Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
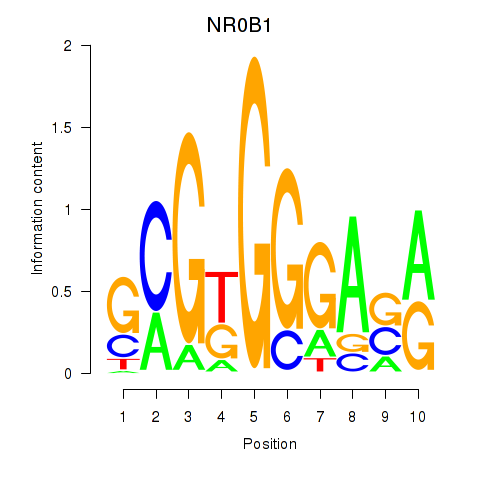
Results for NR0B1
Z-value: 0.57
Transcription factors associated with NR0B1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR0B1
|
ENSG00000169297.8 | NR0B1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR0B1 | hg38_v1_chrX_-_30308333_30308488 | -0.33 | 7.3e-02 | Click! |
Activity profile of NR0B1 motif
Sorted Z-values of NR0B1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR0B1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr13_-_26052009 | 0.70 |
ENST00000319420.4
|
SHISA2
|
shisa family member 2 |
| chr1_-_223364059 | 0.66 |
ENST00000343846.7
ENST00000484758.6 ENST00000344029.6 ENST00000366878.9 ENST00000494793.6 ENST00000681285.1 ENST00000680429.1 ENST00000681669.1 ENST00000681305.1 |
SUSD4
|
sushi domain containing 4 |
| chr7_+_90211686 | 0.65 |
ENST00000287908.7
ENST00000394621.7 ENST00000394626.5 |
STEAP2
|
STEAP2 metalloreductase |
| chr1_-_46132650 | 0.65 |
ENST00000372006.5
ENST00000425892.2 ENST00000420542.5 |
PIK3R3
|
phosphoinositide-3-kinase regulatory subunit 3 |
| chr17_-_7252054 | 0.60 |
ENST00000575783.5
ENST00000573600.5 |
CTDNEP1
|
CTD nuclear envelope phosphatase 1 |
| chr1_-_46132616 | 0.59 |
ENST00000423209.5
ENST00000262741.10 |
PIK3R3
|
phosphoinositide-3-kinase regulatory subunit 3 |
| chr2_+_233636502 | 0.53 |
ENST00000373445.1
|
UGT1A10
|
UDP glucuronosyltransferase family 1 member A10 |
| chr1_+_25819926 | 0.53 |
ENST00000533762.5
ENST00000529116.5 ENST00000474295.5 ENST00000488327.6 ENST00000472643.5 ENST00000374303.7 ENST00000526894.5 ENST00000524618.5 ENST00000374307.9 |
MTFR1L
|
mitochondrial fission regulator 1 like |
| chr1_-_223363337 | 0.46 |
ENST00000608996.5
|
SUSD4
|
sushi domain containing 4 |
| chr1_-_34929574 | 0.44 |
ENST00000373347.6
|
DLGAP3
|
DLG associated protein 3 |
| chr8_-_141000937 | 0.41 |
ENST00000520892.5
|
PTK2
|
protein tyrosine kinase 2 |
| chr7_+_90211830 | 0.40 |
ENST00000394622.6
ENST00000394632.5 ENST00000426158.1 ENST00000402625.6 |
STEAP2
|
STEAP2 metalloreductase |
| chr1_+_25820146 | 0.40 |
ENST00000525713.5
ENST00000374301.7 |
MTFR1L
|
mitochondrial fission regulator 1 like |
| chrX_-_129654946 | 0.40 |
ENST00000429967.3
|
APLN
|
apelin |
| chr10_-_77637558 | 0.39 |
ENST00000372421.10
ENST00000639370.1 ENST00000640773.1 ENST00000638895.1 |
KCNMA1
|
potassium calcium-activated channel subfamily M alpha 1 |
| chr19_-_15934521 | 0.39 |
ENST00000402119.9
|
CYP4F11
|
cytochrome P450 family 4 subfamily F member 11 |
| chr19_-_18606779 | 0.38 |
ENST00000684169.1
ENST00000392386.8 |
CRLF1
|
cytokine receptor like factor 1 |
| chr4_-_151015713 | 0.38 |
ENST00000357115.9
|
LRBA
|
LPS responsive beige-like anchor protein |
| chr10_-_21174187 | 0.38 |
ENST00000417816.2
|
NEBL
|
nebulette |
| chr5_-_138033021 | 0.38 |
ENST00000033079.7
|
FAM13B
|
family with sequence similarity 13 member B |
| chr1_+_18107763 | 0.36 |
ENST00000251296.4
|
IGSF21
|
immunoglobin superfamily member 21 |
| chr17_+_47941562 | 0.36 |
ENST00000225573.5
ENST00000434554.7 ENST00000642017.2 |
PNPO
|
pyridoxamine 5'-phosphate oxidase |
| chr17_-_7251955 | 0.35 |
ENST00000318988.10
|
CTDNEP1
|
CTD nuclear envelope phosphatase 1 |
| chr15_-_52678560 | 0.35 |
ENST00000562351.2
ENST00000261844.11 ENST00000399202.8 ENST00000562135.5 |
FAM214A
|
family with sequence similarity 214 member A |
| chr17_+_57256514 | 0.34 |
ENST00000284073.7
ENST00000674964.1 |
MSI2
|
musashi RNA binding protein 2 |
| chr18_-_5396265 | 0.34 |
ENST00000579951.2
|
EPB41L3
|
erythrocyte membrane protein band 4.1 like 3 |
| chrX_-_112840815 | 0.32 |
ENST00000304758.5
ENST00000371959.9 |
AMOT
|
angiomotin |
| chr2_-_43595963 | 0.32 |
ENST00000405006.8
|
THADA
|
THADA armadillo repeat containing |
| chr1_-_94927079 | 0.31 |
ENST00000370206.9
ENST00000394202.8 |
CNN3
|
calponin 3 |
| chr12_+_56080126 | 0.30 |
ENST00000411731.6
|
ERBB3
|
erb-b2 receptor tyrosine kinase 3 |
| chr3_-_184261547 | 0.30 |
ENST00000296238.4
|
CAMK2N2
|
calcium/calmodulin dependent protein kinase II inhibitor 2 |
| chr15_+_74130551 | 0.30 |
ENST00000453268.3
|
ISLR2
|
immunoglobulin superfamily containing leucine rich repeat 2 |
| chr19_-_15934410 | 0.30 |
ENST00000326742.12
|
CYP4F11
|
cytochrome P450 family 4 subfamily F member 11 |
| chr5_-_42811884 | 0.29 |
ENST00000514985.6
ENST00000511224.5 ENST00000507920.5 ENST00000510965.1 |
SELENOP
|
selenoprotein P |
| chrX_+_131058340 | 0.29 |
ENST00000276211.10
ENST00000370922.5 |
ARHGAP36
|
Rho GTPase activating protein 36 |
| chr15_+_74130243 | 0.29 |
ENST00000561740.5
ENST00000435464.5 |
ISLR2
|
immunoglobulin superfamily containing leucine rich repeat 2 |
| chr7_-_120858303 | 0.29 |
ENST00000415871.5
ENST00000430985.1 |
TSPAN12
|
tetraspanin 12 |
| chr8_+_104223344 | 0.27 |
ENST00000523362.5
|
RIMS2
|
regulating synaptic membrane exocytosis 2 |
| chr7_-_8262668 | 0.26 |
ENST00000446305.1
|
ICA1
|
islet cell autoantigen 1 |
| chr5_+_170353480 | 0.25 |
ENST00000377360.8
|
KCNIP1
|
potassium voltage-gated channel interacting protein 1 |
| chr16_-_800705 | 0.25 |
ENST00000248150.5
|
GNG13
|
G protein subunit gamma 13 |
| chr18_+_46917561 | 0.25 |
ENST00000683218.1
|
KATNAL2
|
katanin catalytic subunit A1 like 2 |
| chr19_+_35031263 | 0.25 |
ENST00000640135.1
ENST00000596348.2 |
SCN1B
|
sodium voltage-gated channel beta subunit 1 |
| chr1_-_23484171 | 0.24 |
ENST00000336689.8
ENST00000437606.6 |
ASAP3
|
ArfGAP with SH3 domain, ankyrin repeat and PH domain 3 |
| chr6_-_11044275 | 0.24 |
ENST00000354666.4
|
ELOVL2
|
ELOVL fatty acid elongase 2 |
| chr20_-_4015518 | 0.24 |
ENST00000545616.2
ENST00000358395.11 |
RNF24
|
ring finger protein 24 |
| chr2_-_61471062 | 0.24 |
ENST00000398571.7
|
USP34
|
ubiquitin specific peptidase 34 |
| chr2_-_152099155 | 0.24 |
ENST00000637309.1
|
CACNB4
|
calcium voltage-gated channel auxiliary subunit beta 4 |
| chr7_-_100889660 | 0.23 |
ENST00000388761.4
|
UFSP1
|
UFM1 specific peptidase 1 (inactive) |
| chr2_-_43595980 | 0.23 |
ENST00000403856.1
ENST00000404790.5 ENST00000405975.7 |
THADA
|
THADA armadillo repeat containing |
| chr14_-_88554898 | 0.23 |
ENST00000556564.6
|
PTPN21
|
protein tyrosine phosphatase non-receptor type 21 |
| chr15_-_77632869 | 0.23 |
ENST00000355300.7
|
LINGO1
|
leucine rich repeat and Ig domain containing 1 |
| chr11_-_111912871 | 0.22 |
ENST00000528628.5
|
CRYAB
|
crystallin alpha B |
| chr12_+_56080155 | 0.22 |
ENST00000267101.8
|
ERBB3
|
erb-b2 receptor tyrosine kinase 3 |
| chr4_-_151015263 | 0.22 |
ENST00000510413.5
ENST00000507224.5 ENST00000651943.2 |
LRBA
|
LPS responsive beige-like anchor protein |
| chr22_+_22895368 | 0.21 |
ENST00000390321.2
|
IGLC1
|
immunoglobulin lambda constant 1 |
| chr1_-_48472166 | 0.21 |
ENST00000371847.8
ENST00000396199.7 |
SPATA6
|
spermatogenesis associated 6 |
| chr2_-_152099023 | 0.21 |
ENST00000201943.10
ENST00000427385.6 ENST00000539935.7 |
CACNB4
|
calcium voltage-gated channel auxiliary subunit beta 4 |
| chr12_+_56079843 | 0.21 |
ENST00000549282.5
ENST00000549061.5 ENST00000683059.1 |
ERBB3
|
erb-b2 receptor tyrosine kinase 3 |
| chr1_-_92486049 | 0.21 |
ENST00000427103.5
|
GFI1
|
growth factor independent 1 transcriptional repressor |
| chr12_-_124567464 | 0.21 |
ENST00000458234.5
|
NCOR2
|
nuclear receptor corepressor 2 |
| chr2_+_27078598 | 0.21 |
ENST00000380320.9
|
EMILIN1
|
elastin microfibril interfacer 1 |
| chr12_-_27780236 | 0.20 |
ENST00000381273.4
|
MANSC4
|
MANSC domain containing 4 |
| chr10_+_84194621 | 0.20 |
ENST00000332904.7
|
CDHR1
|
cadherin related family member 1 |
| chr2_+_54115437 | 0.20 |
ENST00000303536.8
ENST00000394666.7 |
ACYP2
|
acylphosphatase 2 |
| chr17_+_76868410 | 0.19 |
ENST00000301618.8
|
MGAT5B
|
alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase B |
| chr6_+_36196710 | 0.19 |
ENST00000357641.10
|
BRPF3
|
bromodomain and PHD finger containing 3 |
| chr3_+_188154150 | 0.19 |
ENST00000617246.5
|
LPP
|
LIM domain containing preferred translocation partner in lipoma |
| chr20_-_4015389 | 0.19 |
ENST00000336095.10
|
RNF24
|
ring finger protein 24 |
| chr20_-_37178966 | 0.19 |
ENST00000422138.1
|
MROH8
|
maestro heat like repeat family member 8 |
| chr20_-_3173516 | 0.18 |
ENST00000360342.7
ENST00000645462.1 ENST00000337576.7 |
LZTS3
|
leucine zipper tumor suppressor family member 3 |
| chr6_-_29633171 | 0.18 |
ENST00000377034.9
|
GABBR1
|
gamma-aminobutyric acid type B receptor subunit 1 |
| chr17_-_65560296 | 0.18 |
ENST00000585045.1
ENST00000611991.1 |
AXIN2
|
axin 2 |
| chr11_+_64035925 | 0.18 |
ENST00000682287.1
|
FLRT1
|
fibronectin leucine rich transmembrane protein 1 |
| chr2_-_175005357 | 0.18 |
ENST00000409156.7
ENST00000444573.2 ENST00000409900.9 |
CHN1
|
chimerin 1 |
| chr4_-_174829212 | 0.18 |
ENST00000340217.5
ENST00000274093.8 |
GLRA3
|
glycine receptor alpha 3 |
| chr1_-_40665654 | 0.18 |
ENST00000372684.8
|
RIMS3
|
regulating synaptic membrane exocytosis 3 |
| chr8_+_104223320 | 0.18 |
ENST00000339750.3
|
RIMS2
|
regulating synaptic membrane exocytosis 2 |
| chr2_+_85753984 | 0.18 |
ENST00000306279.4
|
ATOH8
|
atonal bHLH transcription factor 8 |
| chr17_-_3668640 | 0.17 |
ENST00000611779.4
|
TAX1BP3
|
Tax1 binding protein 3 |
| chr17_+_7884783 | 0.17 |
ENST00000380358.9
|
CHD3
|
chromodomain helicase DNA binding protein 3 |
| chr9_+_70043840 | 0.17 |
ENST00000377182.5
|
MAMDC2
|
MAM domain containing 2 |
| chr17_-_7251691 | 0.17 |
ENST00000574322.6
|
CTDNEP1
|
CTD nuclear envelope phosphatase 1 |
| chr17_-_79797030 | 0.17 |
ENST00000269385.9
|
CBX8
|
chromobox 8 |
| chr16_-_788329 | 0.17 |
ENST00000563560.1
ENST00000569601.5 ENST00000565809.5 ENST00000007264.7 ENST00000565377.1 ENST00000567114.5 |
RPUSD1
|
RNA pseudouridine synthase domain containing 1 |
| chr5_+_176365455 | 0.17 |
ENST00000310389.6
|
ARL10
|
ADP ribosylation factor like GTPase 10 |
| chr1_+_153728042 | 0.16 |
ENST00000318967.7
ENST00000435409.6 |
INTS3
|
integrator complex subunit 3 |
| chr8_-_101205561 | 0.16 |
ENST00000519744.5
ENST00000311212.9 ENST00000521272.5 ENST00000519882.5 |
ZNF706
|
zinc finger protein 706 |
| chr12_+_40692413 | 0.16 |
ENST00000551295.7
ENST00000547702.5 ENST00000551424.5 |
CNTN1
|
contactin 1 |
| chr6_-_29633056 | 0.16 |
ENST00000377016.8
|
GABBR1
|
gamma-aminobutyric acid type B receptor subunit 1 |
| chr1_-_48472007 | 0.16 |
ENST00000371843.7
|
SPATA6
|
spermatogenesis associated 6 |
| chr19_-_14979848 | 0.16 |
ENST00000594383.2
|
SLC1A6
|
solute carrier family 1 member 6 |
| chr2_+_54115396 | 0.15 |
ENST00000406041.5
|
ACYP2
|
acylphosphatase 2 |
| chr12_+_12611839 | 0.15 |
ENST00000228865.3
|
CREBL2
|
cAMP responsive element binding protein like 2 |
| chrX_-_53422170 | 0.15 |
ENST00000675504.1
|
SMC1A
|
structural maintenance of chromosomes 1A |
| chr2_+_72917489 | 0.15 |
ENST00000258106.11
|
EMX1
|
empty spiracles homeobox 1 |
| chr4_+_42397473 | 0.15 |
ENST00000319234.5
|
SHISA3
|
shisa family member 3 |
| chr9_+_128340646 | 0.15 |
ENST00000372870.5
|
SLC27A4
|
solute carrier family 27 member 4 |
| chr6_-_31897200 | 0.15 |
ENST00000395728.7
ENST00000375528.8 |
EHMT2
|
euchromatic histone lysine methyltransferase 2 |
| chr17_+_31094880 | 0.15 |
ENST00000487476.5
ENST00000356175.7 |
NF1
|
neurofibromin 1 |
| chr7_-_108456378 | 0.15 |
ENST00000613830.4
ENST00000413765.6 ENST00000379028.8 |
NRCAM
|
neuronal cell adhesion molecule |
| chr15_-_78620964 | 0.14 |
ENST00000326828.6
|
CHRNA3
|
cholinergic receptor nicotinic alpha 3 subunit |
| chr19_-_5978133 | 0.14 |
ENST00000340578.10
ENST00000591736.5 ENST00000587479.2 |
RANBP3
|
RAN binding protein 3 |
| chr15_+_74318553 | 0.14 |
ENST00000558821.5
ENST00000268082.4 |
CCDC33
|
coiled-coil domain containing 33 |
| chr16_-_229398 | 0.14 |
ENST00000430864.5
ENST00000293872.13 ENST00000629543.2 ENST00000337351.8 ENST00000397783.5 |
LUC7L
|
LUC7 like |
| chr15_-_64775574 | 0.14 |
ENST00000300069.5
|
RBPMS2
|
RNA binding protein, mRNA processing factor 2 |
| chr12_-_111599069 | 0.14 |
ENST00000673283.1
|
ATXN2
|
ataxin 2 |
| chr1_+_31413806 | 0.13 |
ENST00000536384.2
|
SERINC2
|
serine incorporator 2 |
| chr3_+_181711915 | 0.13 |
ENST00000325404.3
|
SOX2
|
SRY-box transcription factor 2 |
| chr2_-_152098810 | 0.13 |
ENST00000636442.1
ENST00000638005.1 |
CACNB4
|
calcium voltage-gated channel auxiliary subunit beta 4 |
| chr7_+_149715003 | 0.13 |
ENST00000496259.5
ENST00000319551.12 |
KRBA1
|
KRAB-A domain containing 1 |
| chr14_+_73058591 | 0.13 |
ENST00000525161.5
|
RBM25
|
RNA binding motif protein 25 |
| chr22_+_41381923 | 0.13 |
ENST00000266304.9
|
TEF
|
TEF transcription factor, PAR bZIP family member |
| chr17_-_3668557 | 0.12 |
ENST00000225525.4
|
TAX1BP3
|
Tax1 binding protein 3 |
| chr8_+_99013247 | 0.12 |
ENST00000441350.2
ENST00000357162.7 ENST00000358544.7 |
VPS13B
|
vacuolar protein sorting 13 homolog B |
| chr12_-_116276759 | 0.12 |
ENST00000548743.2
|
MED13L
|
mediator complex subunit 13L |
| chr5_-_74640649 | 0.12 |
ENST00000537006.1
|
ENC1
|
ectodermal-neural cortex 1 |
| chr17_+_66964638 | 0.12 |
ENST00000262138.4
|
CACNG4
|
calcium voltage-gated channel auxiliary subunit gamma 4 |
| chr22_-_31140494 | 0.12 |
ENST00000215885.4
|
PLA2G3
|
phospholipase A2 group III |
| chr15_+_90903288 | 0.12 |
ENST00000559717.6
|
MAN2A2
|
mannosidase alpha class 2A member 2 |
| chr19_-_5978078 | 0.12 |
ENST00000592621.5
ENST00000034275.12 ENST00000591092.5 ENST00000591333.5 ENST00000590623.5 ENST00000439268.6 |
RANBP3
|
RAN binding protein 3 |
| chr14_+_58637934 | 0.12 |
ENST00000395153.8
|
DACT1
|
dishevelled binding antagonist of beta catenin 1 |
| chr3_-_11582330 | 0.12 |
ENST00000451674.6
|
VGLL4
|
vestigial like family member 4 |
| chr11_+_35662739 | 0.12 |
ENST00000299413.7
|
TRIM44
|
tripartite motif containing 44 |
| chr1_+_226062704 | 0.11 |
ENST00000366814.3
ENST00000366815.10 ENST00000655399.1 ENST00000667897.1 |
H3-3A
|
H3.3 histone A |
| chr17_+_44308573 | 0.11 |
ENST00000590941.5
ENST00000225441.11 ENST00000426726.8 |
RUNDC3A
|
RUN domain containing 3A |
| chr5_+_132369691 | 0.11 |
ENST00000245407.8
|
SLC22A5
|
solute carrier family 22 member 5 |
| chr12_-_111599097 | 0.11 |
ENST00000389153.10
ENST00000673449.1 ENST00000644883.1 ENST00000647305.1 ENST00000673557.1 |
ATXN2
|
ataxin 2 |
| chr22_+_17369420 | 0.11 |
ENST00000262608.13
ENST00000342247.10 |
CECR2
|
CECR2 histone acetyl-lysine reader |
| chr17_-_3696133 | 0.11 |
ENST00000225328.10
|
P2RX5
|
purinergic receptor P2X 5 |
| chr19_-_38229714 | 0.11 |
ENST00000416611.5
|
DPF1
|
double PHD fingers 1 |
| chr3_-_116445458 | 0.11 |
ENST00000490035.7
|
LSAMP
|
limbic system associated membrane protein |
| chrX_+_55452119 | 0.11 |
ENST00000342972.3
|
MAGEH1
|
MAGE family member H1 |
| chr19_-_38229654 | 0.11 |
ENST00000412732.5
ENST00000456296.5 |
DPF1
|
double PHD fingers 1 |
| chr4_-_76213589 | 0.11 |
ENST00000638603.1
ENST00000452464.6 |
SCARB2
|
scavenger receptor class B member 2 |
| chr1_-_235866916 | 0.11 |
ENST00000389794.7
|
LYST
|
lysosomal trafficking regulator |
| chr12_-_111599331 | 0.10 |
ENST00000608853.5
|
ATXN2
|
ataxin 2 |
| chr16_+_4795378 | 0.10 |
ENST00000588606.5
|
SMIM22
|
small integral membrane protein 22 |
| chr12_+_7189845 | 0.10 |
ENST00000412720.6
ENST00000396637.7 |
PEX5
|
peroxisomal biogenesis factor 5 |
| chr1_+_32886456 | 0.10 |
ENST00000373467.4
|
HPCA
|
hippocalcin |
| chr15_-_29822077 | 0.10 |
ENST00000677774.1
|
TJP1
|
tight junction protein 1 |
| chr10_-_13707536 | 0.10 |
ENST00000632570.1
ENST00000477221.2 |
FRMD4A
|
FERM domain containing 4A |
| chr16_+_4795357 | 0.10 |
ENST00000586005.6
|
SMIM22
|
small integral membrane protein 22 |
| chr16_-_3443446 | 0.10 |
ENST00000301744.7
|
ZNF597
|
zinc finger protein 597 |
| chr17_+_80101562 | 0.10 |
ENST00000302262.8
ENST00000577106.5 ENST00000390015.7 |
GAA
|
alpha glucosidase |
| chr14_+_73058521 | 0.10 |
ENST00000527432.5
ENST00000531500.5 ENST00000261973.12 ENST00000525321.5 ENST00000526754.5 |
RBM25
|
RNA binding motif protein 25 |
| chr20_+_58891704 | 0.10 |
ENST00000371081.5
ENST00000338783.7 |
GNAS
|
GNAS complex locus |
| chr14_+_103334176 | 0.10 |
ENST00000560338.5
ENST00000560763.5 ENST00000216554.8 |
EIF5
|
eukaryotic translation initiation factor 5 |
| chr20_+_47501929 | 0.10 |
ENST00000371997.3
|
NCOA3
|
nuclear receptor coactivator 3 |
| chr17_-_3696198 | 0.10 |
ENST00000345901.7
|
P2RX5
|
purinergic receptor P2X 5 |
| chr7_-_108456321 | 0.10 |
ENST00000379024.8
ENST00000351718.8 |
NRCAM
|
neuronal cell adhesion molecule |
| chr20_+_37179310 | 0.10 |
ENST00000373632.8
ENST00000237530.11 |
RPN2
|
ribophorin II |
| chr20_-_5113067 | 0.10 |
ENST00000342308.10
ENST00000612323.4 ENST00000202834.11 |
TMEM230
|
transmembrane protein 230 |
| chr20_+_47501875 | 0.10 |
ENST00000371998.8
ENST00000372004.7 |
NCOA3
|
nuclear receptor coactivator 3 |
| chr17_-_83051748 | 0.10 |
ENST00000320865.3
|
B3GNTL1
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase like 1 |
| chr12_-_111599273 | 0.10 |
ENST00000672613.1
ENST00000673436.1 ENST00000643669.2 |
ATXN2
|
ataxin 2 |
| chr12_+_7189675 | 0.10 |
ENST00000675855.1
ENST00000434354.6 ENST00000544456.5 ENST00000545574.5 ENST00000420616.6 |
PEX5
|
peroxisomal biogenesis factor 5 |
| chr17_+_79024243 | 0.09 |
ENST00000311661.4
|
C1QTNF1
|
C1q and TNF related 1 |
| chr12_-_57846686 | 0.09 |
ENST00000548823.1
ENST00000398073.7 |
CTDSP2
|
CTD small phosphatase 2 |
| chr12_-_111599503 | 0.09 |
ENST00000535949.5
ENST00000542287.6 ENST00000616825.4 ENST00000550104.5 |
ATXN2
|
ataxin 2 |
| chr10_-_17454582 | 0.09 |
ENST00000377602.5
|
ST8SIA6
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 6 |
| chr2_-_43995950 | 0.09 |
ENST00000683590.1
ENST00000683623.1 ENST00000682779.1 ENST00000682546.1 ENST00000683220.1 ENST00000683125.1 ENST00000683833.1 ENST00000683989.1 ENST00000682480.1 ENST00000260665.12 ENST00000682308.1 ENST00000682885.1 ENST00000409659.6 ENST00000447246.2 |
LRPPRC
|
leucine rich pentatricopeptide repeat containing |
| chrX_-_136767322 | 0.09 |
ENST00000370620.5
|
ARHGEF6
|
Rac/Cdc42 guanine nucleotide exchange factor 6 |
| chr19_+_41845018 | 0.09 |
ENST00000596827.5
|
DMRTC2
|
DMRT like family C2 |
| chr17_-_3696033 | 0.09 |
ENST00000551178.5
ENST00000552276.5 ENST00000547178.5 |
P2RX5
|
purinergic receptor P2X 5 |
| chr7_-_151632330 | 0.09 |
ENST00000418337.6
|
PRKAG2
|
protein kinase AMP-activated non-catalytic subunit gamma 2 |
| chr17_-_7252482 | 0.09 |
ENST00000572043.5
|
CTDNEP1
|
CTD nuclear envelope phosphatase 1 |
| chr15_-_50765656 | 0.09 |
ENST00000261854.10
|
SPPL2A
|
signal peptide peptidase like 2A |
| chr16_-_68236069 | 0.09 |
ENST00000473183.7
ENST00000565858.5 |
ESRP2
|
epithelial splicing regulatory protein 2 |
| chr19_+_41845029 | 0.09 |
ENST00000269945.8
ENST00000596258.5 |
DMRTC2
|
DMRT like family C2 |
| chr22_-_36507022 | 0.09 |
ENST00000216187.10
ENST00000397224.9 ENST00000423980.1 |
FOXRED2
|
FAD dependent oxidoreductase domain containing 2 |
| chr20_+_37179109 | 0.09 |
ENST00000373622.9
|
RPN2
|
ribophorin II |
| chr20_+_44885679 | 0.09 |
ENST00000353703.9
ENST00000372839.7 ENST00000428262.1 ENST00000445830.1 |
YWHAB
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein beta |
| chr1_-_54406385 | 0.09 |
ENST00000610401.5
|
SSBP3
|
single stranded DNA binding protein 3 |
| chr19_-_50333504 | 0.08 |
ENST00000474951.1
|
KCNC3
|
potassium voltage-gated channel subfamily C member 3 |
| chr11_+_64286420 | 0.08 |
ENST00000313074.7
ENST00000542190.5 ENST00000541952.1 |
GPR137
|
G protein-coupled receptor 137 |
| chr8_-_101205455 | 0.08 |
ENST00000520984.5
|
ZNF706
|
zinc finger protein 706 |
| chr5_+_112976757 | 0.08 |
ENST00000389063.3
|
DCP2
|
decapping mRNA 2 |
| chr2_-_164841410 | 0.08 |
ENST00000342193.8
ENST00000375458.6 |
COBLL1
|
cordon-bleu WH2 repeat protein like 1 |
| chr4_-_170026333 | 0.08 |
ENST00000504999.1
|
MFAP3L
|
microfibril associated protein 3 like |
| chr5_-_56952107 | 0.08 |
ENST00000381226.7
ENST00000381199.8 ENST00000381213.7 |
MIER3
|
MIER family member 3 |
| chr17_-_7929793 | 0.08 |
ENST00000303790.3
|
KCNAB3
|
potassium voltage-gated channel subfamily A regulatory beta subunit 3 |
| chrX_+_106920393 | 0.08 |
ENST00000336803.2
|
CLDN2
|
claudin 2 |
| chr19_+_55339867 | 0.08 |
ENST00000255613.8
|
KMT5C
|
lysine methyltransferase 5C |
| chr17_-_67245165 | 0.08 |
ENST00000580168.5
ENST00000358691.10 |
HELZ
|
helicase with zinc finger |
| chr3_+_178536407 | 0.08 |
ENST00000452583.6
|
KCNMB2
|
potassium calcium-activated channel subfamily M regulatory beta subunit 2 |
| chr1_-_154627576 | 0.08 |
ENST00000648311.1
|
ADAR
|
adenosine deaminase RNA specific |
| chr1_-_32336224 | 0.08 |
ENST00000329421.8
|
MARCKSL1
|
MARCKS like 1 |
| chr11_+_57741451 | 0.08 |
ENST00000534355.6
|
SELENOH
|
selenoprotein H |
| chr1_-_15976070 | 0.08 |
ENST00000537142.5
ENST00000375743.9 |
ZBTB17
|
zinc finger and BTB domain containing 17 |
| chr11_+_65027402 | 0.08 |
ENST00000377244.8
ENST00000534637.5 ENST00000524831.5 |
SNX15
|
sorting nexin 15 |
| chr11_-_64745331 | 0.08 |
ENST00000377489.5
ENST00000354024.7 |
RASGRP2
|
RAS guanyl releasing protein 2 |
| chrX_+_134796758 | 0.08 |
ENST00000414371.6
|
PABIR3
|
PABIR family member 3 |
| chr2_+_233636445 | 0.08 |
ENST00000344644.9
|
UGT1A10
|
UDP glucuronosyltransferase family 1 member A10 |
| chr1_+_2556361 | 0.07 |
ENST00000355716.5
|
TNFRSF14
|
TNF receptor superfamily member 14 |
| chr19_-_49119092 | 0.07 |
ENST00000408991.4
|
C19orf73
|
chromosome 19 open reading frame 73 |
| chr5_+_62412755 | 0.07 |
ENST00000325324.11
|
IPO11
|
importin 11 |
| chr12_-_53321544 | 0.07 |
ENST00000394384.7
ENST00000209873.9 |
AAAS
|
aladin WD repeat nucleoporin |
| chrX_-_153926254 | 0.07 |
ENST00000393721.5
ENST00000370028.7 ENST00000350060.10 |
ARHGAP4
|
Rho GTPase activating protein 4 |
| chr6_+_7541612 | 0.07 |
ENST00000418664.2
|
DSP
|
desmoplakin |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0042361 | menaquinone catabolic process(GO:0042361) vitamin K catabolic process(GO:0042377) |
| 0.2 | 1.1 | GO:0098706 | ferric iron import into cell(GO:0097461) ferric iron import across plasma membrane(GO:0098706) |
| 0.1 | 0.5 | GO:1901895 | negative regulation of calcium-transporting ATPase activity(GO:1901895) |
| 0.1 | 0.4 | GO:0043396 | corticotropin-releasing hormone secretion(GO:0043396) regulation of corticotropin-releasing hormone secretion(GO:0043397) |
| 0.1 | 0.4 | GO:0008615 | pyridoxine metabolic process(GO:0008614) pyridoxine biosynthetic process(GO:0008615) vitamin B6 biosynthetic process(GO:0042819) |
| 0.1 | 0.7 | GO:0051552 | flavone metabolic process(GO:0051552) |
| 0.1 | 0.4 | GO:2000672 | negative regulation of motor neuron apoptotic process(GO:2000672) |
| 0.1 | 0.2 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.1 | 0.2 | GO:0072365 | regulation of cellular ketone metabolic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072365) |
| 0.1 | 0.2 | GO:1900111 | positive regulation of histone H3-K9 dimethylation(GO:1900111) |
| 0.1 | 0.2 | GO:0021896 | forebrain astrocyte differentiation(GO:0021896) forebrain astrocyte development(GO:0021897) |
| 0.1 | 0.2 | GO:0001579 | medium-chain fatty acid transport(GO:0001579) |
| 0.1 | 1.2 | GO:0010867 | positive regulation of triglyceride biosynthetic process(GO:0010867) |
| 0.1 | 0.5 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.0 | 0.2 | GO:0021966 | corticospinal neuron axon guidance(GO:0021966) |
| 0.0 | 0.2 | GO:1901094 | regulation of protein tetramerization(GO:1901090) negative regulation of protein tetramerization(GO:1901091) regulation of protein homotetramerization(GO:1901093) negative regulation of protein homotetramerization(GO:1901094) |
| 0.0 | 0.3 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 1.3 | GO:2001275 | positive regulation of glucose import in response to insulin stimulus(GO:2001275) |
| 0.0 | 0.1 | GO:0033364 | mast cell secretory granule organization(GO:0033364) |
| 0.0 | 0.1 | GO:0002086 | diaphragm contraction(GO:0002086) |
| 0.0 | 0.1 | GO:0000961 | negative regulation of mitochondrial RNA catabolic process(GO:0000961) |
| 0.0 | 0.2 | GO:0035624 | receptor transactivation(GO:0035624) |
| 0.0 | 0.2 | GO:0061181 | regulation of chondrocyte development(GO:0061181) |
| 0.0 | 0.6 | GO:0040037 | negative regulation of fibroblast growth factor receptor signaling pathway(GO:0040037) |
| 0.0 | 0.4 | GO:0034465 | response to carbon monoxide(GO:0034465) |
| 0.0 | 0.3 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.0 | 0.3 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.0 | 0.2 | GO:0070778 | L-aspartate transport(GO:0070778) L-aspartate transmembrane transport(GO:0089712) |
| 0.0 | 0.1 | GO:1904219 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.0 | 0.5 | GO:0010603 | regulation of cytoplasmic mRNA processing body assembly(GO:0010603) |
| 0.0 | 0.2 | GO:0098870 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.0 | 0.1 | GO:0060084 | synaptic transmission involved in micturition(GO:0060084) |
| 0.0 | 0.7 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.0 | 0.2 | GO:0019367 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.2 | GO:0072396 | response to cell cycle checkpoint signaling(GO:0072396) response to DNA integrity checkpoint signaling(GO:0072402) response to DNA damage checkpoint signaling(GO:0072423) |
| 0.0 | 0.1 | GO:1900369 | negative regulation of RNA interference(GO:1900369) |
| 0.0 | 0.3 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.0 | 0.1 | GO:1901350 | cell-cell signaling involved in cell-cell junction organization(GO:1901350) |
| 0.0 | 0.3 | GO:0042074 | cell migration involved in gastrulation(GO:0042074) |
| 0.0 | 0.2 | GO:0060012 | synaptic transmission, glycinergic(GO:0060012) |
| 0.0 | 0.1 | GO:0048619 | embryonic hindgut morphogenesis(GO:0048619) |
| 0.0 | 0.1 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.0 | 0.1 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.1 | GO:0060729 | intestinal epithelial structure maintenance(GO:0060729) |
| 0.0 | 0.3 | GO:1902455 | negative regulation of stem cell population maintenance(GO:1902455) |
| 0.0 | 0.1 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.0 | 0.0 | GO:0060214 | endocardium formation(GO:0060214) |
| 0.0 | 0.3 | GO:0051964 | negative regulation of synapse assembly(GO:0051964) |
| 0.0 | 0.1 | GO:1904000 | positive regulation of eating behavior(GO:1904000) regulation of small intestine smooth muscle contraction(GO:1904347) positive regulation of small intestine smooth muscle contraction(GO:1904349) small intestine smooth muscle contraction(GO:1990770) |
| 0.0 | 0.2 | GO:0031118 | rRNA pseudouridine synthesis(GO:0031118) tRNA pseudouridine synthesis(GO:0031119) |
| 0.0 | 0.1 | GO:0019082 | viral protein processing(GO:0019082) regulation of nerve growth factor production(GO:0032903) negative regulation of nerve growth factor production(GO:0032904) dibasic protein processing(GO:0090472) |
| 0.0 | 0.1 | GO:1905123 | regulation of glucosylceramidase activity(GO:1905123) |
| 0.0 | 0.1 | GO:0046642 | negative regulation of alpha-beta T cell proliferation(GO:0046642) |
| 0.0 | 0.1 | GO:0001714 | endodermal cell fate specification(GO:0001714) |
| 0.0 | 0.1 | GO:1902075 | cellular response to salt(GO:1902075) |
| 0.0 | 0.1 | GO:0021999 | neural plate anterior/posterior regionalization(GO:0021999) |
| 0.0 | 0.4 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.0 | 0.1 | GO:2000860 | positive regulation of mineralocorticoid secretion(GO:2000857) positive regulation of aldosterone secretion(GO:2000860) |
| 0.0 | 0.0 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 0.0 | 0.2 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.0 | 0.0 | GO:1901382 | chorionic trophoblast cell proliferation(GO:0097360) regulation of chorionic trophoblast cell proliferation(GO:1901382) |
| 0.0 | 0.9 | GO:0000266 | mitochondrial fission(GO:0000266) |
| 0.0 | 0.1 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.0 | 0.2 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.0 | 0.3 | GO:2000009 | negative regulation of protein localization to cell surface(GO:2000009) |
| 0.0 | 0.4 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.0 | 0.1 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.0 | 0.0 | GO:1905000 | regulation of membrane repolarization during atrial cardiac muscle cell action potential(GO:1905000) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.2 | GO:0071595 | Nem1-Spo7 phosphatase complex(GO:0071595) |
| 0.1 | 0.4 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.1 | 0.4 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 0.1 | 0.3 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.1 | 0.6 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.0 | 1.3 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.2 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.0 | 1.1 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 0.2 | GO:0030893 | meiotic cohesin complex(GO:0030893) |
| 0.0 | 0.1 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.0 | 0.3 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.0 | 0.3 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.0 | 0.2 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.0 | 0.1 | GO:0030289 | protein phosphatase 4 complex(GO:0030289) |
| 0.0 | 0.2 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 0.9 | GO:0005844 | polysome(GO:0005844) |
| 0.0 | 0.3 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 0.3 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) |
| 0.0 | 0.1 | GO:0044327 | dendritic spine head(GO:0044327) |
| 0.0 | 0.1 | GO:1990031 | pinceau fiber(GO:1990031) |
| 0.0 | 0.1 | GO:0030906 | retromer, cargo-selective complex(GO:0030906) |
| 0.0 | 0.2 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.6 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.2 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:0052851 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.1 | 1.3 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.1 | 0.3 | GO:0008427 | calcium-dependent protein kinase inhibitor activity(GO:0008427) |
| 0.1 | 0.2 | GO:0098640 | integrin binding involved in cell-matrix adhesion(GO:0098640) |
| 0.1 | 0.3 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.1 | 0.7 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.1 | 0.4 | GO:0003998 | acylphosphatase activity(GO:0003998) |
| 0.1 | 0.4 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 0.0 | 0.2 | GO:0086062 | voltage-gated sodium channel activity involved in Purkinje myocyte action potential(GO:0086062) |
| 0.0 | 0.4 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.0 | 0.2 | GO:0016934 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.0 | 0.4 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.0 | 0.2 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 0.2 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 0.0 | 0.2 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.0 | 0.3 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.0 | 0.2 | GO:0036033 | mediator complex binding(GO:0036033) |
| 0.0 | 0.1 | GO:0032450 | maltose alpha-glucosidase activity(GO:0032450) |
| 0.0 | 0.3 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 0.3 | GO:0008430 | selenium binding(GO:0008430) |
| 0.0 | 0.1 | GO:0022850 | serotonin-gated cation channel activity(GO:0022850) |
| 0.0 | 0.2 | GO:0102337 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.1 | GO:0015199 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 0.0 | 0.4 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.0 | 0.1 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.0 | 0.4 | GO:0010181 | FMN binding(GO:0010181) |
| 0.0 | 0.1 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.0 | 0.1 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.0 | 0.4 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 0.1 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 0.0 | 0.1 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.0 | 0.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.7 | GO:0016709 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen(GO:0016709) |
| 0.0 | 0.2 | GO:0004579 | dolichyl-diphosphooligosaccharide-protein glycotransferase activity(GO:0004579) |
| 0.0 | 0.3 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.1 | GO:0003726 | double-stranded RNA adenosine deaminase activity(GO:0003726) |
| 0.0 | 0.1 | GO:0042954 | lipoprotein transporter activity(GO:0042954) |
| 0.0 | 0.1 | GO:0016520 | growth hormone-releasing hormone receptor activity(GO:0016520) |
| 0.0 | 0.2 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.0 | 0.1 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.0 | 0.1 | GO:0001025 | RNA polymerase III transcription factor binding(GO:0001025) |
| 0.0 | 0.1 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.0 | 0.0 | GO:0003941 | L-serine ammonia-lyase activity(GO:0003941) |
| 0.0 | 0.1 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.0 | 0.1 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.0 | 0.1 | GO:0010853 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
| 0.0 | 0.1 | GO:0015924 | mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.0 | 0.1 | GO:0050815 | phosphoserine binding(GO:0050815) |
| 0.0 | 0.7 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.3 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.1 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.0 | 0.1 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.0 | 0.0 | GO:0004739 | pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.0 | 0.2 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.3 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.0 | 0.7 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 0.2 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.7 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.3 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.0 | 0.2 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.0 | 0.5 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |