSIS OF P75NTR | Genes involved in Regulated proteolysis of p75NTR |
| 0.1 | 1.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 3.2 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 2.0 | REACTOME ASPARAGINE N LINKED GLYCOSYLATION | Genes involved in Asparagine N-linked glycosylation |
| 0.0 | 3.9 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.5 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.5 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.2 | REACTOME DEFENSINS | Genes involved in Defensins |