Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
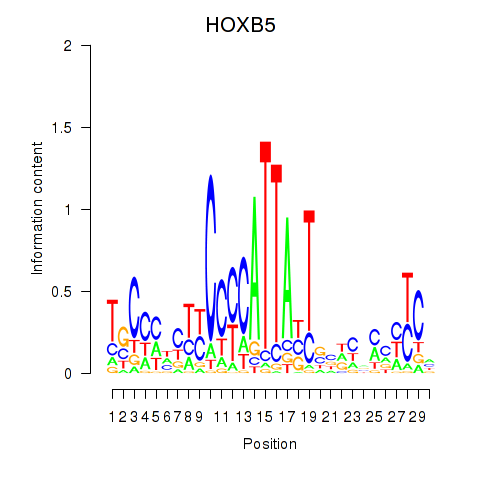
Results for HOXB5
Z-value: 0.63
Transcription factors associated with HOXB5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXB5
|
ENSG00000120075.6 | HOXB5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HOXB5 | hg38_v1_chr17_-_48593748_48593786 | 0.24 | 2.1e-01 | Click! |
Activity profile of HOXB5 motif
Sorted Z-values of HOXB5 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXB5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_-_132085655 | 4.94 |
ENST00000262283.5
|
ENSG00000258417.3
|
novel protein |
| chr19_-_15934410 | 1.28 |
ENST00000326742.12
|
CYP4F11
|
cytochrome P450 family 4 subfamily F member 11 |
| chr19_-_15934521 | 1.15 |
ENST00000402119.9
|
CYP4F11
|
cytochrome P450 family 4 subfamily F member 11 |
| chrX_+_136148440 | 1.13 |
ENST00000627383.2
ENST00000630084.2 |
FHL1
|
four and a half LIM domains 1 |
| chr1_-_16978276 | 1.08 |
ENST00000375534.7
|
MFAP2
|
microfibril associated protein 2 |
| chr19_-_49640092 | 0.94 |
ENST00000246792.4
|
RRAS
|
RAS related |
| chr12_+_56521990 | 0.83 |
ENST00000550726.5
ENST00000542360.1 |
RBMS2
|
RNA binding motif single stranded interacting protein 2 |
| chr12_+_56521951 | 0.83 |
ENST00000552247.6
|
RBMS2
|
RNA binding motif single stranded interacting protein 2 |
| chr15_-_74725370 | 0.82 |
ENST00000567032.5
ENST00000564596.5 ENST00000566503.1 ENST00000395049.8 ENST00000379727.8 ENST00000617691.4 ENST00000395048.6 |
CYP1A1
|
cytochrome P450 family 1 subfamily A member 1 |
| chr16_-_84618041 | 0.77 |
ENST00000564057.1
|
COTL1
|
coactosin like F-actin binding protein 1 |
| chr16_-_84618067 | 0.77 |
ENST00000262428.5
|
COTL1
|
coactosin like F-actin binding protein 1 |
| chr11_-_125778818 | 0.76 |
ENST00000358524.8
|
PATE2
|
prostate and testis expressed 2 |
| chr17_+_48892761 | 0.74 |
ENST00000355938.9
ENST00000393366.7 ENST00000503641.5 ENST00000514808.5 ENST00000506855.1 |
ATP5MC1
|
ATP synthase membrane subunit c locus 1 |
| chr11_-_125778788 | 0.74 |
ENST00000436890.2
|
PATE2
|
prostate and testis expressed 2 |
| chr3_-_195811889 | 0.68 |
ENST00000475231.5
|
MUC4
|
mucin 4, cell surface associated |
| chr12_+_56521798 | 0.62 |
ENST00000262031.10
|
RBMS2
|
RNA binding motif single stranded interacting protein 2 |
| chr10_-_101843765 | 0.58 |
ENST00000370046.5
|
KCNIP2
|
potassium voltage-gated channel interacting protein 2 |
| chr3_-_195811857 | 0.56 |
ENST00000349607.8
ENST00000346145.8 |
MUC4
|
mucin 4, cell surface associated |
| chr3_-_195811916 | 0.56 |
ENST00000463781.8
|
MUC4
|
mucin 4, cell surface associated |
| chr18_-_35497591 | 0.55 |
ENST00000589273.1
ENST00000586489.5 |
INO80C
|
INO80 complex subunit C |
| chr11_-_128522189 | 0.53 |
ENST00000526145.6
|
ETS1
|
ETS proto-oncogene 1, transcription factor |
| chr19_+_49363923 | 0.49 |
ENST00000597546.1
|
DKKL1
|
dickkopf like acrosomal protein 1 |
| chr1_+_4654732 | 0.48 |
ENST00000378190.7
|
AJAP1
|
adherens junctions associated protein 1 |
| chr3_-_165196689 | 0.47 |
ENST00000241274.3
|
SLITRK3
|
SLIT and NTRK like family member 3 |
| chr11_+_63506046 | 0.46 |
ENST00000674247.1
|
LGALS12
|
galectin 12 |
| chr7_-_102611591 | 0.45 |
ENST00000461209.5
|
RASA4
|
RAS p21 protein activator 4 |
| chr1_+_155610218 | 0.41 |
ENST00000649846.1
ENST00000245564.8 ENST00000368341.8 |
MSTO1
|
misato mitochondrial distribution and morphology regulator 1 |
| chr20_+_8789517 | 0.40 |
ENST00000437439.2
|
PLCB1
|
phospholipase C beta 1 |
| chrX_+_47218232 | 0.39 |
ENST00000457458.6
ENST00000522883.1 |
CDK16
|
cyclin dependent kinase 16 |
| chr1_+_157993601 | 0.39 |
ENST00000359209.11
|
KIRREL1
|
kirre like nephrin family adhesion molecule 1 |
| chr3_+_51385282 | 0.38 |
ENST00000528157.7
|
MANF
|
mesencephalic astrocyte derived neurotrophic factor |
| chrX_+_48521817 | 0.37 |
ENST00000446158.5
ENST00000414061.1 |
EBP
|
EBP cholestenol delta-isomerase |
| chrX_+_48521788 | 0.37 |
ENST00000651615.1
ENST00000495186.6 |
ENSG00000286268.1
EBP
|
novel protein EBP cholestenol delta-isomerase |
| chr2_+_17541157 | 0.36 |
ENST00000406397.1
|
VSNL1
|
visinin like 1 |
| chr1_-_161223559 | 0.36 |
ENST00000469730.2
ENST00000463273.5 ENST00000464492.5 ENST00000367990.7 ENST00000470459.6 ENST00000463812.1 ENST00000468465.5 |
APOA2
|
apolipoprotein A2 |
| chr2_+_17540670 | 0.32 |
ENST00000451533.5
ENST00000295156.9 |
VSNL1
|
visinin like 1 |
| chr2_-_46941710 | 0.32 |
ENST00000409207.5
|
MCFD2
|
multiple coagulation factor deficiency 2, ER cargo receptor complex subunit |
| chr1_+_4654601 | 0.31 |
ENST00000378191.5
|
AJAP1
|
adherens junctions associated protein 1 |
| chr2_-_46941760 | 0.28 |
ENST00000444761.6
ENST00000409147.1 |
MCFD2
|
multiple coagulation factor deficiency 2, ER cargo receptor complex subunit |
| chr7_+_142800957 | 0.27 |
ENST00000466254.1
|
TRBC2
|
T cell receptor beta constant 2 |
| chr10_-_101843777 | 0.26 |
ENST00000356640.7
|
KCNIP2
|
potassium voltage-gated channel interacting protein 2 |
| chr11_-_35418966 | 0.25 |
ENST00000531628.2
|
SLC1A2
|
solute carrier family 1 member 2 |
| chr11_-_35419098 | 0.23 |
ENST00000606205.6
ENST00000645303.1 |
SLC1A2
|
solute carrier family 1 member 2 |
| chr21_-_44262216 | 0.20 |
ENST00000270172.7
|
DNMT3L
|
DNA methyltransferase 3 like |
| chr17_-_17237122 | 0.20 |
ENST00000389169.9
ENST00000285071.9 ENST00000417064.1 |
FLCN
|
folliculin |
| chr3_-_127822455 | 0.20 |
ENST00000265052.10
|
MGLL
|
monoglyceride lipase |
| chr1_-_247458105 | 0.20 |
ENST00000641149.1
ENST00000641527.1 |
OR2B11
|
olfactory receptor family 2 subfamily B member 11 |
| chr1_+_146938744 | 0.19 |
ENST00000617931.4
|
NBPF12
|
NBPF member 12 |
| chr12_-_52321395 | 0.19 |
ENST00000293670.3
|
KRT83
|
keratin 83 |
| chr12_+_57591158 | 0.18 |
ENST00000422156.7
ENST00000354947.10 ENST00000540759.6 ENST00000551772.5 ENST00000550465.5 |
PIP4K2C
|
phosphatidylinositol-5-phosphate 4-kinase type 2 gamma |
| chr14_+_22105305 | 0.17 |
ENST00000390453.1
|
TRAV24
|
T cell receptor alpha variable 24 |
| chr16_-_57971086 | 0.17 |
ENST00000564448.5
ENST00000311183.8 |
CNGB1
|
cyclic nucleotide gated channel subunit beta 1 |
| chrM_+_12329 | 0.16 |
ENST00000361567.2
|
MT-ND5
|
mitochondrially encoded NADH:ubiquinone oxidoreductase core subunit 5 |
| chr17_-_44066595 | 0.16 |
ENST00000585388.2
ENST00000293406.8 |
LSM12
|
LSM12 homolog |
| chr19_+_49363730 | 0.15 |
ENST00000596402.1
ENST00000221498.7 |
DKKL1
|
dickkopf like acrosomal protein 1 |
| chr5_+_150190035 | 0.14 |
ENST00000230671.7
ENST00000524041.1 |
SLC6A7
|
solute carrier family 6 member 7 |
| chr6_-_96837460 | 0.14 |
ENST00000229955.4
|
GPR63
|
G protein-coupled receptor 63 |
| chr11_+_4643966 | 0.14 |
ENST00000396952.6
|
OR51E1
|
olfactory receptor family 51 subfamily E member 1 |
| chr14_+_21477177 | 0.14 |
ENST00000448790.7
ENST00000673643.1 ENST00000457430.2 ENST00000673911.1 |
TOX4
|
TOX high mobility group box family member 4 |
| chr6_+_167111789 | 0.14 |
ENST00000400926.5
|
CCR6
|
C-C motif chemokine receptor 6 |
| chr2_-_187565751 | 0.13 |
ENST00000421427.5
|
TFPI
|
tissue factor pathway inhibitor |
| chr1_-_161223408 | 0.13 |
ENST00000491350.1
|
APOA2
|
apolipoprotein A2 |
| chr1_-_206023889 | 0.12 |
ENST00000358184.7
ENST00000360218.3 ENST00000678712.1 ENST00000678498.1 |
CTSE
|
cathepsin E |
| chr11_+_109422174 | 0.12 |
ENST00000327419.7
|
C11orf87
|
chromosome 11 open reading frame 87 |
| chr19_+_13151975 | 0.12 |
ENST00000588173.1
|
IER2
|
immediate early response 2 |
| chr17_+_6776209 | 0.11 |
ENST00000321535.5
|
FBXO39
|
F-box protein 39 |
| chr11_-_57322197 | 0.10 |
ENST00000532437.1
|
TNKS1BP1
|
tankyrase 1 binding protein 1 |
| chr19_-_49763295 | 0.09 |
ENST00000246801.8
|
TSKS
|
testis specific serine kinase substrate |
| chr4_-_151325488 | 0.08 |
ENST00000604030.7
|
SH3D19
|
SH3 domain containing 19 |
| chr5_-_35047935 | 0.07 |
ENST00000510428.1
ENST00000231420.11 |
AGXT2
|
alanine--glyoxylate aminotransferase 2 |
| chr3_+_45945019 | 0.07 |
ENST00000458629.1
ENST00000457814.1 |
CXCR6
|
C-X-C motif chemokine receptor 6 |
| chr7_+_130381092 | 0.07 |
ENST00000484324.1
|
CPA1
|
carboxypeptidase A1 |
| chr1_-_155688475 | 0.07 |
ENST00000405763.7
ENST00000368340.9 ENST00000454523.5 ENST00000368339.9 ENST00000443231.5 ENST00000347088.9 ENST00000361831.9 |
YY1AP1
|
YY1 associated protein 1 |
| chr3_+_130894382 | 0.06 |
ENST00000509662.5
ENST00000328560.12 ENST00000428331.6 ENST00000359644.7 ENST00000422190.6 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr8_-_92095627 | 0.06 |
ENST00000517919.5
ENST00000617740.4 ENST00000613302.4 ENST00000436581.6 ENST00000614812.4 ENST00000519847.5 |
RUNX1T1
|
RUNX1 partner transcriptional co-repressor 1 |
| chr7_+_5282935 | 0.06 |
ENST00000396872.8
ENST00000444741.5 ENST00000297195.8 ENST00000406453.3 |
SLC29A4
|
solute carrier family 29 member 4 |
| chrX_+_14529516 | 0.05 |
ENST00000218075.9
|
GLRA2
|
glycine receptor alpha 2 |
| chr9_-_127937800 | 0.04 |
ENST00000373110.4
ENST00000314392.13 |
DPM2
|
dolichyl-phosphate mannosyltransferase subunit 2, regulatory |
| chr20_-_6054026 | 0.04 |
ENST00000378858.5
|
LRRN4
|
leucine rich repeat neuronal 4 |
| chr20_-_50691536 | 0.03 |
ENST00000327979.8
|
RIPOR3
|
RIPOR family member 3 |
| chr7_+_121328991 | 0.03 |
ENST00000222462.3
|
WNT16
|
Wnt family member 16 |
| chr1_-_155688727 | 0.03 |
ENST00000404643.5
ENST00000359205.9 ENST00000407221.5 ENST00000355499.9 |
YY1AP1
|
YY1 associated protein 1 |
| chr20_-_49568101 | 0.03 |
ENST00000244043.5
|
PTGIS
|
prostaglandin I2 synthase |
| chr12_+_53299682 | 0.03 |
ENST00000267103.10
ENST00000548632.5 |
MYG1
|
MYG1 exonuclease |
| chr19_-_12156710 | 0.02 |
ENST00000455799.1
ENST00000439556.3 |
ZNF625
|
zinc finger protein 625 |
| chr1_-_155688294 | 0.02 |
ENST00000311573.9
|
YY1AP1
|
YY1 associated protein 1 |
| chr6_-_31541937 | 0.02 |
ENST00000456662.5
ENST00000431908.5 ENST00000456976.5 ENST00000428450.5 ENST00000418897.5 ENST00000396172.6 ENST00000419020.1 ENST00000428098.5 |
DDX39B
|
DExD-box helicase 39B |
| chr1_+_40691689 | 0.02 |
ENST00000427410.6
ENST00000447388.7 ENST00000425457.6 ENST00000453631.5 ENST00000456393.6 |
NFYC
|
nuclear transcription factor Y subunit gamma |
| chr8_+_117135259 | 0.01 |
ENST00000519688.5
|
SLC30A8
|
solute carrier family 30 member 8 |
| chr16_-_57971121 | 0.01 |
ENST00000251102.13
|
CNGB1
|
cyclic nucleotide gated channel subunit beta 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.4 | GO:0042377 | menaquinone catabolic process(GO:0042361) vitamin K catabolic process(GO:0042377) |
| 0.3 | 0.8 | GO:0043449 | cellular alkene metabolic process(GO:0043449) |
| 0.2 | 1.1 | GO:0048050 | post-embryonic eye morphogenesis(GO:0048050) |
| 0.1 | 0.8 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 0.1 | 0.7 | GO:0033489 | cholesterol biosynthetic process via desmosterol(GO:0033489) cholesterol biosynthetic process via lathosterol(GO:0033490) |
| 0.1 | 0.5 | GO:0060621 | negative regulation of cholesterol import(GO:0060621) negative regulation of sterol import(GO:2000910) |
| 0.1 | 0.4 | GO:0090427 | activation of meiosis(GO:0090427) |
| 0.1 | 0.8 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.1 | 1.8 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.1 | 0.2 | GO:2000506 | negative regulation of energy homeostasis(GO:2000506) |
| 0.0 | 0.5 | GO:1904996 | positive regulation of leukocyte adhesion to vascular endothelial cell(GO:1904996) |
| 0.0 | 0.1 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 0.0 | 0.5 | GO:0070779 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.0 | 1.5 | GO:0050832 | defense response to fungus(GO:0050832) |
| 0.0 | 0.2 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.0 | 0.2 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 0.0 | 0.9 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.0 | 0.2 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.0 | 0.1 | GO:0009436 | glyoxylate catabolic process(GO:0009436) |
| 0.0 | 0.2 | GO:2000786 | positive regulation of autophagosome assembly(GO:2000786) |
| 0.0 | 0.4 | GO:0048311 | mitochondrion distribution(GO:0048311) |
| 0.0 | 0.1 | GO:0007598 | blood coagulation, extrinsic pathway(GO:0007598) |
| 0.0 | 0.7 | GO:0035774 | positive regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0035774) |
| 0.0 | 0.1 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.0 | 1.1 | GO:0043268 | regulation of membrane depolarization(GO:0003254) positive regulation of potassium ion transport(GO:0043268) |
| 0.0 | 0.4 | GO:0030252 | growth hormone secretion(GO:0030252) |
| 0.0 | 0.1 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | GO:0089717 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.0 | 1.1 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.0 | 0.5 | GO:0034366 | spherical high-density lipoprotein particle(GO:0034366) |
| 0.0 | 0.7 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 0.5 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.0 | GO:0033185 | dolichol-phosphate-mannose synthase complex(GO:0033185) |
| 0.0 | 0.1 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.0 | 1.8 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 0.1 | GO:0097524 | sperm plasma membrane(GO:0097524) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.8 | GO:0030197 | extracellular matrix constituent, lubricant activity(GO:0030197) |
| 0.2 | 0.7 | GO:0047750 | C-8 sterol isomerase activity(GO:0000247) cholestenol delta-isomerase activity(GO:0047750) |
| 0.2 | 0.8 | GO:0070576 | vitamin D 24-hydroxylase activity(GO:0070576) |
| 0.1 | 0.8 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.1 | 0.5 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 0.1 | 0.5 | GO:0030395 | lactose binding(GO:0030395) |
| 0.1 | 0.2 | GO:0016309 | 1-phosphatidylinositol-5-phosphate 4-kinase activity(GO:0016309) |
| 0.1 | 2.4 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.0 | 0.2 | GO:0005223 | intracellular cAMP activated cation channel activity(GO:0005222) intracellular cGMP activated cation channel activity(GO:0005223) |
| 0.0 | 0.5 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.0 | 0.1 | GO:0005298 | proline:sodium symporter activity(GO:0005298) |
| 0.0 | 0.1 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 0.0 | 0.0 | GO:0004582 | dolichyl-phosphate beta-D-mannosyltransferase activity(GO:0004582) |
| 0.0 | 0.1 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.0 | 0.4 | GO:0005521 | lamin binding(GO:0005521) |
| 0.0 | 0.9 | GO:0019003 | GDP binding(GO:0019003) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.4 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.4 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.0 | 1.1 | PID NOTCH PATHWAY | Notch signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 3.3 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.0 | 0.9 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.4 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.0 | 0.5 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.6 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |