Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
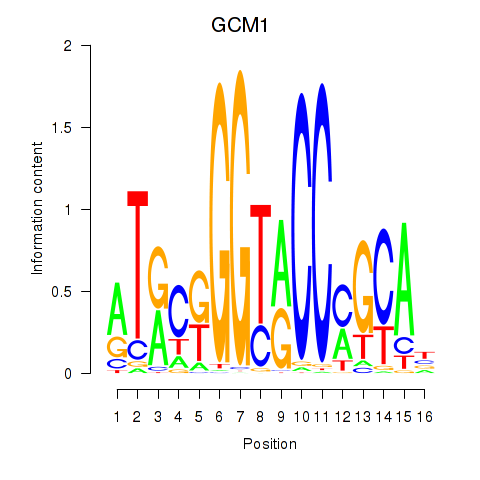
Results for GCM1
Z-value: 0.57
Transcription factors associated with GCM1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
GCM1
|
ENSG00000137270.11 | GCM1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| GCM1 | hg38_v1_chr6_-_53148822_53148846 | 0.05 | 7.9e-01 | Click! |
Activity profile of GCM1 motif
Sorted Z-values of GCM1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of GCM1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_90069662 | 0.74 |
ENST00000390271.2
|
IGKV6D-41
|
immunoglobulin kappa variable 6D-41 (non-functional) |
| chr20_-_1325707 | 0.72 |
ENST00000381812.5
|
SDCBP2
|
syndecan binding protein 2 |
| chr2_+_90021567 | 0.71 |
ENST00000436451.2
|
IGKV6D-21
|
immunoglobulin kappa variable 6D-21 (non-functional) |
| chr2_-_89160329 | 0.69 |
ENST00000390256.2
|
IGKV6-21
|
immunoglobulin kappa variable 6-21 (non-functional) |
| chr20_+_380747 | 0.69 |
ENST00000217233.9
|
TRIB3
|
tribbles pseudokinase 3 |
| chr12_-_121800558 | 0.68 |
ENST00000546227.5
|
RHOF
|
ras homolog family member F, filopodia associated |
| chr20_-_1325746 | 0.68 |
ENST00000339987.7
|
SDCBP2
|
syndecan binding protein 2 |
| chr4_+_4387078 | 0.63 |
ENST00000504171.1
|
NSG1
|
neuronal vesicle trafficking associated 1 |
| chr4_-_8127650 | 0.57 |
ENST00000545242.6
ENST00000676532.1 |
ABLIM2
|
actin binding LIM protein family member 2 |
| chr14_-_24263162 | 0.57 |
ENST00000206765.11
ENST00000544573.5 |
TGM1
|
transglutaminase 1 |
| chr22_+_22594528 | 0.55 |
ENST00000390303.3
|
IGLV3-32
|
immunoglobulin lambda variable 3-32 (non-functional) |
| chr17_-_7590072 | 0.53 |
ENST00000538513.6
ENST00000570788.1 ENST00000250055.3 |
SOX15
|
SRY-box transcription factor 15 |
| chr3_+_10164883 | 0.52 |
ENST00000256458.5
|
IRAK2
|
interleukin 1 receptor associated kinase 2 |
| chr4_-_55636259 | 0.51 |
ENST00000505262.5
ENST00000507338.1 |
NMU
|
neuromedin U |
| chr4_-_8071978 | 0.47 |
ENST00000514025.5
|
ABLIM2
|
actin binding LIM protein family member 2 |
| chr14_+_24070837 | 0.42 |
ENST00000537691.5
ENST00000397016.6 ENST00000560356.5 ENST00000558450.5 |
CPNE6
|
copine 6 |
| chr8_+_25459190 | 0.41 |
ENST00000380665.3
ENST00000330560.8 |
CDCA2
|
cell division cycle associated 2 |
| chr4_-_55636284 | 0.39 |
ENST00000511469.5
ENST00000264218.7 |
NMU
|
neuromedin U |
| chrX_+_71578435 | 0.37 |
ENST00000373696.8
|
GCNA
|
germ cell nuclear acidic peptidase |
| chr2_-_89330429 | 0.37 |
ENST00000620613.1
|
IGKV2-40
|
immunoglobulin kappa variable 2-40 |
| chr19_-_37906588 | 0.36 |
ENST00000447313.7
|
WDR87
|
WD repeat domain 87 |
| chr1_+_55039511 | 0.36 |
ENST00000302118.5
|
PCSK9
|
proprotein convertase subtilisin/kexin type 9 |
| chr11_+_844067 | 0.36 |
ENST00000397406.5
ENST00000409543.6 ENST00000525201.5 |
TSPAN4
|
tetraspanin 4 |
| chr12_-_68933161 | 0.36 |
ENST00000549781.1
ENST00000551568.6 ENST00000548262.5 |
CPM
|
carboxypeptidase M |
| chr19_-_37906646 | 0.32 |
ENST00000303868.5
|
WDR87
|
WD repeat domain 87 |
| chr18_+_23949847 | 0.32 |
ENST00000588004.1
|
LAMA3
|
laminin subunit alpha 3 |
| chr10_-_75235917 | 0.31 |
ENST00000469299.1
ENST00000372538.8 |
COMTD1
|
catechol-O-methyltransferase domain containing 1 |
| chr11_-_8168987 | 0.30 |
ENST00000425599.6
ENST00000531450.1 ENST00000309737.11 ENST00000419822.2 ENST00000335425.7 ENST00000343202.8 |
RIC3
|
RIC3 acetylcholine receptor chaperone |
| chr22_+_43923755 | 0.30 |
ENST00000423180.2
ENST00000216180.8 |
PNPLA3
|
patatin like phospholipase domain containing 3 |
| chr17_+_42761246 | 0.30 |
ENST00000587142.5
ENST00000588576.1 |
RAMP2
|
receptor activity modifying protein 2 |
| chr9_-_101383558 | 0.30 |
ENST00000674556.1
|
BAAT
|
bile acid-CoA:amino acid N-acyltransferase |
| chr16_+_11345429 | 0.29 |
ENST00000576027.1
ENST00000312499.6 ENST00000648619.1 |
RMI2
|
RecQ mediated genome instability 2 |
| chr22_-_20268285 | 0.29 |
ENST00000043402.8
|
RTN4R
|
reticulon 4 receptor |
| chr3_-_48595267 | 0.28 |
ENST00000328333.12
ENST00000681320.1 |
COL7A1
|
collagen type VII alpha 1 chain |
| chr2_+_232456146 | 0.27 |
ENST00000295463.4
|
ALPI
|
alkaline phosphatase, intestinal |
| chr9_+_22446808 | 0.27 |
ENST00000325870.3
|
DMRTA1
|
DMRT like family A1 |
| chr2_+_950836 | 0.27 |
ENST00000308624.10
ENST00000407292.1 |
SNTG2
|
syntrophin gamma 2 |
| chr16_+_68843515 | 0.26 |
ENST00000261778.2
|
TANGO6
|
transport and golgi organization 6 homolog |
| chr17_-_15999589 | 0.26 |
ENST00000486655.5
|
ZSWIM7
|
zinc finger SWIM-type containing 7 |
| chr1_+_67207614 | 0.24 |
ENST00000425614.3
|
IL23R
|
interleukin 23 receptor |
| chr18_+_23689609 | 0.24 |
ENST00000399516.7
|
LAMA3
|
laminin subunit alpha 3 |
| chr4_+_55346213 | 0.23 |
ENST00000679836.1
ENST00000264228.9 ENST00000679707.1 |
SRD5A3
ENSG00000288695.1
|
steroid 5 alpha-reductase 3 novel protein, SRD5A3-RP11-177J6.1 readthrough |
| chr19_+_18340629 | 0.23 |
ENST00000597431.2
|
PGPEP1
|
pyroglutamyl-peptidase I |
| chr3_-_86991135 | 0.22 |
ENST00000398399.7
|
VGLL3
|
vestigial like family member 3 |
| chr5_-_134367144 | 0.22 |
ENST00000265334.9
|
CDKL3
|
cyclin dependent kinase like 3 |
| chr16_-_85688912 | 0.21 |
ENST00000253462.8
|
GINS2
|
GINS complex subunit 2 |
| chr18_+_23689439 | 0.21 |
ENST00000313654.14
|
LAMA3
|
laminin subunit alpha 3 |
| chr17_+_82559340 | 0.21 |
ENST00000531030.5
ENST00000526383.2 |
FOXK2
|
forkhead box K2 |
| chr11_+_124865425 | 0.21 |
ENST00000397801.6
|
ROBO3
|
roundabout guidance receptor 3 |
| chr22_+_22792485 | 0.21 |
ENST00000390314.2
|
IGLV2-11
|
immunoglobulin lambda variable 2-11 |
| chr22_+_22822658 | 0.21 |
ENST00000620395.2
|
IGLV2-8
|
immunoglobulin lambda variable 2-8 |
| chr16_-_74700786 | 0.21 |
ENST00000306247.11
ENST00000575686.1 |
MLKL
|
mixed lineage kinase domain like pseudokinase |
| chr17_+_42761218 | 0.20 |
ENST00000253796.10
|
RAMP2
|
receptor activity modifying protein 2 |
| chr8_+_141128581 | 0.20 |
ENST00000519811.6
|
DENND3
|
DENN domain containing 3 |
| chr17_-_8079903 | 0.20 |
ENST00000649809.1
|
ALOX12B
|
arachidonate 12-lipoxygenase, 12R type |
| chr5_-_180649579 | 0.20 |
ENST00000261937.11
ENST00000502649.5 |
FLT4
|
fms related receptor tyrosine kinase 4 |
| chr11_-_63015804 | 0.20 |
ENST00000535878.5
ENST00000545207.5 |
SLC22A8
|
solute carrier family 22 member 8 |
| chr16_-_70685791 | 0.20 |
ENST00000616026.4
|
MTSS2
|
MTSS I-BAR domain containing 2 |
| chr6_-_30075767 | 0.20 |
ENST00000244360.8
ENST00000376751.8 |
RNF39
|
ring finger protein 39 |
| chr16_-_705726 | 0.20 |
ENST00000397621.6
ENST00000324361.9 |
FBXL16
|
F-box and leucine rich repeat protein 16 |
| chr9_-_112890868 | 0.19 |
ENST00000374228.5
|
SLC46A2
|
solute carrier family 46 member 2 |
| chr4_+_109827963 | 0.19 |
ENST00000317735.7
|
RRH
|
retinal pigment epithelium-derived rhodopsin homolog |
| chr6_+_30061231 | 0.19 |
ENST00000376782.6
ENST00000359374.8 ENST00000376785.2 ENST00000332435.10 |
POLR1H
|
RNA polymerase I subunit H |
| chr8_-_56320098 | 0.19 |
ENST00000303749.8
ENST00000396721.6 |
SDR16C5
|
short chain dehydrogenase/reductase family 16C member 5 |
| chr10_+_79706328 | 0.18 |
ENST00000342531.2
|
NUTM2B
|
NUT family member 2B |
| chr15_-_78641200 | 0.18 |
ENST00000261751.8
|
CHRNB4
|
cholinergic receptor nicotinic beta 4 subunit |
| chr16_-_74700845 | 0.18 |
ENST00000308807.12
ENST00000573267.1 |
MLKL
|
mixed lineage kinase domain like pseudokinase |
| chr10_-_72215903 | 0.18 |
ENST00000526751.5
ENST00000672774.1 ENST00000394919.5 ENST00000394915.7 |
ASCC1
|
activating signal cointegrator 1 complex subunit 1 |
| chr8_+_141128612 | 0.18 |
ENST00000518347.5
ENST00000262585.6 ENST00000520986.5 ENST00000523058.5 ENST00000518668.5 |
DENND3
|
DENN domain containing 3 |
| chr7_-_56034133 | 0.17 |
ENST00000421626.5
|
PSPH
|
phosphoserine phosphatase |
| chr19_+_12791470 | 0.17 |
ENST00000302754.6
|
JUNB
|
JunB proto-oncogene, AP-1 transcription factor subunit |
| chr7_+_124476371 | 0.17 |
ENST00000473520.1
|
SSU72P8
|
SSU72 pseudogene 8 |
| chr4_-_993430 | 0.17 |
ENST00000361661.6
ENST00000622731.4 |
SLC26A1
|
solute carrier family 26 member 1 |
| chr16_+_1972035 | 0.17 |
ENST00000568546.6
ENST00000615855.4 |
TBL3
|
transducin beta like 3 |
| chr17_+_42854078 | 0.16 |
ENST00000591562.1
ENST00000588033.1 |
AOC3
|
amine oxidase copper containing 3 |
| chr22_+_22431949 | 0.16 |
ENST00000390301.3
|
IGLV1-36
|
immunoglobulin lambda variable 1-36 |
| chr3_+_130894382 | 0.16 |
ENST00000509662.5
ENST00000328560.12 ENST00000428331.6 ENST00000359644.7 ENST00000422190.6 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr20_+_31637905 | 0.16 |
ENST00000376075.4
|
COX4I2
|
cytochrome c oxidase subunit 4I2 |
| chr2_-_106194286 | 0.16 |
ENST00000409501.7
ENST00000441952.5 ENST00000457835.5 ENST00000483426.5 ENST00000283148.12 |
UXS1
|
UDP-glucuronate decarboxylase 1 |
| chr8_+_22042790 | 0.16 |
ENST00000359441.4
|
FGF17
|
fibroblast growth factor 17 |
| chr16_-_57479745 | 0.16 |
ENST00000566936.5
ENST00000568617.5 ENST00000567276.5 ENST00000569548.5 ENST00000569250.5 ENST00000564378.5 |
DOK4
|
docking protein 4 |
| chrX_-_85379659 | 0.16 |
ENST00000262753.9
|
POF1B
|
POF1B actin binding protein |
| chr10_-_72216267 | 0.16 |
ENST00000342444.8
ENST00000533958.1 ENST00000672957.1 ENST00000527593.5 ENST00000672940.1 ENST00000530461.5 ENST00000317168.11 ENST00000524829.5 |
ASCC1
|
activating signal cointegrator 1 complex subunit 1 |
| chr17_-_15999634 | 0.16 |
ENST00000472495.5
|
ZSWIM7
|
zinc finger SWIM-type containing 7 |
| chr2_-_171066936 | 0.16 |
ENST00000453628.1
ENST00000434911.6 |
TLK1
|
tousled like kinase 1 |
| chr4_+_55346349 | 0.15 |
ENST00000505210.1
|
SRD5A3
|
steroid 5 alpha-reductase 3 |
| chr22_+_38468036 | 0.15 |
ENST00000409006.3
ENST00000216014.9 |
KDELR3
|
KDEL endoplasmic reticulum protein retention receptor 3 |
| chr12_-_38906269 | 0.15 |
ENST00000550863.1
|
CPNE8
|
copine 8 |
| chr7_-_151210488 | 0.15 |
ENST00000644661.2
|
H2BE1
|
H2B.E variant histone 1 |
| chr19_-_41439528 | 0.14 |
ENST00000301183.15
ENST00000417807.7 ENST00000590641.6 |
DMAC2
|
distal membrane arm assembly complex 2 |
| chr18_+_10526011 | 0.14 |
ENST00000322897.11
|
NAPG
|
NSF attachment protein gamma |
| chr8_+_22579100 | 0.14 |
ENST00000452226.5
ENST00000397760.8 |
PDLIM2
|
PDZ and LIM domain 2 |
| chr3_-_196515315 | 0.14 |
ENST00000397537.3
|
SMCO1
|
single-pass membrane protein with coiled-coil domains 1 |
| chr1_-_11848345 | 0.14 |
ENST00000376476.1
|
NPPA
|
natriuretic peptide A |
| chrX_-_85379703 | 0.14 |
ENST00000373145.3
|
POF1B
|
POF1B actin binding protein |
| chr2_-_24360299 | 0.13 |
ENST00000361999.7
|
ITSN2
|
intersectin 2 |
| chr19_+_11435619 | 0.13 |
ENST00000589126.5
ENST00000588269.1 ENST00000587509.5 ENST00000591462.6 ENST00000592741.5 ENST00000677123.1 ENST00000593101.5 ENST00000587327.5 |
PRKCSH
|
protein kinase C substrate 80K-H |
| chr9_+_127611760 | 0.13 |
ENST00000625363.2
ENST00000626539.3 |
STXBP1
|
syntaxin binding protein 1 |
| chr16_-_70685975 | 0.13 |
ENST00000338779.11
|
MTSS2
|
MTSS I-BAR domain containing 2 |
| chr4_-_48780242 | 0.13 |
ENST00000507711.5
ENST00000358350.9 |
FRYL
|
FRY like transcription coactivator |
| chr19_-_6379058 | 0.13 |
ENST00000597721.1
|
PSPN
|
persephin |
| chr19_+_11547840 | 0.12 |
ENST00000588935.1
|
CNN1
|
calponin 1 |
| chr22_+_22395005 | 0.12 |
ENST00000390298.2
|
IGLV7-43
|
immunoglobulin lambda variable 7-43 |
| chr8_+_22578735 | 0.12 |
ENST00000339162.11
ENST00000308354.11 |
PDLIM2
|
PDZ and LIM domain 2 |
| chr3_-_99876193 | 0.12 |
ENST00000383694.3
|
FILIP1L
|
filamin A interacting protein 1 like |
| chr10_-_103917657 | 0.12 |
ENST00000369764.1
|
STN1
|
STN1 subunit of CST complex |
| chr1_-_44031446 | 0.12 |
ENST00000372310.8
ENST00000466926.1 |
SLC6A9
|
solute carrier family 6 member 9 |
| chr1_-_44031352 | 0.11 |
ENST00000372306.7
ENST00000475075.6 |
SLC6A9
|
solute carrier family 6 member 9 |
| chr17_+_17179527 | 0.11 |
ENST00000579361.1
|
MPRIP
|
myosin phosphatase Rho interacting protein |
| chr19_-_47555856 | 0.11 |
ENST00000391901.7
ENST00000314121.8 |
ZNF541
|
zinc finger protein 541 |
| chr2_-_136118142 | 0.11 |
ENST00000241393.4
|
CXCR4
|
C-X-C motif chemokine receptor 4 |
| chr9_-_135699473 | 0.11 |
ENST00000425225.2
ENST00000298466.9 |
SOHLH1
|
spermatogenesis and oogenesis specific basic helix-loop-helix 1 |
| chr9_+_17579059 | 0.11 |
ENST00000380607.5
|
SH3GL2
|
SH3 domain containing GRB2 like 2, endophilin A1 |
| chr17_-_8630713 | 0.11 |
ENST00000411957.1
ENST00000360416.8 |
MYH10
|
myosin heavy chain 10 |
| chr16_+_10877181 | 0.11 |
ENST00000618207.4
ENST00000618327.4 ENST00000324288.14 ENST00000381835.9 |
CIITA
|
class II major histocompatibility complex transactivator |
| chr5_-_180649613 | 0.11 |
ENST00000393347.7
ENST00000619105.4 |
FLT4
|
fms related receptor tyrosine kinase 4 |
| chr12_-_11022620 | 0.11 |
ENST00000390673.2
|
TAS2R19
|
taste 2 receptor member 19 |
| chr7_-_712437 | 0.11 |
ENST00000360274.8
|
PRKAR1B
|
protein kinase cAMP-dependent type I regulatory subunit beta |
| chr12_+_32399517 | 0.11 |
ENST00000534526.7
|
FGD4
|
FYVE, RhoGEF and PH domain containing 4 |
| chr3_-_99876104 | 0.11 |
ENST00000471562.1
ENST00000495625.2 |
FILIP1L
|
filamin A interacting protein 1 like |
| chr22_+_22758698 | 0.10 |
ENST00000390312.2
|
IGLV2-14
|
immunoglobulin lambda variable 2-14 |
| chr17_-_44503369 | 0.10 |
ENST00000585614.1
ENST00000591680.6 |
GPATCH8
|
G-patch domain containing 8 |
| chr20_+_45406035 | 0.10 |
ENST00000372723.7
ENST00000372722.7 |
DBNDD2
|
dysbindin domain containing 2 |
| chr12_-_4445810 | 0.10 |
ENST00000228837.3
|
FGF6
|
fibroblast growth factor 6 |
| chr1_-_37690481 | 0.10 |
ENST00000464085.5
ENST00000358011.8 ENST00000461359.5 ENST00000486637.2 |
C1orf109
|
chromosome 1 open reading frame 109 |
| chr1_-_11847772 | 0.10 |
ENST00000376480.7
ENST00000610706.1 |
NPPA
|
natriuretic peptide A |
| chr3_+_47380995 | 0.10 |
ENST00000456221.5
ENST00000265562.5 |
PTPN23
|
protein tyrosine phosphatase non-receptor type 23 |
| chr11_-_35418966 | 0.10 |
ENST00000531628.2
|
SLC1A2
|
solute carrier family 1 member 2 |
| chr22_+_21632751 | 0.10 |
ENST00000292779.4
|
CCDC116
|
coiled-coil domain containing 116 |
| chr2_+_117814648 | 0.10 |
ENST00000263239.7
|
DDX18
|
DEAD-box helicase 18 |
| chr7_+_101218146 | 0.09 |
ENST00000305105.3
|
ZNHIT1
|
zinc finger HIT-type containing 1 |
| chr5_+_136049513 | 0.09 |
ENST00000514554.5
|
TGFBI
|
transforming growth factor beta induced |
| chr19_-_12775513 | 0.09 |
ENST00000397668.8
ENST00000587178.1 ENST00000264827.9 |
HOOK2
|
hook microtubule tethering protein 2 |
| chr15_-_45129857 | 0.09 |
ENST00000267803.8
ENST00000613425.4 ENST00000559014.5 ENST00000558851.1 ENST00000559988.5 ENST00000558996.5 ENST00000558422.5 ENST00000559226.5 ENST00000560572.6 ENST00000558326.5 ENST00000558377.5 ENST00000559644.5 |
DUOXA1
|
dual oxidase maturation factor 1 |
| chr22_-_41688799 | 0.09 |
ENST00000469028.2
ENST00000463675.6 ENST00000649479.1 ENST00000469522.1 |
SNU13
|
small nuclear ribonucleoprotein 13 |
| chr19_+_17406099 | 0.09 |
ENST00000634942.2
ENST00000634568.1 ENST00000600514.5 |
BISPR
MVB12A
|
BST2 interferon stimulated positive regulator multivesicular body subunit 12A |
| chrX_+_52184904 | 0.09 |
ENST00000375626.7
ENST00000467526.1 |
MAGED4
|
MAGE family member D4 |
| chr16_+_56565070 | 0.08 |
ENST00000219162.4
|
MT4
|
metallothionein 4 |
| chr11_-_41459592 | 0.08 |
ENST00000528697.6
ENST00000530763.5 |
LRRC4C
|
leucine rich repeat containing 4C |
| chr7_+_7566866 | 0.08 |
ENST00000405785.5
ENST00000340080.9 ENST00000433635.1 |
MIOS
|
meiosis regulator for oocyte development |
| chr20_+_45406162 | 0.08 |
ENST00000357275.6
ENST00000372720.7 |
DBNDD2
|
dysbindin domain containing 2 |
| chr13_-_77918820 | 0.08 |
ENST00000646607.2
|
EDNRB
|
endothelin receptor type B |
| chr17_+_42853232 | 0.08 |
ENST00000617500.4
|
AOC3
|
amine oxidase copper containing 3 |
| chr11_-_35419098 | 0.08 |
ENST00000606205.6
ENST00000645303.1 |
SLC1A2
|
solute carrier family 1 member 2 |
| chrX_-_71068384 | 0.08 |
ENST00000276105.3
ENST00000622259.4 |
SNX12
|
sorting nexin 12 |
| chr4_-_17781613 | 0.07 |
ENST00000265018.4
|
FAM184B
|
family with sequence similarity 184 member B |
| chr22_-_32202451 | 0.07 |
ENST00000248983.8
|
RFPL2
|
ret finger protein like 2 |
| chr18_-_21111778 | 0.07 |
ENST00000399799.3
|
ROCK1
|
Rho associated coiled-coil containing protein kinase 1 |
| chr7_+_93232340 | 0.07 |
ENST00000251739.9
ENST00000544910.5 ENST00000305866.10 ENST00000458530.1 |
VPS50
|
VPS50 subunit of EARP/GARPII complex |
| chr22_+_22697789 | 0.07 |
ENST00000390306.2
|
IGLV2-23
|
immunoglobulin lambda variable 2-23 |
| chr7_+_100373458 | 0.07 |
ENST00000453419.5
ENST00000198536.7 ENST00000394000.6 ENST00000350573.2 |
PILRA
|
paired immunoglobin like type 2 receptor alpha |
| chr3_+_130893959 | 0.07 |
ENST00000508532.5
ENST00000533801.6 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr17_-_19716604 | 0.07 |
ENST00000325411.9
ENST00000350657.9 |
SLC47A2
|
solute carrier family 47 member 2 |
| chr15_+_78340344 | 0.07 |
ENST00000299529.7
|
CRABP1
|
cellular retinoic acid binding protein 1 |
| chr2_+_112482133 | 0.07 |
ENST00000233336.7
|
TTL
|
tubulin tyrosine ligase |
| chr17_-_3696133 | 0.07 |
ENST00000225328.10
|
P2RX5
|
purinergic receptor P2X 5 |
| chrX_-_21658324 | 0.07 |
ENST00000379499.3
|
KLHL34
|
kelch like family member 34 |
| chr17_-_19716573 | 0.07 |
ENST00000433844.4
|
SLC47A2
|
solute carrier family 47 member 2 |
| chr2_-_113235443 | 0.07 |
ENST00000465084.1
|
PAX8
|
paired box 8 |
| chr1_-_26067622 | 0.06 |
ENST00000374272.4
|
TRIM63
|
tripartite motif containing 63 |
| chr7_+_48924559 | 0.06 |
ENST00000650262.1
|
CDC14C
|
cell division cycle 14C |
| chr4_+_75724569 | 0.06 |
ENST00000514213.7
ENST00000264904.8 |
USO1
|
USO1 vesicle transport factor |
| chr9_-_96778053 | 0.06 |
ENST00000375231.5
ENST00000223428.9 |
ZNF510
|
zinc finger protein 510 |
| chr7_+_100126954 | 0.06 |
ENST00000398075.4
|
MBLAC1
|
metallo-beta-lactamase domain containing 1 |
| chr10_+_129467178 | 0.06 |
ENST00000306010.8
ENST00000651593.1 |
MGMT
|
O-6-methylguanine-DNA methyltransferase |
| chr19_-_3547306 | 0.06 |
ENST00000589063.5
ENST00000615073.4 |
MFSD12
|
major facilitator superfamily domain containing 12 |
| chr3_+_130894157 | 0.06 |
ENST00000504948.5
ENST00000513801.5 ENST00000505072.5 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr6_-_136526654 | 0.06 |
ENST00000611373.1
|
MAP7
|
microtubule associated protein 7 |
| chr9_-_120929160 | 0.06 |
ENST00000540010.1
|
TRAF1
|
TNF receptor associated factor 1 |
| chr1_-_8423664 | 0.06 |
ENST00000476556.5
|
RERE
|
arginine-glutamic acid dipeptide repeats |
| chr1_+_54209946 | 0.06 |
ENST00000398219.2
|
MRPL37
|
mitochondrial ribosomal protein L37 |
| chr10_+_96000091 | 0.06 |
ENST00000424464.5
ENST00000410012.6 ENST00000344386.3 |
CC2D2B
|
coiled-coil and C2 domain containing 2B |
| chr2_+_234050679 | 0.06 |
ENST00000373368.5
ENST00000168148.8 |
SPP2
|
secreted phosphoprotein 2 |
| chr11_+_832887 | 0.06 |
ENST00000524748.6
ENST00000397420.9 ENST00000397421.5 ENST00000322008.9 ENST00000525718.6 ENST00000529810.5 ENST00000526693.5 ENST00000525333.5 ENST00000527341.5 |
CD151
|
CD151 molecule (Raph blood group) |
| chr8_-_81841958 | 0.06 |
ENST00000519817.5
ENST00000521773.5 ENST00000523757.5 ENST00000345957.9 |
SNX16
|
sorting nexin 16 |
| chr19_-_18397596 | 0.06 |
ENST00000595840.1
ENST00000339007.4 |
LRRC25
|
leucine rich repeat containing 25 |
| chr14_+_77181780 | 0.06 |
ENST00000298351.5
ENST00000554346.5 |
TMEM63C
|
transmembrane protein 63C |
| chr5_-_134367070 | 0.06 |
ENST00000521118.5
|
CDKL3
|
cyclin dependent kinase like 3 |
| chr17_-_3696198 | 0.06 |
ENST00000345901.7
|
P2RX5
|
purinergic receptor P2X 5 |
| chr12_-_104050112 | 0.05 |
ENST00000547583.1
ENST00000546851.1 ENST00000360814.9 |
GLT8D2
|
glycosyltransferase 8 domain containing 2 |
| chr12_+_8513499 | 0.05 |
ENST00000299665.3
|
CLEC4D
|
C-type lectin domain family 4 member D |
| chr1_+_23743153 | 0.05 |
ENST00000418390.6
|
ELOA
|
elongin A |
| chr11_+_20363685 | 0.05 |
ENST00000530266.5
ENST00000451739.7 ENST00000421577.6 ENST00000443524.6 ENST00000419348.6 |
HTATIP2
|
HIV-1 Tat interactive protein 2 |
| chr20_+_59123381 | 0.05 |
ENST00000637017.1
|
ZNF831
|
zinc finger protein 831 |
| chr11_-_63015831 | 0.05 |
ENST00000430500.6
ENST00000336232.7 |
SLC22A8
|
solute carrier family 22 member 8 |
| chr9_-_136118850 | 0.05 |
ENST00000418388.6
|
TMEM250
|
transmembrane protein 250 |
| chr4_+_186191549 | 0.05 |
ENST00000378802.5
|
CYP4V2
|
cytochrome P450 family 4 subfamily V member 2 |
| chr9_-_136118431 | 0.05 |
ENST00000561457.2
ENST00000448040.2 |
TMEM250
|
transmembrane protein 250 |
| chr15_+_45129933 | 0.05 |
ENST00000321429.8
ENST00000389037.7 ENST00000558322.5 |
DUOX1
|
dual oxidase 1 |
| chr16_+_67029093 | 0.05 |
ENST00000561924.6
|
CBFB
|
core-binding factor subunit beta |
| chr22_+_22195753 | 0.05 |
ENST00000390285.4
|
IGLV6-57
|
immunoglobulin lambda variable 6-57 |
| chr3_+_52287811 | 0.05 |
ENST00000305690.12
ENST00000473032.5 ENST00000436784.7 ENST00000471180.5 |
GLYCTK
|
glycerate kinase |
| chr3_+_46407251 | 0.05 |
ENST00000399036.4
|
CCRL2
|
C-C motif chemokine receptor like 2 |
| chrX_+_154444121 | 0.05 |
ENST00000393600.8
|
FAM50A
|
family with sequence similarity 50 member A |
| chr15_+_41828085 | 0.05 |
ENST00000397299.9
ENST00000408047.5 ENST00000431823.1 ENST00000382448.8 ENST00000342159.6 |
JMJD7
JMJD7-PLA2G4B
|
jumonji domain containing 7 JMJD7-PLA2G4B readthrough |
| chr16_+_72108296 | 0.05 |
ENST00000563819.5
ENST00000567142.2 |
DHX38
|
DEAH-box helicase 38 |
| chr1_+_23743366 | 0.04 |
ENST00000613537.4
|
ELOA
|
elongin A |
| chr11_+_20364119 | 0.04 |
ENST00000532081.1
ENST00000531058.1 |
HTATIP2
|
HIV-1 Tat interactive protein 2 |
| chr18_-_22417910 | 0.04 |
ENST00000391403.4
|
CTAGE1
|
cutaneous T cell lymphoma-associated antigen 1 |
| chr22_+_22588155 | 0.04 |
ENST00000390302.3
|
IGLV2-33
|
immunoglobulin lambda variable 2-33 (non-functional) |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0070318 | positive regulation of G0 to G1 transition(GO:0070318) |
| 0.1 | 0.4 | GO:2000646 | positive regulation of receptor catabolic process(GO:2000646) |
| 0.1 | 0.2 | GO:0061537 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.1 | 0.4 | GO:0016095 | polyprenol catabolic process(GO:0016095) |
| 0.1 | 0.9 | GO:2000252 | negative regulation of feeding behavior(GO:2000252) |
| 0.1 | 0.2 | GO:1902283 | negative regulation of primary amine oxidase activity(GO:1902283) |
| 0.1 | 0.2 | GO:0038155 | interleukin-23-mediated signaling pathway(GO:0038155) |
| 0.1 | 0.6 | GO:0098887 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) |
| 0.1 | 0.2 | GO:1903296 | regulation of glutamate secretion, neurotransmission(GO:1903294) positive regulation of glutamate secretion, neurotransmission(GO:1903296) |
| 0.1 | 0.5 | GO:0097647 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.0 | 0.2 | GO:1902304 | positive regulation of potassium ion export(GO:1902304) |
| 0.0 | 0.6 | GO:0010838 | positive regulation of keratinocyte proliferation(GO:0010838) |
| 0.0 | 0.2 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.0 | 0.1 | GO:2000642 | negative regulation of early endosome to late endosome transport(GO:2000642) |
| 0.0 | 0.4 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) |
| 0.0 | 0.3 | GO:0060179 | male mating behavior(GO:0060179) |
| 0.0 | 0.8 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.7 | GO:0045717 | negative regulation of fatty acid biosynthetic process(GO:0045717) |
| 0.0 | 0.1 | GO:0021592 | fourth ventricle development(GO:0021592) |
| 0.0 | 0.1 | GO:0045348 | positive regulation of MHC class II biosynthetic process(GO:0045348) |
| 0.0 | 0.2 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 0.0 | 0.3 | GO:0019530 | taurine metabolic process(GO:0019530) |
| 0.0 | 0.1 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.0 | 0.1 | GO:0035645 | enteric smooth muscle cell differentiation(GO:0035645) |
| 0.0 | 0.2 | GO:0016199 | axon midline choice point recognition(GO:0016199) |
| 0.0 | 0.1 | GO:0034653 | diterpenoid catabolic process(GO:0016103) terpenoid catabolic process(GO:0016115) retinoic acid catabolic process(GO:0034653) |
| 0.0 | 0.3 | GO:0036155 | acylglycerol acyl-chain remodeling(GO:0036155) |
| 0.0 | 0.2 | GO:0051122 | hepoxilin metabolic process(GO:0051121) hepoxilin biosynthetic process(GO:0051122) |
| 0.0 | 0.3 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 0.0 | 0.3 | GO:0090037 | positive regulation of protein kinase C signaling(GO:0090037) |
| 0.0 | 0.1 | GO:1903347 | negative regulation of bicellular tight junction assembly(GO:1903347) |
| 0.0 | 0.4 | GO:0035510 | DNA dealkylation(GO:0035510) DNA demethylation(GO:0080111) |
| 0.0 | 0.2 | GO:2000609 | regulation of thyroid hormone generation(GO:2000609) |
| 0.0 | 0.2 | GO:0034036 | purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.0 | 0.5 | GO:0034162 | toll-like receptor 9 signaling pathway(GO:0034162) |
| 0.0 | 0.1 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) |
| 0.0 | 0.2 | GO:0060136 | embryonic process involved in female pregnancy(GO:0060136) |
| 0.0 | 0.1 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) |
| 0.0 | 0.1 | GO:0046618 | drug export(GO:0046618) |
| 0.0 | 0.9 | GO:0097484 | dendrite extension(GO:0097484) |
| 0.0 | 0.2 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.0 | 0.2 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.0 | 0.1 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.0 | 0.1 | GO:0042335 | cuticle development(GO:0042335) |
| 0.0 | 2.4 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.2 | GO:0070777 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.0 | 0.1 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.0 | 0.1 | GO:0039663 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.0 | 0.4 | GO:0070207 | protein homotrimerization(GO:0070207) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:1990667 | PCSK9-LDLR complex(GO:1990666) PCSK9-AnxA2 complex(GO:1990667) |
| 0.1 | 0.8 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.1 | 0.5 | GO:1903439 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.1 | 0.2 | GO:0072536 | interleukin-23 receptor complex(GO:0072536) |
| 0.1 | 0.6 | GO:0098845 | postsynaptic endosome(GO:0098845) |
| 0.1 | 0.2 | GO:0000811 | GINS complex(GO:0000811) |
| 0.1 | 0.2 | GO:0097196 | Shu complex(GO:0097196) |
| 0.0 | 0.3 | GO:0030934 | anchoring collagen complex(GO:0030934) |
| 0.0 | 0.2 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 0.1 | GO:1990879 | CST complex(GO:1990879) |
| 0.0 | 0.2 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 0.1 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.0 | 0.2 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.0 | 0.3 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.0 | 0.1 | GO:0070695 | FHF complex(GO:0070695) |
| 0.0 | 0.1 | GO:0001651 | dense fibrillar component(GO:0001651) |
| 0.0 | 0.3 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.0 | 0.1 | GO:0070449 | elongin complex(GO:0070449) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0052815 | medium-chain acyl-CoA hydrolase activity(GO:0052815) long-chain acyl-CoA hydrolase activity(GO:0052816) |
| 0.1 | 0.4 | GO:0033765 | 3-oxo-5-alpha-steroid 4-dehydrogenase activity(GO:0003865) steroid dehydrogenase activity, acting on the CH-CH group of donors(GO:0033765) cholestenone 5-alpha-reductase activity(GO:0047751) |
| 0.1 | 0.4 | GO:0034189 | very-low-density lipoprotein particle binding(GO:0034189) |
| 0.1 | 0.6 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 0.5 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.1 | 0.2 | GO:0052596 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 0.1 | 0.2 | GO:0042020 | interleukin-23 binding(GO:0042019) interleukin-23 receptor activity(GO:0042020) |
| 0.1 | 0.2 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.1 | 0.6 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.0 | 0.1 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.0 | 0.3 | GO:0004465 | lipoprotein lipase activity(GO:0004465) |
| 0.0 | 0.2 | GO:0004052 | arachidonate 12-lipoxygenase activity(GO:0004052) |
| 0.0 | 0.2 | GO:0005111 | type 2 fibroblast growth factor receptor binding(GO:0005111) |
| 0.0 | 0.3 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.0 | 0.7 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.0 | 0.2 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.0 | 0.3 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.0 | 0.3 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.0 | 0.1 | GO:0034512 | box C/D snoRNA binding(GO:0034512) |
| 0.0 | 0.1 | GO:0015307 | drug:proton antiporter activity(GO:0015307) |
| 0.0 | 0.1 | GO:0008267 | poly-glutamine tract binding(GO:0008267) |
| 0.0 | 1.1 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.0 | 0.2 | GO:0035374 | chondroitin sulfate binding(GO:0035374) |
| 0.0 | 0.1 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.0 | 0.2 | GO:0046923 | ER retention sequence binding(GO:0046923) |
| 0.0 | 0.2 | GO:0004931 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.0 | 0.3 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.0 | 0.2 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 0.0 | 0.2 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 0.2 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.2 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.0 | 0.3 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
| 0.0 | 0.1 | GO:0004945 | angiotensin receptor activity(GO:0001595) angiotensin type II receptor activity(GO:0004945) |
| 0.0 | 0.1 | GO:0032027 | myosin light chain binding(GO:0032027) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.8 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 0.3 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.0 | 0.5 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.0 | 0.7 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.0 | 0.4 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.0 | 0.3 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.1 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 0.0 | 0.3 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.2 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 0.3 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.0 | 0.2 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.0 | 0.3 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |