Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
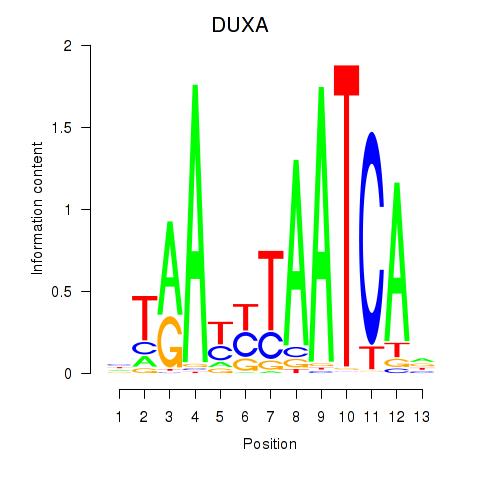
Results for DUXA
Z-value: 0.88
Transcription factors associated with DUXA
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
DUXA
|
ENSG00000258873.3 | DUXA |
Activity profile of DUXA motif
Sorted Z-values of DUXA motif
Network of associatons between targets according to the STRING database.
First level regulatory network of DUXA
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_50968125 | 5.53 |
ENST00000594641.1
|
KLK6
|
kallikrein related peptidase 6 |
| chr19_-_50968775 | 4.15 |
ENST00000391808.5
|
KLK6
|
kallikrein related peptidase 6 |
| chr4_+_68447453 | 3.55 |
ENST00000305363.9
|
TMPRSS11E
|
transmembrane serine protease 11E |
| chr8_-_7781413 | 3.49 |
ENST00000528972.1
|
PRR23D2
|
proline rich 23 domain containing 2 |
| chr8_+_7539627 | 3.44 |
ENST00000533250.2
|
PRR23D1
|
proline rich 23 domain containing 1 |
| chr12_-_57237090 | 2.56 |
ENST00000556732.1
|
NDUFA4L2
|
NDUFA4 mitochondrial complex associated like 2 |
| chr2_+_227813834 | 2.35 |
ENST00000358813.5
ENST00000409189.7 |
CCL20
|
C-C motif chemokine ligand 20 |
| chr22_+_31082860 | 2.32 |
ENST00000619644.4
|
SMTN
|
smoothelin |
| chr11_-_89920428 | 2.26 |
ENST00000605881.5
|
TRIM49D1
|
tripartite motif containing 49D1 |
| chr1_+_17249088 | 2.20 |
ENST00000375460.3
|
PADI3
|
peptidyl arginine deiminase 3 |
| chr15_+_41286011 | 2.08 |
ENST00000661438.1
|
ENSG00000285920.2
|
novel protein |
| chr18_+_63777773 | 2.06 |
ENST00000447428.5
ENST00000546027.5 |
SERPINB7
|
serpin family B member 7 |
| chr11_-_48983826 | 1.95 |
ENST00000649162.1
|
TRIM51GP
|
tripartite motif-containing 51G, pseudogene |
| chr7_-_99679987 | 1.79 |
ENST00000222982.8
ENST00000439761.3 ENST00000339843.6 |
CYP3A5
|
cytochrome P450 family 3 subfamily A member 5 |
| chr2_+_233691607 | 1.78 |
ENST00000373424.5
ENST00000441351.1 |
UGT1A6
|
UDP glucuronosyltransferase family 1 member A6 |
| chr1_+_12773738 | 1.76 |
ENST00000357726.5
|
PRAMEF12
|
PRAME family member 12 |
| chr17_-_41397600 | 1.75 |
ENST00000251645.3
|
KRT31
|
keratin 31 |
| chr11_+_89924064 | 1.71 |
ENST00000623787.3
|
TRIM49D2
|
tripartite motif containing 49D2 |
| chr11_-_28108109 | 1.68 |
ENST00000263181.7
|
KIF18A
|
kinesin family member 18A |
| chr6_-_150025520 | 1.58 |
ENST00000367341.6
ENST00000286380.2 |
RAET1L
|
retinoic acid early transcript 1L |
| chr8_-_124565699 | 1.46 |
ENST00000519168.5
|
MTSS1
|
MTSS I-BAR domain containing 1 |
| chr1_+_13068677 | 1.41 |
ENST00000614839.4
|
PRAMEF25
|
PRAME family member 25 |
| chr15_-_79971164 | 1.40 |
ENST00000335661.6
ENST00000267953.4 ENST00000677151.1 |
BCL2A1
|
BCL2 related protein A1 |
| chr2_-_208129824 | 1.37 |
ENST00000282141.4
|
CRYGC
|
crystallin gamma C |
| chr19_-_7058640 | 1.35 |
ENST00000333843.8
|
MBD3L3
|
methyl-CpG binding domain protein 3 like 3 |
| chr3_-_139539577 | 1.33 |
ENST00000619087.4
|
RBP1
|
retinol binding protein 1 |
| chr19_+_7030578 | 1.32 |
ENST00000329753.5
|
MBD3L5
|
methyl-CpG binding domain protein 3 like 5 |
| chr14_+_20469399 | 1.31 |
ENST00000361505.10
ENST00000553591.1 |
PNP
|
purine nucleoside phosphorylase |
| chr10_+_76318330 | 1.31 |
ENST00000496424.2
|
LRMDA
|
leucine rich melanocyte differentiation associated |
| chr20_+_23491090 | 1.30 |
ENST00000449810.5
ENST00000246012.2 |
CST8
|
cystatin 8 |
| chr19_-_7021431 | 1.28 |
ENST00000636986.2
ENST00000637800.1 |
MBD3L2B
|
methyl-CpG binding domain protein 3 like 2B |
| chr14_+_23630109 | 1.28 |
ENST00000432832.6
|
DHRS2
|
dehydrogenase/reductase 2 |
| chr7_+_124476371 | 1.28 |
ENST00000473520.1
|
SSU72P8
|
SSU72 pseudogene 8 |
| chr12_-_6635938 | 1.26 |
ENST00000329858.9
|
LPAR5
|
lysophosphatidic acid receptor 5 |
| chr1_-_110390989 | 1.23 |
ENST00000369779.9
ENST00000472422.6 |
SLC16A4
|
solute carrier family 16 member 4 |
| chr19_+_7049321 | 1.18 |
ENST00000381393.3
|
MBD3L2
|
methyl-CpG binding domain protein 3 like 2 |
| chr6_+_121437378 | 1.18 |
ENST00000650427.1
ENST00000647564.1 |
GJA1
|
gap junction protein alpha 1 |
| chr14_-_63318933 | 1.15 |
ENST00000621500.2
|
GPHB5
|
glycoprotein hormone subunit beta 5 |
| chr6_+_54018992 | 1.10 |
ENST00000509997.5
|
MLIP
|
muscular LMNA interacting protein |
| chr10_+_11823348 | 1.08 |
ENST00000277570.10
ENST00000622831.4 |
PROSER2
|
proline and serine rich 2 |
| chr1_-_110391041 | 1.06 |
ENST00000369781.8
ENST00000437429.6 ENST00000541986.5 |
SLC16A4
|
solute carrier family 16 member 4 |
| chr9_+_72577369 | 1.06 |
ENST00000651183.1
|
TMC1
|
transmembrane channel like 1 |
| chr12_-_119877300 | 1.04 |
ENST00000392521.7
|
CIT
|
citron rho-interacting serine/threonine kinase |
| chr19_-_7040179 | 1.04 |
ENST00000381394.9
|
MBD3L4
|
methyl-CpG binding domain protein 3 like 4 |
| chr12_-_119877270 | 1.03 |
ENST00000261833.11
ENST00000612548.4 |
CIT
|
citron rho-interacting serine/threonine kinase |
| chr4_+_147732070 | 1.01 |
ENST00000336498.8
|
ARHGAP10
|
Rho GTPase activating protein 10 |
| chr7_+_141790217 | 1.00 |
ENST00000247883.5
|
TAS2R5
|
taste 2 receptor member 5 |
| chr2_-_40453438 | 0.93 |
ENST00000455476.5
|
SLC8A1
|
solute carrier family 8 member A1 |
| chr8_-_90082871 | 0.92 |
ENST00000265431.7
|
CALB1
|
calbindin 1 |
| chr19_-_6393205 | 0.92 |
ENST00000595047.5
|
GTF2F1
|
general transcription factor IIF subunit 1 |
| chr2_-_55334529 | 0.91 |
ENST00000645860.1
ENST00000642563.1 ENST00000647396.1 |
CCDC88A
|
coiled-coil domain containing 88A |
| chr6_-_111567271 | 0.90 |
ENST00000464903.1
ENST00000368730.5 ENST00000368735.1 |
TRAF3IP2
|
TRAF3 interacting protein 2 |
| chr5_+_151025343 | 0.89 |
ENST00000521632.1
|
GPX3
|
glutathione peroxidase 3 |
| chr11_-_4288083 | 0.89 |
ENST00000638166.1
|
SSU72P4
|
SSU72 pseudogene 4 |
| chr13_+_108596152 | 0.88 |
ENST00000356711.7
ENST00000251041.10 |
MYO16
|
myosin XVI |
| chr14_+_22147988 | 0.88 |
ENST00000390457.2
|
TRAV27
|
T cell receptor alpha variable 27 |
| chrX_+_100666854 | 0.86 |
ENST00000640282.1
|
SRPX2
|
sushi repeat containing protein X-linked 2 |
| chr6_-_111567120 | 0.85 |
ENST00000368734.5
|
TRAF3IP2
|
TRAF3 interacting protein 2 |
| chr19_-_6393131 | 0.84 |
ENST00000394456.10
|
GTF2F1
|
general transcription factor IIF subunit 1 |
| chr6_+_31158518 | 0.83 |
ENST00000376255.4
ENST00000376257.8 |
TCF19
|
transcription factor 19 |
| chr9_+_34652167 | 0.82 |
ENST00000441545.7
ENST00000553620.5 |
IL11RA
|
interleukin 11 receptor subunit alpha |
| chr2_-_174764436 | 0.76 |
ENST00000409323.1
ENST00000261007.9 ENST00000348749.9 ENST00000672640.1 |
CHRNA1
|
cholinergic receptor nicotinic alpha 1 subunit |
| chr7_-_93890744 | 0.76 |
ENST00000650573.1
ENST00000222543.11 ENST00000649913.1 ENST00000647793.1 |
TFPI2
|
tissue factor pathway inhibitor 2 |
| chr1_+_13070853 | 0.75 |
ENST00000619661.2
|
PRAMEF25
|
PRAME family member 25 |
| chr6_+_54018910 | 0.71 |
ENST00000514921.5
ENST00000274897.9 ENST00000370877.6 |
MLIP
|
muscular LMNA interacting protein |
| chr11_-_4339244 | 0.71 |
ENST00000524542.2
|
SSU72P7
|
SSU72 pseudogene 7 |
| chr1_-_12898270 | 0.70 |
ENST00000235347.4
|
PRAMEF10
|
PRAME family member 10 |
| chr11_-_4242640 | 0.69 |
ENST00000640805.1
|
SSU72P2
|
SSU72 pseudogene 2 |
| chr14_-_80231052 | 0.68 |
ENST00000557010.5
|
DIO2
|
iodothyronine deiodinase 2 |
| chr13_+_57140918 | 0.67 |
ENST00000377931.1
ENST00000614894.4 |
PRR20A
PRR20C
|
proline rich 20A proline rich 20C |
| chr11_-_89921767 | 0.66 |
ENST00000530311.6
|
TRIM49D1
|
tripartite motif containing 49D1 |
| chrX_+_86714623 | 0.66 |
ENST00000484479.1
|
DACH2
|
dachshund family transcription factor 2 |
| chr18_+_63887698 | 0.66 |
ENST00000457692.5
ENST00000299502.9 ENST00000413956.5 |
SERPINB2
|
serpin family B member 2 |
| chr1_+_153031195 | 0.64 |
ENST00000307098.5
|
SPRR1B
|
small proline rich protein 1B |
| chr5_-_113434978 | 0.64 |
ENST00000390666.4
|
TSSK1B
|
testis specific serine kinase 1B |
| chr1_-_13201409 | 0.63 |
ENST00000625019.3
|
PRAMEF13
|
PRAME family member 13 |
| chr11_+_4233288 | 0.62 |
ENST00000639584.1
|
SSU72P5
|
SSU72 pseudogene 5 |
| chr8_+_69466617 | 0.61 |
ENST00000525061.5
ENST00000260128.8 ENST00000458141.6 |
SULF1
|
sulfatase 1 |
| chr6_+_36029082 | 0.60 |
ENST00000472333.1
|
MAPK14
|
mitogen-activated protein kinase 14 |
| chr13_+_57147488 | 0.59 |
ENST00000377930.1
|
PRR20B
|
proline rich 20B |
| chr10_-_29634964 | 0.59 |
ENST00000375398.6
ENST00000355867.8 |
SVIL
|
supervillin |
| chr11_+_28108248 | 0.59 |
ENST00000406787.7
ENST00000403099.5 ENST00000407364.8 |
METTL15
|
methyltransferase like 15 |
| chr6_-_65707214 | 0.59 |
ENST00000370621.7
ENST00000393380.6 ENST00000503581.6 |
EYS
|
eyes shut homolog |
| chr13_+_32031706 | 0.58 |
ENST00000542859.6
|
FRY
|
FRY microtubule binding protein |
| chr1_+_13060769 | 0.58 |
ENST00000617807.3
|
HNRNPCL3
|
heterogeneous nuclear ribonucleoprotein C like 3 |
| chr3_+_45886501 | 0.57 |
ENST00000395963.2
|
CCR9
|
C-C motif chemokine receptor 9 |
| chr4_+_87608529 | 0.55 |
ENST00000651931.1
|
DSPP
|
dentin sialophosphoprotein |
| chr1_-_173824856 | 0.55 |
ENST00000682279.1
|
CENPL
|
centromere protein L |
| chr11_-_13496018 | 0.55 |
ENST00000529816.1
|
PTH
|
parathyroid hormone |
| chr5_+_154858218 | 0.54 |
ENST00000523698.5
ENST00000517876.5 ENST00000520472.5 |
CNOT8
|
CCR4-NOT transcription complex subunit 8 |
| chr11_+_114400030 | 0.53 |
ENST00000540163.5
|
RBM7
|
RNA binding motif protein 7 |
| chr16_-_21211566 | 0.53 |
ENST00000574091.6
|
ZP2
|
zona pellucida glycoprotein 2 |
| chr1_-_12945416 | 0.53 |
ENST00000415464.6
|
PRAMEF6
|
PRAME family member 6 |
| chr15_+_66453418 | 0.53 |
ENST00000566326.1
|
MAP2K1
|
mitogen-activated protein kinase kinase 1 |
| chr19_-_14674829 | 0.53 |
ENST00000443157.6
ENST00000253673.6 |
ADGRE3
|
adhesion G protein-coupled receptor E3 |
| chr19_-_14674886 | 0.52 |
ENST00000344373.8
ENST00000595472.1 |
ADGRE3
|
adhesion G protein-coupled receptor E3 |
| chr10_-_110304894 | 0.52 |
ENST00000369603.10
|
SMNDC1
|
survival motor neuron domain containing 1 |
| chr1_-_13116854 | 0.52 |
ENST00000621994.3
|
HNRNPCL2
|
heterogeneous nuclear ribonucleoprotein C like 2 |
| chr12_+_56128217 | 0.51 |
ENST00000267113.4
ENST00000394048.10 |
ESYT1
|
extended synaptotagmin 1 |
| chr1_-_12848720 | 0.51 |
ENST00000317869.7
|
HNRNPCL1
|
heterogeneous nuclear ribonucleoprotein C like 1 |
| chr16_-_15056060 | 0.51 |
ENST00000287706.8
ENST00000624579.3 ENST00000622833.4 |
NTAN1
|
N-terminal asparagine amidase |
| chr1_+_13303539 | 0.50 |
ENST00000437300.2
|
PRAMEF33
|
PRAME family member 33 |
| chr12_-_10909562 | 0.50 |
ENST00000390677.2
|
TAS2R13
|
taste 2 receptor member 13 |
| chr19_-_46846138 | 0.49 |
ENST00000597020.5
|
AP2S1
|
adaptor related protein complex 2 subunit sigma 1 |
| chr7_+_93906557 | 0.48 |
ENST00000248572.10
ENST00000429473.1 ENST00000430875.1 ENST00000428834.1 |
GNGT1
|
G protein subunit gamma transducin 1 |
| chr16_+_29995661 | 0.47 |
ENST00000304516.11
|
INO80E
|
INO80 complex subunit E |
| chr12_-_102480604 | 0.47 |
ENST00000392905.7
|
IGF1
|
insulin like growth factor 1 |
| chr12_+_50504970 | 0.47 |
ENST00000301180.10
|
DIP2B
|
disco interacting protein 2 homolog B |
| chr2_+_161136901 | 0.47 |
ENST00000259075.6
ENST00000432002.5 |
TANK
|
TRAF family member associated NFKB activator |
| chr1_+_13171848 | 0.47 |
ENST00000415919.3
|
PRAMEF9
|
PRAME family member 9 |
| chr17_-_10373002 | 0.47 |
ENST00000252172.9
ENST00000418404.8 |
MYH13
|
myosin heavy chain 13 |
| chr1_-_12886201 | 0.47 |
ENST00000235349.6
|
PRAMEF4
|
PRAME family member 4 |
| chr16_-_15055969 | 0.47 |
ENST00000566419.1
ENST00000568320.5 |
NTAN1
|
N-terminal asparagine amidase |
| chr9_-_92878018 | 0.47 |
ENST00000332591.6
ENST00000375495.8 ENST00000395505.6 ENST00000395506.7 |
ZNF484
|
zinc finger protein 484 |
| chr6_-_49636832 | 0.46 |
ENST00000371175.10
ENST00000646272.1 ENST00000646939.1 ENST00000618248.3 ENST00000229810.9 ENST00000646963.1 |
RHAG
|
Rh associated glycoprotein |
| chrX_+_37780049 | 0.46 |
ENST00000378588.5
|
CYBB
|
cytochrome b-245 beta chain |
| chr8_+_52939182 | 0.46 |
ENST00000674939.1
|
NPBWR1
|
neuropeptides B and W receptor 1 |
| chr11_+_68903849 | 0.45 |
ENST00000675615.1
ENST00000255078.8 |
IGHMBP2
|
immunoglobulin mu DNA binding protein 2 |
| chr12_+_12725897 | 0.45 |
ENST00000326765.10
|
APOLD1
|
apolipoprotein L domain containing 1 |
| chr2_+_201129483 | 0.45 |
ENST00000440180.5
|
CFLAR
|
CASP8 and FADD like apoptosis regulator |
| chr3_-_196270540 | 0.44 |
ENST00000419333.5
|
PCYT1A
|
phosphate cytidylyltransferase 1, choline, alpha |
| chr15_+_45252228 | 0.44 |
ENST00000560438.5
ENST00000347644.8 |
SLC28A2
|
solute carrier family 28 member 2 |
| chr6_+_31971831 | 0.44 |
ENST00000375331.7
ENST00000375333.3 |
STK19
|
serine/threonine kinase 19 |
| chr9_-_27005659 | 0.43 |
ENST00000380055.6
|
LRRC19
|
leucine rich repeat containing 19 |
| chr13_+_97953652 | 0.43 |
ENST00000460070.6
ENST00000481455.6 ENST00000261574.10 ENST00000651721.2 ENST00000493281.6 ENST00000463157.6 ENST00000471898.5 ENST00000489058.6 ENST00000481689.6 |
IPO5
|
importin 5 |
| chr11_+_95790459 | 0.43 |
ENST00000325486.9
ENST00000325542.10 ENST00000544522.5 ENST00000541365.5 |
CEP57
|
centrosomal protein 57 |
| chr5_+_150497772 | 0.42 |
ENST00000523767.5
|
NDST1
|
N-deacetylase and N-sulfotransferase 1 |
| chr8_-_48921419 | 0.42 |
ENST00000020945.4
|
SNAI2
|
snail family transcriptional repressor 2 |
| chrX_+_77910656 | 0.41 |
ENST00000343533.9
ENST00000341514.11 ENST00000645454.1 ENST00000642651.1 ENST00000644362.1 |
ATP7A
PGK1
|
ATPase copper transporting alpha phosphoglycerate kinase 1 |
| chr1_+_13254212 | 0.41 |
ENST00000622421.2
|
PRAMEF5
|
PRAME family member 5 |
| chr1_+_13061158 | 0.40 |
ENST00000681473.1
|
HNRNPCL3
|
heterogeneous nuclear ribonucleoprotein C like 3 |
| chr16_-_89719369 | 0.40 |
ENST00000561976.5
|
VPS9D1
|
VPS9 domain containing 1 |
| chr1_-_13165631 | 0.40 |
ENST00000323770.8
|
HNRNPCL4
|
heterogeneous nuclear ribonucleoprotein C like 4 |
| chr6_+_168017873 | 0.39 |
ENST00000351261.4
ENST00000354419.6 |
KIF25
|
kinesin family member 25 |
| chr3_+_160225409 | 0.38 |
ENST00000326474.5
|
C3orf80
|
chromosome 3 open reading frame 80 |
| chr1_+_95151377 | 0.38 |
ENST00000604203.1
|
TLCD4-RWDD3
|
TLCD4-RWDD3 readthrough |
| chr13_+_57154061 | 0.37 |
ENST00000544357.2
|
PRR20C
|
proline rich 20C |
| chr13_+_57160632 | 0.37 |
ENST00000452123.3
|
PRR20D
|
proline rich 20D |
| chr13_+_57167197 | 0.37 |
ENST00000434815.1
|
PRR20E
|
proline rich 20E |
| chr1_-_116667668 | 0.37 |
ENST00000369486.8
ENST00000369483.5 |
IGSF3
|
immunoglobulin superfamily member 3 |
| chr4_-_8428424 | 0.37 |
ENST00000514423.1
ENST00000503233.5 |
ACOX3
|
acyl-CoA oxidase 3, pristanoyl |
| chr11_-_11353241 | 0.37 |
ENST00000528848.3
|
CSNK2A3
|
casein kinase 2 alpha 3 |
| chr16_+_6483728 | 0.37 |
ENST00000675459.1
ENST00000551752.5 |
RBFOX1
|
RNA binding fox-1 homolog 1 |
| chr17_+_36486668 | 0.37 |
ENST00000617429.5
ENST00000620324.4 ENST00000616269.1 |
ZNHIT3
|
zinc finger HIT-type containing 3 |
| chr16_+_29996201 | 0.36 |
ENST00000620599.4
ENST00000563197.6 ENST00000567254.5 ENST00000567705.5 |
INO80E
|
INO80 complex subunit E |
| chr6_-_138499487 | 0.36 |
ENST00000343505.9
|
NHSL1
|
NHS like 1 |
| chr8_-_7486400 | 0.36 |
ENST00000335479.2
|
DEFB106B
|
defensin beta 106B |
| chr5_-_16916400 | 0.36 |
ENST00000513882.5
|
MYO10
|
myosin X |
| chr12_-_122730828 | 0.36 |
ENST00000432564.3
|
HCAR1
|
hydroxycarboxylic acid receptor 1 |
| chr2_-_68871382 | 0.36 |
ENST00000295379.2
|
BMP10
|
bone morphogenetic protein 10 |
| chr3_-_71360753 | 0.36 |
ENST00000648783.1
|
FOXP1
|
forkhead box P1 |
| chr20_-_17558811 | 0.35 |
ENST00000536626.7
ENST00000377868.6 |
BFSP1
|
beaded filament structural protein 1 |
| chr14_-_54441325 | 0.35 |
ENST00000556113.1
ENST00000553660.5 ENST00000216416.9 ENST00000395573.8 ENST00000557690.5 |
CNIH1
|
cornichon family AMPA receptor auxiliary protein 1 |
| chr8_+_7825137 | 0.35 |
ENST00000335186.3
|
DEFB106A
|
defensin beta 106A |
| chr2_-_174764407 | 0.35 |
ENST00000409219.5
ENST00000409542.5 |
CHRNA1
|
cholinergic receptor nicotinic alpha 1 subunit |
| chr2_-_200889266 | 0.35 |
ENST00000443398.5
ENST00000286175.12 ENST00000409449.5 |
PPIL3
|
peptidylprolyl isomerase like 3 |
| chr10_-_32935511 | 0.35 |
ENST00000423113.5
|
ITGB1
|
integrin subunit beta 1 |
| chr1_+_74235377 | 0.35 |
ENST00000326637.8
|
TNNI3K
|
TNNI3 interacting kinase |
| chr12_+_18262730 | 0.34 |
ENST00000675017.1
|
PIK3C2G
|
phosphatidylinositol-4-phosphate 3-kinase catalytic subunit type 2 gamma |
| chr2_+_169066994 | 0.34 |
ENST00000357546.6
ENST00000432060.6 |
DHRS9
|
dehydrogenase/reductase 9 |
| chr12_-_48570046 | 0.34 |
ENST00000301046.6
ENST00000549817.1 |
LALBA
|
lactalbumin alpha |
| chr12_-_48865863 | 0.34 |
ENST00000309739.6
|
RND1
|
Rho family GTPase 1 |
| chr1_-_12947580 | 0.33 |
ENST00000376189.5
|
PRAMEF6
|
PRAME family member 6 |
| chr12_-_54259531 | 0.33 |
ENST00000550411.5
ENST00000439541.6 |
CBX5
|
chromobox 5 |
| chr12_+_8513499 | 0.33 |
ENST00000299665.3
|
CLEC4D
|
C-type lectin domain family 4 member D |
| chr8_-_17002327 | 0.33 |
ENST00000180166.6
|
FGF20
|
fibroblast growth factor 20 |
| chr11_+_56176618 | 0.33 |
ENST00000312298.1
|
OR5J2
|
olfactory receptor family 5 subfamily J member 2 |
| chr3_-_128467248 | 0.33 |
ENST00000319153.3
|
DNAJB8
|
DnaJ heat shock protein family (Hsp40) member B8 |
| chr2_-_200889136 | 0.32 |
ENST00000409361.5
|
PPIL3
|
peptidylprolyl isomerase like 3 |
| chr3_-_125055987 | 0.32 |
ENST00000311127.9
|
HEG1
|
heart development protein with EGF like domains 1 |
| chr5_-_78294656 | 0.32 |
ENST00000519295.5
ENST00000255194.11 |
AP3B1
|
adaptor related protein complex 3 subunit beta 1 |
| chr2_+_196713117 | 0.32 |
ENST00000409270.5
|
CCDC150
|
coiled-coil domain containing 150 |
| chr5_-_79514127 | 0.32 |
ENST00000334082.11
|
HOMER1
|
homer scaffold protein 1 |
| chr6_+_167291329 | 0.32 |
ENST00000366829.2
|
UNC93A
|
unc-93 homolog A |
| chr2_-_200888993 | 0.31 |
ENST00000409264.6
ENST00000392283.9 |
PPIL3
|
peptidylprolyl isomerase like 3 |
| chr12_-_118190510 | 0.31 |
ENST00000540561.5
ENST00000537952.1 ENST00000537822.1 |
TAOK3
|
TAO kinase 3 |
| chrX_+_38006551 | 0.31 |
ENST00000297875.7
|
SYTL5
|
synaptotagmin like 5 |
| chr6_+_167291309 | 0.30 |
ENST00000230256.8
|
UNC93A
|
unc-93 homolog A |
| chrX_+_23667461 | 0.30 |
ENST00000379341.9
ENST00000379331.3 |
PRDX4
|
peroxiredoxin 4 |
| chr16_+_6483813 | 0.30 |
ENST00000675653.1
|
RBFOX1
|
RNA binding fox-1 homolog 1 |
| chr3_-_114199407 | 0.30 |
ENST00000460779.5
|
DRD3
|
dopamine receptor D3 |
| chr12_-_91153149 | 0.30 |
ENST00000550758.1
|
DCN
|
decorin |
| chr5_-_181238195 | 0.30 |
ENST00000509148.1
|
RACK1
|
receptor for activated C kinase 1 |
| chr1_+_13315581 | 0.29 |
ENST00000376152.2
|
PRAMEF15
|
PRAME family member 15 |
| chr17_+_36486629 | 0.29 |
ENST00000619730.4
|
ZNHIT3
|
zinc finger HIT-type containing 3 |
| chr1_+_224356852 | 0.29 |
ENST00000366858.7
ENST00000366857.9 ENST00000465271.6 ENST00000366856.3 |
CNIH4
|
cornichon family AMPA receptor auxiliary protein 4 |
| chr20_-_45670239 | 0.28 |
ENST00000324384.4
ENST00000356562.6 |
WFDC11
|
WAP four-disulfide core domain 11 |
| chr2_+_159733958 | 0.28 |
ENST00000409591.5
|
MARCHF7
|
membrane associated ring-CH-type finger 7 |
| chr3_-_196515315 | 0.28 |
ENST00000397537.3
|
SMCO1
|
single-pass membrane protein with coiled-coil domains 1 |
| chr9_-_92536031 | 0.28 |
ENST00000344604.9
ENST00000375540.5 |
ECM2
|
extracellular matrix protein 2 |
| chr12_+_8509460 | 0.28 |
ENST00000382064.6
|
CLEC4D
|
C-type lectin domain family 4 member D |
| chr4_+_55853639 | 0.28 |
ENST00000381295.7
ENST00000346134.11 ENST00000349598.6 |
EXOC1
|
exocyst complex component 1 |
| chr12_-_118359639 | 0.27 |
ENST00000541786.5
ENST00000419821.6 ENST00000541878.5 |
TAOK3
|
TAO kinase 3 |
| chr3_-_177196451 | 0.27 |
ENST00000430069.5
ENST00000630796.2 ENST00000428970.5 |
TBL1XR1
|
TBL1X receptor 1 |
| chr19_+_52429181 | 0.27 |
ENST00000301085.8
|
ZNF534
|
zinc finger protein 534 |
| chr4_+_146175702 | 0.27 |
ENST00000296581.11
ENST00000649747.1 ENST00000502781.5 |
LSM6
|
LSM6 homolog, U6 small nuclear RNA and mRNA degradation associated |
| chr6_+_26199509 | 0.27 |
ENST00000356530.5
|
H2BC7
|
H2B clustered histone 7 |
| chr16_+_57186281 | 0.27 |
ENST00000564435.5
ENST00000562959.1 ENST00000568505.6 ENST00000394420.9 ENST00000537866.5 |
RSPRY1
|
ring finger and SPRY domain containing 1 |
| chr11_-_73761051 | 0.26 |
ENST00000336083.8
ENST00000536566.5 ENST00000541588.5 ENST00000540771.5 ENST00000310653.10 |
RAB6A
|
RAB6A, member RAS oncogene family |
| chr6_+_29170907 | 0.26 |
ENST00000641417.1
|
OR2J2
|
olfactory receptor family 2 subfamily J member 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.4 | GO:0045362 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 0.5 | 2.2 | GO:0018101 | protein citrullination(GO:0018101) histone citrullination(GO:0036414) |
| 0.5 | 9.7 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.4 | 1.8 | GO:0009822 | alkaloid catabolic process(GO:0009822) |
| 0.4 | 2.1 | GO:0090362 | positive regulation of platelet-derived growth factor production(GO:0090362) |
| 0.3 | 1.2 | GO:0010645 | regulation of cell communication by chemical coupling(GO:0010645) positive regulation of cell communication by chemical coupling(GO:0010652) |
| 0.3 | 1.5 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.2 | 0.4 | GO:1901420 | negative regulation of response to alcohol(GO:1901420) |
| 0.2 | 0.6 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.2 | 0.6 | GO:0002305 | gamma-delta intraepithelial T cell differentiation(GO:0002304) CD8-positive, gamma-delta intraepithelial T cell differentiation(GO:0002305) |
| 0.2 | 4.9 | GO:0006346 | methylation-dependent chromatin silencing(GO:0006346) |
| 0.2 | 1.3 | GO:0072526 | pyridine-containing compound catabolic process(GO:0072526) |
| 0.2 | 0.3 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 0.2 | 1.9 | GO:0052697 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.2 | 0.5 | GO:1904844 | response to L-glutamine(GO:1904844) cellular response to L-glutamine(GO:1904845) |
| 0.2 | 0.9 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.1 | 0.7 | GO:1904075 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.1 | 0.9 | GO:0006982 | response to lipid hydroperoxide(GO:0006982) |
| 0.1 | 0.6 | GO:1904428 | negative regulation of tubulin deacetylation(GO:1904428) |
| 0.1 | 2.0 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.1 | 0.5 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) |
| 0.1 | 0.4 | GO:0071284 | copper ion export(GO:0060003) cellular response to lead ion(GO:0071284) |
| 0.1 | 0.4 | GO:0046603 | negative regulation of mitotic centrosome separation(GO:0046603) |
| 0.1 | 0.5 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.1 | 0.3 | GO:0005989 | lactose metabolic process(GO:0005988) lactose biosynthetic process(GO:0005989) |
| 0.1 | 0.5 | GO:0007343 | egg activation(GO:0007343) |
| 0.1 | 1.1 | GO:0060005 | vestibular reflex(GO:0060005) |
| 0.1 | 1.2 | GO:0002155 | regulation of thyroid hormone mediated signaling pathway(GO:0002155) |
| 0.1 | 0.6 | GO:2001184 | myoblast differentiation involved in skeletal muscle regeneration(GO:0014835) positive regulation of interleukin-12 secretion(GO:2001184) |
| 0.1 | 1.7 | GO:0002138 | retinoic acid biosynthetic process(GO:0002138) |
| 0.1 | 0.3 | GO:0015917 | aminophospholipid transport(GO:0015917) |
| 0.1 | 0.9 | GO:1903944 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 0.3 | GO:0007206 | phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.1 | 0.5 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.1 | 0.9 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
| 0.1 | 0.5 | GO:0090170 | regulation of Golgi inheritance(GO:0090170) |
| 0.1 | 0.3 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
| 0.1 | 0.3 | GO:0032416 | negative regulation of sodium:proton antiporter activity(GO:0032416) musculoskeletal movement, spinal reflex action(GO:0050883) |
| 0.1 | 1.7 | GO:0072520 | seminiferous tubule development(GO:0072520) |
| 0.1 | 0.4 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.1 | 0.9 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.1 | 0.3 | GO:0018125 | peptidyl-cysteine methylation(GO:0018125) |
| 0.1 | 0.6 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.1 | 1.3 | GO:0032968 | positive regulation of transcription elongation from RNA polymerase II promoter(GO:0032968) |
| 0.1 | 1.7 | GO:0002230 | positive regulation of defense response to virus by host(GO:0002230) |
| 0.1 | 0.3 | GO:0051138 | positive regulation of NK T cell differentiation(GO:0051138) |
| 0.1 | 0.4 | GO:0015014 | heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) |
| 0.1 | 3.1 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.1 | 0.2 | GO:0031959 | mineralocorticoid receptor signaling pathway(GO:0031959) positive regulation of cardiac vascular smooth muscle cell differentiation(GO:2000724) |
| 0.1 | 0.4 | GO:0060298 | positive regulation of sarcomere organization(GO:0060298) |
| 0.0 | 0.3 | GO:0007161 | calcium-independent cell-matrix adhesion(GO:0007161) regulation of collagen catabolic process(GO:0010710) |
| 0.0 | 0.2 | GO:0043605 | cellular amide catabolic process(GO:0043605) |
| 0.0 | 0.4 | GO:0015860 | purine nucleoside transmembrane transport(GO:0015860) |
| 0.0 | 0.2 | GO:0042427 | serotonin biosynthetic process(GO:0042427) |
| 0.0 | 0.1 | GO:1901674 | histone H3-K27 acetylation(GO:0043974) regulation of histone H3-K27 acetylation(GO:1901674) |
| 0.0 | 0.2 | GO:1990168 | protein K27-linked deubiquitination(GO:1990167) protein K33-linked deubiquitination(GO:1990168) |
| 0.0 | 0.4 | GO:0006657 | CDP-choline pathway(GO:0006657) |
| 0.0 | 0.7 | GO:0006451 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.0 | 0.2 | GO:0036324 | vascular endothelial growth factor receptor-2 signaling pathway(GO:0036324) |
| 0.0 | 0.2 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 0.0 | 1.3 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.0 | 0.3 | GO:0051343 | positive regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051343) |
| 0.0 | 0.4 | GO:0097475 | motor neuron migration(GO:0097475) |
| 0.0 | 0.2 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.0 | 0.1 | GO:0031247 | actin rod assembly(GO:0031247) |
| 0.0 | 0.4 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.0 | 1.4 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.0 | 1.2 | GO:0007635 | chemosensory behavior(GO:0007635) |
| 0.0 | 0.3 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.0 | 0.1 | GO:1904381 | Golgi apparatus mannose trimming(GO:1904381) |
| 0.0 | 0.3 | GO:0060613 | fat pad development(GO:0060613) |
| 0.0 | 0.1 | GO:0031291 | Ran protein signal transduction(GO:0031291) |
| 0.0 | 0.8 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.0 | 0.1 | GO:1901898 | negative regulation of relaxation of muscle(GO:1901078) negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.0 | 0.1 | GO:1903216 | regulation of protein processing involved in protein targeting to mitochondrion(GO:1903216) negative regulation of protein processing involved in protein targeting to mitochondrion(GO:1903217) |
| 0.0 | 1.3 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.0 | 0.1 | GO:0021763 | subthalamic nucleus development(GO:0021763) prolactin secreting cell differentiation(GO:0060127) superior vena cava morphogenesis(GO:0060578) |
| 0.0 | 0.4 | GO:0070307 | lens fiber cell development(GO:0070307) |
| 0.0 | 0.4 | GO:0032808 | lacrimal gland development(GO:0032808) |
| 0.0 | 0.2 | GO:0051415 | interphase microtubule nucleation by interphase microtubule organizing center(GO:0051415) microtubule nucleation by microtubule organizing center(GO:0051418) |
| 0.0 | 0.1 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.0 | 0.1 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.0 | 0.2 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 0.4 | GO:0086069 | bundle of His cell to Purkinje myocyte communication(GO:0086069) |
| 0.0 | 0.1 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 0.0 | 0.1 | GO:0035022 | positive regulation of Rac protein signal transduction(GO:0035022) |
| 0.0 | 0.3 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.0 | 0.2 | GO:0019919 | peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) |
| 0.0 | 0.1 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.0 | 0.6 | GO:0050962 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.0 | 0.1 | GO:0070172 | positive regulation of tooth mineralization(GO:0070172) |
| 0.0 | 0.7 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.0 | 0.4 | GO:0038003 | opioid receptor signaling pathway(GO:0038003) |
| 0.0 | 0.5 | GO:0032802 | low-density lipoprotein particle receptor catabolic process(GO:0032802) |
| 0.0 | 0.4 | GO:0033540 | fatty acid beta-oxidation using acyl-CoA oxidase(GO:0033540) |
| 0.0 | 0.1 | GO:0030187 | melatonin metabolic process(GO:0030186) melatonin biosynthetic process(GO:0030187) |
| 0.0 | 1.6 | GO:0042267 | natural killer cell mediated cytotoxicity(GO:0042267) |
| 0.0 | 0.2 | GO:0043587 | tongue morphogenesis(GO:0043587) |
| 0.0 | 2.4 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 0.3 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.0 | 0.1 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.0 | 1.8 | GO:1902600 | hydrogen ion transmembrane transport(GO:1902600) |
| 0.0 | 0.5 | GO:0007095 | mitotic G2 DNA damage checkpoint(GO:0007095) |
| 0.0 | 0.2 | GO:0071847 | TNFSF11-mediated signaling pathway(GO:0071847) negative regulation of osteoclast development(GO:2001205) |
| 0.0 | 0.3 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.0 | 0.1 | GO:0048478 | replication fork protection(GO:0048478) |
| 0.0 | 0.5 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.0 | 0.1 | GO:0034721 | histone H3-K4 demethylation, trimethyl-H3-K4-specific(GO:0034721) |
| 0.0 | 0.5 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.0 | 1.0 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.2 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.0 | 0.2 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) |
| 0.0 | 0.1 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.0 | 1.2 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 0.5 | GO:0031055 | chromatin remodeling at centromere(GO:0031055) CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.0 | 0.1 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.0 | 0.0 | GO:0048560 | establishment of anatomical structure orientation(GO:0048560) |
| 0.0 | 1.9 | GO:0006939 | smooth muscle contraction(GO:0006939) |
| 0.0 | 0.6 | GO:0051123 | RNA polymerase II transcriptional preinitiation complex assembly(GO:0051123) |
| 0.0 | 0.6 | GO:0030318 | melanocyte differentiation(GO:0030318) |
| 0.0 | 0.1 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.5 | GO:0010659 | cardiac muscle cell apoptotic process(GO:0010659) |
| 0.0 | 0.3 | GO:0039703 | viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.0 | 0.3 | GO:0051560 | mitochondrial calcium ion homeostasis(GO:0051560) |
| 0.0 | 0.2 | GO:0003334 | keratinocyte development(GO:0003334) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.8 | GO:0005674 | transcription factor TFIIF complex(GO:0005674) |
| 0.2 | 2.6 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.1 | 1.7 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.1 | 0.3 | GO:0034677 | integrin alpha7-beta1 complex(GO:0034677) |
| 0.1 | 0.7 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.1 | 0.4 | GO:1990742 | microvesicle(GO:1990742) |
| 0.1 | 0.2 | GO:0035189 | Rb-E2F complex(GO:0035189) |
| 0.1 | 1.0 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.1 | 0.3 | GO:0098592 | cytoplasmic side of apical plasma membrane(GO:0098592) |
| 0.1 | 0.2 | GO:1990032 | parallel fiber(GO:1990032) |
| 0.1 | 1.1 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.1 | 0.3 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.0 | 0.6 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 1.3 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.0 | 0.2 | GO:0008275 | gamma-tubulin small complex(GO:0008275) |
| 0.0 | 0.9 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 0.0 | 0.4 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.0 | 1.1 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 0.3 | GO:1990630 | IRE1-RACK1-PP2A complex(GO:1990630) |
| 0.0 | 0.5 | GO:0032797 | SMN complex(GO:0032797) |
| 0.0 | 0.8 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 0.5 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.0 | 0.3 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.0 | 0.3 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.3 | GO:0005688 | U6 snRNP(GO:0005688) |
| 0.0 | 0.3 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.0 | 0.4 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.1 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.0 | 0.2 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 0.5 | GO:0030122 | AP-2 adaptor complex(GO:0030122) endolysosome membrane(GO:0036020) |
| 0.0 | 0.5 | GO:0044232 | organelle membrane contact site(GO:0044232) |
| 0.0 | 0.1 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 0.0 | 1.7 | GO:0101003 | ficolin-1-rich granule membrane(GO:0101003) |
| 0.0 | 0.2 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.0 | 2.6 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 0.3 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.4 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.0 | 0.6 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.0 | 0.1 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.0 | 0.1 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.0 | 0.3 | GO:0032982 | myosin filament(GO:0032982) |
| 0.0 | 0.1 | GO:0031298 | replication fork protection complex(GO:0031298) |
| 0.0 | 0.0 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 0.0 | 13.2 | GO:0030141 | secretory granule(GO:0030141) |
| 0.0 | 0.9 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.3 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.2 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 0.3 | GO:0005614 | interstitial matrix(GO:0005614) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.4 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 0.5 | 2.2 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.3 | 1.3 | GO:0002060 | purine nucleobase binding(GO:0002060) |
| 0.3 | 1.2 | GO:0031531 | thyrotropin-releasing hormone receptor binding(GO:0031531) |
| 0.3 | 1.3 | GO:0004090 | carbonyl reductase (NADPH) activity(GO:0004090) |
| 0.2 | 0.9 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.1 | 1.2 | GO:0086075 | gap junction channel activity involved in cardiac conduction electrical coupling(GO:0086075) |
| 0.1 | 0.4 | GO:0043682 | copper-exporting ATPase activity(GO:0004008) phosphoglycerate kinase activity(GO:0004618) copper-transporting ATPase activity(GO:0043682) |
| 0.1 | 0.7 | GO:0004800 | thyroxine 5'-deiodinase activity(GO:0004800) |
| 0.1 | 1.6 | GO:0046703 | natural killer cell lectin-like receptor binding(GO:0046703) |
| 0.1 | 4.7 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.1 | 0.6 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 0.6 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.1 | 1.8 | GO:0019211 | phosphatase activator activity(GO:0019211) |
| 0.1 | 0.9 | GO:0008430 | selenium binding(GO:0008430) |
| 0.1 | 0.4 | GO:0005415 | nucleoside:sodium symporter activity(GO:0005415) |
| 0.1 | 0.4 | GO:0050119 | N-acetylglucosamine deacetylase activity(GO:0050119) |
| 0.1 | 0.9 | GO:0099580 | ion antiporter activity involved in regulation of postsynaptic membrane potential(GO:0099580) |
| 0.1 | 0.3 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.1 | 0.4 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.1 | 0.4 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.1 | 0.3 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.1 | 0.2 | GO:0033142 | progesterone receptor binding(GO:0033142) |
| 0.1 | 0.5 | GO:0035800 | deubiquitinase activator activity(GO:0035800) |
| 0.1 | 0.5 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.1 | 1.1 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.1 | 1.7 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.1 | 0.4 | GO:0016402 | pristanoyl-CoA oxidase activity(GO:0016402) |
| 0.1 | 2.3 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.1 | 1.4 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.1 | 1.8 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.1 | 0.5 | GO:0043141 | ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.1 | 1.3 | GO:0019841 | retinol binding(GO:0019841) |
| 0.1 | 1.7 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 1.9 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 1.7 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 0.2 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 0.1 | 0.6 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.1 | 0.2 | GO:0042284 | sphingolipid delta-4 desaturase activity(GO:0042284) |
| 0.0 | 0.9 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.0 | 0.2 | GO:0004597 | peptide-aspartate beta-dioxygenase activity(GO:0004597) |
| 0.0 | 0.3 | GO:0098639 | collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.0 | 13.5 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.7 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.0 | 0.1 | GO:0046848 | hydroxyapatite binding(GO:0046848) |
| 0.0 | 0.4 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.0 | 0.1 | GO:0008511 | sodium:potassium:chloride symporter activity(GO:0008511) |
| 0.0 | 1.1 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.0 | 0.8 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.3 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.7 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.0 | 0.3 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.0 | 0.3 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.0 | 0.2 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.0 | 0.3 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 0.3 | GO:0031730 | CCR5 chemokine receptor binding(GO:0031730) |
| 0.0 | 1.5 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 0.3 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.0 | 0.9 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.3 | GO:0047035 | testosterone dehydrogenase (NAD+) activity(GO:0047035) |
| 0.0 | 0.1 | GO:0023024 | MHC class I protein complex binding(GO:0023024) |
| 0.0 | 2.0 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.0 | 0.2 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.0 | 3.4 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.2 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 0.0 | 0.5 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.0 | 2.4 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.4 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.0 | 0.4 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.0 | 0.6 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.0 | 2.1 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 0.1 | GO:0004447 | iodide peroxidase activity(GO:0004447) |
| 0.0 | 0.1 | GO:0047708 | biotinidase activity(GO:0047708) |
| 0.0 | 0.2 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.0 | 0.5 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.5 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.0 | 0.1 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.0 | 0.5 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.0 | 0.1 | GO:0070538 | oleic acid binding(GO:0070538) |
| 0.0 | 0.1 | GO:0005168 | neurotrophin TRKA receptor binding(GO:0005168) |
| 0.0 | 0.3 | GO:0035256 | G-protein coupled glutamate receptor binding(GO:0035256) |
| 0.0 | 0.1 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 0.0 | 0.1 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 0.5 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 1.0 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.1 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) collagen receptor activity(GO:0038064) |
| 0.0 | 0.6 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.1 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.0 | 0.4 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.0 | 0.7 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.0 | 0.2 | GO:1990239 | steroid hormone binding(GO:1990239) |
| 0.0 | 0.5 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.7 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.0 | 0.1 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.0 | 0.5 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.0 | 0.3 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.0 | 0.6 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 3.7 | GO:0004721 | phosphoprotein phosphatase activity(GO:0004721) |
| 0.0 | 0.3 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 9.7 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.5 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 1.6 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 0.9 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 2.4 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 1.6 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 1.1 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.0 | 0.7 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.0 | 1.5 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 0.5 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.3 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 0.9 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 1.3 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.1 | 1.8 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.1 | 1.3 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.1 | 1.2 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.1 | 1.1 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 1.0 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.0 | 1.7 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 3.0 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 1.0 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 0.5 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.0 | 0.5 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 1.2 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.9 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 1.1 | REACTOME ADP SIGNALLING THROUGH P2RY1 | Genes involved in ADP signalling through P2Y purinoceptor 1 |
| 0.0 | 0.9 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.8 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.0 | 0.3 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.0 | 0.3 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.3 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.0 | 0.2 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 0.7 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.1 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PIPS AT THE GOLGI MEMBRANE | Genes involved in Synthesis of PIPs at the Golgi membrane |
| 0.0 | 0.4 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.4 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.3 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 0.5 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.0 | 1.0 | REACTOME REGULATION OF APOPTOSIS | Genes involved in Regulation of Apoptosis |
| 0.0 | 0.3 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.2 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.0 | 0.3 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.2 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.1 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.4 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |