Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
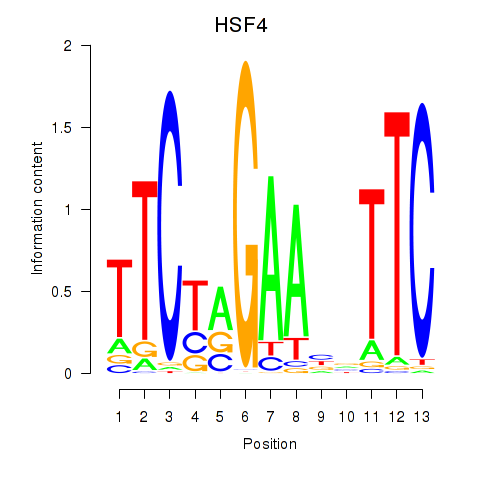
Results for HSF4
Z-value: 0.41
Transcription factors associated with HSF4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HSF4
|
ENSG00000102878.18 | HSF4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HSF4 | hg38_v1_chr16_+_67164730_67164768 | 0.19 | 3.0e-01 | Click! |
Activity profile of HSF4 motif
Sorted Z-values of HSF4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HSF4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_76141240 | 1.74 |
ENST00000322957.7
|
FOXJ1
|
forkhead box J1 |
| chr9_-_135499846 | 1.59 |
ENST00000429260.7
|
C9orf116
|
chromosome 9 open reading frame 116 |
| chr18_-_74147816 | 1.52 |
ENST00000419743.7
ENST00000582526.1 |
FBXO15
|
F-box protein 15 |
| chr19_-_55160668 | 1.43 |
ENST00000588076.1
|
DNAAF3
|
dynein axonemal assembly factor 3 |
| chr1_-_60073733 | 1.13 |
ENST00000450089.6
|
C1orf87
|
chromosome 1 open reading frame 87 |
| chr1_-_60073750 | 1.12 |
ENST00000371201.3
|
C1orf87
|
chromosome 1 open reading frame 87 |
| chr1_-_173669835 | 0.79 |
ENST00000333279.3
|
ANKRD45
|
ankyrin repeat domain 45 |
| chr18_+_46917561 | 0.74 |
ENST00000683218.1
|
KATNAL2
|
katanin catalytic subunit A1 like 2 |
| chr5_+_141223332 | 0.70 |
ENST00000239449.7
ENST00000624896.1 ENST00000624396.1 |
PCDHB14
ENSG00000279983.1
|
protocadherin beta 14 novel protein |
| chr13_-_31161927 | 0.69 |
ENST00000380405.7
|
HSPH1
|
heat shock protein family H (Hsp110) member 1 |
| chr9_+_72149351 | 0.64 |
ENST00000238018.8
|
GDA
|
guanine deaminase |
| chr13_-_31161890 | 0.64 |
ENST00000320027.10
|
HSPH1
|
heat shock protein family H (Hsp110) member 1 |
| chr2_-_237590660 | 0.64 |
ENST00000409576.1
|
RAB17
|
RAB17, member RAS oncogene family |
| chr11_-_123062022 | 0.62 |
ENST00000532182.5
ENST00000524590.5 ENST00000528292.5 ENST00000533540.5 ENST00000534624.6 ENST00000525463.5 |
HSPA8
|
heat shock protein family A (Hsp70) member 8 |
| chr11_-_123062335 | 0.62 |
ENST00000453788.6
ENST00000527387.5 |
HSPA8
|
heat shock protein family A (Hsp70) member 8 |
| chr1_-_26306576 | 0.62 |
ENST00000421827.2
ENST00000374215.5 ENST00000374223.5 ENST00000357089.8 ENST00000314675.11 ENST00000423664.5 ENST00000374221.7 |
UBXN11
|
UBX domain protein 11 |
| chr20_-_36951701 | 0.61 |
ENST00000646673.2
|
SAMHD1
|
SAM and HD domain containing deoxynucleoside triphosphate triphosphohydrolase 1 |
| chr20_-_36951837 | 0.57 |
ENST00000262878.5
|
SAMHD1
|
SAM and HD domain containing deoxynucleoside triphosphate triphosphohydrolase 1 |
| chr19_-_6720641 | 0.56 |
ENST00000245907.11
|
C3
|
complement C3 |
| chr1_+_26921715 | 0.56 |
ENST00000321265.10
|
NUDC
|
nuclear distribution C, dynein complex regulator |
| chr1_+_212608628 | 0.55 |
ENST00000613954.4
ENST00000341491.9 ENST00000366985.5 |
ATF3
|
activating transcription factor 3 |
| chr20_-_36951637 | 0.54 |
ENST00000646066.1
|
SAMHD1
|
SAM and HD domain containing deoxynucleoside triphosphate triphosphohydrolase 1 |
| chr13_-_31162341 | 0.52 |
ENST00000445273.6
ENST00000630972.2 |
HSPH1
|
heat shock protein family H (Hsp110) member 1 |
| chr16_-_18375069 | 0.52 |
ENST00000545114.5
|
NPIPA9
|
nuclear pore complex interacting protein family, member A9 |
| chr20_-_36951665 | 0.52 |
ENST00000643918.1
|
SAMHD1
|
SAM and HD domain containing deoxynucleoside triphosphate triphosphohydrolase 1 |
| chr15_+_78264552 | 0.51 |
ENST00000394852.8
ENST00000343789.7 |
DNAJA4
|
DnaJ heat shock protein family (Hsp40) member A4 |
| chr9_+_33025265 | 0.51 |
ENST00000330899.5
|
DNAJA1
|
DnaJ heat shock protein family (Hsp40) member A1 |
| chr4_+_76011171 | 0.50 |
ENST00000513353.5
ENST00000341029.9 |
ART3
|
ADP-ribosyltransferase 3 (inactive) |
| chr14_-_102087034 | 0.48 |
ENST00000216281.13
ENST00000553585.5 |
HSP90AA1
|
heat shock protein 90 alpha family class A member 1 |
| chr19_+_1354931 | 0.43 |
ENST00000591337.7
|
PWWP3A
|
PWWP domain containing 3A, DNA repair factor |
| chr16_-_9943182 | 0.43 |
ENST00000535259.6
|
GRIN2A
|
glutamate ionotropic receptor NMDA type subunit 2A |
| chr11_-_90223036 | 0.42 |
ENST00000320585.11
|
CHORDC1
|
cysteine and histidine rich domain containing 1 |
| chr11_-_90223059 | 0.42 |
ENST00000457199.6
ENST00000530765.5 |
CHORDC1
|
cysteine and histidine rich domain containing 1 |
| chr19_-_51390528 | 0.40 |
ENST00000570516.1
ENST00000574814.2 |
C19orf84
|
chromosome 19 open reading frame 84 |
| chr7_-_56051544 | 0.39 |
ENST00000395471.7
|
PSPH
|
phosphoserine phosphatase |
| chr10_-_60572599 | 0.38 |
ENST00000503366.5
|
ANK3
|
ankyrin 3 |
| chr3_+_101724602 | 0.37 |
ENST00000341893.8
|
CEP97
|
centrosomal protein 97 |
| chr7_-_56051288 | 0.37 |
ENST00000419984.6
ENST00000413218.5 ENST00000275605.8 ENST00000424596.1 ENST00000421312.5 ENST00000416592.1 |
PSPH
|
phosphoserine phosphatase |
| chr4_-_25863537 | 0.35 |
ENST00000502949.5
ENST00000264868.9 ENST00000513691.1 ENST00000514872.1 |
SEL1L3
|
SEL1L family member 3 |
| chr15_+_41660397 | 0.30 |
ENST00000219905.12
ENST00000566586.6 |
MGA
|
MAX dimerization protein MGA |
| chr9_-_86100123 | 0.30 |
ENST00000388711.7
ENST00000466178.1 |
GOLM1
|
golgi membrane protein 1 |
| chr11_+_112175501 | 0.29 |
ENST00000357685.11
ENST00000361053.8 |
BCO2
|
beta-carotene oxygenase 2 |
| chr4_+_41359599 | 0.27 |
ENST00000513024.5
|
LIMCH1
|
LIM and calponin homology domains 1 |
| chr5_-_10249876 | 0.27 |
ENST00000511437.6
ENST00000280330.12 ENST00000510047.5 |
ATPSCKMT
|
ATP synthase c subunit lysine N-methyltransferase |
| chr11_+_112175526 | 0.27 |
ENST00000532612.5
ENST00000438022.5 |
BCO2
|
beta-carotene oxygenase 2 |
| chr2_+_85539158 | 0.26 |
ENST00000306434.8
|
MAT2A
|
methionine adenosyltransferase 2A |
| chr9_+_72149424 | 0.26 |
ENST00000358399.8
ENST00000376986.5 |
GDA
|
guanine deaminase |
| chr19_+_859654 | 0.24 |
ENST00000592860.2
ENST00000327726.11 |
CFD
|
complement factor D |
| chr8_+_37736667 | 0.23 |
ENST00000518586.5
ENST00000335171.10 ENST00000521644.5 |
ERLIN2
|
ER lipid raft associated 2 |
| chr10_+_129467178 | 0.22 |
ENST00000306010.8
ENST00000651593.1 |
MGMT
|
O-6-methylguanine-DNA methyltransferase |
| chr9_-_86099506 | 0.22 |
ENST00000388712.7
|
GOLM1
|
golgi membrane protein 1 |
| chr20_-_44960348 | 0.22 |
ENST00000372813.4
|
TOMM34
|
translocase of outer mitochondrial membrane 34 |
| chr6_-_76072654 | 0.21 |
ENST00000369950.8
ENST00000611179.4 ENST00000369963.5 |
IMPG1
|
interphotoreceptor matrix proteoglycan 1 |
| chr1_-_244863085 | 0.21 |
ENST00000440865.2
|
HNRNPU
|
heterogeneous nuclear ribonucleoprotein U |
| chr20_+_18813777 | 0.20 |
ENST00000377428.4
|
SCP2D1
|
SCP2 sterol binding domain containing 1 |
| chr2_-_68952880 | 0.20 |
ENST00000481498.1
ENST00000328895.9 |
GKN2
|
gastrokine 2 |
| chr15_+_43800586 | 0.20 |
ENST00000442995.4
ENST00000458412.2 |
HYPK
|
huntingtin interacting protein K |
| chr6_+_44247866 | 0.20 |
ENST00000371554.2
|
HSP90AB1
|
heat shock protein 90 alpha family class B member 1 |
| chr4_-_158723355 | 0.19 |
ENST00000307720.4
|
PPID
|
peptidylprolyl isomerase D |
| chr11_-_86672419 | 0.19 |
ENST00000524826.7
ENST00000532471.1 |
ME3
|
malic enzyme 3 |
| chr16_-_23557331 | 0.18 |
ENST00000563232.1
ENST00000449606.7 |
EARS2
|
glutamyl-tRNA synthetase 2, mitochondrial |
| chr1_+_44213440 | 0.18 |
ENST00000361745.10
ENST00000446292.5 ENST00000440641.5 ENST00000372289.7 ENST00000436069.5 ENST00000437511.5 |
DMAP1
|
DNA methyltransferase 1 associated protein 1 |
| chr1_+_44213487 | 0.18 |
ENST00000315913.9
|
DMAP1
|
DNA methyltransferase 1 associated protein 1 |
| chr18_+_9136757 | 0.17 |
ENST00000262126.9
ENST00000577992.1 |
ANKRD12
|
ankyrin repeat domain 12 |
| chr8_+_142834801 | 0.17 |
ENST00000220940.2
|
GML
|
glycosylphosphatidylinositol anchored molecule like |
| chr3_+_96814552 | 0.16 |
ENST00000470610.6
ENST00000389672.9 |
EPHA6
|
EPH receptor A6 |
| chr15_+_96325935 | 0.15 |
ENST00000421109.6
|
NR2F2
|
nuclear receptor subfamily 2 group F member 2 |
| chr1_-_156282799 | 0.15 |
ENST00000361813.5
|
SMG5
|
SMG5 nonsense mediated mRNA decay factor |
| chr10_-_20897288 | 0.14 |
ENST00000377122.9
|
NEBL
|
nebulette |
| chr6_+_127577168 | 0.14 |
ENST00000329722.8
|
C6orf58
|
chromosome 6 open reading frame 58 |
| chr21_+_42974510 | 0.14 |
ENST00000432907.6
ENST00000291547.10 |
PKNOX1
|
PBX/knotted 1 homeobox 1 |
| chr18_-_46917432 | 0.14 |
ENST00000324794.11
ENST00000545673.5 ENST00000585916.6 |
PIAS2
|
protein inhibitor of activated STAT 2 |
| chr22_+_29268248 | 0.14 |
ENST00000436425.5
ENST00000447973.5 ENST00000397938.7 ENST00000406548.5 ENST00000437155.6 ENST00000415761.5 ENST00000331029.11 |
EWSR1
|
EWS RNA binding protein 1 |
| chr1_+_149782671 | 0.14 |
ENST00000444948.5
ENST00000369168.5 |
FCGR1A
|
Fc fragment of IgG receptor Ia |
| chr18_+_13465009 | 0.14 |
ENST00000593236.1
ENST00000678400.1 |
LDLRAD4
|
low density lipoprotein receptor class A domain containing 4 |
| chr11_+_193078 | 0.14 |
ENST00000342878.3
|
SCGB1C1
|
secretoglobin family 1C member 1 |
| chr5_+_10250272 | 0.13 |
ENST00000280326.9
ENST00000625723.1 |
CCT5
|
chaperonin containing TCP1 subunit 5 |
| chr1_+_154993581 | 0.13 |
ENST00000392487.1
|
LENEP
|
lens epithelial protein |
| chrX_-_153724343 | 0.13 |
ENST00000442093.5
ENST00000345046.12 ENST00000645377.1 ENST00000672675.1 ENST00000647529.1 ENST00000429550.5 |
BCAP31
|
B cell receptor associated protein 31 |
| chr14_-_23071538 | 0.12 |
ENST00000555566.1
ENST00000338631.10 ENST00000557515.5 ENST00000397341.7 |
ACIN1
|
apoptotic chromatin condensation inducer 1 |
| chr16_-_8861799 | 0.12 |
ENST00000610831.4
ENST00000614449.4 |
CARHSP1
|
calcium regulated heat stable protein 1 |
| chr18_+_59899988 | 0.12 |
ENST00000316660.7
ENST00000269518.9 |
PMAIP1
|
phorbol-12-myristate-13-acetate-induced protein 1 |
| chr5_-_140673568 | 0.12 |
ENST00000542735.2
|
DND1
|
DND microRNA-mediated repression inhibitor 1 |
| chr21_-_39349048 | 0.12 |
ENST00000380748.5
ENST00000380749.10 |
HMGN1
|
high mobility group nucleosome binding domain 1 |
| chr5_+_134371561 | 0.12 |
ENST00000265339.7
ENST00000506787.5 ENST00000507277.1 |
UBE2B
|
ubiquitin conjugating enzyme E2 B |
| chr11_-_111910830 | 0.12 |
ENST00000526167.5
ENST00000651650.1 |
CRYAB
|
crystallin alpha B |
| chr11_-_111910790 | 0.12 |
ENST00000533280.6
|
CRYAB
|
crystallin alpha B |
| chr13_-_30617500 | 0.11 |
ENST00000405805.5
|
HMGB1
|
high mobility group box 1 |
| chr2_-_47906437 | 0.11 |
ENST00000403359.8
|
FBXO11
|
F-box protein 11 |
| chr11_+_64186163 | 0.11 |
ENST00000305218.9
ENST00000538945.5 |
STIP1
|
stress induced phosphoprotein 1 |
| chr5_+_57174016 | 0.11 |
ENST00000514387.6
ENST00000506184.7 |
GPBP1
|
GC-rich promoter binding protein 1 |
| chr11_-_111910888 | 0.11 |
ENST00000525823.1
ENST00000528961.6 |
CRYAB
|
crystallin alpha B |
| chr11_-_47848467 | 0.11 |
ENST00000378460.6
|
NUP160
|
nucleoporin 160 |
| chr3_+_130894050 | 0.11 |
ENST00000510168.6
|
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr17_-_63773534 | 0.11 |
ENST00000403162.7
ENST00000582252.1 ENST00000225726.10 |
CCDC47
|
coiled-coil domain containing 47 |
| chr20_+_19886506 | 0.11 |
ENST00000648440.1
|
RIN2
|
Ras and Rab interactor 2 |
| chr22_+_40857076 | 0.10 |
ENST00000614001.1
ENST00000357137.9 |
XPNPEP3
|
X-prolyl aminopeptidase 3 |
| chr13_-_41019289 | 0.10 |
ENST00000239882.7
|
ELF1
|
E74 like ETS transcription factor 1 |
| chr11_+_10751203 | 0.10 |
ENST00000361367.7
|
CTR9
|
CTR9 homolog, Paf1/RNA polymerase II complex component |
| chr14_+_63204436 | 0.10 |
ENST00000316754.8
|
RHOJ
|
ras homolog family member J |
| chrX_-_103093109 | 0.10 |
ENST00000395065.8
|
NXF3
|
nuclear RNA export factor 3 |
| chrX_-_103688033 | 0.10 |
ENST00000434230.5
ENST00000418819.5 ENST00000360458.5 |
MORF4L2
|
mortality factor 4 like 2 |
| chr11_+_107928394 | 0.10 |
ENST00000320578.3
|
RAB39A
|
RAB39A, member RAS oncogene family |
| chr15_-_85794902 | 0.10 |
ENST00000337975.6
|
KLHL25
|
kelch like family member 25 |
| chr20_+_37521206 | 0.10 |
ENST00000346199.3
ENST00000647955.1 ENST00000649451.1 ENST00000649697.1 ENST00000649309.1 |
NNAT
|
neuronatin |
| chrX_-_155334580 | 0.10 |
ENST00000369449.7
ENST00000321926.4 |
CLIC2
|
chloride intracellular channel 2 |
| chr14_-_22919124 | 0.10 |
ENST00000555209.5
ENST00000554256.5 ENST00000557403.5 ENST00000359890.8 ENST00000557549.5 ENST00000555676.5 ENST00000557571.5 ENST00000557464.5 ENST00000554618.5 ENST00000556862.5 ENST00000555722.5 ENST00000346528.9 ENST00000542016.6 ENST00000399922.6 ENST00000557227.5 |
RBM23
|
RNA binding motif protein 23 |
| chr3_+_45026296 | 0.09 |
ENST00000296130.5
|
CLEC3B
|
C-type lectin domain family 3 member B |
| chr2_+_197500371 | 0.09 |
ENST00000409468.1
ENST00000233893.10 |
HSPE1
|
heat shock protein family E (Hsp10) member 1 |
| chr5_-_134371004 | 0.09 |
ENST00000521755.1
ENST00000523054.5 ENST00000518409.1 |
CDKL3
ENSG00000273345.5
|
cyclin dependent kinase like 3 novel transcript |
| chr18_+_13611764 | 0.09 |
ENST00000585931.5
|
LDLRAD4
|
low density lipoprotein receptor class A domain containing 4 |
| chr7_+_56051756 | 0.09 |
ENST00000275603.9
|
CCT6A
|
chaperonin containing TCP1 subunit 6A |
| chr12_+_48727431 | 0.08 |
ENST00000548380.6
|
TEX49
|
testis expressed 49 |
| chr13_+_30617902 | 0.08 |
ENST00000255304.9
ENST00000614860.1 |
USPL1
|
ubiquitin specific peptidase like 1 |
| chr12_-_48682158 | 0.08 |
ENST00000553086.5
ENST00000548304.1 ENST00000550347.5 ENST00000420613.7 ENST00000550931.5 ENST00000550870.1 |
KANSL2
|
KAT8 regulatory NSL complex subunit 2 |
| chr1_+_196888014 | 0.08 |
ENST00000367416.6
ENST00000608469.6 ENST00000251424.8 ENST00000367418.2 |
CFHR4
|
complement factor H related 4 |
| chr1_+_156338455 | 0.08 |
ENST00000368253.6
ENST00000470342.5 ENST00000368254.6 |
TSACC
|
TSSK6 activating cochaperone |
| chr11_-_27700472 | 0.08 |
ENST00000418212.5
ENST00000533246.5 |
BDNF
|
brain derived neurotrophic factor |
| chr11_-_47848313 | 0.08 |
ENST00000530326.5
ENST00000528071.5 |
NUP160
|
nucleoporin 160 |
| chr2_-_208124514 | 0.07 |
ENST00000264376.5
|
CRYGD
|
crystallin gamma D |
| chr4_-_94342826 | 0.07 |
ENST00000295256.10
|
HPGDS
|
hematopoietic prostaglandin D synthase |
| chr19_+_49851205 | 0.07 |
ENST00000601675.5
|
PTOV1
|
PTOV1 extended AT-hook containing adaptor protein |
| chr6_+_44246166 | 0.07 |
ENST00000620073.4
|
HSP90AB1
|
heat shock protein 90 alpha family class B member 1 |
| chr2_+_202265721 | 0.07 |
ENST00000264279.10
|
NOP58
|
NOP58 ribonucleoprotein |
| chr20_-_23750929 | 0.07 |
ENST00000304749.7
|
CST1
|
cystatin SN |
| chr1_+_65309517 | 0.07 |
ENST00000371069.5
|
DNAJC6
|
DnaJ heat shock protein family (Hsp40) member C6 |
| chr1_-_53238485 | 0.07 |
ENST00000371466.4
ENST00000371470.8 |
MAGOH
|
mago homolog, exon junction complex subunit |
| chr9_-_13279407 | 0.06 |
ENST00000546205.5
|
MPDZ
|
multiple PDZ domain crumbs cell polarity complex component |
| chr20_-_23688951 | 0.06 |
ENST00000217423.4
|
CST4
|
cystatin S |
| chr7_+_56051685 | 0.06 |
ENST00000335503.3
|
CCT6A
|
chaperonin containing TCP1 subunit 6A |
| chr19_+_49851136 | 0.06 |
ENST00000391842.5
|
PTOV1
|
PTOV1 extended AT-hook containing adaptor protein |
| chr19_+_49851173 | 0.06 |
ENST00000599732.5
|
PTOV1
|
PTOV1 extended AT-hook containing adaptor protein |
| chr1_+_121087343 | 0.06 |
ENST00000616817.4
ENST00000623603.3 ENST00000369384.9 ENST00000369383.8 ENST00000369178.5 |
FCGR1B
|
Fc fragment of IgG receptor Ib |
| chr22_+_29268316 | 0.06 |
ENST00000414183.6
ENST00000333395.10 ENST00000455726.5 ENST00000332035.10 |
EWSR1
|
EWS RNA binding protein 1 |
| chr11_+_66002754 | 0.06 |
ENST00000527348.1
|
BANF1
|
BAF nuclear assembly factor 1 |
| chr11_+_76381303 | 0.06 |
ENST00000662483.1
ENST00000663165.1 ENST00000529331.2 ENST00000321844.6 ENST00000667240.1 |
GVQW3
|
GVQW motif containing 3 |
| chr19_+_2269520 | 0.06 |
ENST00000602676.6
ENST00000582888.8 |
OAZ1
|
ornithine decarboxylase antizyme 1 |
| chr11_+_64186219 | 0.05 |
ENST00000543847.1
|
STIP1
|
stress induced phosphoprotein 1 |
| chr4_-_83485065 | 0.05 |
ENST00000515303.2
ENST00000321945.12 ENST00000503217.2 |
ABRAXAS1
|
abraxas 1, BRCA1 A complex subunit |
| chr21_+_43728851 | 0.05 |
ENST00000327574.4
|
PDXK
|
pyridoxal kinase |
| chr2_+_197500398 | 0.05 |
ENST00000604458.1
|
HSPE1-MOB4
|
HSPE1-MOB4 readthrough |
| chr1_+_156338619 | 0.05 |
ENST00000481479.5
ENST00000368252.5 ENST00000466306.5 ENST00000368251.1 |
TSACC
|
TSSK6 activating cochaperone |
| chr18_-_21704763 | 0.05 |
ENST00000580981.5
ENST00000289119.7 |
ABHD3
|
abhydrolase domain containing 3, phospholipase |
| chr12_-_76423256 | 0.05 |
ENST00000546946.5
|
OSBPL8
|
oxysterol binding protein like 8 |
| chrX_-_103688090 | 0.05 |
ENST00000433176.6
|
MORF4L2
|
mortality factor 4 like 2 |
| chr17_+_137569 | 0.05 |
ENST00000595228.4
|
SCGB1C2
|
secretoglobin family 1C member 2 |
| chr11_+_66002225 | 0.05 |
ENST00000445560.6
ENST00000530204.1 |
BANF1
|
BAF nuclear assembly factor 1 |
| chr8_+_37736612 | 0.05 |
ENST00000518526.5
ENST00000523887.5 ENST00000648919.1 ENST00000519638.3 |
ERLIN2
|
ER lipid raft associated 2 |
| chr8_-_27614681 | 0.05 |
ENST00000519472.5
ENST00000523589.5 ENST00000522413.5 ENST00000523396.1 ENST00000316403.15 |
CLU
|
clusterin |
| chr12_-_9733292 | 0.05 |
ENST00000621400.4
ENST00000327839.3 |
CLECL1
|
C-type lectin like 1 |
| chr11_+_67240047 | 0.05 |
ENST00000530342.2
|
KDM2A
|
lysine demethylase 2A |
| chr12_+_65169546 | 0.05 |
ENST00000308330.3
|
LEMD3
|
LEM domain containing 3 |
| chr19_+_52274443 | 0.04 |
ENST00000600821.5
ENST00000595149.5 ENST00000595000.5 ENST00000593612.1 |
ZNF766
|
zinc finger protein 766 |
| chr19_+_41003946 | 0.04 |
ENST00000593831.1
|
CYP2B6
|
cytochrome P450 family 2 subfamily B member 6 |
| chr2_-_68252482 | 0.04 |
ENST00000234310.8
|
PPP3R1
|
protein phosphatase 3 regulatory subunit B, alpha |
| chr3_+_1093002 | 0.04 |
ENST00000446702.7
|
CNTN6
|
contactin 6 |
| chr12_+_110502200 | 0.04 |
ENST00000409246.5
ENST00000392672.8 ENST00000409425.5 |
RAD9B
|
RAD9 checkpoint clamp component B |
| chr20_-_23751256 | 0.04 |
ENST00000398402.1
|
CST1
|
cystatin SN |
| chr14_-_23071617 | 0.04 |
ENST00000357481.6
|
ACIN1
|
apoptotic chromatin condensation inducer 1 |
| chr3_+_133400046 | 0.04 |
ENST00000302334.3
|
BFSP2
|
beaded filament structural protein 2 |
| chr12_+_110502307 | 0.04 |
ENST00000409778.7
ENST00000409300.6 |
RAD9B
|
RAD9 checkpoint clamp component B |
| chr20_+_63658286 | 0.04 |
ENST00000360203.11
ENST00000508582.7 ENST00000318100.9 ENST00000356810.5 |
RTEL1
|
regulator of telomere elongation helicase 1 |
| chr17_+_34319427 | 0.04 |
ENST00000394620.2
|
CCL8
|
C-C motif chemokine ligand 8 |
| chr1_-_228425350 | 0.03 |
ENST00000366696.2
|
H3-4
|
H3.4 histone |
| chr7_+_5879827 | 0.03 |
ENST00000416608.5
|
OCM
|
oncomodulin |
| chr2_-_197499857 | 0.03 |
ENST00000428204.6
ENST00000678170.1 ENST00000676933.1 ENST00000678621.1 |
HSPD1
|
heat shock protein family D (Hsp60) member 1 |
| chr3_+_186783567 | 0.03 |
ENST00000323963.10
ENST00000440191.6 |
EIF4A2
|
eukaryotic translation initiation factor 4A2 |
| chr17_+_35760881 | 0.03 |
ENST00000605587.2
ENST00000604830.1 |
C17orf50
|
chromosome 17 open reading frame 50 |
| chr19_+_6135635 | 0.03 |
ENST00000588304.5
ENST00000588722.5 ENST00000588485.6 ENST00000591403.5 ENST00000586696.5 ENST00000681525.1 ENST00000589401.5 |
ACSBG2
|
acyl-CoA synthetase bubblegum family member 2 |
| chr11_+_66002475 | 0.03 |
ENST00000312175.7
ENST00000533166.5 |
BANF1
|
BAF nuclear assembly factor 1 |
| chr2_-_197499826 | 0.03 |
ENST00000439605.2
ENST00000388968.8 ENST00000418022.2 |
HSPD1
|
heat shock protein family D (Hsp60) member 1 |
| chr17_-_42016680 | 0.03 |
ENST00000674497.1
|
DNAJC7
|
DnaJ heat shock protein family (Hsp40) member C7 |
| chrX_+_109536641 | 0.03 |
ENST00000372107.5
|
NXT2
|
nuclear transport factor 2 like export factor 2 |
| chrX_-_154292117 | 0.03 |
ENST00000610938.1
|
TEX28
|
testis expressed 28 |
| chr16_-_8861713 | 0.03 |
ENST00000567554.5
|
CARHSP1
|
calcium regulated heat stable protein 1 |
| chr15_+_78872881 | 0.02 |
ENST00000559930.5
ENST00000426013.7 |
MORF4L1
|
mortality factor 4 like 1 |
| chr3_+_138347648 | 0.02 |
ENST00000614350.4
ENST00000289104.8 |
MRAS
|
muscle RAS oncogene homolog |
| chr3_-_42410558 | 0.02 |
ENST00000441172.1
ENST00000287748.8 |
LYZL4
|
lysozyme like 4 |
| chr11_-_10568571 | 0.02 |
ENST00000529598.1
|
LYVE1
|
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr11_-_111911759 | 0.02 |
ENST00000650687.2
|
CRYAB
|
crystallin alpha B |
| chrX_+_109536832 | 0.02 |
ENST00000372106.6
|
NXT2
|
nuclear transport factor 2 like export factor 2 |
| chr8_-_88327475 | 0.02 |
ENST00000286614.11
|
MMP16
|
matrix metallopeptidase 16 |
| chr16_+_29778252 | 0.02 |
ENST00000400752.6
|
ZG16
|
zymogen granule protein 16 |
| chr7_+_157336961 | 0.02 |
ENST00000429029.6
|
DNAJB6
|
DnaJ heat shock protein family (Hsp40) member B6 |
| chr8_-_119592954 | 0.02 |
ENST00000522167.5
|
ENPP2
|
ectonucleotide pyrophosphatase/phosphodiesterase 2 |
| chr15_+_78872809 | 0.02 |
ENST00000331268.9
|
MORF4L1
|
mortality factor 4 like 1 |
| chr9_-_83978429 | 0.02 |
ENST00000351839.7
|
HNRNPK
|
heterogeneous nuclear ribonucleoprotein K |
| chr11_-_10568650 | 0.02 |
ENST00000256178.8
|
LYVE1
|
lymphatic vessel endothelial hyaluronan receptor 1 |
| chr1_-_182400660 | 0.02 |
ENST00000367565.2
|
TEDDM1
|
transmembrane epididymal protein 1 |
| chr16_-_8861744 | 0.01 |
ENST00000569398.5
ENST00000568968.1 |
CARHSP1
|
calcium regulated heat stable protein 1 |
| chr19_-_42217667 | 0.01 |
ENST00000336034.8
ENST00000596251.6 ENST00000598200.1 ENST00000598727.5 |
DEDD2
|
death effector domain containing 2 |
| chr15_-_58065870 | 0.01 |
ENST00000537372.5
|
ALDH1A2
|
aldehyde dehydrogenase 1 family member A2 |
| chr8_+_61287950 | 0.01 |
ENST00000519846.5
ENST00000325897.5 ENST00000523868.2 ENST00000518592.5 |
CLVS1
|
clavesin 1 |
| chrX_-_140784366 | 0.01 |
ENST00000674533.1
|
CDR1
|
cerebellar degeneration related protein 1 |
| chr11_+_124622853 | 0.01 |
ENST00000441174.8
ENST00000531667.5 |
TBRG1
|
transforming growth factor beta regulator 1 |
| chr12_+_2794961 | 0.01 |
ENST00000001008.6
|
FKBP4
|
FKBP prolyl isomerase 4 |
| chr9_-_135699473 | 0.01 |
ENST00000425225.2
ENST00000298466.9 |
SOHLH1
|
spermatogenesis and oogenesis specific basic helix-loop-helix 1 |
| chr5_+_446139 | 0.00 |
ENST00000315013.9
|
EXOC3
|
exocyst complex component 3 |
| chr12_+_103587266 | 0.00 |
ENST00000388887.7
|
STAB2
|
stabilin 2 |
| chrX_+_123961304 | 0.00 |
ENST00000371160.5
ENST00000435103.5 |
STAG2
|
stromal antigen 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0006147 | guanine catabolic process(GO:0006147) |
| 0.3 | 0.8 | GO:2000298 | regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000298) |
| 0.3 | 2.2 | GO:0046061 | dGTP catabolic process(GO:0006203) dATP catabolic process(GO:0046061) |
| 0.3 | 1.9 | GO:1903750 | regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903750) negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:1903751) |
| 0.2 | 1.2 | GO:1902904 | negative regulation of fibril organization(GO:1902904) chaperone-mediated autophagy translocation complex disassembly(GO:1904764) |
| 0.2 | 1.7 | GO:0035502 | metanephric part of ureteric bud development(GO:0035502) negative regulation of interleukin-6 biosynthetic process(GO:0045409) glomerular parietal epithelial cell differentiation(GO:0072139) |
| 0.2 | 0.6 | GO:0002415 | immunoglobulin transcytosis in epithelial cells mediated by polymeric immunoglobulin receptor(GO:0002415) |
| 0.2 | 0.6 | GO:0002892 | type IIa hypersensitivity(GO:0001794) regulation of type IIa hypersensitivity(GO:0001796) type II hypersensitivity(GO:0002445) regulation of type II hypersensitivity(GO:0002892) |
| 0.2 | 0.6 | GO:0016116 | tetraterpenoid metabolic process(GO:0016108) carotenoid metabolic process(GO:0016116) carotene catabolic process(GO:0016121) xanthophyll metabolic process(GO:0016122) terpene catabolic process(GO:0046247) |
| 0.2 | 0.6 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.1 | 0.5 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.1 | 0.4 | GO:0072658 | maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.1 | 0.3 | GO:1901389 | regulation of transforming growth factor beta activation(GO:1901388) negative regulation of transforming growth factor beta activation(GO:1901389) |
| 0.1 | 0.4 | GO:0033058 | directional locomotion(GO:0033058) |
| 0.1 | 0.3 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 0.8 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.1 | 0.5 | GO:1903748 | negative regulation of establishment of protein localization to mitochondrion(GO:1903748) |
| 0.0 | 1.4 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.0 | 0.3 | GO:0090206 | negative regulation of cholesterol biosynthetic process(GO:0045541) negative regulation of cholesterol metabolic process(GO:0090206) |
| 0.0 | 0.1 | GO:0010520 | regulation of reciprocal meiotic recombination(GO:0010520) positive regulation of meiosis I(GO:0060903) |
| 0.0 | 0.2 | GO:0009956 | radial pattern formation(GO:0009956) |
| 0.0 | 0.1 | GO:0034729 | histone H3-K79 methylation(GO:0034729) |
| 0.0 | 0.2 | GO:0002424 | T cell mediated immune response to tumor cell(GO:0002424) regulation of T cell mediated immune response to tumor cell(GO:0002840) |
| 0.0 | 0.2 | GO:0070127 | tRNA aminoacylation for mitochondrial protein translation(GO:0070127) |
| 0.0 | 0.1 | GO:0031550 | positive regulation of brain-derived neurotrophic factor receptor signaling pathway(GO:0031550) |
| 0.0 | 0.1 | GO:1904154 | positive regulation of retrograde protein transport, ER to cytosol(GO:1904154) |
| 0.0 | 0.4 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.0 | 0.4 | GO:1902018 | negative regulation of cilium assembly(GO:1902018) |
| 0.0 | 0.1 | GO:0030576 | Cajal body organization(GO:0030576) |
| 0.0 | 0.2 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.0 | 0.2 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 0.0 | 0.1 | GO:2000255 | negative regulation of male germ cell proliferation(GO:2000255) |
| 0.0 | 0.2 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.0 | 0.2 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) |
| 0.0 | 0.5 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.2 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.0 | 0.1 | GO:0009443 | pyridoxal 5'-phosphate salvage(GO:0009443) |
| 0.0 | 0.2 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.0 | 0.5 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 0.0 | 0.1 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.0 | 0.1 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.0 | 0.2 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.0 | 0.2 | GO:1904851 | positive regulation of establishment of protein localization to telomere(GO:1904851) |
| 0.0 | 0.1 | GO:0048254 | snoRNA localization(GO:0048254) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.2 | 1.2 | GO:0098575 | lumenal side of lysosomal membrane(GO:0098575) |
| 0.1 | 0.3 | GO:1990913 | sperm head plasma membrane(GO:1990913) ooplasm(GO:1990917) |
| 0.1 | 0.3 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.1 | 2.0 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 0.5 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.0 | 0.4 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.4 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.0 | 0.1 | GO:0042272 | nuclear RNA export factor complex(GO:0042272) |
| 0.0 | 0.1 | GO:0005784 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.0 | 0.1 | GO:0033503 | HULC complex(GO:0033503) |
| 0.0 | 0.2 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.0 | 0.1 | GO:0001652 | granular component(GO:0001652) |
| 0.0 | 0.4 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.0 | 0.5 | GO:1902562 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 0.2 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.0 | 0.2 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.0 | 0.1 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.2 | GO:0008832 | dGTPase activity(GO:0008832) triphosphoric monoester hydrolase activity(GO:0016793) guanyl deoxyribonucleotide binding(GO:0032560) dGTP binding(GO:0032567) |
| 0.3 | 0.9 | GO:0008892 | guanine deaminase activity(GO:0008892) |
| 0.2 | 1.7 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.2 | 0.8 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.1 | 1.9 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.1 | 0.3 | GO:0002135 | CTP binding(GO:0002135) |
| 0.1 | 0.7 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.1 | 0.5 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.1 | 0.5 | GO:0030911 | TPR domain binding(GO:0030911) |
| 0.0 | 0.4 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.0 | 0.2 | GO:0004473 | malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) |
| 0.0 | 0.1 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 0.0 | 0.2 | GO:0009008 | S-methyltransferase activity(GO:0008172) DNA-methyltransferase activity(GO:0009008) |
| 0.0 | 0.1 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.0 | 0.1 | GO:0004667 | prostaglandin-D synthase activity(GO:0004667) |
| 0.0 | 0.1 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.0 | 0.1 | GO:0052739 | phosphatidylserine 1-acylhydrolase activity(GO:0052739) |
| 0.0 | 0.9 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.0 | 0.1 | GO:0010858 | calcium-dependent protein kinase regulator activity(GO:0010858) |
| 0.0 | 0.1 | GO:0008478 | pyridoxal kinase activity(GO:0008478) |
| 0.0 | 0.2 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 0.1 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.5 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 0.6 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.4 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.0 | 1.8 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.9 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.0 | 0.8 | REACTOME INITIAL TRIGGERING OF COMPLEMENT | Genes involved in Initial triggering of complement |
| 0.0 | 1.2 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.0 | 0.8 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 0.5 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.6 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 0.3 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.0 | 0.4 | REACTOME RAS ACTIVATION UOPN CA2 INFUX THROUGH NMDA RECEPTOR | Genes involved in Ras activation uopn Ca2+ infux through NMDA receptor |
| 0.0 | 0.1 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |