Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
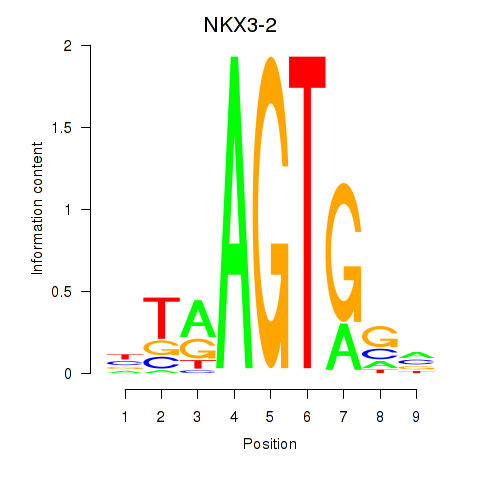
Results for NKX3-2
Z-value: 1.16
Transcription factors associated with NKX3-2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NKX3-2
|
ENSG00000109705.8 | NKX3-2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NKX3-2 | hg38_v1_chr4_-_13544506_13544513 | -0.20 | 2.8e-01 | Click! |
Activity profile of NKX3-2 motif
Sorted Z-values of NKX3-2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NKX3-2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_-_32589833 | 9.47 |
ENST00000360004.5
|
HLA-DRB1
|
major histocompatibility complex, class II, DR beta 1 |
| chr1_-_161367872 | 9.34 |
ENST00000367974.2
|
CFAP126
|
cilia and flagella associated protein 126 |
| chr13_+_42781578 | 7.97 |
ENST00000313851.3
|
FAM216B
|
family with sequence similarity 216 member B |
| chr13_+_42781547 | 7.96 |
ENST00000537894.5
|
FAM216B
|
family with sequence similarity 216 member B |
| chr5_-_150412743 | 7.24 |
ENST00000353334.11
ENST00000009530.12 ENST00000377795.7 |
CD74
|
CD74 molecule |
| chr17_-_76141240 | 6.92 |
ENST00000322957.7
|
FOXJ1
|
forkhead box J1 |
| chr7_-_99971845 | 5.41 |
ENST00000419575.1
|
AZGP1
|
alpha-2-glycoprotein 1, zinc-binding |
| chr9_+_34458752 | 5.22 |
ENST00000614641.4
ENST00000242317.9 ENST00000437363.5 |
DNAI1
|
dynein axonemal intermediate chain 1 |
| chr7_-_99976017 | 4.94 |
ENST00000411734.1
ENST00000292401.9 |
AZGP1
|
alpha-2-glycoprotein 1, zinc-binding |
| chr12_+_49961864 | 4.52 |
ENST00000293599.7
|
AQP5
|
aquaporin 5 |
| chr14_-_106622837 | 4.35 |
ENST00000390628.3
|
IGHV1-58
|
immunoglobulin heavy variable 1-58 |
| chr11_-_119196769 | 4.19 |
ENST00000415318.2
|
CCDC153
|
coiled-coil domain containing 153 |
| chr16_+_57372465 | 3.89 |
ENST00000563383.1
|
CX3CL1
|
C-X3-C motif chemokine ligand 1 |
| chr3_+_186996444 | 3.84 |
ENST00000676633.1
|
ST6GAL1
|
ST6 beta-galactoside alpha-2,6-sialyltransferase 1 |
| chr16_+_57372481 | 3.81 |
ENST00000006053.7
|
CX3CL1
|
C-X3-C motif chemokine ligand 1 |
| chr16_-_21278282 | 3.78 |
ENST00000572914.2
|
CRYM
|
crystallin mu |
| chr7_-_131556602 | 3.77 |
ENST00000322985.9
ENST00000378555.8 |
PODXL
|
podocalyxin like |
| chr5_+_157269317 | 3.69 |
ENST00000618329.4
|
CYFIP2
|
cytoplasmic FMR1 interacting protein 2 |
| chr2_+_98087160 | 3.57 |
ENST00000477737.6
|
VWA3B
|
von Willebrand factor A domain containing 3B |
| chr11_-_8593940 | 3.42 |
ENST00000315204.5
ENST00000396672.5 ENST00000431279.6 ENST00000418597.5 |
STK33
|
serine/threonine kinase 33 |
| chr9_-_110208156 | 3.37 |
ENST00000400613.5
|
C9orf152
|
chromosome 9 open reading frame 152 |
| chr1_-_183653307 | 3.27 |
ENST00000308641.6
|
APOBEC4
|
apolipoprotein B mRNA editing enzyme catalytic polypeptide like 4 |
| chr6_+_32854179 | 3.24 |
ENST00000374859.3
|
PSMB9
|
proteasome 20S subunit beta 9 |
| chr7_-_120858066 | 3.15 |
ENST00000222747.8
|
TSPAN12
|
tetraspanin 12 |
| chr17_+_70075215 | 3.09 |
ENST00000283936.5
ENST00000615244.4 ENST00000392671.6 |
KCNJ16
|
potassium inwardly rectifying channel subfamily J member 16 |
| chr9_-_72953047 | 3.07 |
ENST00000297785.8
ENST00000376939.5 |
ALDH1A1
|
aldehyde dehydrogenase 1 family member A1 |
| chr9_+_68705414 | 2.98 |
ENST00000541509.5
|
PIP5K1B
|
phosphatidylinositol-4-phosphate 5-kinase type 1 beta |
| chr19_+_40991274 | 2.97 |
ENST00000324071.10
|
CYP2B6
|
cytochrome P450 family 2 subfamily B member 6 |
| chr11_-_26572254 | 2.79 |
ENST00000529533.6
|
MUC15
|
mucin 15, cell surface associated |
| chr15_-_43220989 | 2.52 |
ENST00000540029.5
ENST00000441366.7 ENST00000648595.1 |
EPB42
|
erythrocyte membrane protein band 4.2 |
| chr12_-_113221087 | 2.50 |
ENST00000299732.6
ENST00000416617.3 |
IQCD
|
IQ motif containing D |
| chr11_-_26572102 | 2.47 |
ENST00000455601.6
|
MUC15
|
mucin 15, cell surface associated |
| chr6_-_52909666 | 2.46 |
ENST00000370968.5
ENST00000211122.4 |
GSTA3
|
glutathione S-transferase alpha 3 |
| chr1_+_162381703 | 2.21 |
ENST00000458626.4
|
C1orf226
|
chromosome 1 open reading frame 226 |
| chr17_+_70075317 | 2.19 |
ENST00000589377.1
|
KCNJ16
|
potassium inwardly rectifying channel subfamily J member 16 |
| chr1_-_160282458 | 2.19 |
ENST00000485079.1
|
ENSG00000258465.8
|
novel protein |
| chr5_-_55173173 | 2.19 |
ENST00000296733.5
ENST00000322374.10 ENST00000381375.7 |
CDC20B
|
cell division cycle 20B |
| chr3_-_45884685 | 2.18 |
ENST00000684620.1
|
LZTFL1
|
leucine zipper transcription factor like 1 |
| chr5_-_160852200 | 2.17 |
ENST00000327245.10
|
ATP10B
|
ATPase phospholipid transporting 10B (putative) |
| chr10_+_7703300 | 2.11 |
ENST00000358415.9
|
ITIH2
|
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr5_-_176899332 | 2.06 |
ENST00000292432.10
|
HK3
|
hexokinase 3 |
| chr1_+_14924100 | 1.94 |
ENST00000361144.9
|
KAZN
|
kazrin, periplakin interacting protein |
| chr4_-_167234426 | 1.91 |
ENST00000541354.5
ENST00000509854.5 ENST00000512681.5 ENST00000357545.9 ENST00000510741.5 ENST00000510403.5 |
SPOCK3
|
SPARC (osteonectin), cwcv and kazal like domains proteoglycan 3 |
| chr4_-_167234579 | 1.90 |
ENST00000502330.5
ENST00000357154.7 ENST00000421836.6 |
SPOCK3
|
SPARC (osteonectin), cwcv and kazal like domains proteoglycan 3 |
| chr11_-_107018431 | 1.90 |
ENST00000282249.6
|
GUCY1A2
|
guanylate cyclase 1 soluble subunit alpha 2 |
| chr6_-_32763522 | 1.87 |
ENST00000435145.6
ENST00000437316.7 |
HLA-DQB2
|
major histocompatibility complex, class II, DQ beta 2 |
| chr11_-_26572130 | 1.72 |
ENST00000527569.1
|
MUC15
|
mucin 15, cell surface associated |
| chr15_-_93073111 | 1.70 |
ENST00000557420.1
ENST00000542321.6 |
RGMA
|
repulsive guidance molecule BMP co-receptor a |
| chr17_+_60677822 | 1.69 |
ENST00000407086.8
ENST00000589222.5 ENST00000626960.2 ENST00000390652.9 |
BCAS3
|
BCAS3 microtubule associated cell migration factor |
| chr12_+_50925007 | 1.68 |
ENST00000332160.5
|
METTL7A
|
methyltransferase like 7A |
| chr10_-_45672708 | 1.62 |
ENST00000374366.7
ENST00000374371.6 ENST00000344646.10 |
ZFAND4
|
zinc finger AN1-type containing 4 |
| chr15_-_93073706 | 1.62 |
ENST00000425933.6
|
RGMA
|
repulsive guidance molecule BMP co-receptor a |
| chr11_+_36296281 | 1.58 |
ENST00000530639.6
|
PRR5L
|
proline rich 5 like |
| chr7_+_90709816 | 1.56 |
ENST00000436577.3
|
CDK14
|
cyclin dependent kinase 14 |
| chr17_+_55264952 | 1.56 |
ENST00000226067.10
|
HLF
|
HLF transcription factor, PAR bZIP family member |
| chr4_-_167234266 | 1.52 |
ENST00000511269.5
ENST00000506697.5 ENST00000512042.1 |
SPOCK3
|
SPARC (osteonectin), cwcv and kazal like domains proteoglycan 3 |
| chr11_-_59866478 | 1.51 |
ENST00000257264.4
|
TCN1
|
transcobalamin 1 |
| chr7_+_90709530 | 1.44 |
ENST00000406263.5
|
CDK14
|
cyclin dependent kinase 14 |
| chr11_-_5243644 | 1.44 |
ENST00000643122.1
|
HBD
|
hemoglobin subunit delta |
| chr3_-_112499457 | 1.39 |
ENST00000334529.10
|
BTLA
|
B and T lymphocyte associated |
| chr11_-_73142308 | 1.38 |
ENST00000409418.9
|
FCHSD2
|
FCH and double SH3 domains 2 |
| chr2_-_24972032 | 1.34 |
ENST00000534855.5
|
DNAJC27
|
DnaJ heat shock protein family (Hsp40) member C27 |
| chr14_-_23301474 | 1.32 |
ENST00000561437.1
ENST00000559942.5 ENST00000560913.1 ENST00000559314.5 ENST00000558058.5 |
PPP1R3E
|
protein phosphatase 1 regulatory subunit 3E |
| chr16_+_2969548 | 1.29 |
ENST00000572687.1
|
PAQR4
|
progestin and adipoQ receptor family member 4 |
| chr12_-_9999176 | 1.29 |
ENST00000298527.10
ENST00000348658.4 |
CLEC1B
|
C-type lectin domain family 1 member B |
| chr13_+_102656933 | 1.28 |
ENST00000650757.1
|
TPP2
|
tripeptidyl peptidase 2 |
| chr7_-_101321723 | 1.27 |
ENST00000498704.6
ENST00000517481.5 ENST00000437644.2 ENST00000315322.10 ENST00000621899.4 |
IFT22
|
intraflagellar transport 22 |
| chrX_+_48802156 | 1.26 |
ENST00000643374.1
ENST00000644068.1 ENST00000441703.6 ENST00000643934.1 ENST00000489352.5 |
HDAC6
|
histone deacetylase 6 |
| chr1_-_216805367 | 1.24 |
ENST00000360012.7
|
ESRRG
|
estrogen related receptor gamma |
| chr19_-_50909328 | 1.23 |
ENST00000431178.2
|
KLK4
|
kallikrein related peptidase 4 |
| chr14_+_20688756 | 1.22 |
ENST00000397990.5
ENST00000555597.1 |
ANG
RNASE4
|
angiogenin ribonuclease A family member 4 |
| chr18_-_55586092 | 1.21 |
ENST00000563888.6
ENST00000540999.5 ENST00000627685.2 |
TCF4
|
transcription factor 4 |
| chr11_+_66278160 | 1.20 |
ENST00000311445.7
ENST00000528852.5 |
CNIH2
|
cornichon family AMPA receptor auxiliary protein 2 |
| chr3_+_169911566 | 1.20 |
ENST00000428432.6
ENST00000335556.7 |
SAMD7
|
sterile alpha motif domain containing 7 |
| chr3_+_121894379 | 1.19 |
ENST00000489711.6
|
SLC15A2
|
solute carrier family 15 member 2 |
| chr2_-_24971900 | 1.18 |
ENST00000264711.7
|
DNAJC27
|
DnaJ heat shock protein family (Hsp40) member C27 |
| chr1_-_217076889 | 1.17 |
ENST00000493748.5
ENST00000463665.5 |
ESRRG
|
estrogen related receptor gamma |
| chr11_-_8594140 | 1.16 |
ENST00000534493.5
ENST00000422559.6 |
STK33
|
serine/threonine kinase 33 |
| chr10_-_5977492 | 1.16 |
ENST00000530685.5
ENST00000397255.7 ENST00000379971.5 ENST00000528354.5 ENST00000397250.6 ENST00000429135.2 |
IL15RA
|
interleukin 15 receptor subunit alpha |
| chr3_+_179653032 | 1.14 |
ENST00000680587.1
ENST00000681064.1 ENST00000263966.8 ENST00000681358.1 ENST00000679749.1 |
USP13
|
ubiquitin specific peptidase 13 |
| chr19_+_17226662 | 1.10 |
ENST00000598068.5
|
OCEL1
|
occludin/ELL domain containing 1 |
| chr1_-_204494752 | 1.10 |
ENST00000684373.1
|
PIK3C2B
|
phosphatidylinositol-4-phosphate 3-kinase catalytic subunit type 2 beta |
| chr11_+_119107335 | 1.10 |
ENST00000648610.2
ENST00000336702.7 |
C2CD2L
|
C2CD2 like |
| chr1_+_164559173 | 1.08 |
ENST00000420696.7
|
PBX1
|
PBX homeobox 1 |
| chr4_+_48483324 | 1.05 |
ENST00000273861.5
|
SLC10A4
|
solute carrier family 10 member 4 |
| chr9_+_112486819 | 1.05 |
ENST00000337530.11
|
KIAA1958
|
KIAA1958 |
| chr19_+_16888991 | 1.03 |
ENST00000248076.4
|
F2RL3
|
F2R like thrombin or trypsin receptor 3 |
| chr3_+_51943244 | 1.01 |
ENST00000498510.2
|
PARP3
|
poly(ADP-ribose) polymerase family member 3 |
| chr1_+_211259932 | 1.01 |
ENST00000367005.8
|
RCOR3
|
REST corepressor 3 |
| chr4_+_17577487 | 1.01 |
ENST00000606142.5
|
LAP3
|
leucine aminopeptidase 3 |
| chr12_+_56080126 | 1.01 |
ENST00000411731.6
|
ERBB3
|
erb-b2 receptor tyrosine kinase 3 |
| chr9_-_122213874 | 1.00 |
ENST00000482062.1
|
LHX6
|
LIM homeobox 6 |
| chr12_-_22544409 | 0.98 |
ENST00000536386.5
ENST00000396028.6 ENST00000545552.5 ENST00000446597.6 ENST00000333957.8 |
C2CD5
|
C2 calcium dependent domain containing 5 |
| chr21_-_42395943 | 0.97 |
ENST00000398405.5
|
TMPRSS3
|
transmembrane serine protease 3 |
| chr12_-_14929116 | 0.95 |
ENST00000540097.1
|
ERP27
|
endoplasmic reticulum protein 27 |
| chr12_+_56080155 | 0.93 |
ENST00000267101.8
|
ERBB3
|
erb-b2 receptor tyrosine kinase 3 |
| chr9_-_122213903 | 0.93 |
ENST00000464484.3
|
LHX6
|
LIM homeobox 6 |
| chr3_+_132597260 | 0.92 |
ENST00000249887.3
|
ACKR4
|
atypical chemokine receptor 4 |
| chr16_-_66696680 | 0.92 |
ENST00000330687.8
ENST00000563952.1 ENST00000394106.7 |
CMTM4
|
CKLF like MARVEL transmembrane domain containing 4 |
| chr6_-_31730198 | 0.91 |
ENST00000375787.6
|
DDAH2
|
dimethylarginine dimethylaminohydrolase 2 |
| chr10_-_27240505 | 0.91 |
ENST00000375888.5
ENST00000676732.1 |
ACBD5
|
acyl-CoA binding domain containing 5 |
| chr5_+_141430565 | 0.91 |
ENST00000613314.1
|
PCDHGA12
|
protocadherin gamma subfamily A, 12 |
| chr2_-_157874976 | 0.91 |
ENST00000682025.1
ENST00000683487.1 ENST00000682300.1 ENST00000683441.1 ENST00000684595.1 ENST00000683426.1 ENST00000683820.1 ENST00000263640.7 |
ACVR1
|
activin A receptor type 1 |
| chr22_+_31753867 | 0.90 |
ENST00000535622.6
ENST00000645693.1 ENST00000642974.1 ENST00000645711.1 ENST00000644331.1 ENST00000645560.1 ENST00000437411.6 ENST00000433147.2 ENST00000646755.1 ENST00000382112.8 ENST00000647438.1 ENST00000400248.7 ENST00000456178.6 ENST00000646969.1 ENST00000642696.1 ENST00000647343.1 ENST00000400242.8 ENST00000651528.2 ENST00000645015.1 ENST00000645564.1 ENST00000646465.1 ENST00000400249.7 |
DEPDC5
|
DEP domain containing 5, GATOR1 subcomplex subunit |
| chr6_-_31729478 | 0.86 |
ENST00000436437.2
|
DDAH2
|
dimethylarginine dimethylaminohydrolase 2 |
| chr17_+_18476737 | 0.86 |
ENST00000581545.5
ENST00000582333.5 ENST00000328114.11 ENST00000583322.5 ENST00000584941.5 |
LGALS9C
|
galectin 9C |
| chr17_-_42745025 | 0.85 |
ENST00000592492.5
ENST00000585893.5 ENST00000593214.5 ENST00000590078.5 ENST00000428826.7 ENST00000586382.5 ENST00000415827.6 ENST00000592743.5 ENST00000586089.5 |
EZH1
|
enhancer of zeste 1 polycomb repressive complex 2 subunit |
| chr6_-_31729785 | 0.84 |
ENST00000416410.6
|
DDAH2
|
dimethylarginine dimethylaminohydrolase 2 |
| chr6_+_28141830 | 0.84 |
ENST00000330236.7
|
ZKSCAN8
|
zinc finger with KRAB and SCAN domains 8 |
| chr6_-_116060859 | 0.84 |
ENST00000606080.2
|
FRK
|
fyn related Src family tyrosine kinase |
| chr11_+_57741451 | 0.82 |
ENST00000534355.6
|
SELENOH
|
selenoprotein H |
| chr20_-_57265738 | 0.81 |
ENST00000433911.1
|
BMP7
|
bone morphogenetic protein 7 |
| chr12_-_122227491 | 0.79 |
ENST00000475784.1
ENST00000645606.1 |
ENSG00000284934.2
|
novel protein |
| chr5_+_73626158 | 0.78 |
ENST00000296794.10
ENST00000545377.5 ENST00000509848.5 ENST00000513042.7 |
ARHGEF28
|
Rho guanine nucleotide exchange factor 28 |
| chr17_-_39688016 | 0.78 |
ENST00000579146.5
ENST00000300658.9 ENST00000378011.8 ENST00000429199.6 |
PGAP3
|
post-GPI attachment to proteins phospholipase 3 |
| chr8_-_142669947 | 0.77 |
ENST00000612905.2
ENST00000615982.4 ENST00000503272.1 ENST00000571961.7 |
JRK
|
Jrk helix-turn-helix protein |
| chr5_+_56815534 | 0.77 |
ENST00000399503.4
|
MAP3K1
|
mitogen-activated protein kinase kinase kinase 1 |
| chr20_+_54475647 | 0.76 |
ENST00000395939.5
|
DOK5
|
docking protein 5 |
| chr15_+_99105071 | 0.76 |
ENST00000328642.11
ENST00000594047.2 ENST00000336292.11 |
SYNM
|
synemin |
| chr20_+_2295994 | 0.76 |
ENST00000381458.6
|
TGM3
|
transglutaminase 3 |
| chr14_-_87993159 | 0.75 |
ENST00000393568.8
ENST00000261304.7 |
GALC
|
galactosylceramidase |
| chr9_+_113349514 | 0.75 |
ENST00000374183.5
|
BSPRY
|
B-box and SPRY domain containing |
| chr14_-_67600132 | 0.75 |
ENST00000558493.1
ENST00000561272.5 |
PIGH
|
phosphatidylinositol glycan anchor biosynthesis class H |
| chr10_-_27240850 | 0.75 |
ENST00000426079.5
ENST00000677248.1 ENST00000375897.7 |
ACBD5
|
acyl-CoA binding domain containing 5 |
| chr20_+_15196834 | 0.74 |
ENST00000402914.5
|
MACROD2
|
mono-ADP ribosylhydrolase 2 |
| chr20_+_54475584 | 0.74 |
ENST00000262593.10
|
DOK5
|
docking protein 5 |
| chr2_-_24793382 | 0.72 |
ENST00000328379.6
|
PTRHD1
|
peptidyl-tRNA hydrolase domain containing 1 |
| chr2_-_85328262 | 0.72 |
ENST00000282120.6
|
TGOLN2
|
trans-golgi network protein 2 |
| chr16_+_5071806 | 0.70 |
ENST00000684335.1
ENST00000684190.1 ENST00000586840.1 ENST00000262374.10 |
ALG1
|
ALG1 chitobiosyldiphosphodolichol beta-mannosyltransferase |
| chrX_-_72306962 | 0.69 |
ENST00000246139.9
|
CITED1
|
Cbp/p300 interacting transactivator with Glu/Asp rich carboxy-terminal domain 1 |
| chr3_-_112499358 | 0.69 |
ENST00000383680.4
|
BTLA
|
B and T lymphocyte associated |
| chr20_-_2841109 | 0.69 |
ENST00000356872.7
ENST00000439542.1 |
PCED1A
|
PC-esterase domain containing 1A |
| chr14_+_50312311 | 0.69 |
ENST00000426751.7
ENST00000311459.12 ENST00000672419.1 ENST00000672910.1 ENST00000557421.7 ENST00000245448.11 ENST00000554204.7 |
DMAC2L
|
distal membrane arm assembly complex 2 like |
| chr5_+_126423176 | 0.68 |
ENST00000542322.5
ENST00000544396.5 |
GRAMD2B
|
GRAM domain containing 2B |
| chr15_-_43266857 | 0.68 |
ENST00000349114.8
ENST00000220420.10 |
TGM5
|
transglutaminase 5 |
| chr5_+_170353480 | 0.68 |
ENST00000377360.8
|
KCNIP1
|
potassium voltage-gated channel interacting protein 1 |
| chr15_-_37101205 | 0.68 |
ENST00000338564.9
ENST00000558313.5 ENST00000340545.9 |
MEIS2
|
Meis homeobox 2 |
| chr1_+_160127672 | 0.67 |
ENST00000447527.1
|
ATP1A2
|
ATPase Na+/K+ transporting subunit alpha 2 |
| chr20_+_21126074 | 0.66 |
ENST00000619189.5
|
KIZ
|
kizuna centrosomal protein |
| chr12_+_55681647 | 0.66 |
ENST00000614691.1
|
METTL7B
|
methyltransferase like 7B |
| chr3_-_39192584 | 0.65 |
ENST00000340369.4
ENST00000421646.1 ENST00000396251.1 |
XIRP1
|
xin actin binding repeat containing 1 |
| chr19_+_54105923 | 0.65 |
ENST00000420296.1
|
NDUFA3
|
NADH:ubiquinone oxidoreductase subunit A3 |
| chr10_+_96129707 | 0.64 |
ENST00000316045.9
|
ZNF518A
|
zinc finger protein 518A |
| chr12_+_55681711 | 0.63 |
ENST00000394252.4
|
METTL7B
|
methyltransferase like 7B |
| chr18_+_75210755 | 0.63 |
ENST00000322038.5
|
TSHZ1
|
teashirt zinc finger homeobox 1 |
| chr1_-_202889099 | 0.63 |
ENST00000367262.4
|
RABIF
|
RAB interacting factor |
| chr15_+_90352239 | 0.62 |
ENST00000354377.8
ENST00000379090.9 |
ZNF774
|
zinc finger protein 774 |
| chr18_+_26226417 | 0.62 |
ENST00000269142.10
|
TAF4B
|
TATA-box binding protein associated factor 4b |
| chr21_-_5163695 | 0.62 |
ENST00000623998.1
|
ENSG00000280433.1
|
novel protein, similar to trafficking protein particle complex 10 TRAPPC10 |
| chr3_-_51974001 | 0.61 |
ENST00000489595.6
ENST00000461108.5 ENST00000395008.6 ENST00000361143.10 ENST00000525795.1 ENST00000488257.2 |
PCBP4
ABHD14B
ENSG00000272762.5
|
poly(rC) binding protein 4 abhydrolase domain containing 14B novel transcript |
| chr2_+_119223815 | 0.61 |
ENST00000393106.6
ENST00000393110.7 ENST00000409811.5 ENST00000393107.2 |
STEAP3
|
STEAP3 metalloreductase |
| chr1_+_158289916 | 0.60 |
ENST00000368170.8
|
CD1C
|
CD1c molecule |
| chr11_+_34621109 | 0.60 |
ENST00000450654.6
|
EHF
|
ETS homologous factor |
| chr10_+_129467178 | 0.60 |
ENST00000306010.8
ENST00000651593.1 |
MGMT
|
O-6-methylguanine-DNA methyltransferase |
| chrX_+_48801949 | 0.60 |
ENST00000376610.6
ENST00000462730.5 ENST00000376619.6 ENST00000465269.5 ENST00000334136.11 ENST00000476625.5 ENST00000646703.1 |
HDAC6
|
histone deacetylase 6 |
| chr17_+_39688079 | 0.60 |
ENST00000578199.5
|
ERBB2
|
erb-b2 receptor tyrosine kinase 2 |
| chrX_+_22136552 | 0.59 |
ENST00000682888.1
ENST00000684356.1 |
PHEX
|
phosphate regulating endopeptidase homolog X-linked |
| chr2_+_86441341 | 0.58 |
ENST00000312912.10
ENST00000409064.5 |
KDM3A
|
lysine demethylase 3A |
| chr1_+_35268663 | 0.58 |
ENST00000314607.11
|
ZMYM4
|
zinc finger MYM-type containing 4 |
| chrX_+_103628692 | 0.57 |
ENST00000372626.7
|
TCEAL1
|
transcription elongation factor A like 1 |
| chr3_+_52420955 | 0.57 |
ENST00000465863.1
|
PHF7
|
PHD finger protein 7 |
| chr11_+_63938971 | 0.56 |
ENST00000539656.5
ENST00000377793.9 |
NAA40
|
N-alpha-acetyltransferase 40, NatD catalytic subunit |
| chr12_-_122227449 | 0.56 |
ENST00000650715.1
|
DIABLO
|
diablo IAP-binding mitochondrial protein |
| chr20_-_2840623 | 0.56 |
ENST00000360652.7
ENST00000448755.5 |
PCED1A
|
PC-esterase domain containing 1A |
| chr10_-_48251757 | 0.55 |
ENST00000305531.3
|
FRMPD2
|
FERM and PDZ domain containing 2 |
| chr10_-_27240743 | 0.55 |
ENST00000677901.1
ENST00000677960.1 ENST00000677440.1 ENST00000396271.8 ENST00000677141.1 ENST00000677311.1 ENST00000677667.1 ENST00000677200.1 ENST00000676997.1 ENST00000676511.1 |
ACBD5
|
acyl-CoA binding domain containing 5 |
| chr11_-_77820706 | 0.54 |
ENST00000440064.2
ENST00000528095.5 ENST00000308488.11 |
RSF1
|
remodeling and spacing factor 1 |
| chr10_-_125816596 | 0.54 |
ENST00000368786.5
|
UROS
|
uroporphyrinogen III synthase |
| chr11_+_77821125 | 0.54 |
ENST00000526415.5
ENST00000393427.6 ENST00000527134.5 ENST00000304716.12 ENST00000630098.2 |
AAMDC
|
adipogenesis associated Mth938 domain containing |
| chr10_+_100535927 | 0.54 |
ENST00000299163.7
|
HIF1AN
|
hypoxia inducible factor 1 subunit alpha inhibitor |
| chr6_-_47042306 | 0.54 |
ENST00000371253.7
|
ADGRF1
|
adhesion G protein-coupled receptor F1 |
| chr11_-_120138104 | 0.54 |
ENST00000341846.10
|
TRIM29
|
tripartite motif containing 29 |
| chr7_-_128775793 | 0.52 |
ENST00000249389.3
|
OPN1SW
|
opsin 1, short wave sensitive |
| chr19_-_6459735 | 0.51 |
ENST00000334510.9
ENST00000301454.9 |
SLC25A23
|
solute carrier family 25 member 23 |
| chr11_+_77821105 | 0.50 |
ENST00000532481.5
|
AAMDC
|
adipogenesis associated Mth938 domain containing |
| chr5_+_34929524 | 0.50 |
ENST00000648817.1
|
DNAJC21
|
DnaJ heat shock protein family (Hsp40) member C21 |
| chr19_-_44304968 | 0.50 |
ENST00000591609.1
ENST00000589799.5 ENST00000291182.9 ENST00000650576.1 ENST00000589248.5 |
ZNF235
|
zinc finger protein 235 |
| chr11_-_102530738 | 0.50 |
ENST00000260227.5
|
MMP7
|
matrix metallopeptidase 7 |
| chr2_+_96266211 | 0.50 |
ENST00000488633.2
|
CIAO1
|
cytosolic iron-sulfur assembly component 1 |
| chr19_+_840991 | 0.50 |
ENST00000234347.10
|
PRTN3
|
proteinase 3 |
| chr12_+_123973197 | 0.49 |
ENST00000392404.7
ENST00000337815.9 ENST00000538932.6 ENST00000618862.2 ENST00000389727.8 |
ZNF664
ENSG00000274874.2
RFLNA
|
zinc finger protein 664 novel protein refilin A |
| chr3_+_184186023 | 0.49 |
ENST00000429586.6
ENST00000292808.5 |
ABCF3
|
ATP binding cassette subfamily F member 3 |
| chr11_+_77821187 | 0.49 |
ENST00000525409.5
|
AAMDC
|
adipogenesis associated Mth938 domain containing |
| chr2_-_79159747 | 0.48 |
ENST00000409839.7
ENST00000305165.3 ENST00000393878.5 ENST00000490901.2 |
REG3A
|
regenerating family member 3 alpha |
| chr11_+_26188836 | 0.48 |
ENST00000672621.1
|
ANO3
|
anoctamin 3 |
| chr11_+_61203512 | 0.48 |
ENST00000325558.11
|
PGA3
|
pepsinogen A3 |
| chr19_+_40807112 | 0.47 |
ENST00000595621.1
ENST00000595051.1 |
EGLN2
|
egl-9 family hypoxia inducible factor 2 |
| chr2_+_231707650 | 0.47 |
ENST00000409321.5
|
PTMA
|
prothymosin alpha |
| chr20_-_62926469 | 0.47 |
ENST00000354665.8
ENST00000370368.5 ENST00000395340.5 ENST00000395343.6 |
DIDO1
|
death inducer-obliterator 1 |
| chr4_+_88007624 | 0.46 |
ENST00000237596.7
|
PKD2
|
polycystin 2, transient receptor potential cation channel |
| chr19_-_10333512 | 0.46 |
ENST00000617231.5
ENST00000611074.4 ENST00000615032.4 |
RAVER1
|
ribonucleoprotein, PTB binding 1 |
| chr15_-_52652031 | 0.46 |
ENST00000546305.6
|
FAM214A
|
family with sequence similarity 214 member A |
| chr6_+_29306626 | 0.46 |
ENST00000377160.4
|
OR14J1
|
olfactory receptor family 14 subfamily J member 1 |
| chr11_-_128867268 | 0.46 |
ENST00000392665.6
ENST00000392666.6 |
KCNJ1
|
potassium inwardly rectifying channel subfamily J member 1 |
| chr1_+_205257180 | 0.46 |
ENST00000330675.12
|
TMCC2
|
transmembrane and coiled-coil domain family 2 |
| chr22_-_17773976 | 0.46 |
ENST00000317361.11
|
BID
|
BH3 interacting domain death agonist |
| chr7_+_73692596 | 0.45 |
ENST00000453316.1
|
BUD23
|
BUD23 rRNA methyltransferase and ribosome maturation factor |
| chr1_+_107141022 | 0.45 |
ENST00000370067.5
ENST00000370068.6 |
NTNG1
|
netrin G1 |
| chr9_+_112486954 | 0.45 |
ENST00000536272.5
|
KIAA1958
|
KIAA1958 |
| chr13_-_40666600 | 0.44 |
ENST00000379561.6
|
FOXO1
|
forkhead box O1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.6 | 7.7 | GO:0051040 | regulation of calcium-independent cell-cell adhesion(GO:0051040) |
| 2.4 | 7.2 | GO:0035691 | macrophage migration inhibitory factor signaling pathway(GO:0035691) |
| 2.4 | 9.5 | GO:0002399 | MHC class II protein complex assembly(GO:0002399) |
| 1.1 | 4.5 | GO:0015670 | carbon dioxide transport(GO:0015670) |
| 0.9 | 7.4 | GO:0035502 | metanephric part of ureteric bud development(GO:0035502) |
| 0.7 | 2.2 | GO:1903567 | negative regulation of protein localization to cilium(GO:1903565) regulation of protein localization to ciliary membrane(GO:1903567) negative regulation of protein localization to ciliary membrane(GO:1903568) |
| 0.6 | 2.6 | GO:0033634 | positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) |
| 0.6 | 1.9 | GO:0070846 | misfolded protein transport(GO:0070843) polyubiquitinated protein transport(GO:0070844) polyubiquitinated misfolded protein transport(GO:0070845) Hsp90 deacetylation(GO:0070846) |
| 0.5 | 2.5 | GO:0045204 | MAPK export from nucleus(GO:0045204) |
| 0.4 | 3.1 | GO:0006001 | fructose catabolic process(GO:0006001) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) |
| 0.4 | 3.7 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.4 | 1.2 | GO:1902683 | regulation of receptor localization to synapse(GO:1902683) |
| 0.3 | 0.9 | GO:0061445 | endocardial cell fate commitment(GO:0060957) endocardial cushion cell fate commitment(GO:0061445) |
| 0.3 | 3.8 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.3 | 1.1 | GO:0044313 | protein K6-linked deubiquitination(GO:0044313) |
| 0.3 | 0.8 | GO:0010808 | positive regulation of synaptic vesicle priming(GO:0010808) |
| 0.3 | 5.2 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.3 | 1.0 | GO:1990166 | protein localization to site of double-strand break(GO:1990166) |
| 0.3 | 0.8 | GO:0031443 | fast-twitch skeletal muscle fiber contraction(GO:0031443) |
| 0.3 | 0.8 | GO:0006683 | galactosylceramide catabolic process(GO:0006683) |
| 0.2 | 2.2 | GO:0030242 | pexophagy(GO:0030242) |
| 0.2 | 4.9 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.2 | 0.7 | GO:1900920 | regulation of amino acid uptake involved in synaptic transmission(GO:0051941) regulation of glutamate uptake involved in transmission of nerve impulse(GO:0051946) regulation of L-glutamate import(GO:1900920) |
| 0.2 | 8.2 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.2 | 0.8 | GO:1905069 | nephrogenic mesenchyme morphogenesis(GO:0072134) allantois development(GO:1905069) |
| 0.2 | 3.8 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.2 | 0.6 | GO:0046292 | formaldehyde metabolic process(GO:0046292) |
| 0.2 | 2.5 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.2 | 1.2 | GO:0032431 | diacylglycerol biosynthetic process(GO:0006651) activation of phospholipase A2 activity(GO:0032431) |
| 0.2 | 0.5 | GO:0097274 | urea homeostasis(GO:0097274) |
| 0.2 | 1.5 | GO:0051386 | regulation of neurotrophin TRK receptor signaling pathway(GO:0051386) |
| 0.2 | 0.5 | GO:0061182 | negative regulation of chondrocyte development(GO:0061182) |
| 0.2 | 0.5 | GO:0070476 | RNA (guanine-N7)-methylation(GO:0036265) rRNA (guanine-N7)-methylation(GO:0070476) |
| 0.2 | 3.3 | GO:0048681 | negative regulation of axon regeneration(GO:0048681) |
| 0.1 | 0.6 | GO:1904383 | response to sodium phosphate(GO:1904383) |
| 0.1 | 1.9 | GO:0021853 | cerebral cortex GABAergic interneuron migration(GO:0021853) interneuron migration(GO:1904936) |
| 0.1 | 0.5 | GO:0071418 | cellular response to amine stimulus(GO:0071418) |
| 0.1 | 0.4 | GO:0033037 | polysaccharide localization(GO:0033037) |
| 0.1 | 1.5 | GO:0006824 | cobalt ion transport(GO:0006824) |
| 0.1 | 3.0 | GO:0019373 | epoxygenase P450 pathway(GO:0019373) |
| 0.1 | 1.0 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.1 | 1.2 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.1 | 2.6 | GO:0000052 | citrulline metabolic process(GO:0000052) |
| 0.1 | 6.8 | GO:0010107 | potassium ion import(GO:0010107) |
| 0.1 | 0.7 | GO:0071105 | response to interleukin-11(GO:0071105) |
| 0.1 | 0.4 | GO:1902617 | response to fluoride(GO:1902617) |
| 0.1 | 0.5 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) negative regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061428) |
| 0.1 | 0.4 | GO:0016259 | selenocysteine metabolic process(GO:0016259) |
| 0.1 | 0.4 | GO:0009405 | pathogenesis(GO:0009405) |
| 0.1 | 0.6 | GO:0060023 | soft palate development(GO:0060023) |
| 0.1 | 0.3 | GO:0071529 | cementum mineralization(GO:0071529) |
| 0.1 | 1.6 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.1 | 0.7 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.1 | 0.6 | GO:0098706 | ferric iron import into cell(GO:0097461) ferric iron import across plasma membrane(GO:0098706) |
| 0.1 | 0.5 | GO:0097029 | mature conventional dendritic cell differentiation(GO:0097029) |
| 0.1 | 0.2 | GO:0006478 | peptidyl-tyrosine sulfation(GO:0006478) |
| 0.1 | 3.1 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.1 | 0.5 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.1 | 1.8 | GO:0097186 | amelogenesis(GO:0097186) |
| 0.1 | 0.6 | GO:0033088 | negative regulation of immature T cell proliferation in thymus(GO:0033088) |
| 0.1 | 2.5 | GO:1901685 | glutathione derivative metabolic process(GO:1901685) glutathione derivative biosynthetic process(GO:1901687) |
| 0.1 | 0.3 | GO:0048073 | regulation of eye pigmentation(GO:0048073) |
| 0.1 | 0.2 | GO:0002036 | regulation of L-glutamate transport(GO:0002036) |
| 0.1 | 5.2 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 0.8 | GO:0010668 | ectodermal cell differentiation(GO:0010668) |
| 0.1 | 1.0 | GO:0060155 | platelet dense granule organization(GO:0060155) |
| 0.1 | 1.4 | GO:0015669 | gas transport(GO:0015669) oxygen transport(GO:0015671) |
| 0.1 | 1.7 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.1 | 0.6 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 0.1 | 1.9 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.1 | 0.5 | GO:0061591 | calcium activated phospholipid scrambling(GO:0061588) calcium activated phosphatidylcholine scrambling(GO:0061590) calcium activated galactosylceramide scrambling(GO:0061591) |
| 0.1 | 0.3 | GO:0034343 | type III interferon production(GO:0034343) regulation of type III interferon production(GO:0034344) |
| 0.1 | 0.3 | GO:0015853 | adenine transport(GO:0015853) |
| 0.1 | 0.2 | GO:0048867 | stem cell fate determination(GO:0048867) |
| 0.1 | 0.6 | GO:0048007 | antigen processing and presentation of lipid antigen via MHC class Ib(GO:0048003) antigen processing and presentation, exogenous lipid antigen via MHC class Ib(GO:0048007) |
| 0.1 | 0.2 | GO:0051987 | positive regulation of attachment of spindle microtubules to kinetochore(GO:0051987) |
| 0.1 | 0.2 | GO:1902463 | protein localization to cell leading edge(GO:1902463) |
| 0.0 | 0.2 | GO:0051257 | meiotic metaphase I plate congression(GO:0043060) meiotic spindle midzone assembly(GO:0051257) meiotic metaphase plate congression(GO:0051311) |
| 0.0 | 0.9 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.0 | 1.5 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.0 | 0.6 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) |
| 0.0 | 1.1 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 0.1 | GO:0071469 | detection of mechanical stimulus involved in sensory perception of touch(GO:0050976) cellular response to alkaline pH(GO:0071469) |
| 0.0 | 0.6 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.0 | 1.0 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.0 | 0.5 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.0 | 0.2 | GO:0031438 | negative regulation of mRNA cleavage(GO:0031438) negative regulation of mRNA endonucleolytic cleavage involved in unfolded protein response(GO:1904721) |
| 0.0 | 0.8 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 8.8 | GO:0044782 | cilium organization(GO:0044782) |
| 0.0 | 5.1 | GO:1904668 | positive regulation of ubiquitin protein ligase activity(GO:1904668) |
| 0.0 | 1.4 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.0 | 1.3 | GO:0046835 | carbohydrate phosphorylation(GO:0046835) |
| 0.0 | 1.3 | GO:0030220 | platelet formation(GO:0030220) platelet morphogenesis(GO:0036344) |
| 0.0 | 0.3 | GO:0016139 | glycoside catabolic process(GO:0016139) |
| 0.0 | 1.1 | GO:0010971 | positive regulation of G2/M transition of mitotic cell cycle(GO:0010971) |
| 0.0 | 0.5 | GO:0010918 | positive regulation of mitochondrial membrane potential(GO:0010918) |
| 0.0 | 0.2 | GO:0033313 | meiotic cell cycle checkpoint(GO:0033313) |
| 0.0 | 0.3 | GO:2000795 | negative regulation of epithelial cell proliferation involved in lung morphogenesis(GO:2000795) |
| 0.0 | 0.5 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 0.6 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.0 | 0.7 | GO:0070734 | histone H3-K27 methylation(GO:0070734) |
| 0.0 | 0.7 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.0 | 1.1 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.0 | 0.5 | GO:0031163 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.0 | 1.4 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 1.7 | GO:0048384 | retinoic acid receptor signaling pathway(GO:0048384) |
| 0.0 | 2.9 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.0 | 0.8 | GO:0005980 | glycogen catabolic process(GO:0005980) |
| 0.0 | 1.4 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.0 | 0.4 | GO:0044849 | estrous cycle(GO:0044849) |
| 0.0 | 0.1 | GO:0035928 | rRNA import into mitochondrion(GO:0035928) |
| 0.0 | 0.3 | GO:0030050 | vesicle transport along actin filament(GO:0030050) |
| 0.0 | 0.3 | GO:0070070 | proton-transporting V-type ATPase complex assembly(GO:0070070) vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.0 | 0.2 | GO:0090625 | mRNA cleavage involved in gene silencing by siRNA(GO:0090625) |
| 0.0 | 0.6 | GO:0032098 | regulation of appetite(GO:0032098) |
| 0.0 | 0.1 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.0 | 0.2 | GO:0031087 | deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.0 | 0.1 | GO:0006049 | UDP-N-acetylglucosamine catabolic process(GO:0006049) |
| 0.0 | 2.2 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 2.2 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.0 | 0.6 | GO:0050427 | purine ribonucleoside bisphosphate metabolic process(GO:0034035) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) |
| 0.0 | 0.8 | GO:0035774 | positive regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0035774) |
| 0.0 | 0.1 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.0 | 0.3 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.0 | 0.3 | GO:0016246 | RNA interference(GO:0016246) |
| 0.0 | 2.6 | GO:0018210 | peptidyl-threonine phosphorylation(GO:0018107) peptidyl-threonine modification(GO:0018210) |
| 0.0 | 0.4 | GO:0006783 | heme biosynthetic process(GO:0006783) |
| 0.0 | 0.2 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.0 | 0.1 | GO:0045076 | regulation of interleukin-2 biosynthetic process(GO:0045076) |
| 0.0 | 0.8 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.0 | 0.2 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.0 | 0.8 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 18.9 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.5 | 5.2 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.4 | 3.0 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.4 | 3.2 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.3 | 1.1 | GO:0098592 | cytoplasmic side of apical plasma membrane(GO:0098592) |
| 0.3 | 3.8 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.2 | 0.8 | GO:0071817 | MMXD complex(GO:0071817) |
| 0.2 | 1.2 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 0.2 | 0.9 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.1 | 3.0 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.1 | 0.4 | GO:0008623 | CHRAC(GO:0008623) |
| 0.1 | 2.1 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.1 | 0.8 | GO:0044305 | calyx of Held(GO:0044305) |
| 0.1 | 1.4 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.1 | 0.3 | GO:1990876 | cytoplasmic side of nuclear pore(GO:1990876) |
| 0.1 | 0.8 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.1 | 0.5 | GO:0071458 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) |
| 0.1 | 0.2 | GO:0034272 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.1 | 1.3 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 9.5 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.1 | 0.5 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.1 | 0.8 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.1 | 2.2 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 1.6 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 0.3 | GO:0043291 | RAVE complex(GO:0043291) |
| 0.1 | 0.7 | GO:0031010 | ISWI-type complex(GO:0031010) |
| 0.1 | 3.8 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 3.1 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 4.2 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 7.1 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 0.2 | GO:0005715 | late recombination nodule(GO:0005715) |
| 0.0 | 0.5 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.0 | 1.1 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.6 | GO:0097486 | multivesicular body lumen(GO:0097486) |
| 0.0 | 0.4 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.0 | 7.4 | GO:0034705 | potassium channel complex(GO:0034705) |
| 0.0 | 0.3 | GO:0065010 | extracellular membrane-bounded organelle(GO:0065010) |
| 0.0 | 3.7 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.2 | GO:0000221 | vacuolar proton-transporting V-type ATPase, V1 domain(GO:0000221) |
| 0.0 | 0.7 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.0 | 0.6 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.0 | 1.2 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.2 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.0 | 0.7 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 2.2 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) microbody part(GO:0044438) peroxisomal part(GO:0044439) |
| 0.0 | 1.0 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 0.7 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 0.6 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.0 | 0.3 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.0 | 0.2 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.0 | 1.6 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 0.1 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.0 | 0.1 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.0 | 3.3 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.3 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.0 | 0.1 | GO:0031595 | nuclear proteasome complex(GO:0031595) |
| 0.0 | 2.1 | GO:1904813 | ficolin-1-rich granule(GO:0101002) ficolin-1-rich granule lumen(GO:1904813) |
| 0.0 | 0.2 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.0 | 0.6 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.2 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.0 | 0.1 | GO:0005958 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) |
| 0.0 | 0.4 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.4 | GO:0035580 | specific granule lumen(GO:0035580) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 7.2 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 1.0 | 3.0 | GO:0052810 | 1-phosphatidylinositol-5-kinase activity(GO:0052810) |
| 0.9 | 3.8 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.6 | 3.8 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.6 | 1.7 | GO:0010698 | acetyltransferase activator activity(GO:0010698) |
| 0.5 | 3.1 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.5 | 9.8 | GO:0008320 | protein transmembrane transporter activity(GO:0008320) |
| 0.4 | 4.0 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.4 | 5.2 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.4 | 1.9 | GO:0042903 | tubulin deacetylase activity(GO:0042903) |
| 0.3 | 2.6 | GO:0016403 | dimethylargininase activity(GO:0016403) |
| 0.3 | 9.2 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.3 | 3.3 | GO:1990459 | transferrin receptor binding(GO:1990459) |
| 0.3 | 5.4 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.3 | 2.4 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.3 | 1.3 | GO:0008240 | tripeptidyl-peptidase activity(GO:0008240) |
| 0.3 | 0.8 | GO:0004336 | galactosylceramidase activity(GO:0004336) |
| 0.3 | 4.5 | GO:0015250 | water channel activity(GO:0015250) |
| 0.2 | 2.2 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) |
| 0.2 | 2.2 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.2 | 1.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.2 | 1.2 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 0.2 | 4.9 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.2 | 1.4 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.2 | 1.2 | GO:0015198 | oligopeptide transporter activity(GO:0015198) |
| 0.2 | 7.4 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.2 | 1.1 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 0.2 | 1.9 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.1 | 3.0 | GO:0008391 | arachidonic acid monooxygenase activity(GO:0008391) arachidonic acid epoxygenase activity(GO:0008392) |
| 0.1 | 1.3 | GO:0004340 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.1 | 0.6 | GO:1990189 | peptide-serine-N-acetyltransferase activity(GO:1990189) |
| 0.1 | 0.5 | GO:0004852 | uroporphyrinogen-III synthase activity(GO:0004852) |
| 0.1 | 0.9 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.1 | 1.0 | GO:0003910 | DNA ligase (ATP) activity(GO:0003910) |
| 0.1 | 2.1 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.1 | 3.1 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 0.5 | GO:0048763 | calcium-induced calcium release activity(GO:0048763) |
| 0.1 | 0.8 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.1 | 1.0 | GO:0019826 | oxygen sensor activity(GO:0019826) |
| 0.1 | 3.2 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.1 | 1.1 | GO:0001727 | lipid kinase activity(GO:0001727) 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.1 | 1.1 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.1 | 0.7 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.1 | 1.4 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.1 | 0.6 | GO:0008823 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.1 | 0.4 | GO:0004583 | dolichyl-phosphate-glucose-glycolipid alpha-glucosyltransferase activity(GO:0004583) |
| 0.1 | 0.3 | GO:0015207 | ATP:ADP antiporter activity(GO:0005471) adenine transmembrane transporter activity(GO:0015207) |
| 0.1 | 0.8 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.1 | 0.2 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 0.1 | 0.2 | GO:0102008 | cytosolic dipeptidase activity(GO:0102008) |
| 0.1 | 0.5 | GO:0016435 | rRNA (guanine) methyltransferase activity(GO:0016435) |
| 0.1 | 0.6 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.1 | 0.8 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.1 | 0.6 | GO:0009008 | DNA-methyltransferase activity(GO:0009008) |
| 0.1 | 2.2 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.1 | 1.4 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.1 | 0.6 | GO:0030882 | lipid antigen binding(GO:0030882) endogenous lipid antigen binding(GO:0030883) exogenous lipid antigen binding(GO:0030884) |
| 0.1 | 0.4 | GO:0050659 | N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase activity(GO:0050659) |
| 0.1 | 0.8 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.1 | 0.2 | GO:0050146 | nucleoside phosphotransferase activity(GO:0050146) |
| 0.1 | 3.0 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 0.6 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.1 | 1.2 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.1 | 0.3 | GO:0046624 | sphingolipid transporter activity(GO:0046624) |
| 0.1 | 0.2 | GO:0031780 | corticotropin hormone receptor binding(GO:0031780) type 5 melanocortin receptor binding(GO:0031783) |
| 0.1 | 0.7 | GO:1990239 | steroid hormone binding(GO:1990239) |
| 0.1 | 0.5 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.0 | 0.9 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.0 | 2.9 | GO:0016814 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amidines(GO:0016814) |
| 0.0 | 2.5 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.3 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.0 | 0.7 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.0 | 0.2 | GO:0097677 | STAT family protein binding(GO:0097677) |
| 0.0 | 0.1 | GO:0097604 | temperature-gated cation channel activity(GO:0097604) |
| 0.0 | 0.7 | GO:0050693 | LBD domain binding(GO:0050693) |
| 0.0 | 0.6 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.0 | 0.8 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.0 | 0.5 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.1 | GO:0004906 | interferon-gamma receptor activity(GO:0004906) |
| 0.0 | 0.4 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.0 | 0.2 | GO:0060698 | endoribonuclease inhibitor activity(GO:0060698) |
| 0.0 | 0.9 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.0 | 0.3 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.0 | 0.1 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 1.5 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.0 | 0.6 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.0 | 0.2 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.0 | 0.2 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.0 | 0.9 | GO:0019956 | chemokine binding(GO:0019956) |
| 0.0 | 0.7 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.0 | 1.6 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.0 | 0.2 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.5 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 0.1 | GO:0003827 | alpha-1,3-mannosylglycoprotein 2-beta-N-acetylglucosaminyltransferase activity(GO:0003827) |
| 0.0 | 0.8 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.4 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.0 | 0.5 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 0.2 | GO:0032407 | MutSalpha complex binding(GO:0032407) |
| 0.0 | 0.5 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 0.1 | GO:0004677 | DNA-dependent protein kinase activity(GO:0004677) |
| 0.0 | 0.5 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.5 | GO:0005549 | odorant binding(GO:0005549) |
| 0.0 | 0.1 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 0.0 | 7.3 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.2 | GO:0004887 | thyroid hormone receptor activity(GO:0004887) |
| 0.0 | 0.5 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 3.9 | GO:0008168 | methyltransferase activity(GO:0008168) |
| 0.0 | 0.1 | GO:0004117 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) |
| 0.0 | 0.1 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.7 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 4.4 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 3.0 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.1 | 5.6 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.1 | 1.0 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 3.8 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.1 | 3.7 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.8 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.0 | 2.5 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 6.8 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.5 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.0 | 3.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 0.8 | PID UPA UPAR PATHWAY | Urokinase-type plasminogen activator (uPA) and uPAR-mediated signaling |
| 0.0 | 0.9 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.0 | 0.4 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 0.7 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 1.6 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 1.2 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 0.1 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.0 | 1.1 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 2.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 9.5 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.3 | 10.9 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.3 | 4.5 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.2 | 3.1 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.2 | 3.0 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.1 | 5.4 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.1 | 1.4 | REACTOME INWARDLY RECTIFYING K CHANNELS | Genes involved in Inwardly rectifying K+ channels |
| 0.1 | 7.7 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 4.1 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.1 | 2.6 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 2.0 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 2.8 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 3.3 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 2.9 | REACTOME CDK MEDIATED PHOSPHORYLATION AND REMOVAL OF CDC6 | Genes involved in CDK-mediated phosphorylation and removal of Cdc6 |
| 0.0 | 6.3 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.6 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 0.5 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 1.0 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.0 | 1.0 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 1.1 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 1.1 | REACTOME TRAF6 MEDIATED NFKB ACTIVATION | Genes involved in TRAF6 mediated NF-kB activation |
| 0.0 | 0.8 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 0.6 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.0 | 0.3 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.0 | 0.8 | REACTOME TRANSFERRIN ENDOCYTOSIS AND RECYCLING | Genes involved in Transferrin endocytosis and recycling |
| 0.0 | 1.7 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.5 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 0.6 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.0 | 0.5 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 0.6 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.0 | 0.9 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 1.5 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.2 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.3 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.8 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.2 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.0 | 0.7 | REACTOME COSTIMULATION BY THE CD28 FAMILY | Genes involved in Costimulation by the CD28 family |
| 0.0 | 0.4 | REACTOME PIP3 ACTIVATES AKT SIGNALING | Genes involved in PIP3 activates AKT signaling |
| 0.0 | 0.4 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |