Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
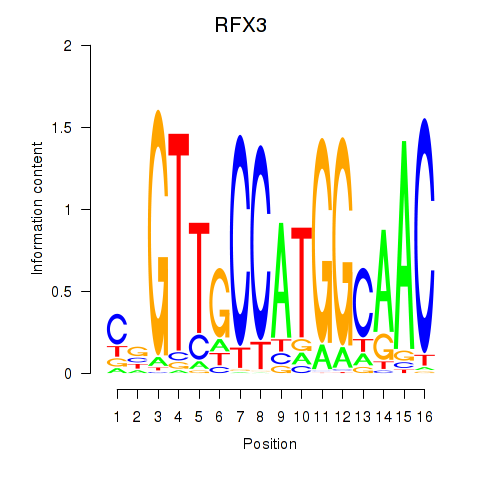
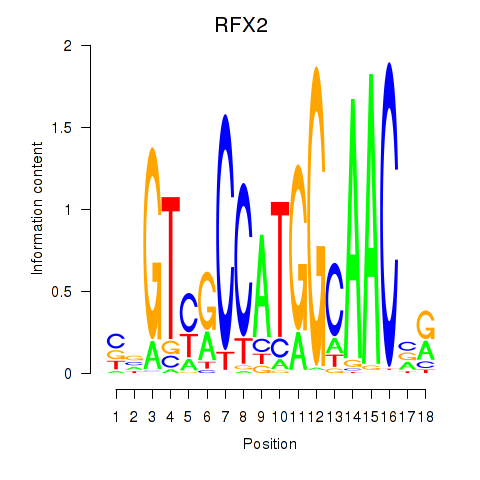
Results for RFX3_RFX2
Z-value: 4.51
Transcription factors associated with RFX3_RFX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
RFX3
|
ENSG00000080298.16 | RFX3 |
|
RFX2
|
ENSG00000087903.13 | RFX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RFX2 | hg38_v1_chr19_-_6110463_6110541 | 0.56 | 4.1e-19 | Click! |
Activity profile of RFX3_RFX2 motif
Sorted Z-values of RFX3_RFX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of RFX3_RFX2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 31.5 | 126.1 | GO:1904448 | negative regulation of gamma-aminobutyric acid secretion(GO:0014053) aspartate secretion(GO:0061528) regulation of aspartate secretion(GO:1904448) positive regulation of aspartate secretion(GO:1904450) |
| 30.4 | 182.5 | GO:0010734 | protein glutathionylation(GO:0010731) regulation of protein glutathionylation(GO:0010732) negative regulation of protein glutathionylation(GO:0010734) |
| 17.0 | 169.8 | GO:0036309 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 16.7 | 116.7 | GO:0070560 | protein secretion by platelet(GO:0070560) |
| 15.5 | 62.1 | GO:1903070 | negative regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903070) |
| 14.9 | 44.6 | GO:1903031 | regulation of microtubule plus-end binding(GO:1903031) positive regulation of microtubule plus-end binding(GO:1903033) |
| 13.2 | 39.6 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 13.0 | 65.2 | GO:0093001 | glycolysis from storage polysaccharide through glucose-1-phosphate(GO:0093001) |
| 13.0 | 90.7 | GO:1904098 | regulation of protein O-linked glycosylation(GO:1904098) positive regulation of protein O-linked glycosylation(GO:1904100) |
| 12.9 | 38.7 | GO:0051040 | regulation of calcium-independent cell-cell adhesion(GO:0051040) |
| 11.8 | 35.3 | GO:0001757 | somite specification(GO:0001757) |
| 11.5 | 172.6 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 11.4 | 34.2 | GO:0042357 | thiamine diphosphate metabolic process(GO:0042357) |
| 10.9 | 32.8 | GO:0050760 | negative regulation of thymidylate synthase biosynthetic process(GO:0050760) |
| 10.7 | 32.0 | GO:1902490 | regulation of sperm capacitation(GO:1902490) |
| 10.5 | 31.5 | GO:0034499 | late endosome to Golgi transport(GO:0034499) |
| 10.4 | 41.8 | GO:0035359 | negative regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035359) |
| 10.4 | 31.2 | GO:0015847 | putrescine transport(GO:0015847) positive regulation of polyamine transmembrane transport(GO:1902269) |
| 10.4 | 93.3 | GO:1903912 | negative regulation of endoplasmic reticulum stress-induced eIF2 alpha phosphorylation(GO:1903912) |
| 10.2 | 81.7 | GO:0002318 | myeloid progenitor cell differentiation(GO:0002318) |
| 9.3 | 74.4 | GO:0034316 | negative regulation of Arp2/3 complex-mediated actin nucleation(GO:0034316) |
| 9.3 | 250.5 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 9.1 | 172.7 | GO:0038166 | angiotensin-activated signaling pathway(GO:0038166) |
| 8.8 | 26.4 | GO:1903644 | regulation of chaperone-mediated protein folding(GO:1903644) |
| 8.7 | 43.3 | GO:0044375 | regulation of peroxisome size(GO:0044375) |
| 8.6 | 25.8 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 8.2 | 40.9 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 8.1 | 81.2 | GO:0034454 | microtubule anchoring at centrosome(GO:0034454) |
| 8.0 | 152.2 | GO:0034389 | lipid particle organization(GO:0034389) |
| 7.5 | 45.1 | GO:0006398 | mRNA 3'-end processing by stem-loop binding and cleavage(GO:0006398) |
| 7.4 | 22.2 | GO:1903348 | positive regulation of bicellular tight junction assembly(GO:1903348) |
| 7.4 | 44.1 | GO:0097338 | response to clozapine(GO:0097338) |
| 7.1 | 49.5 | GO:0051013 | microtubule severing(GO:0051013) |
| 6.7 | 6.7 | GO:0036466 | synaptic vesicle recycling via endosome(GO:0036466) |
| 6.6 | 19.9 | GO:1900220 | semaphorin-plexin signaling pathway involved in bone trabecula morphogenesis(GO:1900220) |
| 6.5 | 58.5 | GO:0007288 | sperm axoneme assembly(GO:0007288) |
| 6.4 | 153.1 | GO:0002021 | response to dietary excess(GO:0002021) |
| 6.4 | 127.4 | GO:0051823 | regulation of synapse structural plasticity(GO:0051823) |
| 6.4 | 70.0 | GO:0097113 | AMPA glutamate receptor clustering(GO:0097113) glutamate receptor clustering(GO:0097688) |
| 6.3 | 44.2 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 6.2 | 18.7 | GO:0018094 | protein polyglycylation(GO:0018094) |
| 6.2 | 37.0 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 5.6 | 16.7 | GO:0055048 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 5.5 | 16.5 | GO:1990519 | pyrimidine nucleotide transport(GO:0006864) mitochondrial pyrimidine nucleotide import(GO:1990519) |
| 5.5 | 16.5 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 5.1 | 30.5 | GO:0038016 | insulin receptor internalization(GO:0038016) |
| 5.1 | 15.2 | GO:0061590 | calcium activated phospholipid scrambling(GO:0061588) calcium activated phosphatidylcholine scrambling(GO:0061590) calcium activated galactosylceramide scrambling(GO:0061591) |
| 4.9 | 19.7 | GO:0098904 | regulation of AV node cell action potential(GO:0098904) |
| 4.8 | 14.5 | GO:0071034 | CUT catabolic process(GO:0071034) CUT metabolic process(GO:0071043) |
| 4.8 | 24.0 | GO:1900425 | negative regulation of defense response to bacterium(GO:1900425) |
| 4.8 | 19.0 | GO:0090063 | positive regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070426) positive regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070434) positive regulation of microtubule nucleation(GO:0090063) |
| 4.4 | 17.7 | GO:1902857 | positive regulation of nonmotile primary cilium assembly(GO:1902857) |
| 4.4 | 30.9 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 4.4 | 13.2 | GO:0099403 | negative regulation of maintenance of sister chromatid cohesion(GO:0034092) negative regulation of maintenance of mitotic sister chromatid cohesion(GO:0034183) protein auto-ADP-ribosylation(GO:0070213) maintenance of mitotic sister chromatid cohesion, telomeric(GO:0099403) mitotic sister chromatid cohesion, telomeric(GO:0099404) regulation of telomeric DNA binding(GO:1904742) regulation of maintenance of mitotic sister chromatid cohesion, telomeric(GO:1904907) negative regulation of maintenance of mitotic sister chromatid cohesion, telomeric(GO:1904908) |
| 4.4 | 13.1 | GO:0039022 | pronephric nephron morphogenesis(GO:0039007) pronephric nephron tubule morphogenesis(GO:0039008) pronephric duct development(GO:0039022) pronephric duct morphogenesis(GO:0039023) Kupffer's vesicle development(GO:0070121) |
| 4.2 | 96.3 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 4.2 | 66.9 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 4.1 | 37.1 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 3.9 | 11.6 | GO:1900060 | glucosylceramide biosynthetic process(GO:0006679) negative regulation of sphingolipid biosynthetic process(GO:0090155) negative regulation of ceramide biosynthetic process(GO:1900060) |
| 3.8 | 26.9 | GO:0030643 | cellular phosphate ion homeostasis(GO:0030643) cellular trivalent inorganic anion homeostasis(GO:0072502) |
| 3.8 | 15.3 | GO:1902775 | mitochondrial ribosome assembly(GO:0061668) mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 3.8 | 15.1 | GO:0080154 | regulation of fertilization(GO:0080154) |
| 3.8 | 165.9 | GO:2000300 | regulation of synaptic vesicle exocytosis(GO:2000300) |
| 3.8 | 33.8 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 3.7 | 22.2 | GO:0036159 | inner dynein arm assembly(GO:0036159) |
| 3.6 | 36.0 | GO:0032510 | endosome to lysosome transport via multivesicular body sorting pathway(GO:0032510) |
| 3.6 | 10.8 | GO:0090187 | positive regulation of pancreatic juice secretion(GO:0090187) |
| 3.6 | 35.7 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 3.4 | 41.2 | GO:0003406 | retinal pigment epithelium development(GO:0003406) |
| 3.4 | 13.4 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 3.3 | 53.1 | GO:0071027 | nuclear RNA surveillance(GO:0071027) nuclear mRNA surveillance(GO:0071028) |
| 3.3 | 16.4 | GO:0060718 | chorionic trophoblast cell differentiation(GO:0060718) |
| 3.2 | 9.7 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 3.1 | 15.7 | GO:1904925 | positive regulation of macromitophagy(GO:1901526) positive regulation of mitophagy in response to mitochondrial depolarization(GO:1904925) |
| 3.1 | 6.1 | GO:0021798 | neuroblast differentiation(GO:0014016) forebrain dorsal/ventral pattern formation(GO:0021798) |
| 3.0 | 18.2 | GO:0048549 | positive regulation of pinocytosis(GO:0048549) |
| 2.7 | 69.5 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 2.7 | 100.8 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 2.6 | 13.2 | GO:0051594 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 2.6 | 15.7 | GO:0061737 | leukotriene signaling pathway(GO:0061737) |
| 2.6 | 33.9 | GO:1901978 | positive regulation of cell cycle checkpoint(GO:1901978) |
| 2.6 | 12.8 | GO:0046167 | glycerol-3-phosphate biosynthetic process(GO:0046167) |
| 2.5 | 93.7 | GO:0048665 | neuron fate specification(GO:0048665) |
| 2.5 | 7.5 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 2.5 | 5.0 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 2.4 | 65.3 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 2.3 | 25.6 | GO:0035414 | negative regulation of catenin import into nucleus(GO:0035414) |
| 2.3 | 18.2 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 2.2 | 167.6 | GO:0060997 | dendritic spine morphogenesis(GO:0060997) |
| 2.2 | 17.5 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 2.2 | 61.1 | GO:0007141 | male meiosis I(GO:0007141) |
| 2.1 | 12.7 | GO:0006287 | base-excision repair, gap-filling(GO:0006287) |
| 2.1 | 37.5 | GO:0060117 | auditory receptor cell development(GO:0060117) |
| 2.1 | 72.9 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 2.1 | 39.5 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 2.0 | 18.4 | GO:0060267 | positive regulation of respiratory burst(GO:0060267) |
| 2.0 | 10.1 | GO:2000382 | positive regulation of mesoderm development(GO:2000382) |
| 2.0 | 41.3 | GO:0050775 | positive regulation of dendrite morphogenesis(GO:0050775) |
| 2.0 | 60.8 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 2.0 | 17.6 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 1.9 | 3.8 | GO:0031587 | positive regulation of inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0031587) |
| 1.9 | 35.8 | GO:1901663 | ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 1.9 | 26.3 | GO:0035082 | axoneme assembly(GO:0035082) |
| 1.8 | 27.7 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 1.8 | 5.5 | GO:0021684 | cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 1.7 | 83.3 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 1.7 | 15.3 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 1.7 | 15.3 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 1.7 | 37.3 | GO:0090036 | regulation of protein kinase C signaling(GO:0090036) |
| 1.6 | 4.8 | GO:0018874 | benzoate metabolic process(GO:0018874) butyrate metabolic process(GO:0019605) |
| 1.6 | 14.3 | GO:0015886 | heme transport(GO:0015886) |
| 1.6 | 81.8 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 1.6 | 18.7 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 1.5 | 1.5 | GO:0032425 | regulation of mismatch repair(GO:0032423) positive regulation of mismatch repair(GO:0032425) |
| 1.4 | 37.6 | GO:0035428 | hexose transmembrane transport(GO:0035428) glucose transmembrane transport(GO:1904659) |
| 1.4 | 5.7 | GO:0090069 | regulation of ribosome biogenesis(GO:0090069) regulation of rRNA processing(GO:2000232) |
| 1.4 | 11.4 | GO:1902412 | regulation of mitotic cytokinesis(GO:1902412) |
| 1.3 | 22.6 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 1.3 | 4.0 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 1.3 | 13.2 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 1.3 | 2.6 | GO:0018103 | protein C-linked glycosylation(GO:0018103) peptidyl-tryptophan modification(GO:0018211) protein C-linked glycosylation via tryptophan(GO:0018317) protein C-linked glycosylation via 2'-alpha-mannosyl-L-tryptophan(GO:0018406) |
| 1.2 | 18.5 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 1.2 | 17.0 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) |
| 1.2 | 20.6 | GO:0032516 | positive regulation of phosphoprotein phosphatase activity(GO:0032516) |
| 1.2 | 3.6 | GO:0010968 | regulation of microtubule nucleation(GO:0010968) |
| 1.2 | 3.6 | GO:0030573 | bile acid catabolic process(GO:0030573) |
| 1.2 | 11.7 | GO:2000189 | positive regulation of cholesterol homeostasis(GO:2000189) |
| 1.1 | 42.2 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 1.1 | 4.6 | GO:0030821 | negative regulation of cyclic nucleotide catabolic process(GO:0030806) negative regulation of cAMP catabolic process(GO:0030821) |
| 1.1 | 34.6 | GO:0007340 | acrosome reaction(GO:0007340) |
| 1.1 | 55.8 | GO:0043268 | positive regulation of potassium ion transport(GO:0043268) |
| 1.1 | 8.5 | GO:1903546 | protein localization to photoreceptor outer segment(GO:1903546) |
| 1.1 | 4.3 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 1.1 | 5.3 | GO:0042360 | vitamin E metabolic process(GO:0042360) |
| 1.0 | 11.3 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 1.0 | 14.0 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 1.0 | 3.9 | GO:0070945 | neutrophil mediated killing of gram-negative bacterium(GO:0070945) |
| 1.0 | 48.5 | GO:0071480 | cellular response to gamma radiation(GO:0071480) |
| 1.0 | 2.9 | GO:0070343 | white fat cell proliferation(GO:0070343) regulation of white fat cell proliferation(GO:0070350) |
| 1.0 | 211.3 | GO:0060271 | cilium morphogenesis(GO:0060271) |
| 0.9 | 6.6 | GO:1903750 | regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903750) negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903751) negative regulation of p38MAPK cascade(GO:1903753) |
| 0.9 | 55.5 | GO:0002437 | inflammatory response to antigenic stimulus(GO:0002437) |
| 0.9 | 8.0 | GO:0006704 | glucocorticoid biosynthetic process(GO:0006704) |
| 0.9 | 12.0 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.8 | 12.7 | GO:0031573 | intra-S DNA damage checkpoint(GO:0031573) |
| 0.8 | 13.5 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.8 | 32.6 | GO:0003091 | renal water homeostasis(GO:0003091) |
| 0.8 | 47.6 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.8 | 3.9 | GO:0021937 | cerebellar Purkinje cell-granule cell precursor cell signaling involved in regulation of granule cell precursor cell proliferation(GO:0021937) |
| 0.8 | 33.7 | GO:0008038 | neuron recognition(GO:0008038) |
| 0.8 | 34.3 | GO:0017156 | calcium ion regulated exocytosis(GO:0017156) |
| 0.8 | 8.4 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.8 | 4.5 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.7 | 15.5 | GO:0061049 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.7 | 9.5 | GO:0034315 | regulation of Arp2/3 complex-mediated actin nucleation(GO:0034315) |
| 0.7 | 20.2 | GO:0090083 | regulation of inclusion body assembly(GO:0090083) |
| 0.7 | 17.1 | GO:0006700 | C21-steroid hormone biosynthetic process(GO:0006700) |
| 0.7 | 19.0 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.7 | 47.9 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.7 | 2.8 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.7 | 6.2 | GO:0006335 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.7 | 12.8 | GO:0034656 | nucleobase-containing small molecule catabolic process(GO:0034656) |
| 0.7 | 30.6 | GO:0032570 | response to progesterone(GO:0032570) |
| 0.7 | 12.6 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.6 | 27.6 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.6 | 14.1 | GO:0060292 | long term synaptic depression(GO:0060292) |
| 0.6 | 4.3 | GO:0045007 | depurination(GO:0045007) |
| 0.6 | 1.8 | GO:0006478 | peptidyl-tyrosine sulfation(GO:0006478) |
| 0.6 | 38.9 | GO:0030049 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.6 | 37.8 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.6 | 6.5 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.6 | 20.3 | GO:0048168 | regulation of neuronal synaptic plasticity(GO:0048168) |
| 0.6 | 54.3 | GO:0005977 | glycogen metabolic process(GO:0005977) |
| 0.6 | 4.6 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.6 | 10.1 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.5 | 18.2 | GO:0071425 | hematopoietic stem cell proliferation(GO:0071425) |
| 0.5 | 16.4 | GO:0035024 | negative regulation of Rho protein signal transduction(GO:0035024) |
| 0.5 | 17.8 | GO:0022400 | regulation of rhodopsin mediated signaling pathway(GO:0022400) |
| 0.5 | 37.1 | GO:0045646 | regulation of erythrocyte differentiation(GO:0045646) |
| 0.5 | 2.5 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.5 | 16.2 | GO:0060996 | dendritic spine development(GO:0060996) |
| 0.5 | 8.1 | GO:0006228 | UTP biosynthetic process(GO:0006228) |
| 0.5 | 6.0 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.5 | 19.1 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.5 | 10.8 | GO:2000171 | negative regulation of dendrite development(GO:2000171) |
| 0.4 | 15.2 | GO:0048147 | negative regulation of fibroblast proliferation(GO:0048147) |
| 0.4 | 21.4 | GO:0051155 | positive regulation of striated muscle cell differentiation(GO:0051155) |
| 0.4 | 4.7 | GO:0006112 | energy reserve metabolic process(GO:0006112) |
| 0.4 | 26.7 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.4 | 13.9 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.4 | 39.5 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.4 | 9.1 | GO:0043518 | negative regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043518) |
| 0.4 | 13.4 | GO:0001578 | microtubule bundle formation(GO:0001578) |
| 0.4 | 5.8 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.3 | 6.6 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.3 | 22.1 | GO:0009060 | aerobic respiration(GO:0009060) |
| 0.3 | 0.7 | GO:1902612 | regulation of anti-Mullerian hormone signaling pathway(GO:1902612) negative regulation of anti-Mullerian hormone signaling pathway(GO:1902613) anti-Mullerian hormone signaling pathway(GO:1990262) |
| 0.3 | 10.4 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.3 | 20.7 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.3 | 14.8 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.3 | 5.6 | GO:0001975 | response to amphetamine(GO:0001975) |
| 0.3 | 24.5 | GO:0032092 | positive regulation of protein binding(GO:0032092) |
| 0.3 | 12.4 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.3 | 26.3 | GO:1903955 | positive regulation of protein targeting to mitochondrion(GO:1903955) |
| 0.2 | 7.9 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.2 | 7.8 | GO:0006379 | mRNA cleavage(GO:0006379) |
| 0.2 | 1.8 | GO:0009597 | detection of virus(GO:0009597) |
| 0.2 | 6.8 | GO:0007602 | phototransduction(GO:0007602) |
| 0.2 | 4.0 | GO:0006359 | regulation of transcription from RNA polymerase III promoter(GO:0006359) |
| 0.2 | 51.6 | GO:0051056 | regulation of small GTPase mediated signal transduction(GO:0051056) |
| 0.2 | 9.7 | GO:0007193 | adenylate cyclase-inhibiting G-protein coupled receptor signaling pathway(GO:0007193) |
| 0.2 | 1.3 | GO:0002053 | positive regulation of mesenchymal cell proliferation(GO:0002053) |
| 0.2 | 15.5 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.1 | 25.2 | GO:0007264 | small GTPase mediated signal transduction(GO:0007264) |
| 0.1 | 1.5 | GO:0034067 | protein localization to Golgi apparatus(GO:0034067) |
| 0.1 | 4.3 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.1 | 0.7 | GO:0046325 | negative regulation of glucose import(GO:0046325) |
| 0.1 | 44.6 | GO:0007283 | spermatogenesis(GO:0007283) male gamete generation(GO:0048232) |
| 0.1 | 5.5 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.1 | 22.5 | GO:0043401 | steroid hormone mediated signaling pathway(GO:0043401) |
| 0.1 | 0.3 | GO:0019918 | peptidyl-arginine methylation, to symmetrical-dimethyl arginine(GO:0019918) |
| 0.1 | 1.1 | GO:0060219 | camera-type eye photoreceptor cell differentiation(GO:0060219) |
| 0.1 | 24.1 | GO:0007411 | axon guidance(GO:0007411) |
| 0.1 | 8.7 | GO:0006970 | response to osmotic stress(GO:0006970) |
| 0.1 | 8.7 | GO:0014066 | regulation of phosphatidylinositol 3-kinase signaling(GO:0014066) |
| 0.1 | 1.6 | GO:0042059 | negative regulation of epidermal growth factor receptor signaling pathway(GO:0042059) |
| 0.1 | 3.2 | GO:0014047 | glutamate secretion(GO:0014047) |
| 0.1 | 22.9 | GO:0001558 | regulation of cell growth(GO:0001558) |
| 0.1 | 1.2 | GO:0045668 | negative regulation of osteoblast differentiation(GO:0045668) |
| 0.1 | 4.8 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.1 | 1.3 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.1 | 3.0 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.1 | 6.8 | GO:0002244 | hematopoietic progenitor cell differentiation(GO:0002244) |
| 0.1 | 0.2 | GO:1901895 | negative regulation of calcium-transporting ATPase activity(GO:1901895) |
| 0.1 | 9.4 | GO:0043523 | regulation of neuron apoptotic process(GO:0043523) |
| 0.1 | 1.8 | GO:0030042 | actin filament depolymerization(GO:0030042) |
| 0.1 | 11.9 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.0 | 5.9 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.8 | GO:1901661 | quinone metabolic process(GO:1901661) |
| 0.0 | 0.9 | GO:0043149 | contractile actin filament bundle assembly(GO:0030038) stress fiber assembly(GO:0043149) |
| 0.0 | 0.5 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 31.5 | 126.1 | GO:0032144 | 4-aminobutyrate transaminase complex(GO:0032144) |
| 25.7 | 77.2 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 18.8 | 18.8 | GO:0002199 | zona pellucida receptor complex(GO:0002199) |
| 15.3 | 61.1 | GO:0097123 | cyclin A1-CDK2 complex(GO:0097123) |
| 12.0 | 96.3 | GO:0000138 | Golgi trans cisterna(GO:0000138) |
| 11.3 | 33.9 | GO:0044609 | DBIRD complex(GO:0044609) |
| 11.3 | 33.8 | GO:0016939 | kinesin II complex(GO:0016939) |
| 10.9 | 65.2 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 10.5 | 63.2 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 10.3 | 61.7 | GO:0002177 | manchette(GO:0002177) |
| 10.2 | 61.1 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 10.1 | 182.5 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 9.8 | 49.2 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 8.6 | 68.5 | GO:0034464 | BBSome(GO:0034464) |
| 6.1 | 30.6 | GO:0036128 | CatSper complex(GO:0036128) |
| 5.6 | 72.9 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 5.6 | 16.7 | GO:0061673 | cortical microtubule(GO:0055028) mitotic spindle astral microtubule(GO:0061673) |
| 5.5 | 105.4 | GO:0036038 | MKS complex(GO:0036038) |
| 5.0 | 70.0 | GO:0098839 | postsynaptic density membrane(GO:0098839) |
| 4.7 | 116.7 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 4.6 | 37.0 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 4.6 | 78.4 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 4.2 | 72.0 | GO:0033010 | paranodal junction(GO:0033010) |
| 4.1 | 213.0 | GO:0016235 | aggresome(GO:0016235) |
| 4.0 | 55.6 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 3.9 | 35.3 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 3.8 | 60.3 | GO:0035102 | PRC1 complex(GO:0035102) |
| 3.6 | 18.2 | GO:1990031 | pinceau fiber(GO:1990031) |
| 3.6 | 32.8 | GO:1990761 | growth cone lamellipodium(GO:1990761) |
| 3.3 | 175.1 | GO:0031430 | M band(GO:0031430) |
| 3.1 | 91.6 | GO:0034451 | centriolar satellite(GO:0034451) |
| 3.0 | 15.2 | GO:0070847 | core mediator complex(GO:0070847) |
| 2.9 | 26.4 | GO:0001520 | outer dense fiber(GO:0001520) |
| 2.8 | 38.9 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 2.6 | 21.1 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 2.6 | 68.1 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 2.6 | 18.2 | GO:0044352 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 2.3 | 4.5 | GO:0032797 | SMN complex(GO:0032797) |
| 2.3 | 225.4 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 2.2 | 42.3 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 2.2 | 158.5 | GO:0005930 | axoneme(GO:0005930) ciliary plasm(GO:0097014) |
| 2.0 | 35.6 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 1.9 | 11.4 | GO:0005818 | astral microtubule(GO:0000235) aster(GO:0005818) |
| 1.9 | 11.3 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 1.9 | 26.1 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 1.8 | 9.2 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 1.8 | 19.9 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 1.7 | 199.0 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 1.7 | 11.9 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 1.6 | 77.5 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 1.6 | 22.9 | GO:0001673 | male germ cell nucleus(GO:0001673) |
| 1.6 | 16.1 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 1.5 | 43.3 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 1.5 | 13.4 | GO:0097427 | microtubule bundle(GO:0097427) |
| 1.5 | 16.1 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 1.5 | 90.2 | GO:0036064 | ciliary basal body(GO:0036064) |
| 1.4 | 25.6 | GO:0016342 | catenin complex(GO:0016342) |
| 1.3 | 7.5 | GO:0044214 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 1.2 | 12.0 | GO:0005655 | nucleolar ribonuclease P complex(GO:0005655) |
| 1.2 | 14.1 | GO:0005883 | neurofilament(GO:0005883) |
| 1.1 | 14.5 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 1.0 | 12.4 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 1.0 | 130.2 | GO:0043204 | perikaryon(GO:0043204) |
| 0.9 | 6.0 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.8 | 11.6 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.7 | 136.0 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.7 | 81.5 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.7 | 47.8 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.7 | 50.7 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.7 | 15.7 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.7 | 74.4 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.7 | 40.0 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.6 | 31.8 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.6 | 145.3 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.6 | 217.2 | GO:0005874 | microtubule(GO:0005874) |
| 0.5 | 1.6 | GO:0033565 | ESCRT-0 complex(GO:0033565) |
| 0.5 | 44.4 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.5 | 21.0 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.5 | 43.0 | GO:0005811 | lipid particle(GO:0005811) |
| 0.5 | 17.1 | GO:0098793 | presynapse(GO:0098793) |
| 0.5 | 5.9 | GO:0097486 | multivesicular body lumen(GO:0097486) |
| 0.5 | 54.8 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.4 | 26.1 | GO:0005776 | autophagosome(GO:0005776) |
| 0.4 | 15.4 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.4 | 102.4 | GO:0005938 | cell cortex(GO:0005938) |
| 0.3 | 7.1 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.3 | 3.3 | GO:0097546 | ciliary base(GO:0097546) |
| 0.3 | 18.5 | GO:0005814 | centriole(GO:0005814) |
| 0.3 | 29.6 | GO:0044309 | dendritic spine(GO:0043197) neuron spine(GO:0044309) |
| 0.3 | 5.5 | GO:0032809 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.3 | 6.3 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.3 | 3.2 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.3 | 212.8 | GO:0043005 | neuron projection(GO:0043005) |
| 0.3 | 25.0 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.3 | 1.3 | GO:0000120 | RNA polymerase I transcription factor complex(GO:0000120) |
| 0.2 | 4.1 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.2 | 19.6 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.2 | 13.6 | GO:0000922 | spindle pole(GO:0000922) |
| 0.2 | 33.7 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.2 | 4.0 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.2 | 13.3 | GO:0015030 | Cajal body(GO:0015030) |
| 0.2 | 36.8 | GO:0005929 | cilium(GO:0005929) |
| 0.2 | 26.0 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.2 | 4.0 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.1 | 3.8 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.1 | 13.5 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 3.3 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.1 | 3.0 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 7.8 | GO:0005903 | brush border(GO:0005903) |
| 0.1 | 19.7 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.1 | 80.7 | GO:0005615 | extracellular space(GO:0005615) |
| 0.1 | 7.9 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.1 | 39.0 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 30.5 | GO:0031226 | intrinsic component of plasma membrane(GO:0031226) |
| 0.0 | 0.6 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.0 | 2.9 | GO:0042995 | cell projection(GO:0042995) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 31.5 | 126.1 | GO:0032145 | 4-aminobutyrate transaminase activity(GO:0003867) succinate-semialdehyde dehydrogenase binding(GO:0032145) (S)-3-amino-2-methylpropionate transaminase activity(GO:0047298) |
| 13.3 | 172.6 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 13.3 | 53.1 | GO:0034353 | RNA pyrophosphohydrolase activity(GO:0034353) |
| 13.2 | 39.7 | GO:0008321 | Ral guanyl-nucleotide exchange factor activity(GO:0008321) |
| 13.0 | 90.7 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 10.4 | 41.8 | GO:0035473 | lipase binding(GO:0035473) |
| 10.4 | 31.2 | GO:0015203 | polyamine transmembrane transporter activity(GO:0015203) putrescine transmembrane transporter activity(GO:0015489) |
| 10.2 | 40.9 | GO:0061628 | H3K27me3 modified histone binding(GO:0061628) |
| 9.7 | 174.3 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 9.3 | 74.4 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 9.3 | 37.1 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 8.9 | 62.1 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 8.8 | 70.0 | GO:0031811 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 8.1 | 65.2 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 7.7 | 77.2 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 7.7 | 38.3 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 7.1 | 49.5 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 6.5 | 25.8 | GO:0022865 | transmembrane electron transfer carrier(GO:0022865) |
| 6.2 | 18.7 | GO:0070735 | protein-glycine ligase activity(GO:0070735) |
| 5.7 | 28.7 | GO:0070740 | tubulin-glutamic acid ligase activity(GO:0070740) |
| 5.7 | 17.1 | GO:0030156 | benzodiazepine receptor binding(GO:0030156) |
| 5.6 | 28.2 | GO:0005119 | smoothened binding(GO:0005119) |
| 5.5 | 16.5 | GO:0015218 | pyrimidine nucleotide transmembrane transporter activity(GO:0015218) |
| 5.0 | 309.6 | GO:0030507 | spectrin binding(GO:0030507) |
| 4.6 | 18.3 | GO:0050815 | phosphoserine binding(GO:0050815) |
| 4.5 | 36.3 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 4.5 | 121.9 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 4.2 | 37.5 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 4.1 | 44.6 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 3.9 | 11.6 | GO:0047322 | [hydroxymethylglutaryl-CoA reductase (NADPH)] kinase activity(GO:0047322) [acetyl-CoA carboxylase] kinase activity(GO:0050405) |
| 3.8 | 30.5 | GO:0015235 | cobalamin transporter activity(GO:0015235) |
| 3.7 | 11.1 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) |
| 3.5 | 228.0 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 3.1 | 15.7 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 2.9 | 17.6 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 2.8 | 19.7 | GO:0071253 | connexin binding(GO:0071253) |
| 2.7 | 124.5 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 2.7 | 8.0 | GO:0003845 | 11-beta-hydroxysteroid dehydrogenase [NAD(P)] activity(GO:0003845) |
| 2.7 | 95.9 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 2.6 | 12.8 | GO:0004370 | glycerol kinase activity(GO:0004370) |
| 2.5 | 15.3 | GO:0042285 | UDP-xylosyltransferase activity(GO:0035252) xylosyltransferase activity(GO:0042285) |
| 2.5 | 32.2 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 2.4 | 33.6 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 2.3 | 25.7 | GO:0004645 | phosphorylase activity(GO:0004645) |
| 2.3 | 41.9 | GO:0048156 | tau protein binding(GO:0048156) |
| 2.2 | 6.5 | GO:0031862 | prostanoid receptor binding(GO:0031862) |
| 2.1 | 33.0 | GO:0034452 | dynactin binding(GO:0034452) |
| 2.1 | 88.6 | GO:0003785 | actin monomer binding(GO:0003785) |
| 2.1 | 37.0 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 2.0 | 75.7 | GO:0070840 | dynein complex binding(GO:0070840) |
| 2.0 | 14.1 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 2.0 | 6.0 | GO:0003943 | N-acetylgalactosamine-4-sulfatase activity(GO:0003943) |
| 1.9 | 65.3 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 1.9 | 22.6 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 1.8 | 40.6 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 1.8 | 10.8 | GO:0034191 | apolipoprotein A-I receptor binding(GO:0034191) |
| 1.8 | 44.2 | GO:0044548 | S100 protein binding(GO:0044548) |
| 1.6 | 15.5 | GO:0005549 | odorant binding(GO:0005549) |
| 1.5 | 44.9 | GO:0071949 | FAD binding(GO:0071949) |
| 1.5 | 9.2 | GO:0070568 | guanylyltransferase activity(GO:0070568) |
| 1.5 | 11.9 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 1.5 | 32.6 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 1.5 | 13.2 | GO:0004396 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 1.5 | 19.1 | GO:0038191 | neuropilin binding(GO:0038191) |
| 1.4 | 12.7 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 1.4 | 147.1 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 1.4 | 62.5 | GO:0030552 | cAMP binding(GO:0030552) |
| 1.3 | 5.2 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 1.3 | 37.6 | GO:0005355 | glucose transmembrane transporter activity(GO:0005355) |
| 1.3 | 63.5 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 1.3 | 139.1 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 1.3 | 41.5 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 1.2 | 19.9 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 1.2 | 84.7 | GO:0016279 | lysine N-methyltransferase activity(GO:0016278) protein-lysine N-methyltransferase activity(GO:0016279) |
| 1.2 | 163.2 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 1.2 | 9.5 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 1.2 | 4.7 | GO:0004360 | glutamine-fructose-6-phosphate transaminase (isomerizing) activity(GO:0004360) |
| 1.2 | 69.5 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 1.2 | 34.9 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 1.1 | 5.6 | GO:0016427 | tRNA (cytosine) methyltransferase activity(GO:0016427) |
| 1.1 | 18.7 | GO:0031432 | titin binding(GO:0031432) |
| 1.1 | 4.3 | GO:0032406 | MutLbeta complex binding(GO:0032406) MutSbeta complex binding(GO:0032408) |
| 1.0 | 15.2 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 1.0 | 12.0 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 1.0 | 9.7 | GO:0097506 | uracil DNA N-glycosylase activity(GO:0004844) deaminated base DNA N-glycosylase activity(GO:0097506) |
| 1.0 | 31.8 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.9 | 38.7 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.9 | 4.6 | GO:0005298 | proline:sodium symporter activity(GO:0005298) |
| 0.9 | 4.5 | GO:0043141 | ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.9 | 28.1 | GO:0031418 | L-ascorbic acid binding(GO:0031418) |
| 0.9 | 42.5 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.9 | 12.9 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.9 | 16.2 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.8 | 110.4 | GO:0005496 | steroid binding(GO:0005496) |
| 0.8 | 13.2 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.8 | 16.4 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.8 | 23.8 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.8 | 6.8 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.7 | 137.1 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.7 | 18.7 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.7 | 15.7 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.7 | 54.4 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.7 | 17.0 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.6 | 16.9 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.6 | 177.1 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.6 | 16.2 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.6 | 9.7 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.6 | 1.8 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 0.6 | 6.6 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.6 | 19.0 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.6 | 6.1 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.5 | 9.1 | GO:0016594 | glycine binding(GO:0016594) |
| 0.5 | 18.0 | GO:0004033 | aldo-keto reductase (NADP) activity(GO:0004033) |
| 0.5 | 39.5 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.5 | 2.5 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.5 | 4.8 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.5 | 10.2 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.5 | 17.7 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.5 | 10.4 | GO:0004812 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.4 | 22.2 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.4 | 7.9 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.4 | 2.8 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.4 | 4.8 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.4 | 10.1 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.4 | 22.5 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.4 | 16.5 | GO:0001046 | core promoter sequence-specific DNA binding(GO:0001046) |
| 0.3 | 8.5 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.3 | 22.8 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.3 | 21.6 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.3 | 30.0 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.3 | 3.0 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.3 | 29.6 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.3 | 47.7 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.3 | 4.4 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.3 | 11.7 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.3 | 7.1 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.3 | 5.5 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.3 | 3.0 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.3 | 1.3 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) inositol-1,3,4-trisphosphate 4-phosphatase activity(GO:0017161) phosphatidylinositol-4,5-bisphosphate 4-phosphatase activity(GO:0034597) inositol-3,4-bisphosphate 4-phosphatase activity(GO:0052828) |
| 0.3 | 5.3 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.3 | 15.8 | GO:0004004 | ATP-dependent RNA helicase activity(GO:0004004) |
| 0.3 | 4.6 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.2 | 80.0 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.2 | 7.1 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.2 | 0.7 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 0.2 | 11.6 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.2 | 5.9 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.2 | 32.4 | GO:0005178 | integrin binding(GO:0005178) |
| 0.2 | 18.5 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.2 | 3.6 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.2 | 6.1 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.2 | 1.8 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.2 | 12.4 | GO:0005525 | GTP binding(GO:0005525) |
| 0.1 | 1.8 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.1 | 2.6 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.1 | 4.1 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 0.8 | GO:0003960 | NADPH:quinone reductase activity(GO:0003960) |
| 0.1 | 51.6 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.1 | 1.1 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.1 | 14.8 | GO:0035091 | phosphatidylinositol binding(GO:0035091) |
| 0.1 | 0.3 | GO:0035243 | protein-arginine omega-N symmetric methyltransferase activity(GO:0035243) |
| 0.1 | 2.9 | GO:0032451 | demethylase activity(GO:0032451) |
| 0.1 | 138.0 | GO:0003700 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
| 0.1 | 13.5 | GO:0008201 | heparin binding(GO:0008201) |
| 0.1 | 1.2 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.1 | 2.7 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.1 | 1.4 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 4.7 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 1.7 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.0 | 15.6 | GO:0004930 | G-protein coupled receptor activity(GO:0004930) |
| 0.0 | 4.6 | GO:0000287 | magnesium ion binding(GO:0000287) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.0 | 250.2 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 4.1 | 70.0 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 3.6 | 217.1 | ST ADRENERGIC | Adrenergic Pathway |
| 1.8 | 21.3 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 1.6 | 117.0 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 1.4 | 68.3 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 1.1 | 4.4 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 1.0 | 60.4 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.7 | 44.2 | PID FGF PATHWAY | FGF signaling pathway |
| 0.5 | 9.7 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.4 | 18.3 | PID MYC PATHWAY | C-MYC pathway |
| 0.4 | 18.2 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.4 | 20.2 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.4 | 24.9 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.3 | 17.7 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.3 | 19.9 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.3 | 15.4 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.3 | 14.8 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.2 | 11.9 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.2 | 5.0 | PID MET PATHWAY | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) |
| 0.2 | 31.4 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.2 | 4.6 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.2 | 6.4 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.2 | 11.0 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.2 | 10.8 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.2 | 11.3 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.2 | 12.3 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.2 | 13.2 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.1 | 6.3 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 36.2 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.1 | 2.5 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.1 | 23.3 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 1.6 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.1 | 0.7 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 1.2 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.0 | 263.9 | REACTOME RAS ACTIVATION UOPN CA2 INFUX THROUGH NMDA RECEPTOR | Genes involved in Ras activation uopn Ca2+ infux through NMDA receptor |
| 6.0 | 172.6 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 3.0 | 45.1 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 2.7 | 106.8 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 2.5 | 44.2 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 2.2 | 68.9 | REACTOME POST NMDA RECEPTOR ACTIVATION EVENTS | Genes involved in Post NMDA receptor activation events |
| 2.1 | 61.1 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 1.9 | 16.7 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 1.2 | 14.5 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 1.1 | 60.2 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 1.1 | 18.2 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.9 | 57.6 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.9 | 50.5 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.9 | 24.1 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.8 | 19.9 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.8 | 30.9 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.7 | 14.0 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.7 | 89.8 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.6 | 9.7 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.6 | 11.6 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.6 | 10.0 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.6 | 38.7 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.6 | 12.6 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.6 | 102.1 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.6 | 24.9 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.5 | 4.6 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.5 | 7.5 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.5 | 9.1 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.5 | 7.2 | REACTOME PROCESSING OF CAPPED INTRONLESS PRE MRNA | Genes involved in Processing of Capped Intronless Pre-mRNA |
| 0.4 | 18.3 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.4 | 24.7 | REACTOME TRIGLYCERIDE BIOSYNTHESIS | Genes involved in Triglyceride Biosynthesis |
| 0.4 | 58.4 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.3 | 128.6 | REACTOME GENERIC TRANSCRIPTION PATHWAY | Genes involved in Generic Transcription Pathway |
| 0.3 | 7.7 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.3 | 18.0 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.3 | 3.6 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.3 | 24.0 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.3 | 5.3 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.3 | 6.2 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.3 | 11.5 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.3 | 19.7 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.3 | 3.3 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.2 | 2.7 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.2 | 8.0 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.2 | 4.4 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.2 | 7.8 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.2 | 4.6 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.2 | 15.5 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.1 | 5.3 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.1 | 2.2 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 1.6 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.1 | 0.7 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.0 | 1.3 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 4.8 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.0 | 0.6 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.0 | 0.2 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.0 | 1.4 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |