Project
GNF SymAtlas + NCI-60 cancer cell lines, human (Su, 2004; Ross, 2000)
Navigation
Downloads
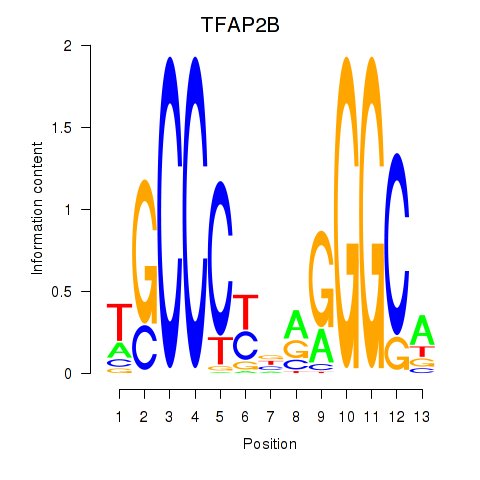
Results for TFAP2B
Z-value: 1.22
Transcription factors associated with TFAP2B
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TFAP2B
|
ENSG00000008196.13 | TFAP2B |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TFAP2B | hg38_v1_chr6_+_50818857_50818877 | 0.27 | 6.5e-05 | Click! |
Activity profile of TFAP2B motif
Sorted Z-values of TFAP2B motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TFAP2B
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.0 | 19.8 | GO:0042663 | regulation of endodermal cell fate specification(GO:0042663) |
| 4.7 | 14.2 | GO:0046586 | regulation of calcium-dependent cell-cell adhesion(GO:0046586) |
| 3.5 | 17.4 | GO:1904501 | glial cell fate determination(GO:0007403) canonical Wnt signaling pathway involved in positive regulation of cardiac outflow tract cell proliferation(GO:0061324) regulation of chromatin-mediated maintenance of transcription(GO:1904499) positive regulation of chromatin-mediated maintenance of transcription(GO:1904501) regulation of euchromatin binding(GO:1904793) |
| 3.1 | 9.2 | GO:0071461 | cellular response to redox state(GO:0071461) |
| 2.8 | 11.0 | GO:0060024 | rhythmic synaptic transmission(GO:0060024) |
| 2.7 | 8.1 | GO:0009720 | detection of hormone stimulus(GO:0009720) |
| 2.3 | 11.3 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 2.1 | 10.6 | GO:0044027 | hypermethylation of CpG island(GO:0044027) |
| 1.9 | 9.4 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 1.9 | 3.7 | GO:0003308 | negative regulation of Wnt signaling pathway involved in heart development(GO:0003308) |
| 1.9 | 24.2 | GO:2001199 | negative regulation of dendritic cell differentiation(GO:2001199) |
| 1.9 | 5.6 | GO:0016256 | N-glycan processing to lysosome(GO:0016256) |
| 1.8 | 23.4 | GO:2000623 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 1.5 | 9.0 | GO:0021966 | corticospinal neuron axon guidance(GO:0021966) |
| 1.5 | 4.5 | GO:0072365 | regulation of cellular ketone metabolic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072365) |
| 1.5 | 4.4 | GO:0071603 | endothelial cell-cell adhesion(GO:0071603) |
| 1.5 | 5.8 | GO:0072137 | condensed mesenchymal cell proliferation(GO:0072137) |
| 1.4 | 6.9 | GO:2000672 | negative regulation of motor neuron apoptotic process(GO:2000672) |
| 1.3 | 10.3 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 1.3 | 3.8 | GO:0048817 | negative regulation of transforming growth factor beta2 production(GO:0032912) negative regulation of hair follicle maturation(GO:0048817) regulation of melanosome transport(GO:1902908) |
| 1.2 | 2.5 | GO:0072312 | metanephric glomerular epithelium development(GO:0072244) metanephric glomerular visceral epithelial cell differentiation(GO:0072248) metanephric glomerular visceral epithelial cell development(GO:0072249) metanephric glomerular epithelial cell differentiation(GO:0072312) metanephric glomerular epithelial cell development(GO:0072313) |
| 1.2 | 4.9 | GO:0002384 | hepatic immune response(GO:0002384) |
| 1.2 | 3.6 | GO:0060152 | peroxisome localization(GO:0060151) microtubule-based peroxisome localization(GO:0060152) |
| 1.2 | 3.5 | GO:1900163 | regulation of phospholipid scramblase activity(GO:1900161) positive regulation of phospholipid scramblase activity(GO:1900163) regulation of glucosylceramide catabolic process(GO:2000752) positive regulation of glucosylceramide catabolic process(GO:2000753) regulation of sphingomyelin catabolic process(GO:2000754) positive regulation of sphingomyelin catabolic process(GO:2000755) |
| 1.1 | 6.7 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 1.1 | 5.3 | GO:0046167 | glycerol-3-phosphate biosynthetic process(GO:0046167) |
| 1.1 | 2.1 | GO:0060492 | foregut regionalization(GO:0060423) lung field specification(GO:0060424) lung induction(GO:0060492) |
| 1.0 | 5.2 | GO:0006546 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 1.0 | 3.1 | GO:0060448 | dichotomous subdivision of terminal units involved in lung branching(GO:0060448) |
| 1.0 | 5.1 | GO:0044856 | plasma membrane raft distribution(GO:0044855) plasma membrane raft localization(GO:0044856) plasma membrane raft polarization(GO:0044858) regulation of plasma membrane raft polarization(GO:1903906) |
| 1.0 | 4.0 | GO:1904924 | negative regulation of mitophagy in response to mitochondrial depolarization(GO:1904924) |
| 1.0 | 3.8 | GO:0099640 | axo-dendritic protein transport(GO:0099640) |
| 0.9 | 5.5 | GO:0045079 | negative regulation of chemokine biosynthetic process(GO:0045079) |
| 0.9 | 2.7 | GO:1903774 | positive regulation of viral budding via host ESCRT complex(GO:1903774) |
| 0.9 | 13.3 | GO:1901978 | positive regulation of cell cycle checkpoint(GO:1901978) |
| 0.9 | 3.5 | GO:1904381 | Golgi apparatus mannose trimming(GO:1904381) |
| 0.8 | 4.1 | GO:0045872 | positive regulation of rhodopsin gene expression(GO:0045872) |
| 0.8 | 4.1 | GO:0071504 | response to heparin(GO:0071503) cellular response to heparin(GO:0071504) |
| 0.8 | 2.5 | GO:0002818 | intracellular defense response(GO:0002818) |
| 0.8 | 2.4 | GO:0021913 | regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification(GO:0021913) |
| 0.8 | 3.1 | GO:2000595 | optic nerve formation(GO:0021634) optic chiasma development(GO:0061360) regulation of optic nerve formation(GO:2000595) positive regulation of optic nerve formation(GO:2000597) |
| 0.8 | 6.8 | GO:0002934 | desmosome organization(GO:0002934) |
| 0.7 | 3.6 | GO:0045906 | negative regulation of vasoconstriction(GO:0045906) |
| 0.7 | 0.7 | GO:0060129 | thyroid-stimulating hormone-secreting cell differentiation(GO:0060129) |
| 0.7 | 2.1 | GO:0014707 | branchiomeric skeletal muscle development(GO:0014707) |
| 0.7 | 2.7 | GO:0051586 | glial cell-derived neurotrophic factor secretion(GO:0044467) positive regulation of dopamine uptake involved in synaptic transmission(GO:0051586) positive regulation of catecholamine uptake involved in synaptic transmission(GO:0051944) regulation of glial cell-derived neurotrophic factor secretion(GO:1900166) positive regulation of glial cell-derived neurotrophic factor secretion(GO:1900168) |
| 0.7 | 4.1 | GO:0035105 | sterol regulatory element binding protein import into nucleus(GO:0035105) |
| 0.7 | 3.3 | GO:1901911 | diadenosine polyphosphate catabolic process(GO:0015961) diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 0.7 | 12.0 | GO:0040015 | negative regulation of multicellular organism growth(GO:0040015) |
| 0.7 | 2.6 | GO:0018199 | peptidyl-glutamine modification(GO:0018199) |
| 0.6 | 1.9 | GO:0045082 | positive regulation of interleukin-10 biosynthetic process(GO:0045082) |
| 0.6 | 2.5 | GO:0001826 | inner cell mass cell differentiation(GO:0001826) |
| 0.6 | 1.8 | GO:0042418 | epinephrine biosynthetic process(GO:0042418) |
| 0.6 | 1.7 | GO:0021658 | rhombomere morphogenesis(GO:0021593) rhombomere 3 morphogenesis(GO:0021658) |
| 0.6 | 1.2 | GO:0021555 | midbrain-hindbrain boundary morphogenesis(GO:0021555) |
| 0.6 | 4.0 | GO:0001999 | renal response to blood flow involved in circulatory renin-angiotensin regulation of systemic arterial blood pressure(GO:0001999) renin secretion into blood stream(GO:0002001) |
| 0.6 | 2.2 | GO:0035407 | histone H3-T11 phosphorylation(GO:0035407) |
| 0.6 | 4.5 | GO:0072553 | terminal button organization(GO:0072553) |
| 0.6 | 2.8 | GO:1901162 | primary amino compound biosynthetic process(GO:1901162) |
| 0.6 | 3.9 | GO:0000720 | pyrimidine dimer repair by nucleotide-excision repair(GO:0000720) positive regulation of t-circle formation(GO:1904431) |
| 0.5 | 7.7 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.5 | 2.1 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 0.5 | 1.6 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.5 | 1.6 | GO:0043606 | histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 0.5 | 1.6 | GO:2000977 | regulation of forebrain neuron differentiation(GO:2000977) |
| 0.5 | 1.5 | GO:0097535 | lymphoid lineage cell migration(GO:0097534) lymphoid lineage cell migration into thymus(GO:0097535) regulation of positive thymic T cell selection(GO:1902232) |
| 0.5 | 1.5 | GO:0019482 | purine nucleobase catabolic process(GO:0006145) pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) beta-alanine metabolic process(GO:0019482) thymine metabolic process(GO:0019859) |
| 0.5 | 4.9 | GO:0071421 | manganese ion transmembrane transport(GO:0071421) |
| 0.5 | 3.4 | GO:0033590 | response to cobalamin(GO:0033590) cellular response to erythropoietin(GO:0036018) |
| 0.5 | 1.4 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.5 | 1.8 | GO:0072369 | regulation of lipid transport by positive regulation of transcription from RNA polymerase II promoter(GO:0072369) |
| 0.5 | 1.4 | GO:0009078 | alanine metabolic process(GO:0006522) alanine catabolic process(GO:0006524) pyruvate family amino acid metabolic process(GO:0009078) pyruvate family amino acid catabolic process(GO:0009080) |
| 0.5 | 1.8 | GO:0003409 | optic cup structural organization(GO:0003409) |
| 0.4 | 4.5 | GO:1900121 | negative regulation of receptor binding(GO:1900121) |
| 0.4 | 2.2 | GO:0003069 | age-dependent response to oxidative stress(GO:0001306) age-dependent response to reactive oxygen species(GO:0001315) regulation of systemic arterial blood pressure by acetylcholine(GO:0003068) vasodilation by acetylcholine involved in regulation of systemic arterial blood pressure(GO:0003069) regulation of systemic arterial blood pressure by neurotransmitter(GO:0003070) age-dependent general metabolic decline(GO:0007571) |
| 0.4 | 12.5 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.4 | 3.1 | GO:0031339 | negative regulation of vesicle fusion(GO:0031339) |
| 0.4 | 2.6 | GO:0021937 | cerebellar Purkinje cell-granule cell precursor cell signaling involved in regulation of granule cell precursor cell proliferation(GO:0021937) |
| 0.4 | 2.2 | GO:0090385 | phagosome-lysosome fusion(GO:0090385) |
| 0.4 | 1.7 | GO:0072301 | posterior mesonephric tubule development(GO:0072166) negative regulation of metanephric glomerulus development(GO:0072299) regulation of metanephric glomerular mesangial cell proliferation(GO:0072301) negative regulation of metanephric glomerular mesangial cell proliferation(GO:0072302) |
| 0.4 | 1.3 | GO:0097477 | spinal cord motor neuron migration(GO:0097476) lateral motor column neuron migration(GO:0097477) |
| 0.4 | 1.3 | GO:0046959 | habituation(GO:0046959) |
| 0.4 | 3.3 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.4 | 5.8 | GO:0060033 | anatomical structure regression(GO:0060033) |
| 0.4 | 5.0 | GO:0002115 | store-operated calcium entry(GO:0002115) |
| 0.4 | 4.5 | GO:0070257 | positive regulation of mucus secretion(GO:0070257) |
| 0.4 | 1.2 | GO:0014063 | negative regulation of serotonin secretion(GO:0014063) |
| 0.4 | 2.7 | GO:2000304 | positive regulation of sphingolipid biosynthetic process(GO:0090154) positive regulation of ceramide biosynthetic process(GO:2000304) |
| 0.4 | 11.8 | GO:0030252 | growth hormone secretion(GO:0030252) |
| 0.4 | 0.8 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.4 | 1.1 | GO:1903526 | negative regulation of membrane tubulation(GO:1903526) |
| 0.4 | 1.8 | GO:0038165 | oncostatin-M-mediated signaling pathway(GO:0038165) |
| 0.4 | 2.5 | GO:0043585 | nose morphogenesis(GO:0043585) |
| 0.4 | 1.4 | GO:0010193 | response to ozone(GO:0010193) |
| 0.3 | 2.8 | GO:0060371 | regulation of atrial cardiac muscle cell membrane depolarization(GO:0060371) |
| 0.3 | 1.7 | GO:0060718 | chorionic trophoblast cell differentiation(GO:0060718) |
| 0.3 | 1.0 | GO:0008078 | mesodermal cell migration(GO:0008078) axial mesoderm morphogenesis(GO:0048319) |
| 0.3 | 15.1 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.3 | 2.0 | GO:0021521 | ventral spinal cord interneuron specification(GO:0021521) cell fate specification involved in pattern specification(GO:0060573) |
| 0.3 | 3.0 | GO:0097500 | receptor localization to nonmotile primary cilium(GO:0097500) |
| 0.3 | 1.3 | GO:0072423 | response to cell cycle checkpoint signaling(GO:0072396) response to DNA integrity checkpoint signaling(GO:0072402) response to DNA damage checkpoint signaling(GO:0072423) |
| 0.3 | 3.9 | GO:0035634 | response to stilbenoid(GO:0035634) |
| 0.3 | 1.9 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.3 | 6.8 | GO:0051255 | spindle midzone assembly(GO:0051255) |
| 0.3 | 5.4 | GO:0001977 | renal system process involved in regulation of blood volume(GO:0001977) |
| 0.3 | 5.9 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.3 | 1.2 | GO:0019230 | proprioception(GO:0019230) |
| 0.3 | 2.8 | GO:0060744 | positive regulation of exit from mitosis(GO:0031536) negative regulation of mammary gland epithelial cell proliferation(GO:0033600) thelarche(GO:0042695) mammary gland branching involved in thelarche(GO:0060744) regulation of branching involved in mammary gland duct morphogenesis(GO:0060762) |
| 0.3 | 2.1 | GO:0045007 | depurination(GO:0045007) |
| 0.3 | 6.4 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.3 | 0.6 | GO:0042320 | regulation of circadian sleep/wake cycle, REM sleep(GO:0042320) |
| 0.3 | 0.9 | GO:0050916 | sensory perception of sweet taste(GO:0050916) |
| 0.3 | 2.7 | GO:0046940 | nucleoside monophosphate phosphorylation(GO:0046940) |
| 0.3 | 7.2 | GO:0048935 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.3 | 5.1 | GO:0050862 | positive regulation of T cell receptor signaling pathway(GO:0050862) |
| 0.3 | 2.1 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.3 | 3.3 | GO:0051967 | negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.3 | 1.8 | GO:0061052 | negative regulation of cell growth involved in cardiac muscle cell development(GO:0061052) |
| 0.3 | 0.9 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.3 | 2.8 | GO:0010533 | regulation of activation of Janus kinase activity(GO:0010533) |
| 0.3 | 4.7 | GO:0043922 | negative regulation by host of viral transcription(GO:0043922) |
| 0.3 | 1.4 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.3 | 2.5 | GO:0043353 | enucleate erythrocyte differentiation(GO:0043353) |
| 0.3 | 7.7 | GO:0051412 | response to corticosterone(GO:0051412) |
| 0.3 | 0.8 | GO:0035752 | lysosomal lumen pH elevation(GO:0035752) |
| 0.3 | 2.5 | GO:0030807 | positive regulation of cyclic nucleotide catabolic process(GO:0030807) positive regulation of cAMP catabolic process(GO:0030822) |
| 0.3 | 40.5 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.3 | 1.0 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.3 | 2.6 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.2 | 7.1 | GO:0007202 | activation of phospholipase C activity(GO:0007202) |
| 0.2 | 2.7 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.2 | 4.7 | GO:0090026 | positive regulation of monocyte chemotaxis(GO:0090026) |
| 0.2 | 10.8 | GO:0033006 | regulation of mast cell activation involved in immune response(GO:0033006) regulation of mast cell degranulation(GO:0043304) |
| 0.2 | 1.6 | GO:0003072 | regulation of blood vessel size by renin-angiotensin(GO:0002034) renal control of peripheral vascular resistance involved in regulation of systemic arterial blood pressure(GO:0003072) negative regulation of plasminogen activation(GO:0010757) |
| 0.2 | 6.7 | GO:0097503 | sialylation(GO:0097503) |
| 0.2 | 3.1 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.2 | 0.9 | GO:0060750 | epithelial cell proliferation involved in mammary gland duct elongation(GO:0060750) branch elongation involved in mammary gland duct branching(GO:0060751) |
| 0.2 | 2.2 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.2 | 1.7 | GO:0021984 | adenohypophysis development(GO:0021984) |
| 0.2 | 2.6 | GO:0098914 | membrane repolarization during atrial cardiac muscle cell action potential(GO:0098914) |
| 0.2 | 0.9 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.2 | 3.5 | GO:0032464 | positive regulation of protein homooligomerization(GO:0032464) |
| 0.2 | 1.5 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 0.2 | 1.6 | GO:0015870 | acetylcholine transport(GO:0015870) |
| 0.2 | 4.5 | GO:0042554 | superoxide anion generation(GO:0042554) |
| 0.2 | 8.3 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.2 | 1.7 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.2 | 0.3 | GO:0002125 | maternal aggressive behavior(GO:0002125) |
| 0.2 | 5.3 | GO:0048536 | spleen development(GO:0048536) |
| 0.2 | 0.9 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.2 | 4.7 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.2 | 4.7 | GO:0001945 | lymph vessel development(GO:0001945) |
| 0.2 | 11.7 | GO:0001954 | positive regulation of cell-matrix adhesion(GO:0001954) |
| 0.2 | 3.8 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.2 | 5.5 | GO:0046676 | negative regulation of insulin secretion(GO:0046676) |
| 0.1 | 2.5 | GO:0006646 | phosphatidylethanolamine biosynthetic process(GO:0006646) |
| 0.1 | 5.0 | GO:0032731 | positive regulation of interleukin-1 beta production(GO:0032731) |
| 0.1 | 1.2 | GO:1903377 | negative regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903377) |
| 0.1 | 2.0 | GO:1990118 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 0.4 | GO:2000653 | regulation of genetic imprinting(GO:2000653) |
| 0.1 | 8.5 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.1 | 1.4 | GO:0042090 | interleukin-12 biosynthetic process(GO:0042090) regulation of interleukin-12 biosynthetic process(GO:0045075) |
| 0.1 | 2.1 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.1 | 2.3 | GO:0035162 | embryonic hemopoiesis(GO:0035162) |
| 0.1 | 1.6 | GO:0001542 | ovulation from ovarian follicle(GO:0001542) |
| 0.1 | 0.4 | GO:0001923 | B-1 B cell differentiation(GO:0001923) |
| 0.1 | 2.9 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.1 | 4.2 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.1 | 11.8 | GO:0042102 | positive regulation of T cell proliferation(GO:0042102) |
| 0.1 | 0.1 | GO:0018963 | phthalate metabolic process(GO:0018963) |
| 0.1 | 2.3 | GO:1904776 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.1 | 1.1 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.1 | 0.3 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.1 | 1.3 | GO:0038129 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) ERBB3 signaling pathway(GO:0038129) |
| 0.1 | 0.7 | GO:0051552 | flavone metabolic process(GO:0051552) |
| 0.1 | 1.0 | GO:0021511 | spinal cord patterning(GO:0021511) |
| 0.1 | 4.0 | GO:0048147 | negative regulation of fibroblast proliferation(GO:0048147) |
| 0.1 | 1.3 | GO:0048742 | regulation of skeletal muscle fiber development(GO:0048742) |
| 0.1 | 6.0 | GO:0006826 | iron ion transport(GO:0006826) |
| 0.1 | 4.3 | GO:0051646 | mitochondrion localization(GO:0051646) |
| 0.1 | 4.1 | GO:0032418 | lysosome localization(GO:0032418) |
| 0.1 | 2.4 | GO:0010458 | exit from mitosis(GO:0010458) |
| 0.1 | 2.5 | GO:0000042 | protein targeting to Golgi(GO:0000042) |
| 0.1 | 1.7 | GO:1903020 | positive regulation of glycoprotein metabolic process(GO:1903020) |
| 0.1 | 1.0 | GO:0035020 | regulation of Rac protein signal transduction(GO:0035020) |
| 0.1 | 2.3 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.1 | 1.8 | GO:0006359 | regulation of transcription from RNA polymerase III promoter(GO:0006359) |
| 0.1 | 1.4 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.1 | 0.9 | GO:0031284 | receptor guanylyl cyclase signaling pathway(GO:0007168) positive regulation of guanylate cyclase activity(GO:0031284) |
| 0.1 | 1.1 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 0.1 | 1.6 | GO:0006743 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.1 | 0.2 | GO:0046666 | retinal cell programmed cell death(GO:0046666) |
| 0.1 | 1.6 | GO:0008228 | opsonization(GO:0008228) |
| 0.1 | 0.7 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.1 | 1.1 | GO:0060056 | mammary gland involution(GO:0060056) |
| 0.1 | 1.1 | GO:0048484 | enteric nervous system development(GO:0048484) |
| 0.1 | 1.2 | GO:1990035 | calcium ion import across plasma membrane(GO:0098703) calcium ion import into cell(GO:1990035) |
| 0.1 | 0.8 | GO:1901407 | regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901407) |
| 0.1 | 2.9 | GO:1904837 | beta-catenin-TCF complex assembly(GO:1904837) |
| 0.1 | 0.5 | GO:1901376 | mycotoxin metabolic process(GO:0043385) aflatoxin metabolic process(GO:0046222) organic heteropentacyclic compound metabolic process(GO:1901376) |
| 0.1 | 1.1 | GO:0045907 | positive regulation of vasoconstriction(GO:0045907) |
| 0.1 | 1.5 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.1 | 2.7 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.1 | 2.1 | GO:0002755 | MyD88-dependent toll-like receptor signaling pathway(GO:0002755) |
| 0.1 | 2.3 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.1 | 0.4 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.1 | 1.7 | GO:0060236 | regulation of mitotic spindle organization(GO:0060236) |
| 0.1 | 1.6 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.1 | 0.7 | GO:0044829 | positive regulation by host of viral genome replication(GO:0044829) |
| 0.1 | 0.6 | GO:0034465 | response to carbon monoxide(GO:0034465) smooth muscle contraction involved in micturition(GO:0060083) |
| 0.1 | 0.2 | GO:2000819 | regulation of nucleotide-excision repair(GO:2000819) |
| 0.1 | 6.9 | GO:0070268 | cornification(GO:0070268) |
| 0.1 | 0.5 | GO:0043117 | positive regulation of vascular permeability(GO:0043117) |
| 0.1 | 6.2 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.1 | 0.9 | GO:0032957 | inositol trisphosphate metabolic process(GO:0032957) |
| 0.1 | 3.7 | GO:0050434 | positive regulation of viral transcription(GO:0050434) |
| 0.1 | 4.2 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.1 | 4.8 | GO:0006687 | glycosphingolipid metabolic process(GO:0006687) |
| 0.1 | 1.1 | GO:0090022 | regulation of neutrophil chemotaxis(GO:0090022) |
| 0.1 | 1.2 | GO:0010667 | negative regulation of cardiac muscle cell apoptotic process(GO:0010667) |
| 0.1 | 0.8 | GO:0032196 | transposition(GO:0032196) |
| 0.1 | 7.9 | GO:0006626 | protein targeting to mitochondrion(GO:0006626) |
| 0.1 | 1.7 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.8 | GO:0032228 | regulation of synaptic transmission, GABAergic(GO:0032228) |
| 0.0 | 0.6 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.6 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 2.6 | GO:0045600 | positive regulation of fat cell differentiation(GO:0045600) |
| 0.0 | 26.2 | GO:0042119 | neutrophil activation involved in immune response(GO:0002283) neutrophil activation(GO:0042119) neutrophil degranulation(GO:0043312) |
| 0.0 | 1.4 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 0.0 | 1.2 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.0 | 1.9 | GO:0006893 | Golgi to plasma membrane transport(GO:0006893) |
| 0.0 | 0.4 | GO:0031065 | positive regulation of histone deacetylation(GO:0031065) |
| 0.0 | 1.2 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.0 | 1.6 | GO:0001755 | neural crest cell migration(GO:0001755) |
| 0.0 | 1.7 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.0 | 1.8 | GO:0035019 | somatic stem cell population maintenance(GO:0035019) |
| 0.0 | 1.3 | GO:0043030 | regulation of macrophage activation(GO:0043030) |
| 0.0 | 5.3 | GO:0030217 | T cell differentiation(GO:0030217) |
| 0.0 | 0.4 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.0 | 1.1 | GO:0009409 | response to cold(GO:0009409) |
| 0.0 | 2.6 | GO:1900181 | negative regulation of protein localization to nucleus(GO:1900181) |
| 0.0 | 7.3 | GO:0060070 | canonical Wnt signaling pathway(GO:0060070) |
| 0.0 | 3.4 | GO:0002062 | chondrocyte differentiation(GO:0002062) |
| 0.0 | 0.3 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.0 | 0.3 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.0 | 0.3 | GO:0070932 | histone H3 deacetylation(GO:0070932) |
| 0.0 | 0.3 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.0 | 2.0 | GO:0006306 | DNA alkylation(GO:0006305) DNA methylation(GO:0006306) |
| 0.0 | 1.0 | GO:0003382 | epithelial cell morphogenesis(GO:0003382) |
| 0.0 | 0.5 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.0 | 0.8 | GO:0061418 | regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061418) |
| 0.0 | 0.5 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.0 | 1.4 | GO:1901264 | carbohydrate derivative transport(GO:1901264) |
| 0.0 | 0.3 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 7.7 | GO:0016485 | protein processing(GO:0016485) |
| 0.0 | 1.6 | GO:0007127 | meiosis I(GO:0007127) |
| 0.0 | 1.3 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.0 | 0.7 | GO:0060711 | labyrinthine layer development(GO:0060711) |
| 0.0 | 0.1 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.0 | 2.5 | GO:0010389 | regulation of G2/M transition of mitotic cell cycle(GO:0010389) |
| 0.0 | 0.1 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.0 | 2.5 | GO:0007187 | G-protein coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger(GO:0007187) |
| 0.0 | 1.8 | GO:0031047 | gene silencing by RNA(GO:0031047) |
| 0.0 | 1.4 | GO:0006024 | aminoglycan biosynthetic process(GO:0006023) glycosaminoglycan biosynthetic process(GO:0006024) |
| 0.0 | 0.2 | GO:0010800 | positive regulation of peptidyl-threonine phosphorylation(GO:0010800) |
| 0.0 | 2.3 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 0.4 | GO:0048477 | oogenesis(GO:0048477) |
| 0.0 | 0.2 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.4 | 22.2 | GO:0036284 | tubulobulbar complex(GO:0036284) |
| 4.4 | 13.3 | GO:0044609 | DBIRD complex(GO:0044609) |
| 2.9 | 17.4 | GO:0034750 | Scrib-APC-beta-catenin complex(GO:0034750) |
| 2.3 | 6.9 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 1.4 | 15.5 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 1.3 | 13.3 | GO:0044194 | cytolytic granule(GO:0044194) |
| 1.2 | 12.8 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 1.0 | 4.9 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 0.9 | 23.4 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.9 | 4.4 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.8 | 2.5 | GO:0043260 | laminin-11 complex(GO:0043260) |
| 0.8 | 14.4 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.8 | 9.8 | GO:0035749 | myelin sheath adaxonal region(GO:0035749) |
| 0.7 | 8.8 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.6 | 6.3 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 0.6 | 4.4 | GO:0071664 | catenin-TCF7L2 complex(GO:0071664) |
| 0.5 | 3.7 | GO:0032021 | NELF complex(GO:0032021) |
| 0.5 | 3.9 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) ERCC4-ERCC1 complex(GO:0070522) |
| 0.5 | 3.4 | GO:0043196 | varicosity(GO:0043196) |
| 0.5 | 2.4 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.5 | 1.9 | GO:0033257 | Bcl3/NF-kappaB2 complex(GO:0033257) |
| 0.4 | 1.3 | GO:0070701 | mucus layer(GO:0070701) |
| 0.4 | 4.2 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.4 | 1.9 | GO:0072558 | NLRP1 inflammasome complex(GO:0072558) |
| 0.4 | 4.1 | GO:0005638 | lamin filament(GO:0005638) |
| 0.4 | 9.0 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.4 | 38.1 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.4 | 4.3 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.3 | 5.1 | GO:0030478 | actin cap(GO:0030478) |
| 0.3 | 1.9 | GO:0032279 | axonemal microtubule(GO:0005879) asymmetric synapse(GO:0032279) |
| 0.3 | 1.6 | GO:0060200 | clathrin-sculpted acetylcholine transport vesicle(GO:0060200) clathrin-sculpted acetylcholine transport vesicle membrane(GO:0060201) |
| 0.3 | 1.6 | GO:0070826 | paraferritin complex(GO:0070826) |
| 0.3 | 2.2 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.3 | 4.5 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.3 | 2.1 | GO:0060091 | kinocilium(GO:0060091) |
| 0.3 | 37.5 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.3 | 11.9 | GO:0045095 | keratin filament(GO:0045095) |
| 0.3 | 3.6 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.3 | 6.8 | GO:0030057 | desmosome(GO:0030057) |
| 0.2 | 5.8 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.2 | 1.6 | GO:0030868 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.2 | 3.5 | GO:0005922 | connexon complex(GO:0005922) |
| 0.2 | 6.7 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.2 | 1.3 | GO:0030893 | meiotic cohesin complex(GO:0030893) |
| 0.2 | 2.2 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.2 | 16.2 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.2 | 3.7 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.2 | 2.1 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.2 | 2.7 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.2 | 2.5 | GO:0030137 | COPI-coated vesicle(GO:0030137) |
| 0.2 | 1.0 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.2 | 1.4 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.2 | 1.7 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.1 | 2.1 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 1.6 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.1 | 6.8 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.1 | 1.2 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.1 | 1.0 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.1 | 3.3 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.1 | 1.8 | GO:0097433 | dense body(GO:0097433) |
| 0.1 | 10.4 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 2.6 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 0.8 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.1 | 0.9 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.1 | 0.7 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.1 | 3.1 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.1 | 0.8 | GO:0030891 | VCB complex(GO:0030891) Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.1 | 2.3 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.1 | 0.7 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.1 | 0.9 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.1 | 0.4 | GO:0005943 | phosphatidylinositol 3-kinase complex, class IA(GO:0005943) |
| 0.1 | 51.0 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.1 | 1.5 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.1 | 1.7 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.1 | 0.4 | GO:0032449 | CBM complex(GO:0032449) |
| 0.1 | 14.7 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.1 | 0.4 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.1 | 1.9 | GO:0071437 | invadopodium(GO:0071437) |
| 0.1 | 2.4 | GO:0042629 | mast cell granule(GO:0042629) |
| 0.1 | 9.7 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 0.4 | GO:0033061 | DNA recombinase mediator complex(GO:0033061) Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.1 | 4.5 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 3.3 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 3.6 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 21.4 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.9 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.0 | 1.1 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 3.6 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 7.1 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 1.1 | GO:0044665 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.0 | 2.7 | GO:0098878 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.0 | 1.3 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.0 | 3.6 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 1.1 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 0.5 | GO:0097486 | multivesicular body lumen(GO:0097486) |
| 0.0 | 1.3 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.0 | 0.4 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.0 | 2.5 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 3.6 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 0.4 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 4.7 | GO:0031514 | motile cilium(GO:0031514) |
| 0.0 | 0.9 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.0 | 2.2 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.0 | 4.0 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 1.3 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 0.6 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 3.1 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 0.3 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.0 | 1.0 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 2.5 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 2.1 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 0.1 | GO:0036396 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 0.0 | 5.4 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.0 | 0.6 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 0.2 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 8.2 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 0.6 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 1.5 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 1.1 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.1 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.0 | 1.5 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.0 | 0.8 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 1.3 | GO:0031225 | anchored component of membrane(GO:0031225) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.3 | 9.8 | GO:0051717 | inositol-1,3,4,5-tetrakisphosphate 3-phosphatase activity(GO:0051717) |
| 2.2 | 10.8 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 2.0 | 8.1 | GO:0034041 | sterol-transporting ATPase activity(GO:0034041) |
| 1.8 | 5.4 | GO:0003845 | 11-beta-hydroxysteroid dehydrogenase [NAD(P)] activity(GO:0003845) |
| 1.8 | 9.0 | GO:0086062 | voltage-gated sodium channel activity involved in Purkinje myocyte action potential(GO:0086062) |
| 1.8 | 5.3 | GO:0030350 | iron-responsive element binding(GO:0030350) |
| 1.7 | 5.2 | GO:0004047 | aminomethyltransferase activity(GO:0004047) |
| 1.6 | 4.9 | GO:0070119 | ciliary neurotrophic factor binding(GO:0070119) |
| 1.4 | 5.6 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 1.3 | 8.0 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 1.3 | 19.8 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 1.3 | 24.5 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 1.3 | 3.8 | GO:0008426 | protein kinase C inhibitor activity(GO:0008426) |
| 1.2 | 4.8 | GO:0004773 | steryl-sulfatase activity(GO:0004773) |
| 1.1 | 9.2 | GO:0005094 | Rho GDP-dissociation inhibitor activity(GO:0005094) |
| 1.1 | 6.9 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 1.1 | 4.5 | GO:0004768 | stearoyl-CoA 9-desaturase activity(GO:0004768) acyl-CoA desaturase activity(GO:0016215) |
| 1.1 | 23.4 | GO:0008494 | translation activator activity(GO:0008494) |
| 1.1 | 5.3 | GO:0004370 | glycerol kinase activity(GO:0004370) |
| 1.0 | 5.1 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 0.9 | 2.8 | GO:0086059 | voltage-gated calcium channel activity involved SA node cell action potential(GO:0086059) |
| 0.9 | 3.6 | GO:0047288 | monosialoganglioside sialyltransferase activity(GO:0047288) |
| 0.9 | 4.3 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.8 | 2.5 | GO:0004566 | beta-glucuronidase activity(GO:0004566) |
| 0.8 | 8.9 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.8 | 2.3 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 0.8 | 3.0 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.7 | 9.4 | GO:0008199 | acid phosphatase activity(GO:0003993) ferric iron binding(GO:0008199) |
| 0.7 | 2.1 | GO:0008534 | oxidized purine nucleobase lesion DNA N-glycosylase activity(GO:0008534) |
| 0.7 | 2.7 | GO:0016230 | sphingomyelin phosphodiesterase activator activity(GO:0016230) |
| 0.7 | 8.8 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.7 | 3.3 | GO:0052840 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) inositol diphosphate tetrakisphosphate diphosphatase activity(GO:0052840) inositol bisdiphosphate tetrakisphosphate diphosphatase activity(GO:0052841) inositol diphosphate pentakisphosphate diphosphatase activity(GO:0052842) inositol-1-diphosphate-2,3,4,5,6-pentakisphosphate diphosphatase activity(GO:0052843) inositol-3-diphosphate-1,2,4,5,6-pentakisphosphate diphosphatase activity(GO:0052844) inositol-5-diphosphate-1,2,3,4,6-pentakisphosphate diphosphatase activity(GO:0052845) inositol-1,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 1-diphosphatase activity(GO:0052846) inositol-1,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 5-diphosphatase activity(GO:0052847) inositol-3,5-bisdiphosphate-2,3,4,6-tetrakisphosphate 5-diphosphatase activity(GO:0052848) |
| 0.7 | 3.3 | GO:0005298 | proline:sodium symporter activity(GO:0005298) |
| 0.7 | 5.9 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.6 | 7.1 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.6 | 4.5 | GO:0019826 | oxygen sensor activity(GO:0019826) |
| 0.6 | 2.5 | GO:0004306 | ethanolamine-phosphate cytidylyltransferase activity(GO:0004306) |
| 0.6 | 2.2 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.6 | 5.0 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.5 | 2.7 | GO:0046899 | nucleoside triphosphate adenylate kinase activity(GO:0046899) |
| 0.5 | 5.7 | GO:0035240 | dopamine binding(GO:0035240) |
| 0.5 | 4.9 | GO:0005384 | manganese ion transmembrane transporter activity(GO:0005384) |
| 0.5 | 3.9 | GO:1990599 | 3' overhang single-stranded DNA endodeoxyribonuclease activity(GO:1990599) |
| 0.5 | 15.9 | GO:0004467 | long-chain fatty acid-CoA ligase activity(GO:0004467) |
| 0.5 | 11.8 | GO:0005024 | transforming growth factor beta-activated receptor activity(GO:0005024) |
| 0.4 | 3.5 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.4 | 3.1 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.4 | 8.7 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.4 | 1.2 | GO:0015275 | stretch-activated, cation-selective, calcium channel activity(GO:0015275) |
| 0.4 | 4.1 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.4 | 1.6 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.4 | 1.6 | GO:0005277 | acetylcholine transmembrane transporter activity(GO:0005277) acetate ester transmembrane transporter activity(GO:1901375) |
| 0.4 | 11.0 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.4 | 2.6 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 0.4 | 14.4 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.4 | 5.1 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.4 | 11.5 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.4 | 1.4 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 0.3 | 1.7 | GO:0044729 | hemi-methylated DNA-binding(GO:0044729) |
| 0.3 | 1.4 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.3 | 6.4 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.3 | 1.0 | GO:0052739 | phosphatidylserine 1-acylhydrolase activity(GO:0052739) |
| 0.3 | 4.5 | GO:0003857 | 3-hydroxyacyl-CoA dehydrogenase activity(GO:0003857) |
| 0.3 | 1.6 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.3 | 0.9 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.3 | 0.9 | GO:0052827 | inositol tetrakisphosphate 1-kinase activity(GO:0047325) inositol tetrakisphosphate kinase activity(GO:0051765) inositol-1,3,4-trisphosphate 6-kinase activity(GO:0052725) inositol-1,3,4-trisphosphate 5-kinase activity(GO:0052726) inositol-1,3,4,5,6-pentakisphosphate 1-phosphatase activity(GO:0052825) inositol pentakisphosphate phosphatase activity(GO:0052827) inositol-1,3,4,6-tetrakisphosphate 6-phosphatase activity(GO:0052830) inositol-1,3,4,6-tetrakisphosphate 1-phosphatase activity(GO:0052831) inositol-3,4,6-trisphosphate 1-kinase activity(GO:0052835) |
| 0.3 | 9.7 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.3 | 7.8 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.3 | 7.8 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.3 | 11.6 | GO:0019956 | chemokine binding(GO:0019956) |
| 0.3 | 8.0 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.2 | 7.0 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.2 | 0.7 | GO:0055077 | gap junction hemi-channel activity(GO:0055077) |
| 0.2 | 2.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.2 | 2.8 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.2 | 0.9 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) |
| 0.2 | 2.2 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.2 | 0.9 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.2 | 3.6 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.2 | 1.3 | GO:0036033 | mediator complex binding(GO:0036033) |
| 0.2 | 3.3 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.2 | 2.2 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.2 | 2.1 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.2 | 4.2 | GO:0005112 | Notch binding(GO:0005112) |
| 0.2 | 5.8 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.2 | 2.0 | GO:0015386 | sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.2 | 10.9 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.2 | 2.9 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.2 | 3.8 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.2 | 3.1 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.2 | 0.9 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.2 | 2.5 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.2 | 4.1 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.2 | 3.4 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.2 | 5.2 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.2 | 12.4 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.2 | 0.8 | GO:0004803 | transposase activity(GO:0004803) |
| 0.2 | 1.1 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.2 | 33.0 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 0.9 | GO:0038052 | RNA polymerase II transcription factor activity, estrogen-activated sequence-specific DNA binding(GO:0038052) |
| 0.1 | 1.4 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.1 | 27.8 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.1 | 1.3 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.1 | 2.0 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.1 | 0.8 | GO:0016807 | cysteine-type carboxypeptidase activity(GO:0016807) cysteine-type exopeptidase activity(GO:0070004) |
| 0.1 | 1.1 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.1 | 1.3 | GO:0043125 | ErbB-3 class receptor binding(GO:0043125) |
| 0.1 | 2.7 | GO:0005003 | ephrin receptor activity(GO:0005003) |
| 0.1 | 4.5 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.1 | 0.3 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 0.1 | 4.7 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.1 | 1.9 | GO:0008392 | arachidonic acid monooxygenase activity(GO:0008391) arachidonic acid epoxygenase activity(GO:0008392) |
| 0.1 | 0.5 | GO:0036435 | K48-linked polyubiquitin binding(GO:0036435) |
| 0.1 | 1.9 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 1.5 | GO:0043548 | phosphatidylinositol 3-kinase binding(GO:0043548) |
| 0.1 | 1.8 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.1 | 1.4 | GO:0005351 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) |
| 0.1 | 4.7 | GO:0042379 | chemokine receptor binding(GO:0042379) |
| 0.1 | 42.8 | GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding(GO:0000978) |
| 0.1 | 2.8 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 2.5 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.1 | 16.5 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.1 | 2.2 | GO:0004298 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.1 | 6.9 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.1 | 0.7 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.1 | 16.8 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.1 | 2.8 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.1 | 8.3 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.1 | 0.7 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.1 | 1.7 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.1 | 0.6 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.1 | 1.3 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.1 | 1.4 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.1 | 1.1 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.1 | 7.3 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 2.1 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.1 | 1.1 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 1.7 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 1.7 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 1.4 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.1 | 1.7 | GO:0070566 | adenylyltransferase activity(GO:0070566) |
| 0.1 | 3.5 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.1 | 1.6 | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor(GO:0016628) |
| 0.0 | 0.5 | GO:0004118 | cGMP-stimulated cyclic-nucleotide phosphodiesterase activity(GO:0004118) |
| 0.0 | 0.4 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.0 | 2.7 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.4 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.0 | 1.4 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 1.6 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.0 | 1.6 | GO:0051183 | vitamin transporter activity(GO:0051183) |
| 0.0 | 2.0 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.0 | 2.3 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 7.0 | GO:0008168 | methyltransferase activity(GO:0008168) |
| 0.0 | 3.9 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 0.6 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.2 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.0 | 1.1 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 7.2 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 4.7 | GO:0000977 | RNA polymerase II regulatory region sequence-specific DNA binding(GO:0000977) |
| 0.0 | 1.8 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.4 | GO:0016405 | CoA-ligase activity(GO:0016405) |
| 0.0 | 10.1 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 1.1 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.0 | 0.6 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.0 | 0.7 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 3.2 | GO:0000981 | RNA polymerase II transcription factor activity, sequence-specific DNA binding(GO:0000981) |
| 0.0 | 0.1 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.0 | 2.0 | GO:0002020 | protease binding(GO:0002020) |
| 0.0 | 0.3 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.2 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.0 | GO:0051373 | FATZ binding(GO:0051373) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 20.9 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.6 | 3.1 | PID PI3KCI AKT PATHWAY | Class I PI3K signaling events mediated by Akt |
| 0.4 | 7.1 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.3 | 29.5 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.3 | 23.6 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.3 | 7.1 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.3 | 26.1 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.3 | 6.9 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.3 | 4.9 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.3 | 13.9 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.2 | 3.0 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.2 | 4.4 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.2 | 2.6 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.2 | 3.6 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.2 | 2.2 | PID FOXO PATHWAY | FoxO family signaling |
| 0.2 | 5.1 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.2 | 2.5 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.2 | 11.9 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.1 | 4.3 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.1 | 6.9 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 5.2 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.1 | 1.3 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.1 | 15.1 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.1 | 2.5 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.1 | 1.9 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.1 | 11.2 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.1 | 5.7 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 24.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 2.6 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.1 | 5.8 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.1 | 3.4 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.1 | 9.3 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.1 | 3.1 | ST G ALPHA I PATHWAY | G alpha i Pathway |
| 0.1 | 8.0 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.1 | 7.0 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.1 | 2.6 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.1 | 1.5 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 2.3 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.1 | 2.2 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.1 | 1.3 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 4.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.1 | 2.4 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.1 | 3.9 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.1 | 1.4 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 1.7 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.1 | 1.9 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.1 | 1.1 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 2.2 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.1 | 2.2 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.1 | 1.2 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.1 | 1.5 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.1 | 2.7 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.1 | 19.6 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.1 | 1.8 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.1 | 0.8 | PID EPO PATHWAY | EPO signaling pathway |
| 0.0 | 1.7 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 1.8 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 0.4 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 1.6 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 0.9 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 1.1 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 1.8 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.9 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 1.4 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 1.0 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 5.7 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.8 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 1.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 1.5 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 1.2 | 32.3 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.8 | 9.8 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.8 | 17.4 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.7 | 23.4 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.4 | 4.7 | REACTOME REGULATION OF INSULIN SECRETION BY GLUCAGON LIKE PEPTIDE1 | Genes involved in Regulation of Insulin Secretion by Glucagon-like Peptide-1 |
| 0.4 | 4.1 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.4 | 4.8 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.4 | 2.5 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.3 | 6.2 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.3 | 8.1 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.3 | 5.7 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.3 | 10.6 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.3 | 2.2 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.3 | 4.9 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.3 | 2.7 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.3 | 10.0 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.3 | 4.5 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.3 | 7.0 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.3 | 12.3 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.3 | 39.1 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.2 | 8.2 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.2 | 1.7 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.2 | 0.8 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.2 | 12.4 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.2 | 9.0 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.2 | 8.9 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.2 | 3.5 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.2 | 5.5 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.2 | 2.2 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.2 | 4.1 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.2 | 2.5 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.1 | 6.2 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.1 | 4.0 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 1.7 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 3.1 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.1 | 2.5 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 3.9 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.1 | 5.4 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.1 | 2.1 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.1 | 1.4 | REACTOME CDC6 ASSOCIATION WITH THE ORC ORIGIN COMPLEX | Genes involved in CDC6 association with the ORC:origin complex |
| 0.1 | 4.4 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 1.4 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.1 | 1.7 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.1 | 1.5 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.1 | 5.4 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.1 | 2.5 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.1 | 1.8 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 6.0 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.1 | 2.1 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.1 | 1.0 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 1.4 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.1 | 1.4 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.1 | 2.3 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.1 | 9.2 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.1 | 1.3 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.1 | 5.5 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 0.7 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.1 | 1.1 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.1 | 1.9 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.1 | 1.6 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 0.1 | 1.7 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.1 | 3.5 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.1 | 10.5 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.1 | 1.3 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 2.1 | REACTOME TRAF6 MEDIATED NFKB ACTIVATION | Genes involved in TRAF6 mediated NF-kB activation |
| 0.1 | 1.1 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.1 | 2.7 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 3.6 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.1 | 7.7 | REACTOME G ALPHA Q SIGNALLING EVENTS | Genes involved in G alpha (q) signalling events |
| 0.1 | 2.8 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.9 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 2.1 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 2.7 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 2.1 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 2.7 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.6 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.4 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |
| 0.0 | 0.7 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 1.3 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 1.4 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.2 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.0 | 0.1 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.0 | 3.4 | REACTOME GENERIC TRANSCRIPTION PATHWAY | Genes involved in Generic Transcription Pathway |
| 0.0 | 0.5 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.5 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |