Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
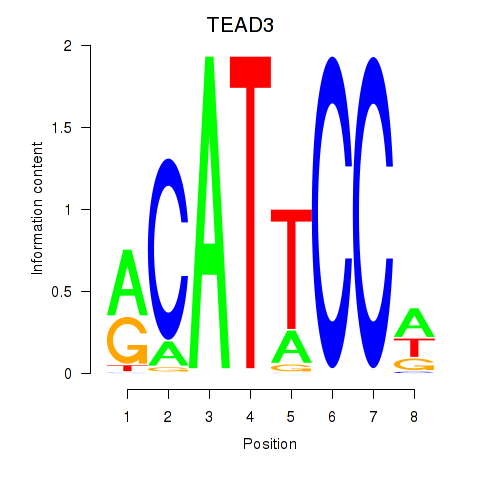
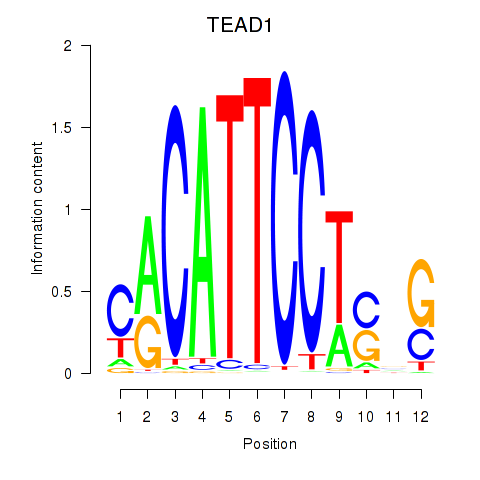
Results for TEAD3_TEAD1
Z-value: 2.66
Transcription factors associated with TEAD3_TEAD1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TEAD3
|
ENSG00000007866.22 | TEAD3 |
|
TEAD1
|
ENSG00000187079.20 | TEAD1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TEAD3 | hg38_v1_chr6_-_35497042_35497117 | 0.52 | 1.0e-16 | Click! |
| TEAD1 | hg38_v1_chr11_+_12674397_12674442 | 0.47 | 1.5e-13 | Click! |
Activity profile of TEAD3_TEAD1 motif
Sorted Z-values of TEAD3_TEAD1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TEAD3_TEAD1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 60.1 | 240.4 | GO:1903609 | negative regulation of peptidyl-tyrosine autophosphorylation(GO:1900085) negative regulation of inward rectifier potassium channel activity(GO:1903609) |
| 30.9 | 92.6 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 21.7 | 108.4 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 16.9 | 50.6 | GO:0052047 | interaction with other organism via secreted substance involved in symbiotic interaction(GO:0052047) |
| 14.0 | 69.8 | GO:0003065 | positive regulation of heart rate by epinephrine(GO:0003065) |
| 13.6 | 40.9 | GO:0010752 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) regulation of cGMP-mediated signaling(GO:0010752) |
| 12.0 | 47.9 | GO:0009956 | radial pattern formation(GO:0009956) |
| 9.7 | 48.5 | GO:0044111 | development involved in symbiotic interaction(GO:0044111) |
| 9.1 | 9.1 | GO:0072190 | ureter urothelium development(GO:0072190) ureter morphogenesis(GO:0072197) |
| 8.7 | 34.9 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 8.6 | 25.9 | GO:1901491 | axial mesoderm morphogenesis(GO:0048319) negative regulation of lymphangiogenesis(GO:1901491) |
| 6.3 | 69.8 | GO:0021730 | trigeminal sensory nucleus development(GO:0021730) principal sensory nucleus of trigeminal nerve development(GO:0021740) negative regulation of epithelial cell proliferation involved in lung morphogenesis(GO:2000795) |
| 6.3 | 31.4 | GO:0038165 | oncostatin-M-mediated signaling pathway(GO:0038165) |
| 6.0 | 18.0 | GO:2000661 | positive regulation of interleukin-1-mediated signaling pathway(GO:2000661) |
| 5.8 | 23.3 | GO:0014724 | regulation of twitch skeletal muscle contraction(GO:0014724) |
| 4.4 | 22.1 | GO:0007161 | calcium-independent cell-matrix adhesion(GO:0007161) |
| 4.4 | 39.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 4.1 | 20.3 | GO:2000542 | negative regulation of gastrulation(GO:2000542) |
| 3.8 | 26.8 | GO:0090191 | negative regulation of branching involved in ureteric bud morphogenesis(GO:0090191) |
| 3.7 | 22.4 | GO:0048549 | positive regulation of pinocytosis(GO:0048549) |
| 3.6 | 10.7 | GO:1904582 | regulation of intracellular mRNA localization(GO:1904580) positive regulation of intracellular mRNA localization(GO:1904582) |
| 3.5 | 10.6 | GO:0060957 | endocardial cell fate commitment(GO:0060957) endocardial cushion cell fate commitment(GO:0061445) |
| 3.4 | 161.7 | GO:0035329 | hippo signaling(GO:0035329) |
| 3.4 | 13.4 | GO:0006238 | CMP salvage(GO:0006238) CMP biosynthetic process(GO:0009224) CMP metabolic process(GO:0046035) |
| 3.3 | 56.4 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 3.3 | 46.2 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 3.2 | 12.7 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 3.1 | 18.6 | GO:0033088 | negative regulation of immature T cell proliferation in thymus(GO:0033088) |
| 3.0 | 9.1 | GO:0060435 | branchiomeric skeletal muscle development(GO:0014707) bronchiole development(GO:0060435) metanephric glomerulus morphogenesis(GO:0072275) metanephric glomerulus vasculature morphogenesis(GO:0072276) metanephric glomerular capillary formation(GO:0072277) |
| 2.9 | 14.4 | GO:0042986 | positive regulation of amyloid precursor protein biosynthetic process(GO:0042986) |
| 2.8 | 8.3 | GO:1905000 | regulation of membrane repolarization during atrial cardiac muscle cell action potential(GO:1905000) |
| 2.7 | 60.3 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 2.7 | 8.0 | GO:2000850 | negative regulation of steroid hormone secretion(GO:2000832) negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 2.6 | 13.0 | GO:0042360 | vitamin E metabolic process(GO:0042360) |
| 2.5 | 7.4 | GO:0043311 | positive regulation of eosinophil degranulation(GO:0043311) positive regulation of eosinophil activation(GO:1902568) |
| 2.4 | 4.7 | GO:0032972 | regulation of muscle filament sliding speed(GO:0032972) |
| 2.2 | 2.2 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 2.1 | 4.3 | GO:0061386 | closure of optic fissure(GO:0061386) |
| 2.1 | 12.4 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 2.0 | 18.3 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 2.0 | 32.4 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 1.9 | 33.0 | GO:0034356 | NAD biosynthesis via nicotinamide riboside salvage pathway(GO:0034356) |
| 1.8 | 24.7 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 1.7 | 12.0 | GO:0060686 | negative regulation of prostatic bud formation(GO:0060686) |
| 1.7 | 8.5 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 1.7 | 13.5 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 1.7 | 5.0 | GO:0060873 | subpallium cell proliferation in forebrain(GO:0022012) lateral ganglionic eminence cell proliferation(GO:0022018) lambdoid suture morphogenesis(GO:0060366) sagittal suture morphogenesis(GO:0060367) anterior semicircular canal development(GO:0060873) lateral semicircular canal development(GO:0060875) |
| 1.7 | 49.6 | GO:0002089 | lens morphogenesis in camera-type eye(GO:0002089) |
| 1.6 | 4.9 | GO:1903465 | vacuolar phosphate transport(GO:0007037) positive regulation of mitotic cell cycle DNA replication(GO:1903465) positive regulation of parathyroid hormone secretion(GO:2000830) |
| 1.6 | 20.7 | GO:0060712 | spongiotrophoblast layer development(GO:0060712) |
| 1.6 | 53.8 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 1.5 | 9.2 | GO:0008218 | bioluminescence(GO:0008218) |
| 1.4 | 21.7 | GO:0035878 | nail development(GO:0035878) |
| 1.4 | 5.7 | GO:0048205 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 1.4 | 5.7 | GO:0010585 | glutamine secretion(GO:0010585) L-glutamine import(GO:0036229) L-glutamine import into cell(GO:1903803) |
| 1.4 | 15.2 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
| 1.4 | 8.2 | GO:0035105 | sterol regulatory element binding protein import into nucleus(GO:0035105) |
| 1.4 | 24.6 | GO:0060856 | establishment of blood-brain barrier(GO:0060856) |
| 1.3 | 22.5 | GO:0030497 | fatty acid elongation(GO:0030497) very long-chain fatty acid biosynthetic process(GO:0042761) |
| 1.3 | 28.7 | GO:1903846 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 1.3 | 3.8 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 1.2 | 9.9 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 1.2 | 6.0 | GO:0034351 | regulation of glial cell apoptotic process(GO:0034350) negative regulation of glial cell apoptotic process(GO:0034351) |
| 1.2 | 47.0 | GO:0009303 | rRNA transcription(GO:0009303) |
| 1.1 | 122.0 | GO:0070527 | platelet aggregation(GO:0070527) |
| 1.1 | 9.0 | GO:0015014 | heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) |
| 1.1 | 1.1 | GO:1905069 | allantois development(GO:1905069) |
| 1.1 | 17.6 | GO:0001946 | lymphangiogenesis(GO:0001946) |
| 1.1 | 4.3 | GO:0061107 | seminal vesicle development(GO:0061107) |
| 1.1 | 40.2 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 1.1 | 3.3 | GO:0045362 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 1.1 | 3.2 | GO:0002159 | desmosome assembly(GO:0002159) |
| 1.1 | 3.2 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 1.1 | 7.5 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 1.1 | 4.3 | GO:0050916 | sensory perception of sweet taste(GO:0050916) sensory perception of umami taste(GO:0050917) |
| 1.1 | 34.0 | GO:0022011 | myelination in peripheral nervous system(GO:0022011) peripheral nervous system axon ensheathment(GO:0032292) |
| 1.0 | 8.3 | GO:1902897 | regulation of postsynaptic density protein 95 clustering(GO:1902897) |
| 1.0 | 5.1 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 1.0 | 5.9 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 1.0 | 1.0 | GO:0032971 | regulation of muscle filament sliding(GO:0032971) |
| 1.0 | 8.7 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.9 | 5.7 | GO:0002225 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antimicrobial humoral response(GO:0002760) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.9 | 10.8 | GO:0051964 | negative regulation of synapse assembly(GO:0051964) |
| 0.9 | 2.7 | GO:0036071 | N-glycan fucosylation(GO:0036071) |
| 0.9 | 3.5 | GO:0019072 | viral genome packaging(GO:0019072) viral RNA genome packaging(GO:0019074) |
| 0.9 | 5.1 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.8 | 1.7 | GO:0060535 | trachea cartilage morphogenesis(GO:0060535) |
| 0.8 | 2.5 | GO:0046533 | negative regulation of photoreceptor cell differentiation(GO:0046533) |
| 0.8 | 5.7 | GO:1905247 | positive regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902961) positive regulation of aspartic-type peptidase activity(GO:1905247) |
| 0.8 | 4.8 | GO:0003228 | atrial cardiac muscle tissue development(GO:0003228) atrial cardiac muscle tissue morphogenesis(GO:0055009) |
| 0.8 | 3.1 | GO:0060480 | lung goblet cell differentiation(GO:0060480) |
| 0.8 | 3.8 | GO:0097070 | ductus arteriosus closure(GO:0097070) |
| 0.8 | 9.1 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.8 | 8.3 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.7 | 74.1 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.7 | 17.8 | GO:0009954 | proximal/distal pattern formation(GO:0009954) |
| 0.7 | 12.4 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.7 | 10.8 | GO:0071285 | cellular response to lithium ion(GO:0071285) |
| 0.7 | 4.9 | GO:0072502 | cellular phosphate ion homeostasis(GO:0030643) cellular trivalent inorganic anion homeostasis(GO:0072502) |
| 0.7 | 1.4 | GO:0061577 | calcium ion transmembrane transport via high voltage-gated calcium channel(GO:0061577) |
| 0.7 | 11.0 | GO:0035313 | wound healing, spreading of epidermal cells(GO:0035313) |
| 0.7 | 2.0 | GO:0007538 | primary sex determination(GO:0007538) |
| 0.6 | 2.5 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 0.6 | 2.5 | GO:0019918 | peptidyl-arginine methylation, to symmetrical-dimethyl arginine(GO:0019918) |
| 0.6 | 9.8 | GO:0007097 | nuclear migration(GO:0007097) |
| 0.6 | 3.0 | GO:0030174 | regulation of DNA-dependent DNA replication initiation(GO:0030174) |
| 0.6 | 4.8 | GO:2000582 | regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.6 | 23.5 | GO:2000249 | regulation of actin cytoskeleton reorganization(GO:2000249) |
| 0.6 | 29.4 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.6 | 5.6 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.5 | 1.6 | GO:1990451 | cellular stress response to acidic pH(GO:1990451) |
| 0.5 | 2.0 | GO:0043324 | eye pigment biosynthetic process(GO:0006726) eye pigment metabolic process(GO:0042441) pigment metabolic process involved in developmental pigmentation(GO:0043324) pigment metabolic process involved in pigmentation(GO:0043474) |
| 0.5 | 5.5 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.5 | 9.4 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.5 | 1.9 | GO:0045917 | positive regulation of complement activation(GO:0045917) positive regulation of protein activation cascade(GO:2000259) |
| 0.4 | 0.9 | GO:0040023 | establishment of nucleus localization(GO:0040023) |
| 0.4 | 4.8 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.4 | 14.3 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.4 | 7.7 | GO:0032354 | response to follicle-stimulating hormone(GO:0032354) |
| 0.4 | 1.7 | GO:0045578 | negative regulation of B cell differentiation(GO:0045578) negative regulation of follicle-stimulating hormone secretion(GO:0046882) |
| 0.4 | 1.3 | GO:0006711 | estrogen catabolic process(GO:0006711) |
| 0.4 | 5.2 | GO:0015986 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.4 | 5.9 | GO:2000178 | negative regulation of neural precursor cell proliferation(GO:2000178) |
| 0.4 | 7.8 | GO:0014037 | Schwann cell differentiation(GO:0014037) |
| 0.4 | 7.8 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.4 | 2.3 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.4 | 0.8 | GO:0055062 | phosphate ion homeostasis(GO:0055062) trivalent inorganic anion homeostasis(GO:0072506) |
| 0.4 | 1.8 | GO:0015742 | alpha-ketoglutarate transport(GO:0015742) |
| 0.4 | 4.3 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.3 | 2.1 | GO:0006477 | protein sulfation(GO:0006477) |
| 0.3 | 4.5 | GO:0014809 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 0.3 | 4.7 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.3 | 1.0 | GO:0060754 | positive regulation of mast cell chemotaxis(GO:0060754) |
| 0.3 | 0.3 | GO:0060298 | positive regulation of sarcomere organization(GO:0060298) |
| 0.3 | 7.0 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.3 | 36.5 | GO:0030449 | regulation of complement activation(GO:0030449) |
| 0.3 | 7.2 | GO:2000675 | negative regulation of type B pancreatic cell apoptotic process(GO:2000675) |
| 0.3 | 1.5 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.3 | 1.2 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 0.3 | 3.5 | GO:0007172 | signal complex assembly(GO:0007172) |
| 0.3 | 1.2 | GO:0002879 | positive regulation of acute inflammatory response to non-antigenic stimulus(GO:0002879) |
| 0.3 | 1.2 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.3 | 31.0 | GO:0070268 | cornification(GO:0070268) |
| 0.3 | 25.8 | GO:0002062 | chondrocyte differentiation(GO:0002062) |
| 0.3 | 4.4 | GO:0007096 | regulation of exit from mitosis(GO:0007096) mitotic spindle midzone assembly(GO:0051256) |
| 0.3 | 6.9 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.3 | 1.4 | GO:0003418 | growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.3 | 5.7 | GO:0003416 | endochondral bone growth(GO:0003416) |
| 0.3 | 2.1 | GO:0032099 | negative regulation of response to food(GO:0032096) negative regulation of appetite(GO:0032099) |
| 0.3 | 14.2 | GO:0048747 | muscle fiber development(GO:0048747) |
| 0.3 | 1.3 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.3 | 1.8 | GO:0001945 | lymph vessel development(GO:0001945) |
| 0.3 | 0.8 | GO:1901301 | regulation of cargo loading into COPII-coated vesicle(GO:1901301) |
| 0.3 | 1.0 | GO:0000415 | negative regulation of histone H3-K36 methylation(GO:0000415) specification of axis polarity(GO:0065001) |
| 0.3 | 0.8 | GO:0060059 | embryonic retina morphogenesis in camera-type eye(GO:0060059) |
| 0.3 | 18.5 | GO:0042771 | intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:0042771) |
| 0.2 | 1.5 | GO:0060346 | bone trabecula formation(GO:0060346) |
| 0.2 | 0.7 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 0.2 | 6.6 | GO:0098743 | cell aggregation(GO:0098743) |
| 0.2 | 111.4 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.2 | 2.4 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.2 | 0.9 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.2 | 31.8 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.2 | 6.7 | GO:0030252 | growth hormone secretion(GO:0030252) |
| 0.2 | 40.8 | GO:0031032 | actomyosin structure organization(GO:0031032) |
| 0.2 | 6.0 | GO:0000717 | nucleotide-excision repair, DNA duplex unwinding(GO:0000717) |
| 0.2 | 1.8 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.2 | 0.6 | GO:0071206 | establishment of protein localization to juxtaparanode region of axon(GO:0071206) |
| 0.2 | 0.6 | GO:0072526 | pyridine-containing compound catabolic process(GO:0072526) |
| 0.2 | 2.2 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.2 | 6.6 | GO:0070830 | bicellular tight junction assembly(GO:0070830) |
| 0.2 | 1.3 | GO:0048295 | positive regulation of isotype switching to IgE isotypes(GO:0048295) |
| 0.2 | 15.6 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.2 | 17.0 | GO:0042058 | regulation of epidermal growth factor receptor signaling pathway(GO:0042058) |
| 0.2 | 2.8 | GO:0043403 | skeletal muscle tissue regeneration(GO:0043403) |
| 0.2 | 2.4 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) |
| 0.2 | 2.9 | GO:0031424 | keratinization(GO:0031424) |
| 0.2 | 4.0 | GO:0034204 | lipid translocation(GO:0034204) phospholipid translocation(GO:0045332) |
| 0.2 | 1.5 | GO:0014051 | gamma-aminobutyric acid secretion(GO:0014051) |
| 0.2 | 2.3 | GO:0006702 | androgen biosynthetic process(GO:0006702) |
| 0.2 | 0.5 | GO:0044828 | negative regulation by host of viral genome replication(GO:0044828) |
| 0.1 | 0.9 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.1 | 2.5 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.1 | 3.2 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.1 | 20.6 | GO:0000187 | activation of MAPK activity(GO:0000187) |
| 0.1 | 2.3 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 0.1 | 3.3 | GO:0002042 | cell migration involved in sprouting angiogenesis(GO:0002042) |
| 0.1 | 9.6 | GO:0032755 | positive regulation of interleukin-6 production(GO:0032755) |
| 0.1 | 16.2 | GO:0021987 | cerebral cortex development(GO:0021987) |
| 0.1 | 6.3 | GO:0008589 | regulation of smoothened signaling pathway(GO:0008589) |
| 0.1 | 22.1 | GO:0001649 | osteoblast differentiation(GO:0001649) |
| 0.1 | 1.4 | GO:0070935 | 3'-UTR-mediated mRNA stabilization(GO:0070935) |
| 0.1 | 2.8 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.1 | 1.2 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.1 | 0.3 | GO:0060605 | tube lumen cavitation(GO:0060605) salivary gland cavitation(GO:0060662) |
| 0.1 | 1.0 | GO:0019375 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.1 | 3.6 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.1 | 1.8 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.1 | 0.7 | GO:0030220 | platelet formation(GO:0030220) |
| 0.1 | 0.5 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.1 | 0.5 | GO:0006878 | cellular copper ion homeostasis(GO:0006878) |
| 0.1 | 1.9 | GO:0010761 | fibroblast migration(GO:0010761) |
| 0.1 | 4.2 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.1 | 0.5 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.1 | 2.2 | GO:0006825 | copper ion transport(GO:0006825) |
| 0.1 | 3.1 | GO:0048384 | retinoic acid receptor signaling pathway(GO:0048384) |
| 0.1 | 3.2 | GO:0032465 | regulation of cytokinesis(GO:0032465) |
| 0.1 | 7.7 | GO:0007052 | mitotic spindle organization(GO:0007052) |
| 0.1 | 1.9 | GO:0051893 | regulation of focal adhesion assembly(GO:0051893) regulation of cell-substrate junction assembly(GO:0090109) |
| 0.1 | 0.3 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.1 | 7.1 | GO:0002576 | platelet degranulation(GO:0002576) |
| 0.1 | 2.1 | GO:0014075 | response to amine(GO:0014075) |
| 0.1 | 1.8 | GO:0000002 | mitochondrial genome maintenance(GO:0000002) |
| 0.1 | 0.5 | GO:0008272 | sulfate transport(GO:0008272) |
| 0.1 | 5.4 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.1 | 1.6 | GO:0043949 | regulation of cAMP-mediated signaling(GO:0043949) |
| 0.0 | 0.4 | GO:0000492 | box C/D snoRNP assembly(GO:0000492) |
| 0.0 | 0.3 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 4.9 | GO:0035335 | peptidyl-tyrosine dephosphorylation(GO:0035335) |
| 0.0 | 0.2 | GO:0009437 | carnitine metabolic process(GO:0009437) |
| 0.0 | 0.3 | GO:1903358 | regulation of Golgi organization(GO:1903358) |
| 0.0 | 2.4 | GO:0072583 | clathrin-mediated endocytosis(GO:0072583) |
| 0.0 | 3.8 | GO:0006661 | phosphatidylinositol biosynthetic process(GO:0006661) |
| 0.0 | 3.4 | GO:0035383 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 1.4 | GO:0007098 | centrosome cycle(GO:0007098) |
| 0.0 | 1.1 | GO:0030879 | mammary gland development(GO:0030879) |
| 0.0 | 0.0 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.0 | 0.1 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.1 | GO:0046013 | T cell homeostatic proliferation(GO:0001777) regulation of T cell homeostatic proliferation(GO:0046013) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 33.5 | 100.6 | GO:0030485 | smooth muscle contractile fiber(GO:0030485) |
| 15.4 | 92.6 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 14.0 | 238.0 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 11.6 | 34.9 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 8.5 | 127.5 | GO:0030478 | actin cap(GO:0030478) |
| 7.7 | 92.4 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 7.4 | 22.1 | GO:0034677 | integrin alpha7-beta1 complex(GO:0034677) |
| 7.3 | 21.9 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) |
| 6.9 | 48.5 | GO:0044352 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 6.8 | 20.3 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 5.1 | 5.1 | GO:0034665 | integrin alpha1-beta1 complex(GO:0034665) |
| 5.0 | 69.8 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 4.5 | 72.5 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 4.0 | 19.8 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 3.7 | 11.0 | GO:0071062 | alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 3.3 | 69.8 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 2.8 | 112.5 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 2.8 | 55.3 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 2.6 | 10.6 | GO:0048179 | activin receptor complex(GO:0048179) |
| 2.6 | 7.7 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 2.4 | 4.7 | GO:1990584 | cardiac Troponin complex(GO:1990584) |
| 2.0 | 20.4 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 1.7 | 8.3 | GO:0031523 | Myb complex(GO:0031523) |
| 1.3 | 5.1 | GO:0071942 | XPC complex(GO:0071942) |
| 1.2 | 3.7 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 1.2 | 14.1 | GO:0043256 | laminin complex(GO:0043256) |
| 1.1 | 5.7 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 1.0 | 30.9 | GO:0031143 | pseudopodium(GO:0031143) |
| 1.0 | 3.0 | GO:0097135 | X chromosome(GO:0000805) cyclin E2-CDK2 complex(GO:0097135) |
| 0.9 | 21.7 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.8 | 20.2 | GO:0036379 | striated muscle thin filament(GO:0005865) myofilament(GO:0036379) |
| 0.8 | 27.7 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.8 | 17.6 | GO:0005652 | nuclear lamina(GO:0005652) |
| 0.8 | 5.7 | GO:0097209 | epidermal lamellar body(GO:0097209) |
| 0.8 | 9.1 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.8 | 6.8 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.7 | 2.9 | GO:0045180 | basal cortex(GO:0045180) |
| 0.7 | 9.8 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.7 | 44.1 | GO:0016459 | myosin complex(GO:0016459) |
| 0.7 | 12.7 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.6 | 90.0 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.6 | 4.4 | GO:0060091 | kinocilium(GO:0060091) |
| 0.6 | 46.2 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.6 | 61.7 | GO:0005901 | caveola(GO:0005901) |
| 0.6 | 1.7 | GO:0043512 | inhibin complex(GO:0043511) inhibin A complex(GO:0043512) |
| 0.5 | 2.1 | GO:0005960 | glycine cleavage complex(GO:0005960) |
| 0.5 | 5.2 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.5 | 20.2 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.4 | 1.2 | GO:0036026 | protein C inhibitor-TMPRSS7 complex(GO:0036024) protein C inhibitor-TMPRSS11E complex(GO:0036025) protein C inhibitor-PLAT complex(GO:0036026) protein C inhibitor-PLAU complex(GO:0036027) protein C inhibitor-thrombin complex(GO:0036028) protein C inhibitor-KLK3 complex(GO:0036029) protein C inhibitor-plasma kallikrein complex(GO:0036030) serine protease inhibitor complex(GO:0097180) protein C inhibitor-coagulation factor V complex(GO:0097181) protein C inhibitor-coagulation factor Xa complex(GO:0097182) protein C inhibitor-coagulation factor XI complex(GO:0097183) |
| 0.4 | 83.2 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.4 | 4.5 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.4 | 1.9 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.4 | 4.6 | GO:0090533 | cation-transporting ATPase complex(GO:0090533) |
| 0.3 | 15.6 | GO:0016235 | aggresome(GO:0016235) |
| 0.3 | 2.8 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.3 | 9.7 | GO:0043034 | costamere(GO:0043034) |
| 0.3 | 88.1 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.3 | 25.1 | GO:0005811 | lipid particle(GO:0005811) |
| 0.3 | 30.9 | GO:0005903 | brush border(GO:0005903) |
| 0.3 | 55.7 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.3 | 27.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.3 | 9.7 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.3 | 4.9 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.2 | 2.2 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.2 | 3.5 | GO:0043186 | P granule(GO:0043186) pole plasm(GO:0045495) germ plasm(GO:0060293) |
| 0.2 | 18.7 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.2 | 3.0 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.2 | 2.0 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.2 | 10.3 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.2 | 8.3 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.2 | 18.9 | GO:0005604 | basement membrane(GO:0005604) |
| 0.2 | 15.1 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.2 | 0.8 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.2 | 2.5 | GO:0034709 | methylosome(GO:0034709) |
| 0.2 | 4.9 | GO:0030132 | clathrin coat of coated pit(GO:0030132) |
| 0.2 | 30.5 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.2 | 2.7 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.2 | 0.5 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.2 | 1.1 | GO:0099634 | postsynaptic specialization membrane(GO:0099634) |
| 0.2 | 3.8 | GO:0005902 | microvillus(GO:0005902) |
| 0.1 | 244.0 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.1 | 0.9 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.1 | 55.0 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.1 | 0.5 | GO:0002945 | cyclin K-CDK13 complex(GO:0002945) |
| 0.1 | 3.3 | GO:0042641 | actomyosin(GO:0042641) |
| 0.1 | 5.9 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 1.0 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 1.5 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 0.4 | GO:0005726 | perichromatin fibrils(GO:0005726) pre-snoRNP complex(GO:0070761) |
| 0.1 | 0.9 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.1 | 5.5 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.1 | 5.1 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.1 | 2.1 | GO:0031228 | intrinsic component of Golgi membrane(GO:0031228) |
| 0.1 | 16.2 | GO:0005769 | early endosome(GO:0005769) |
| 0.1 | 1.5 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 4.6 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 1.2 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 2.2 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 2.2 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 40.5 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 3.9 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.0 | 2.5 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 0.5 | GO:0031904 | endosome lumen(GO:0031904) |
| 0.0 | 3.6 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 0.6 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.0 | 0.7 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.0 | 1.6 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 0.3 | GO:0045177 | apical part of cell(GO:0045177) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 60.1 | 240.4 | GO:0070320 | inward rectifier potassium channel inhibitor activity(GO:0070320) |
| 14.1 | 56.3 | GO:0042134 | rRNA primary transcript binding(GO:0042134) |
| 13.6 | 40.9 | GO:0070052 | collagen V binding(GO:0070052) |
| 9.7 | 48.5 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 9.3 | 27.8 | GO:0004923 | leukemia inhibitory factor receptor activity(GO:0004923) |
| 7.5 | 22.5 | GO:0080023 | 3R-hydroxyacyl-CoA dehydratase activity(GO:0080023) |
| 6.5 | 39.3 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 5.5 | 148.9 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 4.8 | 99.8 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 4.5 | 18.0 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 4.3 | 98.3 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 3.6 | 3.6 | GO:0004924 | oncostatin-M receptor activity(GO:0004924) |
| 3.4 | 27.2 | GO:0098639 | collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 3.0 | 12.0 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 3.0 | 9.0 | GO:0050509 | N-acetylglucosaminyl-proteoglycan 4-beta-glucuronosyltransferase activity(GO:0050509) |
| 2.8 | 5.6 | GO:0010465 | nerve growth factor receptor activity(GO:0010465) |
| 2.8 | 55.8 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 2.5 | 63.7 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 2.5 | 29.5 | GO:0038132 | neuregulin binding(GO:0038132) |
| 2.2 | 34.9 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 1.9 | 13.4 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 1.9 | 16.8 | GO:0043184 | vascular endothelial growth factor receptor 2 binding(GO:0043184) |
| 1.9 | 18.6 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 1.8 | 54.8 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 1.8 | 10.6 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 1.7 | 1.7 | GO:0061629 | titin binding(GO:0031432) RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 1.7 | 8.3 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 1.6 | 20.4 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 1.5 | 18.4 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 1.3 | 127.9 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 1.3 | 49.1 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 1.2 | 23.6 | GO:0031005 | filamin binding(GO:0031005) |
| 1.2 | 3.7 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 1.2 | 7.4 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 1.2 | 18.3 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 1.2 | 4.6 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 1.1 | 12.4 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 1.1 | 28.5 | GO:0005003 | ephrin receptor activity(GO:0005003) |
| 1.1 | 29.4 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 1.1 | 3.3 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 1.1 | 4.3 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 1.0 | 4.1 | GO:0017018 | myosin phosphatase activity(GO:0017018) |
| 1.0 | 26.1 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 1.0 | 3.9 | GO:0004348 | glucosylceramidase activity(GO:0004348) |
| 1.0 | 3.8 | GO:0016404 | 15-hydroxyprostaglandin dehydrogenase (NAD+) activity(GO:0016404) |
| 0.9 | 27.2 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.9 | 14.4 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.9 | 2.7 | GO:0046921 | glycoprotein 6-alpha-L-fucosyltransferase activity(GO:0008424) alpha-(1->6)-fucosyltransferase activity(GO:0046921) |
| 0.8 | 50.6 | GO:0016504 | peptidase activator activity(GO:0016504) |
| 0.8 | 7.3 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.8 | 2.4 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.8 | 3.9 | GO:0051998 | carboxyl-O-methyltransferase activity(GO:0010340) protein carboxyl O-methyltransferase activity(GO:0051998) |
| 0.7 | 7.2 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.7 | 5.7 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.7 | 67.7 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.7 | 31.9 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.6 | 12.1 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.6 | 11.4 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.6 | 13.3 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.6 | 5.7 | GO:0015321 | sodium-dependent phosphate transmembrane transporter activity(GO:0015321) |
| 0.6 | 10.6 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.6 | 8.0 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.6 | 4.7 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.6 | 34.3 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.6 | 2.3 | GO:0004769 | steroid delta-isomerase activity(GO:0004769) |
| 0.6 | 10.7 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.6 | 1.7 | GO:1904928 | coreceptor activity involved in canonical Wnt signaling pathway(GO:1904928) |
| 0.6 | 8.3 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.5 | 11.7 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.5 | 1.6 | GO:0030171 | voltage-gated proton channel activity(GO:0030171) |
| 0.5 | 8.4 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.5 | 4.9 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.5 | 5.7 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.4 | 9.4 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.4 | 2.1 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 0.4 | 2.9 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.4 | 2.5 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 0.4 | 26.0 | GO:0017022 | myosin binding(GO:0017022) |
| 0.4 | 25.2 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.4 | 4.3 | GO:0072542 | protein phosphatase activator activity(GO:0072542) |
| 0.4 | 8.8 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.4 | 4.6 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.4 | 3.0 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.4 | 3.5 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.3 | 1.7 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.3 | 6.7 | GO:0005024 | transforming growth factor beta-activated receptor activity(GO:0005024) |
| 0.3 | 2.3 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 0.3 | 3.6 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.3 | 22.4 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.3 | 4.8 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.3 | 1.2 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.3 | 34.1 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.3 | 29.7 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.3 | 5.7 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.3 | 2.4 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.3 | 2.1 | GO:0001517 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) |
| 0.3 | 32.4 | GO:0019838 | growth factor binding(GO:0019838) |
| 0.3 | 1.2 | GO:0047708 | biotinidase activity(GO:0047708) |
| 0.3 | 16.4 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.3 | 4.8 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.3 | 3.5 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.3 | 6.0 | GO:0004697 | protein kinase C activity(GO:0004697) |
| 0.3 | 6.4 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.3 | 0.8 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.3 | 29.3 | GO:0008170 | N-methyltransferase activity(GO:0008170) |
| 0.3 | 0.8 | GO:0090555 | phosphatidylethanolamine-translocating ATPase activity(GO:0090555) |
| 0.2 | 7.0 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.2 | 1.0 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.2 | 1.7 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.2 | 0.9 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.2 | 21.6 | GO:0032947 | protein complex scaffold(GO:0032947) |
| 0.2 | 14.3 | GO:0019003 | GDP binding(GO:0019003) |
| 0.2 | 1.3 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) |
| 0.2 | 5.2 | GO:0046961 | ATPase activity, coupled to transmembrane movement of ions, rotational mechanism(GO:0044769) proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.2 | 1.8 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.2 | 53.2 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.2 | 19.5 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.2 | 4.8 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.2 | 1.9 | GO:0033612 | receptor serine/threonine kinase binding(GO:0033612) |
| 0.2 | 3.0 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.2 | 3.0 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.2 | 4.6 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.2 | 4.0 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.2 | 67.2 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.1 | 2.7 | GO:0015101 | organic cation transmembrane transporter activity(GO:0015101) |
| 0.1 | 0.4 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.1 | 2.3 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 1.6 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.1 | 3.2 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.1 | 2.1 | GO:0016594 | glycine binding(GO:0016594) |
| 0.1 | 3.2 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 0.4 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.1 | 2.3 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.1 | 2.8 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.1 | 0.5 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.1 | 6.4 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.1 | 83.4 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.1 | 2.4 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.1 | 4.4 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.1 | 9.3 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.1 | 1.1 | GO:0043495 | protein anchor(GO:0043495) |
| 0.1 | 6.4 | GO:0004004 | ATP-dependent RNA helicase activity(GO:0004004) |
| 0.1 | 0.5 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.1 | 0.3 | GO:0004582 | dolichyl-phosphate beta-D-mannosyltransferase activity(GO:0004582) |
| 0.1 | 1.0 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.1 | 0.3 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 0.3 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.1 | 5.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 0.5 | GO:0008271 | secondary active sulfate transmembrane transporter activity(GO:0008271) |
| 0.1 | 1.0 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.1 | 2.5 | GO:0008066 | glutamate receptor activity(GO:0008066) |
| 0.1 | 2.0 | GO:0004879 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) transcription factor activity, direct ligand regulated sequence-specific DNA binding(GO:0098531) |
| 0.1 | 1.2 | GO:0005537 | mannose binding(GO:0005537) |
| 0.1 | 1.2 | GO:0086008 | voltage-gated potassium channel activity involved in cardiac muscle cell action potential repolarization(GO:0086008) |
| 0.1 | 0.6 | GO:0004645 | phosphorylase activity(GO:0004645) |
| 0.1 | 5.4 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.1 | 6.6 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 4.9 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.5 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.0 | 1.3 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.9 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 4.7 | GO:0035257 | nuclear hormone receptor binding(GO:0035257) |
| 0.0 | 1.7 | GO:0005507 | copper ion binding(GO:0005507) |
| 0.0 | 0.3 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.0 | 1.5 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 0.2 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.0 | 0.3 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 0.9 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.4 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 0.6 | GO:0043621 | protein self-association(GO:0043621) |
| 0.0 | 6.1 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 3.8 | GO:0045296 | cadherin binding(GO:0045296) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.5 | 241.3 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 2.6 | 35.8 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 2.4 | 167.5 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 1.5 | 117.7 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 1.4 | 111.7 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 1.2 | 17.7 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 1.2 | 111.9 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 1.1 | 15.2 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 1.1 | 25.9 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 1.1 | 22.2 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.9 | 27.1 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.8 | 47.0 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.8 | 22.6 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.8 | 7.9 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.8 | 27.2 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.7 | 34.9 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.7 | 7.0 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.7 | 18.6 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.6 | 43.4 | PID INTEGRIN1 PATHWAY | Beta1 integrin cell surface interactions |
| 0.6 | 116.8 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.6 | 10.6 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.6 | 36.8 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.6 | 50.7 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.5 | 18.0 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.5 | 13.5 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.5 | 2.8 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.5 | 21.3 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.4 | 9.1 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.3 | 57.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.3 | 10.1 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.2 | 3.0 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.2 | 3.9 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.2 | 15.5 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.2 | 10.6 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.2 | 9.1 | PID FGF PATHWAY | FGF signaling pathway |
| 0.2 | 2.2 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.2 | 7.3 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
| 0.2 | 3.2 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.2 | 3.7 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.2 | 2.3 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.2 | 8.1 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.1 | 7.4 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.1 | 6.0 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.1 | 1.6 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.1 | 4.2 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.1 | 3.1 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.1 | 2.5 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.1 | 4.0 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.1 | 3.3 | PID NFAT 3PATHWAY | Role of Calcineurin-dependent NFAT signaling in lymphocytes |
| 0.1 | 11.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 16.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.1 | 3.3 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.5 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 0.2 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.0 | 0.5 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 2.1 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.7 | 269.5 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 7.4 | 405.9 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 4.6 | 97.5 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 4.4 | 47.9 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 4.1 | 110.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 2.7 | 42.5 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 2.6 | 62.2 | REACTOME P130CAS LINKAGE TO MAPK SIGNALING FOR INTEGRINS | Genes involved in p130Cas linkage to MAPK signaling for integrins |
| 1.8 | 60.8 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 1.1 | 18.2 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.9 | 14.3 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.9 | 28.0 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.7 | 24.6 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.7 | 35.8 | REACTOME MUSCLE CONTRACTION | Genes involved in Muscle contraction |
| 0.7 | 26.4 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.7 | 6.0 | REACTOME P75NTR RECRUITS SIGNALLING COMPLEXES | Genes involved in p75NTR recruits signalling complexes |
| 0.6 | 10.7 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.6 | 22.8 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.6 | 25.2 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.6 | 13.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.6 | 18.2 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.5 | 13.0 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.5 | 5.7 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.5 | 11.9 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.4 | 7.1 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.4 | 9.4 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.4 | 9.6 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.4 | 18.0 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.4 | 41.7 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.3 | 36.3 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.3 | 13.4 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.3 | 3.0 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.3 | 7.9 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.3 | 16.0 | REACTOME SIGNAL TRANSDUCTION BY L1 | Genes involved in Signal transduction by L1 |
| 0.3 | 2.4 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.3 | 17.3 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.3 | 25.3 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.2 | 4.8 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.2 | 7.5 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.2 | 5.2 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.2 | 1.7 | REACTOME PEPTIDE HORMONE BIOSYNTHESIS | Genes involved in Peptide hormone biosynthesis |
| 0.2 | 4.4 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.2 | 2.3 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.2 | 11.3 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.2 | 28.1 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.2 | 5.1 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.1 | 2.4 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.1 | 2.5 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 3.6 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.1 | 3.2 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.1 | 1.9 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.1 | 9.7 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.1 | 3.9 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 1.4 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 2.8 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.1 | 2.4 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.1 | 7.7 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 1.4 | REACTOME REGULATION OF MRNA STABILITY BY PROTEINS THAT BIND AU RICH ELEMENTS | Genes involved in Regulation of mRNA Stability by Proteins that Bind AU-rich Elements |
| 0.1 | 6.6 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 0.3 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.1 | 1.4 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.1 | 5.4 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.1 | 0.8 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.1 | 0.9 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.1 | 2.5 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 2.2 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 0.4 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.4 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 0.6 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 5.2 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.0 | 0.9 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 1.1 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.1 | REACTOME P53 INDEPENDENT G1 S DNA DAMAGE CHECKPOINT | Genes involved in p53-Independent G1/S DNA damage checkpoint |
| 0.0 | 0.2 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.0 | 0.5 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
| 0.0 | 0.5 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 2.3 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 0.2 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |