Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
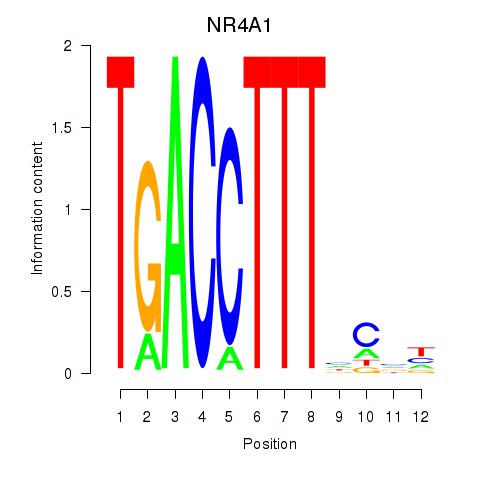
Results for NR4A1
Z-value: 0.64
Transcription factors associated with NR4A1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR4A1
|
ENSG00000123358.20 | NR4A1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR4A1 | hg38_v1_chr12_+_52051402_52051459 | 0.37 | 4.5e-02 | Click! |
Activity profile of NR4A1 motif
Sorted Z-values of NR4A1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR4A1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_+_63652663 | 7.57 |
ENST00000343837.8
ENST00000469440.5 |
SNTN
|
sentan, cilia apical structure protein |
| chr9_-_34397800 | 3.81 |
ENST00000297623.7
|
C9orf24
|
chromosome 9 open reading frame 24 |
| chr11_+_73950985 | 3.27 |
ENST00000339764.6
|
DNAJB13
|
DnaJ heat shock protein family (Hsp40) member B13 |
| chr18_-_27143024 | 2.40 |
ENST00000581714.5
|
CHST9
|
carbohydrate sulfotransferase 9 |
| chr20_+_33168148 | 1.71 |
ENST00000354932.6
|
BPIFA2
|
BPI fold containing family A member 2 |
| chr1_+_212565334 | 1.47 |
ENST00000366981.8
ENST00000366987.6 |
ATF3
|
activating transcription factor 3 |
| chr9_-_72953047 | 1.28 |
ENST00000297785.8
ENST00000376939.5 |
ALDH1A1
|
aldehyde dehydrogenase 1 family member A1 |
| chr1_-_111200633 | 1.27 |
ENST00000357640.9
|
DENND2D
|
DENN domain containing 2D |
| chr1_+_61404076 | 1.12 |
ENST00000357977.5
|
NFIA
|
nuclear factor I A |
| chr3_+_184337591 | 1.09 |
ENST00000383847.7
|
FAM131A
|
family with sequence similarity 131 member A |
| chr3_-_115147237 | 1.02 |
ENST00000357258.8
|
ZBTB20
|
zinc finger and BTB domain containing 20 |
| chr3_-_115147277 | 0.98 |
ENST00000675478.1
|
ZBTB20
|
zinc finger and BTB domain containing 20 |
| chr2_-_24972032 | 0.93 |
ENST00000534855.5
|
DNAJC27
|
DnaJ heat shock protein family (Hsp40) member C27 |
| chrX_-_109625161 | 0.85 |
ENST00000372101.3
|
KCNE5
|
potassium voltage-gated channel subfamily E regulatory subunit 5 |
| chr6_-_152168349 | 0.83 |
ENST00000539504.5
|
SYNE1
|
spectrin repeat containing nuclear envelope protein 1 |
| chr12_+_109139397 | 0.83 |
ENST00000377854.9
ENST00000377848.7 |
ACACB
|
acetyl-CoA carboxylase beta |
| chr5_+_73813518 | 0.82 |
ENST00000296799.8
|
ARHGEF28
|
Rho guanine nucleotide exchange factor 28 |
| chr6_-_152168291 | 0.81 |
ENST00000354674.5
|
SYNE1
|
spectrin repeat containing nuclear envelope protein 1 |
| chr1_+_15736251 | 0.79 |
ENST00000294454.6
|
SLC25A34
|
solute carrier family 25 member 34 |
| chr2_-_24971900 | 0.79 |
ENST00000264711.7
|
DNAJC27
|
DnaJ heat shock protein family (Hsp40) member C27 |
| chr16_+_53207981 | 0.76 |
ENST00000565803.2
|
CHD9
|
chromodomain helicase DNA binding protein 9 |
| chr4_+_70242583 | 0.66 |
ENST00000304954.3
|
CSN3
|
casein kappa |
| chr19_+_13023958 | 0.66 |
ENST00000587760.5
ENST00000585575.5 |
NFIX
|
nuclear factor I X |
| chr10_-_73591330 | 0.66 |
ENST00000451492.5
ENST00000681793.1 ENST00000680396.1 ENST00000413442.5 |
USP54
|
ubiquitin specific peptidase 54 |
| chr1_+_174799895 | 0.65 |
ENST00000489615.5
|
RABGAP1L
|
RAB GTPase activating protein 1 like |
| chr14_-_77028663 | 0.60 |
ENST00000238647.5
|
IRF2BPL
|
interferon regulatory factor 2 binding protein like |
| chr17_+_75633420 | 0.58 |
ENST00000375215.3
|
SMIM5
|
small integral membrane protein 5 |
| chr7_-_105876575 | 0.56 |
ENST00000318724.8
ENST00000419735.8 |
ATXN7L1
|
ataxin 7 like 1 |
| chr6_+_31927486 | 0.55 |
ENST00000442278.6
|
C2
|
complement C2 |
| chr14_-_35121950 | 0.53 |
ENST00000554361.5
ENST00000261475.10 |
PPP2R3C
|
protein phosphatase 2 regulatory subunit B''gamma |
| chr19_+_17305801 | 0.53 |
ENST00000602206.1
ENST00000252602.2 |
MRPL34
|
mitochondrial ribosomal protein L34 |
| chr2_-_177392673 | 0.52 |
ENST00000447413.1
ENST00000397057.6 ENST00000456746.5 ENST00000464747.5 |
ENSG00000213963.6
NFE2L2
|
novel transcript nuclear factor, erythroid 2 like 2 |
| chr7_+_12570712 | 0.50 |
ENST00000417018.1
ENST00000297029.10 |
SCIN
|
scinderin |
| chr2_-_206159410 | 0.49 |
ENST00000457011.5
ENST00000440274.5 ENST00000432169.5 ENST00000233190.11 |
NDUFS1
|
NADH:ubiquinone oxidoreductase core subunit S1 |
| chr3_+_101724602 | 0.49 |
ENST00000341893.8
|
CEP97
|
centrosomal protein 97 |
| chr12_-_16606102 | 0.49 |
ENST00000537304.6
|
LMO3
|
LIM domain only 3 |
| chr12_-_16605939 | 0.47 |
ENST00000541295.5
ENST00000535535.5 |
LMO3
|
LIM domain only 3 |
| chr20_+_13008919 | 0.47 |
ENST00000399002.7
ENST00000434210.5 |
SPTLC3
|
serine palmitoyltransferase long chain base subunit 3 |
| chr7_-_105876477 | 0.47 |
ENST00000478915.1
|
ATXN7L1
|
ataxin 7 like 1 |
| chr12_-_16606795 | 0.46 |
ENST00000447609.5
|
LMO3
|
LIM domain only 3 |
| chr18_-_55302613 | 0.45 |
ENST00000561831.7
|
TCF4
|
transcription factor 4 |
| chr6_+_69232406 | 0.42 |
ENST00000238918.12
|
ADGRB3
|
adhesion G protein-coupled receptor B3 |
| chr19_+_6372757 | 0.41 |
ENST00000245812.8
|
ALKBH7
|
alkB homolog 7 |
| chr6_-_116060859 | 0.41 |
ENST00000606080.2
|
FRK
|
fyn related Src family tyrosine kinase |
| chr20_+_11917859 | 0.39 |
ENST00000618296.4
ENST00000378226.7 |
BTBD3
|
BTB domain containing 3 |
| chr11_+_3855629 | 0.39 |
ENST00000526596.2
ENST00000300737.8 ENST00000616714.4 |
STIM1
|
stromal interaction molecule 1 |
| chr14_-_95157270 | 0.37 |
ENST00000674628.1
ENST00000529720.1 ENST00000393063.6 ENST00000532939.3 |
DICER1
|
dicer 1, ribonuclease III |
| chr10_-_17129786 | 0.36 |
ENST00000377833.10
|
CUBN
|
cubilin |
| chr3_+_138621207 | 0.35 |
ENST00000464668.5
|
FAIM
|
Fas apoptotic inhibitory molecule |
| chr15_+_43692886 | 0.34 |
ENST00000434505.5
ENST00000411750.5 |
CKMT1A
|
creatine kinase, mitochondrial 1A |
| chr6_+_63636481 | 0.34 |
ENST00000509330.5
|
PHF3
|
PHD finger protein 3 |
| chr4_-_73223082 | 0.33 |
ENST00000509867.6
|
ANKRD17
|
ankyrin repeat domain 17 |
| chr3_+_138621225 | 0.32 |
ENST00000479848.1
|
FAIM
|
Fas apoptotic inhibitory molecule |
| chr19_+_49210734 | 0.31 |
ENST00000597316.1
|
TRPM4
|
transient receptor potential cation channel subfamily M member 4 |
| chr7_-_73522278 | 0.30 |
ENST00000404251.1
ENST00000339594.9 |
BAZ1B
|
bromodomain adjacent to zinc finger domain 1B |
| chr2_+_233307806 | 0.30 |
ENST00000447536.5
ENST00000409110.6 |
SAG
|
S-antigen visual arrestin |
| chr1_+_13749408 | 0.29 |
ENST00000343137.8
ENST00000503842.5 ENST00000407521.7 ENST00000505823.5 |
PRDM2
|
PR/SET domain 2 |
| chr20_+_17699942 | 0.29 |
ENST00000427254.1
ENST00000377805.7 |
BANF2
|
BANF family member 2 |
| chr7_-_37448845 | 0.29 |
ENST00000310758.9
|
ELMO1
|
engulfment and cell motility 1 |
| chr6_-_152637351 | 0.29 |
ENST00000367255.10
|
SYNE1
|
spectrin repeat containing nuclear envelope protein 1 |
| chr8_+_122781621 | 0.29 |
ENST00000314393.6
|
ZHX2
|
zinc fingers and homeoboxes 2 |
| chr17_-_7219813 | 0.28 |
ENST00000399510.8
ENST00000648172.8 |
DLG4
|
discs large MAGUK scaffold protein 4 |
| chr10_+_99782628 | 0.28 |
ENST00000648689.1
ENST00000647814.1 |
ABCC2
|
ATP binding cassette subfamily C member 2 |
| chr18_-_5197241 | 0.27 |
ENST00000434239.4
|
AKAIN1
|
A-kinase anchor inhibitor 1 |
| chr18_+_58196736 | 0.27 |
ENST00000675221.1
|
NEDD4L
|
NEDD4 like E3 ubiquitin protein ligase |
| chr3_+_29280945 | 0.27 |
ENST00000434693.6
|
RBMS3
|
RNA binding motif single stranded interacting protein 3 |
| chr10_-_59906509 | 0.26 |
ENST00000263102.7
|
CCDC6
|
coiled-coil domain containing 6 |
| chr17_-_68291116 | 0.25 |
ENST00000327268.8
ENST00000580666.6 |
SLC16A6
|
solute carrier family 16 member 6 |
| chr14_-_95157482 | 0.25 |
ENST00000343455.8
|
DICER1
|
dicer 1, ribonuclease III |
| chr6_+_31927683 | 0.24 |
ENST00000456570.5
|
ENSG00000244255.5
|
novel complement component 2 (C2) and complement factor B (CFB) protein |
| chr4_-_56387422 | 0.24 |
ENST00000513376.5
ENST00000205214.11 ENST00000502617.1 ENST00000451613.5 ENST00000602986.5 |
AASDH
|
aminoadipate-semialdehyde dehydrogenase |
| chr19_-_11529094 | 0.23 |
ENST00000588998.5
ENST00000586149.1 |
ECSIT
|
ECSIT signaling integrator |
| chr20_+_17699960 | 0.23 |
ENST00000246090.6
|
BANF2
|
BANF family member 2 |
| chr20_-_45857196 | 0.23 |
ENST00000457981.5
ENST00000426915.1 ENST00000217455.9 |
ACOT8
|
acyl-CoA thioesterase 8 |
| chr15_-_42491105 | 0.22 |
ENST00000565380.5
ENST00000564754.7 |
ZNF106
|
zinc finger protein 106 |
| chr19_-_11529116 | 0.21 |
ENST00000592312.5
ENST00000590480.1 ENST00000585318.5 ENST00000270517.12 ENST00000252440.11 ENST00000417981.6 |
ECSIT
|
ECSIT signaling integrator |
| chr1_+_228208054 | 0.21 |
ENST00000284548.16
ENST00000422127.5 |
OBSCN
|
obscurin, cytoskeletal calmodulin and titin-interacting RhoGEF |
| chr16_-_81198766 | 0.21 |
ENST00000526632.5
|
PKD1L2
|
polycystin 1 like 2 (gene/pseudogene) |
| chr5_-_74866779 | 0.21 |
ENST00000510496.5
|
FAM169A
|
family with sequence similarity 169 member A |
| chr20_+_38962299 | 0.20 |
ENST00000373325.6
ENST00000373323.8 ENST00000252011.8 ENST00000615559.1 |
DHX35
ENSG00000277581.1
|
DEAH-box helicase 35 novel transcript, sense intronic to DHX35 |
| chr4_-_88823165 | 0.20 |
ENST00000508369.5
|
FAM13A
|
family with sequence similarity 13 member A |
| chr22_-_50578417 | 0.18 |
ENST00000312108.12
ENST00000395650.6 |
CPT1B
|
carnitine palmitoyltransferase 1B |
| chr4_-_88823306 | 0.18 |
ENST00000395002.6
|
FAM13A
|
family with sequence similarity 13 member A |
| chr4_-_86358487 | 0.17 |
ENST00000641066.1
ENST00000512689.6 ENST00000641803.1 ENST00000640970.1 ENST00000641983.1 |
MAPK10
|
mitogen-activated protein kinase 10 |
| chr16_-_31150058 | 0.16 |
ENST00000569305.1
ENST00000268281.9 ENST00000418068.6 |
PRSS36
|
serine protease 36 |
| chr3_+_14947680 | 0.16 |
ENST00000435454.5
ENST00000323373.10 |
NR2C2
|
nuclear receptor subfamily 2 group C member 2 |
| chr1_+_40247926 | 0.15 |
ENST00000372766.4
|
TMCO2
|
transmembrane and coiled-coil domains 2 |
| chr3_-_44477639 | 0.15 |
ENST00000396077.8
|
ZNF445
|
zinc finger protein 445 |
| chr20_+_58891981 | 0.14 |
ENST00000488652.6
ENST00000476935.6 ENST00000492907.6 ENST00000603546.2 |
GNAS
|
GNAS complex locus |
| chr1_+_228208024 | 0.14 |
ENST00000570156.7
ENST00000680850.1 |
OBSCN
|
obscurin, cytoskeletal calmodulin and titin-interacting RhoGEF |
| chr3_+_14947568 | 0.14 |
ENST00000413118.5
ENST00000425241.5 |
NR2C2
|
nuclear receptor subfamily 2 group C member 2 |
| chr4_-_88823214 | 0.13 |
ENST00000513837.5
ENST00000503556.5 |
FAM13A
|
family with sequence similarity 13 member A |
| chr17_+_42998264 | 0.13 |
ENST00000589037.5
|
RPL27
|
ribosomal protein L27 |
| chr15_+_75043263 | 0.13 |
ENST00000563393.1
|
PPCDC
|
phosphopantothenoylcysteine decarboxylase |
| chr2_+_176157293 | 0.12 |
ENST00000683222.1
|
HOXD3
|
homeobox D3 |
| chr9_-_136245802 | 0.12 |
ENST00000358701.10
|
QSOX2
|
quiescin sulfhydryl oxidase 2 |
| chr6_+_31927703 | 0.12 |
ENST00000418949.6
ENST00000299367.10 ENST00000383177.7 ENST00000477310.1 |
C2
ENSG00000244255.5
|
complement C2 novel complement component 2 (C2) and complement factor B (CFB) protein |
| chr16_+_31355165 | 0.12 |
ENST00000562918.5
ENST00000268296.9 |
ITGAX
|
integrin subunit alpha X |
| chr18_+_46174014 | 0.12 |
ENST00000619301.4
ENST00000615052.5 |
C18orf25
|
chromosome 18 open reading frame 25 |
| chr17_+_42998379 | 0.12 |
ENST00000253788.12
ENST00000589913.6 |
RPL27
|
ribosomal protein L27 |
| chr12_+_53985138 | 0.12 |
ENST00000303460.5
|
HOXC10
|
homeobox C10 |
| chr15_+_36594868 | 0.12 |
ENST00000566807.5
ENST00000643612.1 ENST00000567389.5 ENST00000562877.5 |
CDIN1
|
CDAN1 interacting nuclease 1 |
| chr10_-_126670686 | 0.11 |
ENST00000488181.3
|
C10orf90
|
chromosome 10 open reading frame 90 |
| chr8_+_132775340 | 0.11 |
ENST00000622263.4
ENST00000220847.8 ENST00000395379.5 ENST00000395386.7 ENST00000485595.5 ENST00000337920.8 |
PHF20L1
|
PHD finger protein 20 like 1 |
| chr7_-_6009072 | 0.10 |
ENST00000642456.1
ENST00000642292.1 ENST00000441476.6 |
PMS2
|
PMS1 homolog 2, mismatch repair system component |
| chr3_+_113747022 | 0.10 |
ENST00000273398.8
ENST00000496747.5 ENST00000475322.1 |
ATP6V1A
|
ATPase H+ transporting V1 subunit A |
| chr20_-_44187153 | 0.10 |
ENST00000372980.4
|
JPH2
|
junctophilin 2 |
| chr6_-_33803980 | 0.09 |
ENST00000507738.1
ENST00000266003.9 ENST00000430124.7 |
MLN
|
motilin |
| chr20_+_45857607 | 0.08 |
ENST00000255152.3
|
ZSWIM3
|
zinc finger SWIM-type containing 3 |
| chr3_+_184320283 | 0.08 |
ENST00000428387.5
ENST00000434061.6 |
EIF4G1
|
eukaryotic translation initiation factor 4 gamma 1 |
| chr10_+_35195843 | 0.07 |
ENST00000488741.5
ENST00000474931.5 ENST00000468236.5 ENST00000344351.5 ENST00000490511.1 |
CREM
|
cAMP responsive element modulator |
| chr12_+_57941499 | 0.07 |
ENST00000300145.4
|
ATP23
|
ATP23 metallopeptidase and ATP synthase assembly factor homolog |
| chr2_-_43995950 | 0.07 |
ENST00000683590.1
ENST00000683623.1 ENST00000682779.1 ENST00000682546.1 ENST00000683220.1 ENST00000683125.1 ENST00000683833.1 ENST00000683989.1 ENST00000682480.1 ENST00000260665.12 ENST00000682308.1 ENST00000682885.1 ENST00000409659.6 ENST00000447246.2 |
LRPPRC
|
leucine rich pentatricopeptide repeat containing |
| chr17_+_7219857 | 0.06 |
ENST00000583312.5
ENST00000350303.9 ENST00000584103.5 ENST00000356839.10 ENST00000579886.2 |
ACADVL
|
acyl-CoA dehydrogenase very long chain |
| chr3_+_184319677 | 0.06 |
ENST00000441154.5
|
EIF4G1
|
eukaryotic translation initiation factor 4 gamma 1 |
| chr2_-_43995999 | 0.06 |
ENST00000683213.1
ENST00000409946.6 |
LRPPRC
|
leucine rich pentatricopeptide repeat containing |
| chr14_-_95157890 | 0.06 |
ENST00000526495.6
|
DICER1
|
dicer 1, ribonuclease III |
| chr19_-_5622768 | 0.05 |
ENST00000252542.9
|
SAFB2
|
scaffold attachment factor B2 |
| chr17_-_31297231 | 0.05 |
ENST00000247271.5
|
OMG
|
oligodendrocyte myelin glycoprotein |
| chr7_-_6009019 | 0.05 |
ENST00000382321.5
ENST00000265849.12 |
PMS2
|
PMS1 homolog 2, mismatch repair system component |
| chr1_+_165631199 | 0.05 |
ENST00000367888.8
ENST00000367889.8 ENST00000367885.5 ENST00000367884.6 |
MGST3
|
microsomal glutathione S-transferase 3 |
| chr7_+_6009222 | 0.04 |
ENST00000400479.6
ENST00000223029.8 ENST00000395236.2 |
AIMP2
|
aminoacyl tRNA synthetase complex interacting multifunctional protein 2 |
| chr13_-_32538683 | 0.04 |
ENST00000674456.1
ENST00000504114.5 |
N4BP2L2
|
NEDD4 binding protein 2 like 2 |
| chr18_+_46174055 | 0.03 |
ENST00000615553.1
|
C18orf25
|
chromosome 18 open reading frame 25 |
| chr3_-_157503339 | 0.03 |
ENST00000392833.6
|
VEPH1
|
ventricular zone expressed PH domain containing 1 |
| chr3_-_157503375 | 0.03 |
ENST00000362010.7
|
VEPH1
|
ventricular zone expressed PH domain containing 1 |
| chr17_+_42998779 | 0.03 |
ENST00000586277.5
|
RPL27
|
ribosomal protein L27 |
| chr7_+_121328991 | 0.03 |
ENST00000222462.3
|
WNT16
|
Wnt family member 16 |
| chr4_-_40630588 | 0.03 |
ENST00000514014.1
|
RBM47
|
RNA binding motif protein 47 |
| chr17_+_1742836 | 0.02 |
ENST00000324015.7
ENST00000450523.6 ENST00000453723.5 ENST00000453066.6 ENST00000382061.5 |
SERPINF2
|
serpin family F member 2 |
| chr17_-_68291168 | 0.02 |
ENST00000582867.1
|
SLC16A6
|
solute carrier family 16 member 6 |
| chr2_-_27380715 | 0.02 |
ENST00000323703.11
ENST00000436006.1 |
ZNF513
|
zinc finger protein 513 |
| chr11_+_278398 | 0.02 |
ENST00000534750.6
|
NLRP6
|
NLR family pyrin domain containing 6 |
| chr11_+_116829898 | 0.01 |
ENST00000227667.8
ENST00000375345.3 |
APOC3
|
apolipoprotein C3 |
| chr8_-_119673368 | 0.01 |
ENST00000427067.6
|
ENPP2
|
ectonucleotide pyrophosphatase/phosphodiesterase 2 |
| chr5_-_127073467 | 0.01 |
ENST00000607731.5
ENST00000509733.7 ENST00000296662.10 ENST00000535381.6 |
C5orf63
|
chromosome 5 open reading frame 63 |
| chr6_+_30626842 | 0.00 |
ENST00000318999.11
ENST00000376485.8 ENST00000330083.6 ENST00000319027.9 ENST00000376483.8 ENST00000329992.12 |
ATAT1
|
alpha tubulin acetyltransferase 1 |
| chr17_-_2266131 | 0.00 |
ENST00000570606.5
ENST00000354901.8 |
SMG6
|
SMG6 nonsense mediated mRNA decay factor |
| chr3_-_157503574 | 0.00 |
ENST00000494677.5
ENST00000468233.5 |
VEPH1
|
ventricular zone expressed PH domain containing 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 3.3 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 0.5 | 1.5 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.3 | 1.7 | GO:0045204 | MAPK export from nucleus(GO:0045204) |
| 0.2 | 0.7 | GO:0036404 | conversion of ds siRNA to ss siRNA involved in RNA interference(GO:0033168) conversion of ds siRNA to ss siRNA(GO:0036404) |
| 0.2 | 1.4 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.2 | 1.3 | GO:0061624 | fructose catabolic process(GO:0006001) fructose catabolic process to hydroxyacetone phosphate and glyceraldehyde-3-phosphate(GO:0061624) |
| 0.2 | 1.9 | GO:0090292 | nuclear matrix anchoring at nuclear membrane(GO:0090292) |
| 0.2 | 0.8 | GO:2001295 | malonyl-CoA biosynthetic process(GO:2001295) |
| 0.1 | 0.5 | GO:1903788 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) regulation of glutathione biosynthetic process(GO:1903786) positive regulation of glutathione biosynthetic process(GO:1903788) |
| 0.1 | 0.4 | GO:0099558 | maintenance of synapse structure(GO:0099558) |
| 0.1 | 0.3 | GO:1903949 | positive regulation of atrial cardiac muscle cell action potential(GO:1903949) |
| 0.1 | 0.4 | GO:1902445 | regulation of mitochondrial membrane permeability involved in programmed necrotic cell death(GO:1902445) |
| 0.1 | 0.3 | GO:0016999 | antibiotic metabolic process(GO:0016999) detoxification of mercury ion(GO:0050787) |
| 0.1 | 0.8 | GO:0060372 | regulation of atrial cardiac muscle cell membrane repolarization(GO:0060372) |
| 0.1 | 0.3 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.1 | 0.4 | GO:1901341 | activation of store-operated calcium channel activity(GO:0032237) positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.1 | 0.7 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.1 | 2.4 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.1 | 0.6 | GO:0046543 | development of secondary female sexual characteristics(GO:0046543) |
| 0.1 | 0.5 | GO:0002759 | regulation of antimicrobial humoral response(GO:0002759) |
| 0.0 | 0.5 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.0 | 1.1 | GO:0072189 | ureter development(GO:0072189) |
| 0.0 | 2.0 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.0 | 0.1 | GO:0000961 | negative regulation of mitochondrial RNA catabolic process(GO:0000961) |
| 0.0 | 0.5 | GO:0046520 | sphingoid biosynthetic process(GO:0046520) |
| 0.0 | 0.0 | GO:0003408 | optic cup formation involved in camera-type eye development(GO:0003408) |
| 0.0 | 0.3 | GO:0036309 | protein localization to M-band(GO:0036309) |
| 0.0 | 0.5 | GO:1902018 | negative regulation of cilium assembly(GO:1902018) |
| 0.0 | 0.4 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.0 | 0.3 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.0 | 0.3 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.0 | 0.1 | GO:0019482 | beta-alanine metabolic process(GO:0019482) |
| 0.0 | 0.1 | GO:0021615 | glossopharyngeal nerve morphogenesis(GO:0021615) |
| 0.0 | 1.7 | GO:0019730 | antimicrobial humoral response(GO:0019730) |
| 0.0 | 0.3 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.3 | GO:0016188 | synaptic vesicle maturation(GO:0016188) |
| 0.0 | 0.1 | GO:0002191 | cap-dependent translational initiation(GO:0002191) |
| 0.0 | 0.2 | GO:0016559 | peroxisome fission(GO:0016559) |
| 0.0 | 0.3 | GO:0008340 | determination of adult lifespan(GO:0008340) |
| 0.0 | 0.2 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.0 | 0.2 | GO:0016446 | somatic hypermutation of immunoglobulin genes(GO:0016446) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.2 | 0.7 | GO:0033167 | ARC complex(GO:0033167) |
| 0.1 | 1.9 | GO:0034993 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.1 | 0.4 | GO:0045272 | plasma membrane respiratory chain complex I(GO:0045272) |
| 0.1 | 0.4 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.1 | 0.5 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.0 | 0.4 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 0.4 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.0 | 3.3 | GO:0005930 | axoneme(GO:0005930) ciliary plasm(GO:0097014) |
| 0.0 | 0.3 | GO:0098839 | postsynaptic density membrane(GO:0098839) |
| 0.0 | 0.3 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.2 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.3 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.0 | 0.1 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.0 | 0.1 | GO:0005958 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) |
| 0.0 | 7.2 | GO:0005929 | cilium(GO:0005929) |
| 0.0 | 0.3 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.0 | 0.3 | GO:0034706 | sodium channel complex(GO:0034706) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.4 | GO:0047756 | chondroitin 4-sulfotransferase activity(GO:0047756) |
| 0.2 | 1.3 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.2 | 0.8 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.1 | 0.7 | GO:0004530 | deoxyribonuclease I activity(GO:0004530) |
| 0.1 | 0.5 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.1 | 0.2 | GO:0052815 | medium-chain acyl-CoA hydrolase activity(GO:0052815) long-chain acyl-CoA hydrolase activity(GO:0052816) |
| 0.1 | 0.3 | GO:0002046 | opsin binding(GO:0002046) |
| 0.1 | 1.9 | GO:0005521 | lamin binding(GO:0005521) |
| 0.0 | 1.7 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 0.3 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.0 | 0.3 | GO:0031811 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 0.0 | 0.8 | GO:0086008 | voltage-gated potassium channel activity involved in cardiac muscle cell action potential repolarization(GO:0086008) |
| 0.0 | 0.2 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) |
| 0.0 | 0.4 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.0 | 0.1 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.0 | 0.4 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 1.3 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.1 | GO:0004677 | DNA-dependent protein kinase activity(GO:0004677) |
| 0.0 | 0.3 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.0 | 0.2 | GO:0032138 | single base insertion or deletion binding(GO:0032138) |
| 0.0 | 2.8 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 0.5 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.0 | 3.3 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.4 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 0.4 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.3 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.0 | 0.2 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.0 | 0.1 | GO:0004466 | long-chain-acyl-CoA dehydrogenase activity(GO:0004466) |
| 0.0 | 0.3 | GO:0019870 | potassium channel inhibitor activity(GO:0019870) sodium channel inhibitor activity(GO:0019871) |
| 0.0 | 0.5 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.1 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.5 | PID AP1 PATHWAY | AP-1 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.1 | 2.4 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 0.4 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.0 | 1.5 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 0.7 | REACTOME INITIAL TRIGGERING OF COMPLEMENT | Genes involved in Initial triggering of complement |
| 0.0 | 1.0 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.0 | 1.6 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 0.7 | REACTOME MICRORNA MIRNA BIOGENESIS | Genes involved in MicroRNA (miRNA) Biogenesis |
| 0.0 | 0.4 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.2 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |