Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
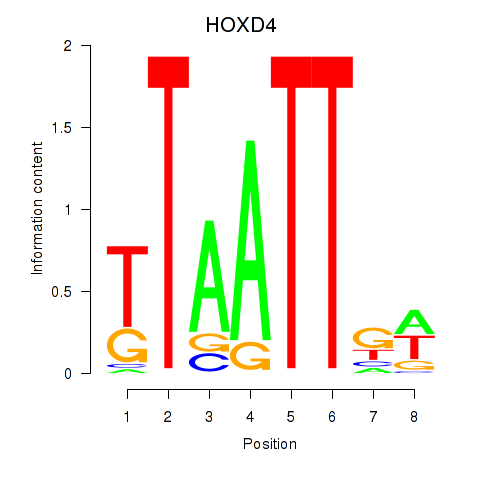
Results for HOXD4
Z-value: 0.47
Transcription factors associated with HOXD4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXD4
|
ENSG00000170166.6 | HOXD4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HOXD4 | hg38_v1_chr2_+_176151543_176151559 | -0.15 | 4.3e-01 | Click! |
Activity profile of HOXD4 motif
Sorted Z-values of HOXD4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXD4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_-_7940444 | 0.61 |
ENST00000378789.4
|
HAO1
|
hydroxyacid oxidase 1 |
| chr17_+_9576627 | 0.58 |
ENST00000396219.7
ENST00000352665.10 |
CFAP52
|
cilia and flagella associated protein 52 |
| chr7_+_92057602 | 0.57 |
ENST00000491695.2
|
AKAP9
|
A-kinase anchoring protein 9 |
| chr13_+_42781578 | 0.53 |
ENST00000313851.3
|
FAM216B
|
family with sequence similarity 216 member B |
| chr18_+_63702958 | 0.53 |
ENST00000544088.6
|
SERPINB11
|
serpin family B member 11 |
| chr12_+_9827517 | 0.52 |
ENST00000537723.5
|
KLRF1
|
killer cell lectin like receptor F1 |
| chr17_-_10549694 | 0.47 |
ENST00000622564.4
|
MYH2
|
myosin heavy chain 2 |
| chr3_+_63652663 | 0.45 |
ENST00000343837.8
ENST00000469440.5 |
SNTN
|
sentan, cilia apical structure protein |
| chr12_-_39340963 | 0.44 |
ENST00000552961.5
|
KIF21A
|
kinesin family member 21A |
| chr2_-_158380960 | 0.43 |
ENST00000409187.5
|
CCDC148
|
coiled-coil domain containing 148 |
| chr7_-_122699108 | 0.40 |
ENST00000340112.3
|
RNF133
|
ring finger protein 133 |
| chr7_-_25228485 | 0.39 |
ENST00000222674.2
|
NPVF
|
neuropeptide VF precursor |
| chr4_+_68447453 | 0.38 |
ENST00000305363.9
|
TMPRSS11E
|
transmembrane serine protease 11E |
| chr13_+_23570370 | 0.38 |
ENST00000403372.6
ENST00000248484.9 |
TNFRSF19
|
TNF receptor superfamily member 19 |
| chr3_+_159069252 | 0.37 |
ENST00000640015.1
ENST00000476809.7 ENST00000485419.7 |
IQCJ-SCHIP1
|
IQCJ-SCHIP1 readthrough |
| chr17_-_10549612 | 0.36 |
ENST00000532183.6
ENST00000397183.6 ENST00000420805.1 |
MYH2
|
myosin heavy chain 2 |
| chr2_-_55296361 | 0.36 |
ENST00000647547.1
|
CCDC88A
|
coiled-coil domain containing 88A |
| chr8_-_71361860 | 0.34 |
ENST00000303824.11
ENST00000645451.1 |
EYA1
|
EYA transcriptional coactivator and phosphatase 1 |
| chr6_-_75363003 | 0.33 |
ENST00000370020.1
|
FILIP1
|
filamin A interacting protein 1 |
| chr8_-_101790934 | 0.32 |
ENST00000523645.5
ENST00000520346.1 ENST00000220931.11 ENST00000522448.5 ENST00000522951.5 ENST00000522252.5 ENST00000519098.5 |
NCALD
|
neurocalcin delta |
| chr16_-_2329687 | 0.29 |
ENST00000567910.1
|
ABCA3
|
ATP binding cassette subfamily A member 3 |
| chr17_-_10549652 | 0.29 |
ENST00000245503.10
|
MYH2
|
myosin heavy chain 2 |
| chr6_-_49744378 | 0.29 |
ENST00000371159.8
ENST00000263045.9 |
CRISP3
|
cysteine rich secretory protein 3 |
| chr6_-_87095059 | 0.28 |
ENST00000369582.6
ENST00000610310.3 ENST00000630630.2 ENST00000627148.3 ENST00000625577.1 |
CGA
|
glycoprotein hormones, alpha polypeptide |
| chr6_-_49744434 | 0.28 |
ENST00000433368.6
ENST00000354620.4 |
CRISP3
|
cysteine rich secretory protein 3 |
| chr4_-_68245683 | 0.27 |
ENST00000332644.6
|
TMPRSS11B
|
transmembrane serine protease 11B |
| chr10_-_96359273 | 0.25 |
ENST00000393871.5
ENST00000419479.5 ENST00000393870.3 |
OPALIN
|
oligodendrocytic myelin paranodal and inner loop protein |
| chr4_+_73481737 | 0.24 |
ENST00000226355.5
|
AFM
|
afamin |
| chr3_+_97764728 | 0.24 |
ENST00000463745.6
|
ARL6
|
ADP ribosylation factor like GTPase 6 |
| chr2_+_190180835 | 0.24 |
ENST00000340623.4
|
C2orf88
|
chromosome 2 open reading frame 88 |
| chr13_-_23433676 | 0.23 |
ENST00000682547.1
ENST00000455470.6 ENST00000382292.9 |
SACS
|
sacsin molecular chaperone |
| chr1_+_182450132 | 0.23 |
ENST00000294854.13
|
RGSL1
|
regulator of G protein signaling like 1 |
| chr5_+_70025247 | 0.22 |
ENST00000380751.9
ENST00000380750.8 ENST00000503931.5 ENST00000506542.1 |
SERF1B
|
small EDRK-rich factor 1B |
| chr4_-_129093454 | 0.22 |
ENST00000281142.10
ENST00000511426.5 |
SCLT1
|
sodium channel and clathrin linker 1 |
| chr4_-_152679984 | 0.22 |
ENST00000304385.8
ENST00000504064.1 |
TMEM154
|
transmembrane protein 154 |
| chr12_-_56741535 | 0.22 |
ENST00000647707.1
|
ENSG00000285625.1
|
novel protein |
| chr5_+_69565122 | 0.21 |
ENST00000507595.1
|
GTF2H2C
|
GTF2H2 family member C |
| chr12_-_86256267 | 0.21 |
ENST00000620241.4
|
MGAT4C
|
MGAT4 family member C |
| chr4_-_129093377 | 0.21 |
ENST00000506368.5
ENST00000439369.6 ENST00000503215.5 |
SCLT1
|
sodium channel and clathrin linker 1 |
| chr8_-_71362054 | 0.21 |
ENST00000340726.8
|
EYA1
|
EYA transcriptional coactivator and phosphatase 1 |
| chr22_+_41092869 | 0.20 |
ENST00000674155.1
|
EP300
|
E1A binding protein p300 |
| chr2_+_108377947 | 0.20 |
ENST00000272452.7
|
SULT1C4
|
sulfotransferase family 1C member 4 |
| chr4_-_174263080 | 0.19 |
ENST00000615392.4
|
FBXO8
|
F-box protein 8 |
| chr3_+_97821984 | 0.19 |
ENST00000389622.7
|
CRYBG3
|
crystallin beta-gamma domain containing 3 |
| chr2_+_190180930 | 0.19 |
ENST00000443551.2
|
C2orf88
|
chromosome 2 open reading frame 88 |
| chr10_-_17129786 | 0.19 |
ENST00000377833.10
|
CUBN
|
cubilin |
| chr5_-_111512473 | 0.18 |
ENST00000296632.8
ENST00000512160.5 ENST00000509887.5 |
STARD4
|
StAR related lipid transfer domain containing 4 |
| chrX_+_77910656 | 0.18 |
ENST00000343533.9
ENST00000341514.11 ENST00000645454.1 ENST00000642651.1 ENST00000644362.1 |
ATP7A
PGK1
|
ATPase copper transporting alpha phosphoglycerate kinase 1 |
| chr10_+_94089067 | 0.17 |
ENST00000371375.1
ENST00000675218.1 |
PLCE1
|
phospholipase C epsilon 1 |
| chr18_-_21703688 | 0.17 |
ENST00000584464.1
ENST00000578270.5 |
ABHD3
|
abhydrolase domain containing 3, phospholipase |
| chr5_+_31193739 | 0.16 |
ENST00000514738.5
|
CDH6
|
cadherin 6 |
| chr13_-_61415508 | 0.16 |
ENST00000409204.4
|
PCDH20
|
protocadherin 20 |
| chr4_-_69639642 | 0.15 |
ENST00000604629.6
ENST00000604021.1 |
UGT2A2
|
UDP glucuronosyltransferase family 2 member A2 |
| chr19_-_44401575 | 0.15 |
ENST00000591679.5
ENST00000544719.6 ENST00000614994.5 ENST00000585868.1 ENST00000588212.1 |
ZNF285
ENSG00000267173.1
|
zinc finger protein 285 novel protein |
| chr13_+_108629605 | 0.14 |
ENST00000457511.7
|
MYO16
|
myosin XVI |
| chr1_-_150765785 | 0.14 |
ENST00000680311.1
ENST00000681728.1 ENST00000680288.1 |
CTSS
|
cathepsin S |
| chr6_-_44313306 | 0.14 |
ENST00000244571.5
|
AARS2
|
alanyl-tRNA synthetase 2, mitochondrial |
| chr11_-_40294089 | 0.14 |
ENST00000278198.2
|
LRRC4C
|
leucine rich repeat containing 4C |
| chrX_+_37990773 | 0.14 |
ENST00000341016.5
|
H2AP
|
H2A.P histone |
| chr13_-_23433735 | 0.14 |
ENST00000423156.2
ENST00000683210.1 ENST00000682775.1 ENST00000684497.1 ENST00000682944.1 ENST00000683489.1 ENST00000684385.1 ENST00000683680.1 |
SACS
|
sacsin molecular chaperone |
| chr1_-_150765735 | 0.14 |
ENST00000679898.1
ENST00000448301.7 ENST00000680664.1 ENST00000679512.1 ENST00000368985.8 ENST00000679582.1 |
CTSS
|
cathepsin S |
| chr5_-_22853320 | 0.13 |
ENST00000504376.6
ENST00000382254.6 |
CDH12
|
cadherin 12 |
| chr8_+_75539893 | 0.13 |
ENST00000674002.1
|
HNF4G
|
hepatocyte nuclear factor 4 gamma |
| chr20_+_8789517 | 0.13 |
ENST00000437439.2
|
PLCB1
|
phospholipase C beta 1 |
| chr5_+_31193678 | 0.13 |
ENST00000265071.3
|
CDH6
|
cadherin 6 |
| chr18_+_24460655 | 0.13 |
ENST00000426880.2
|
HRH4
|
histamine receptor H4 |
| chr1_-_100894818 | 0.12 |
ENST00000370114.8
|
EXTL2
|
exostosin like glycosyltransferase 2 |
| chr4_+_127632926 | 0.12 |
ENST00000335251.11
|
INTU
|
inturned planar cell polarity protein |
| chr11_+_66011994 | 0.12 |
ENST00000312134.3
|
CST6
|
cystatin E/M |
| chr17_-_37542361 | 0.12 |
ENST00000614196.1
|
SYNRG
|
synergin gamma |
| chr4_-_142305935 | 0.12 |
ENST00000511838.5
|
INPP4B
|
inositol polyphosphate-4-phosphatase type II B |
| chr1_-_100894775 | 0.12 |
ENST00000416479.1
ENST00000370113.7 |
EXTL2
|
exostosin like glycosyltransferase 2 |
| chr3_+_142623386 | 0.11 |
ENST00000337777.7
ENST00000497199.5 |
PLS1
|
plastin 1 |
| chr7_-_22193728 | 0.11 |
ENST00000620335.4
|
RAPGEF5
|
Rap guanine nucleotide exchange factor 5 |
| chr2_+_71453538 | 0.11 |
ENST00000258104.8
|
DYSF
|
dysferlin |
| chrX_+_56563569 | 0.11 |
ENST00000338222.7
|
UBQLN2
|
ubiquilin 2 |
| chr4_+_158210479 | 0.11 |
ENST00000504569.5
ENST00000509278.5 ENST00000514558.5 ENST00000503200.5 ENST00000296529.11 |
TMEM144
|
transmembrane protein 144 |
| chr10_+_84424919 | 0.11 |
ENST00000543283.2
ENST00000494586.5 |
CCSER2
|
coiled-coil serine rich protein 2 |
| chr19_-_13953302 | 0.11 |
ENST00000585607.1
ENST00000538517.6 ENST00000587458.1 ENST00000538371.6 |
PODNL1
|
podocan like 1 |
| chrX_-_23884017 | 0.11 |
ENST00000633372.1
|
APOO
|
apolipoprotein O |
| chr9_+_117704382 | 0.10 |
ENST00000646089.1
ENST00000355622.8 |
ENSG00000285082.2
TLR4
|
novel protein toll like receptor 4 |
| chr15_+_76336755 | 0.10 |
ENST00000290759.9
|
ISL2
|
ISL LIM homeobox 2 |
| chr2_+_27663441 | 0.10 |
ENST00000326019.10
ENST00000613058.4 |
SLC4A1AP
|
solute carrier family 4 member 1 adaptor protein |
| chr9_+_273026 | 0.10 |
ENST00000682249.1
ENST00000453981.5 ENST00000487230.5 ENST00000469391.5 |
DOCK8
|
dedicator of cytokinesis 8 |
| chr4_-_84966637 | 0.10 |
ENST00000295888.9
|
WDFY3
|
WD repeat and FYVE domain containing 3 |
| chr12_+_112906777 | 0.10 |
ENST00000452357.7
ENST00000445409.7 |
OAS1
|
2'-5'-oligoadenylate synthetase 1 |
| chr12_+_12785652 | 0.10 |
ENST00000356591.5
|
APOLD1
|
apolipoprotein L domain containing 1 |
| chr6_-_53148822 | 0.10 |
ENST00000259803.8
|
GCM1
|
glial cells missing transcription factor 1 |
| chr6_+_46793379 | 0.10 |
ENST00000230588.9
ENST00000611727.2 |
MEP1A
|
meprin A subunit alpha |
| chr10_-_72217126 | 0.10 |
ENST00000317126.8
|
ASCC1
|
activating signal cointegrator 1 complex subunit 1 |
| chr9_-_21368057 | 0.10 |
ENST00000449498.2
|
IFNA13
|
interferon alpha 13 |
| chr19_-_15815664 | 0.10 |
ENST00000641419.1
|
OR10H1
|
olfactory receptor family 10 subfamily H member 1 |
| chr19_+_48325323 | 0.09 |
ENST00000596315.5
|
EMP3
|
epithelial membrane protein 3 |
| chr4_+_168497066 | 0.09 |
ENST00000261509.10
|
PALLD
|
palladin, cytoskeletal associated protein |
| chr11_-_129192198 | 0.09 |
ENST00000310343.13
|
ARHGAP32
|
Rho GTPase activating protein 32 |
| chr19_+_48325522 | 0.09 |
ENST00000594198.1
ENST00000270221.11 ENST00000597279.5 ENST00000593437.1 |
EMP3
|
epithelial membrane protein 3 |
| chr6_-_26250625 | 0.09 |
ENST00000618052.2
|
H3C7
|
H3 clustered histone 7 |
| chr5_-_35047935 | 0.09 |
ENST00000510428.1
ENST00000231420.11 |
AGXT2
|
alanine--glyoxylate aminotransferase 2 |
| chr17_-_28406160 | 0.09 |
ENST00000618626.1
ENST00000612814.5 |
SLC46A1
|
solute carrier family 46 member 1 |
| chr15_+_36594868 | 0.08 |
ENST00000566807.5
ENST00000643612.1 ENST00000567389.5 ENST00000562877.5 |
CDIN1
|
CDAN1 interacting nuclease 1 |
| chr7_-_22193824 | 0.08 |
ENST00000401957.6
|
RAPGEF5
|
Rap guanine nucleotide exchange factor 5 |
| chr1_+_149782671 | 0.08 |
ENST00000444948.5
ENST00000369168.5 |
FCGR1A
|
Fc fragment of IgG receptor Ia |
| chr9_-_21368962 | 0.08 |
ENST00000610660.1
|
IFNA13
|
interferon alpha 13 |
| chr3_+_4680617 | 0.08 |
ENST00000648212.1
|
ITPR1
|
inositol 1,4,5-trisphosphate receptor type 1 |
| chr6_+_42746958 | 0.08 |
ENST00000614467.4
|
BICRAL
|
BRD4 interacting chromatin remodeling complex associated protein like |
| chr2_+_167187364 | 0.08 |
ENST00000672671.1
|
XIRP2
|
xin actin binding repeat containing 2 |
| chr15_+_92900189 | 0.08 |
ENST00000626874.2
ENST00000627622.1 ENST00000629346.2 ENST00000628375.2 ENST00000420239.7 ENST00000394196.9 |
CHD2
|
chromodomain helicase DNA binding protein 2 |
| chr10_+_52128343 | 0.08 |
ENST00000672084.1
|
PRKG1
|
protein kinase cGMP-dependent 1 |
| chr2_-_165204042 | 0.07 |
ENST00000283254.12
ENST00000453007.1 |
SCN3A
|
sodium voltage-gated channel alpha subunit 3 |
| chr18_+_24460630 | 0.07 |
ENST00000256906.5
|
HRH4
|
histamine receptor H4 |
| chr8_+_42697339 | 0.07 |
ENST00000289957.3
ENST00000534391.1 |
CHRNB3
|
cholinergic receptor nicotinic beta 3 subunit |
| chr5_-_179617581 | 0.07 |
ENST00000523449.5
|
HNRNPH1
|
heterogeneous nuclear ribonucleoprotein H1 |
| chr7_+_77798832 | 0.07 |
ENST00000415251.6
ENST00000275575.11 |
PHTF2
|
putative homeodomain transcription factor 2 |
| chr4_-_137532452 | 0.07 |
ENST00000412923.6
ENST00000511115.5 ENST00000344876.9 ENST00000507846.5 ENST00000510305.5 ENST00000611581.1 |
PCDH18
|
protocadherin 18 |
| chr3_+_15003912 | 0.07 |
ENST00000406272.6
ENST00000617312.4 |
NR2C2
|
nuclear receptor subfamily 2 group C member 2 |
| chr22_-_28712136 | 0.07 |
ENST00000464581.6
|
CHEK2
|
checkpoint kinase 2 |
| chr11_-_129192291 | 0.06 |
ENST00000682385.1
|
ARHGAP32
|
Rho GTPase activating protein 32 |
| chr5_+_69415065 | 0.06 |
ENST00000647531.1
ENST00000645446.1 ENST00000325631.10 ENST00000454295.6 |
MARVELD2
|
MARVEL domain containing 2 |
| chr8_-_42843201 | 0.06 |
ENST00000529779.1
ENST00000345117.2 ENST00000254250.7 |
THAP1
|
THAP domain containing 1 |
| chr6_+_28349907 | 0.06 |
ENST00000252211.7
ENST00000341464.9 ENST00000377255.3 |
ZKSCAN3
|
zinc finger with KRAB and SCAN domains 3 |
| chr15_+_92900338 | 0.06 |
ENST00000625990.3
|
CHD2
|
chromodomain helicase DNA binding protein 2 |
| chr7_+_77798750 | 0.06 |
ENST00000416283.6
ENST00000422959.6 ENST00000307305.12 ENST00000424760.5 |
PHTF2
|
putative homeodomain transcription factor 2 |
| chr1_+_115029823 | 0.06 |
ENST00000256592.3
|
TSHB
|
thyroid stimulating hormone subunit beta |
| chr4_+_40197023 | 0.06 |
ENST00000381799.10
|
RHOH
|
ras homolog family member H |
| chr7_+_138460238 | 0.06 |
ENST00000343526.9
|
TRIM24
|
tripartite motif containing 24 |
| chr2_+_223957431 | 0.06 |
ENST00000258383.4
|
MRPL44
|
mitochondrial ribosomal protein L44 |
| chr1_+_28438104 | 0.06 |
ENST00000633167.1
ENST00000373836.4 |
PHACTR4
|
phosphatase and actin regulator 4 |
| chr5_+_141966820 | 0.06 |
ENST00000513019.5
ENST00000394519.5 ENST00000356143.5 |
RNF14
|
ring finger protein 14 |
| chr4_+_168497044 | 0.06 |
ENST00000505667.6
|
PALLD
|
palladin, cytoskeletal associated protein |
| chr1_-_113871665 | 0.06 |
ENST00000528414.5
ENST00000460620.5 ENST00000359785.10 ENST00000420377.6 ENST00000525799.1 ENST00000538253.5 |
PTPN22
|
protein tyrosine phosphatase non-receptor type 22 |
| chr20_-_31390483 | 0.06 |
ENST00000376315.2
|
DEFB119
|
defensin beta 119 |
| chr15_+_21651844 | 0.06 |
ENST00000623441.1
|
OR4N4C
|
olfactory receptor family 4 subfamily N member 4C |
| chr18_-_24311495 | 0.05 |
ENST00000357041.8
|
OSBPL1A
|
oxysterol binding protein like 1A |
| chr3_+_111998915 | 0.05 |
ENST00000478951.6
|
TAGLN3
|
transgelin 3 |
| chr6_-_136526177 | 0.05 |
ENST00000617204.4
|
MAP7
|
microtubule associated protein 7 |
| chr10_-_14330879 | 0.05 |
ENST00000357447.7
|
FRMD4A
|
FERM domain containing 4A |
| chr1_-_246193727 | 0.05 |
ENST00000391836.3
|
SMYD3
|
SET and MYND domain containing 3 |
| chrX_-_13817027 | 0.05 |
ENST00000493677.5
ENST00000355135.6 ENST00000316715.9 |
GPM6B
|
glycoprotein M6B |
| chr22_-_28711931 | 0.05 |
ENST00000434810.5
ENST00000456369.5 |
CHEK2
|
checkpoint kinase 2 |
| chr7_-_29146436 | 0.05 |
ENST00000396276.7
|
CPVL
|
carboxypeptidase vitellogenic like |
| chr3_-_18438767 | 0.05 |
ENST00000454909.6
|
SATB1
|
SATB homeobox 1 |
| chr4_-_129093548 | 0.05 |
ENST00000503401.1
|
SCLT1
|
sodium channel and clathrin linker 1 |
| chr2_-_162838728 | 0.05 |
ENST00000328032.8
ENST00000332142.10 |
KCNH7
|
potassium voltage-gated channel subfamily H member 7 |
| chr8_-_19013649 | 0.05 |
ENST00000440756.4
|
PSD3
|
pleckstrin and Sec7 domain containing 3 |
| chr1_-_92486916 | 0.05 |
ENST00000294702.6
|
GFI1
|
growth factor independent 1 transcriptional repressor |
| chr9_-_94640130 | 0.05 |
ENST00000414122.1
|
FBP1
|
fructose-bisphosphatase 1 |
| chr1_+_210328244 | 0.04 |
ENST00000541565.5
ENST00000413764.6 |
HHAT
|
hedgehog acyltransferase |
| chr1_+_86914616 | 0.04 |
ENST00000370550.10
ENST00000370551.8 |
HS2ST1
|
heparan sulfate 2-O-sulfotransferase 1 |
| chr7_-_137343752 | 0.04 |
ENST00000393083.2
|
PTN
|
pleiotrophin |
| chr7_-_29146527 | 0.04 |
ENST00000265394.10
|
CPVL
|
carboxypeptidase vitellogenic like |
| chr12_-_118190510 | 0.04 |
ENST00000540561.5
ENST00000537952.1 ENST00000537822.1 |
TAOK3
|
TAO kinase 3 |
| chr20_-_31390580 | 0.04 |
ENST00000339144.3
ENST00000376321.4 |
DEFB119
|
defensin beta 119 |
| chr17_-_41047267 | 0.04 |
ENST00000542137.1
ENST00000391419.3 |
KRTAP2-1
|
keratin associated protein 2-1 |
| chr2_+_186590022 | 0.04 |
ENST00000261023.8
ENST00000374907.7 |
ITGAV
|
integrin subunit alpha V |
| chr4_-_39977836 | 0.04 |
ENST00000303538.13
ENST00000503396.5 |
PDS5A
|
PDS5 cohesin associated factor A |
| chr4_-_151227881 | 0.03 |
ENST00000652233.1
ENST00000514152.5 |
SH3D19
|
SH3 domain containing 19 |
| chr2_-_174597728 | 0.03 |
ENST00000409891.5
ENST00000410117.5 |
WIPF1
|
WAS/WASL interacting protein family member 1 |
| chr5_-_59430600 | 0.03 |
ENST00000636120.1
|
PDE4D
|
phosphodiesterase 4D |
| chr2_-_190062721 | 0.03 |
ENST00000260950.5
|
MSTN
|
myostatin |
| chr16_+_24537693 | 0.03 |
ENST00000564314.5
ENST00000567686.5 |
RBBP6
|
RB binding protein 6, ubiquitin ligase |
| chr5_-_180072086 | 0.03 |
ENST00000261947.4
|
RNF130
|
ring finger protein 130 |
| chr8_-_69834970 | 0.03 |
ENST00000260126.9
|
SLCO5A1
|
solute carrier organic anion transporter family member 5A1 |
| chr1_+_67166448 | 0.03 |
ENST00000347310.10
|
IL23R
|
interleukin 23 receptor |
| chr21_-_29061351 | 0.03 |
ENST00000432178.5
|
CCT8
|
chaperonin containing TCP1 subunit 8 |
| chr14_+_50533026 | 0.03 |
ENST00000441560.6
|
ATL1
|
atlastin GTPase 1 |
| chr4_-_142305826 | 0.03 |
ENST00000514525.1
|
INPP4B
|
inositol polyphosphate-4-phosphatase type II B |
| chr13_+_77741160 | 0.03 |
ENST00000314070.9
ENST00000351546.7 |
SLAIN1
|
SLAIN motif family member 1 |
| chr7_-_105269007 | 0.02 |
ENST00000357311.7
|
SRPK2
|
SRSF protein kinase 2 |
| chr2_+_170178136 | 0.02 |
ENST00000409044.7
ENST00000408978.9 |
MYO3B
|
myosin IIIB |
| chr12_+_15956585 | 0.02 |
ENST00000526530.1
|
DERA
|
deoxyribose-phosphate aldolase |
| chr5_-_62403506 | 0.02 |
ENST00000680062.1
|
DIMT1
|
DIMT1 rRNA methyltransferase and ribosome maturation factor |
| chr20_-_45101112 | 0.02 |
ENST00000306117.5
ENST00000537075.3 |
KCNS1
|
potassium voltage-gated channel modifier subfamily S member 1 |
| chr6_-_136526472 | 0.02 |
ENST00000454590.5
ENST00000432797.6 |
MAP7
|
microtubule associated protein 7 |
| chr2_-_174597795 | 0.02 |
ENST00000679041.1
|
WIPF1
|
WAS/WASL interacting protein family member 1 |
| chr3_-_57292676 | 0.02 |
ENST00000389601.3
ENST00000487349.6 |
ASB14
|
ankyrin repeat and SOCS box containing 14 |
| chr17_-_75848641 | 0.02 |
ENST00000586257.5
|
WBP2
|
WW domain binding protein 2 |
| chr14_+_22598224 | 0.02 |
ENST00000428304.6
ENST00000542041.1 ENST00000216327.10 |
ABHD4
|
abhydrolase domain containing 4, N-acyl phospholipase B |
| chrX_-_93673558 | 0.02 |
ENST00000475430.2
ENST00000373079.4 |
NAP1L3
|
nucleosome assembly protein 1 like 3 |
| chr1_+_150067820 | 0.02 |
ENST00000419023.3
ENST00000644526.1 |
VPS45
|
vacuolar protein sorting 45 homolog |
| chrX_-_120575783 | 0.02 |
ENST00000680673.1
|
CUL4B
|
cullin 4B |
| chr7_-_77798359 | 0.02 |
ENST00000257663.4
|
TMEM60
|
transmembrane protein 60 |
| chr1_+_197268222 | 0.02 |
ENST00000367400.8
ENST00000638467.1 ENST00000367399.6 |
CRB1
|
crumbs cell polarity complex component 1 |
| chr1_+_84164962 | 0.01 |
ENST00000614872.4
ENST00000394839.6 |
PRKACB
|
protein kinase cAMP-activated catalytic subunit beta |
| chr8_+_49909783 | 0.01 |
ENST00000518864.5
|
SNTG1
|
syntrophin gamma 1 |
| chr3_-_197260369 | 0.01 |
ENST00000658155.1
ENST00000453607.5 |
DLG1
|
discs large MAGUK scaffold protein 1 |
| chr3_+_160225409 | 0.01 |
ENST00000326474.5
|
C3orf80
|
chromosome 3 open reading frame 80 |
| chr13_+_19633642 | 0.01 |
ENST00000361479.10
|
MPHOSPH8
|
M-phase phosphoprotein 8 |
| chr7_-_137343688 | 0.01 |
ENST00000348225.7
|
PTN
|
pleiotrophin |
| chr1_+_150067668 | 0.01 |
ENST00000611412.4
ENST00000644510.2 ENST00000643611.1 |
VPS45
|
vacuolar protein sorting 45 homolog |
| chr4_+_70592253 | 0.01 |
ENST00000322937.10
ENST00000613447.4 |
AMBN
|
ameloblastin |
| chr6_-_26108125 | 0.00 |
ENST00000338379.6
|
H1-6
|
H1.6 linker histone, cluster member |
| chr20_-_10420737 | 0.00 |
ENST00000649912.1
|
ENSG00000285723.1
|
novel protein |
| chr10_-_29736956 | 0.00 |
ENST00000674475.1
|
SVIL
|
supervillin |
| chr2_-_70248454 | 0.00 |
ENST00000416149.6
ENST00000282574.8 |
TIA1
|
TIA1 cytotoxic granule associated RNA binding protein |
| chr2_-_74392025 | 0.00 |
ENST00000440727.1
ENST00000409240.5 |
DCTN1
|
dynactin subunit 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0034769 | basement membrane disassembly(GO:0034769) |
| 0.1 | 0.2 | GO:0010273 | detoxification of copper ion(GO:0010273) stress response to copper ion(GO:1990169) |
| 0.1 | 0.6 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 0.1 | 1.2 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.1 | 0.2 | GO:1904253 | positive regulation of bile acid biosynthetic process(GO:0070859) positive regulation of bile acid metabolic process(GO:1904253) |
| 0.1 | 0.5 | GO:0072513 | positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.1 | 0.2 | GO:0046909 | intermembrane transport(GO:0046909) protein transport from ciliary membrane to plasma membrane(GO:1903445) |
| 0.1 | 0.4 | GO:1903566 | positive regulation of protein localization to cilium(GO:1903566) |
| 0.0 | 0.6 | GO:0051661 | maintenance of centrosome location(GO:0051661) |
| 0.0 | 0.2 | GO:0018076 | N-terminal peptidyl-lysine acetylation(GO:0018076) |
| 0.0 | 0.2 | GO:0019276 | UDP-N-acetylgalactosamine metabolic process(GO:0019276) |
| 0.0 | 0.1 | GO:1903926 | signal transduction involved in intra-S DNA damage checkpoint(GO:0072428) response to bisphenol A(GO:1903925) cellular response to bisphenol A(GO:1903926) |
| 0.0 | 0.1 | GO:1900186 | negative regulation of clathrin-mediated endocytosis(GO:1900186) |
| 0.0 | 0.4 | GO:0032277 | negative regulation of gonadotropin secretion(GO:0032277) |
| 0.0 | 0.1 | GO:0032707 | negative regulation of interleukin-23 production(GO:0032707) |
| 0.0 | 0.1 | GO:0021524 | visceral motor neuron differentiation(GO:0021524) |
| 0.0 | 0.1 | GO:0090427 | activation of meiosis(GO:0090427) |
| 0.0 | 0.1 | GO:0060221 | retinal rod cell differentiation(GO:0060221) |
| 0.0 | 0.2 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.0 | 0.1 | GO:0060018 | astrocyte fate commitment(GO:0060018) |
| 0.0 | 0.1 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.0 | 0.1 | GO:0009436 | glyoxylate catabolic process(GO:0009436) |
| 0.0 | 0.3 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.0 | 0.1 | GO:0070127 | tRNA aminoacylation for mitochondrial protein translation(GO:0070127) |
| 0.0 | 0.1 | GO:0061386 | closure of optic fissure(GO:0061386) negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.0 | 0.4 | GO:0090084 | negative regulation of inclusion body assembly(GO:0090084) |
| 0.0 | 0.0 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.0 | 0.1 | GO:0051958 | methotrexate transport(GO:0051958) |
| 0.0 | 0.2 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.0 | 0.0 | GO:0045715 | negative regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045715) |
| 0.0 | 0.1 | GO:0070433 | negative regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070425) regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070432) negative regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070433) positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 0.0 | 0.0 | GO:0005986 | sucrose biosynthetic process(GO:0005986) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:0005826 | actomyosin contractile ring(GO:0005826) |
| 0.1 | 0.6 | GO:0044307 | dendritic branch(GO:0044307) |
| 0.1 | 0.3 | GO:0097232 | lamellar body membrane(GO:0097232) alveolar lamellar body membrane(GO:0097233) |
| 0.0 | 0.3 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.0 | 0.4 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.0 | 0.4 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.0 | 0.5 | GO:0071439 | clathrin complex(GO:0071439) |
| 0.0 | 0.2 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.0 | 0.2 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.0 | 0.1 | GO:0034274 | Atg12-Atg5-Atg16 complex(GO:0034274) |
| 0.0 | 0.2 | GO:0000439 | core TFIIH complex(GO:0000439) |
| 0.0 | 0.1 | GO:1990357 | terminal web(GO:1990357) |
| 0.0 | 0.1 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 0.0 | GO:0034686 | integrin alphav-beta8 complex(GO:0034686) |
| 0.0 | 0.1 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.0 | 0.1 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0003973 | (S)-2-hydroxy-acid oxidase activity(GO:0003973) very-long-chain-(S)-2-hydroxy-acid oxidase activity(GO:0052852) long-chain-(S)-2-hydroxy-long-chain-acid oxidase activity(GO:0052853) medium-chain-(S)-2-hydroxy-acid oxidase activity(GO:0052854) |
| 0.1 | 0.2 | GO:0008431 | vitamin E binding(GO:0008431) |
| 0.1 | 0.2 | GO:0035248 | alpha-1,4-N-acetylgalactosaminyltransferase activity(GO:0035248) |
| 0.1 | 0.2 | GO:0004618 | copper-exporting ATPase activity(GO:0004008) phosphoglycerate kinase activity(GO:0004618) copper-transporting ATPase activity(GO:0043682) |
| 0.1 | 0.2 | GO:0052739 | phosphatidylserine 1-acylhydrolase activity(GO:0052739) |
| 0.0 | 0.5 | GO:0032393 | MHC class I receptor activity(GO:0032393) |
| 0.0 | 1.1 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.0 | 0.2 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.0 | 0.9 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.0 | 0.1 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) inositol-1,3,4-trisphosphate 4-phosphatase activity(GO:0017161) phosphatidylinositol-4,5-bisphosphate 4-phosphatase activity(GO:0034597) inositol-3,4-bisphosphate 4-phosphatase activity(GO:0052828) |
| 0.0 | 0.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.0 | 0.1 | GO:0008453 | alanine-glyoxylate transaminase activity(GO:0008453) |
| 0.0 | 0.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.1 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.0 | 0.4 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.0 | 0.2 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.0 | 0.1 | GO:0015350 | methotrexate transporter activity(GO:0015350) |
| 0.0 | 0.4 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.0 | 0.2 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.0 | 0.1 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 0.0 | 0.1 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.0 | 0.0 | GO:0042132 | fructose 1,6-bisphosphate 1-phosphatase activity(GO:0042132) |
| 0.0 | 0.1 | GO:0003831 | beta-N-acetylglucosaminylglycopeptide beta-1,4-galactosyltransferase activity(GO:0003831) |
| 0.0 | 0.4 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.1 | GO:0032296 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.1 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.0 | 0.3 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.0 | 0.1 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 0.6 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.0 | 0.3 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.3 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.4 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |