Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
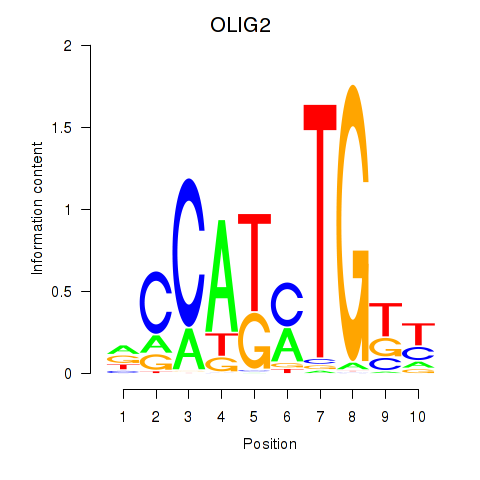
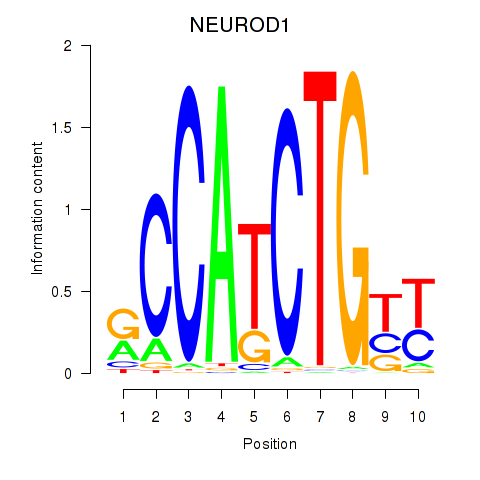
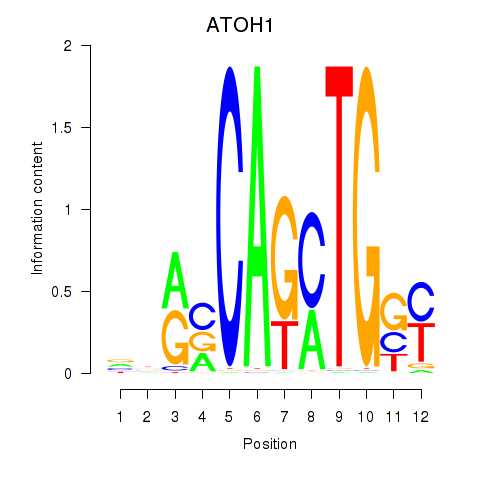
Results for OLIG2_NEUROD1_ATOH1
Z-value: 0.63
Transcription factors associated with OLIG2_NEUROD1_ATOH1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
OLIG2
|
ENSG00000205927.5 | OLIG2 |
|
NEUROD1
|
ENSG00000162992.5 | NEUROD1 |
|
ATOH1
|
ENSG00000172238.6 | ATOH1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NEUROD1 | hg38_v1_chr2_-_181680490_181680525 | 0.39 | 3.3e-02 | Click! |
| OLIG2 | hg38_v1_chr21_+_33025927_33025942 | -0.15 | 4.3e-01 | Click! |
| ATOH1 | hg38_v1_chr4_+_93828746_93828758 | -0.14 | 4.6e-01 | Click! |
Activity profile of OLIG2_NEUROD1_ATOH1 motif
Sorted Z-values of OLIG2_NEUROD1_ATOH1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of OLIG2_NEUROD1_ATOH1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_+_128411715 | 1.78 |
ENST00000420034.5
ENST00000372842.5 |
CERCAM
|
cerebral endothelial cell adhesion molecule |
| chr1_-_205422050 | 1.76 |
ENST00000367153.9
|
LEMD1
|
LEM domain containing 1 |
| chr8_-_134510182 | 1.19 |
ENST00000521673.5
|
ZFAT
|
zinc finger and AT-hook domain containing |
| chr8_-_41665200 | 1.16 |
ENST00000335651.6
|
ANK1
|
ankyrin 1 |
| chr4_-_115113614 | 0.89 |
ENST00000264363.7
|
NDST4
|
N-deacetylase and N-sulfotransferase 4 |
| chrX_+_150983350 | 0.88 |
ENST00000455596.5
ENST00000448905.6 |
HMGB3
|
high mobility group box 3 |
| chr11_-_2149603 | 0.86 |
ENST00000643349.1
|
ENSG00000284779.2
|
novel protein |
| chr6_-_65707214 | 0.85 |
ENST00000370621.7
ENST00000393380.6 ENST00000503581.6 |
EYS
|
eyes shut homolog |
| chr12_+_107318395 | 0.79 |
ENST00000420571.6
ENST00000280758.10 |
BTBD11
|
BTB domain containing 11 |
| chr1_-_16978276 | 0.78 |
ENST00000375534.7
|
MFAP2
|
microfibril associated protein 2 |
| chrX_+_136197020 | 0.78 |
ENST00000370676.7
|
FHL1
|
four and a half LIM domains 1 |
| chrX_+_136197039 | 0.74 |
ENST00000370683.6
|
FHL1
|
four and a half LIM domains 1 |
| chr15_+_66453418 | 0.73 |
ENST00000566326.1
|
MAP2K1
|
mitogen-activated protein kinase kinase 1 |
| chrX_+_136196750 | 0.72 |
ENST00000539015.5
|
FHL1
|
four and a half LIM domains 1 |
| chr19_-_38253238 | 0.70 |
ENST00000587515.5
|
PPP1R14A
|
protein phosphatase 1 regulatory inhibitor subunit 14A |
| chr5_+_136058849 | 0.70 |
ENST00000508076.5
|
TGFBI
|
transforming growth factor beta induced |
| chr4_-_115113822 | 0.70 |
ENST00000613194.4
|
NDST4
|
N-deacetylase and N-sulfotransferase 4 |
| chr7_-_41703062 | 0.67 |
ENST00000242208.5
|
INHBA
|
inhibin subunit beta A |
| chr5_-_181205182 | 0.67 |
ENST00000274773.12
|
TRIM7
|
tripartite motif containing 7 |
| chr14_-_74612226 | 0.66 |
ENST00000261978.9
|
LTBP2
|
latent transforming growth factor beta binding protein 2 |
| chr6_+_54083423 | 0.66 |
ENST00000460844.6
ENST00000370876.6 |
MLIP
|
muscular LMNA interacting protein |
| chr11_+_43942627 | 0.65 |
ENST00000617612.3
|
C11orf96
|
chromosome 11 open reading frame 96 |
| chr8_+_53851786 | 0.64 |
ENST00000297313.8
ENST00000344277.10 |
RGS20
|
regulator of G protein signaling 20 |
| chr17_-_38853629 | 0.62 |
ENST00000378096.3
ENST00000479035.7 ENST00000394332.5 ENST00000394333.5 ENST00000577407.5 |
RPL23
|
ribosomal protein L23 |
| chr12_+_4273751 | 0.61 |
ENST00000675880.1
ENST00000261254.8 |
CCND2
|
cyclin D2 |
| chr7_+_2647703 | 0.59 |
ENST00000403167.5
|
TTYH3
|
tweety family member 3 |
| chr11_-_66907891 | 0.59 |
ENST00000393955.6
|
PC
|
pyruvate carboxylase |
| chr19_-_11339573 | 0.59 |
ENST00000222120.8
|
RAB3D
|
RAB3D, member RAS oncogene family |
| chr9_+_113536497 | 0.57 |
ENST00000462143.5
|
RGS3
|
regulator of G protein signaling 3 |
| chr6_+_31715339 | 0.56 |
ENST00000375824.1
ENST00000375825.7 |
LY6G6D
|
lymphocyte antigen 6 family member G6D |
| chr8_-_48921419 | 0.55 |
ENST00000020945.4
|
SNAI2
|
snail family transcriptional repressor 2 |
| chr12_+_40692413 | 0.55 |
ENST00000551295.7
ENST00000547702.5 ENST00000551424.5 |
CNTN1
|
contactin 1 |
| chr19_+_34994778 | 0.55 |
ENST00000599564.5
|
GRAMD1A
|
GRAM domain containing 1A |
| chr21_+_29130630 | 0.55 |
ENST00000399926.5
ENST00000399928.6 |
MAP3K7CL
|
MAP3K7 C-terminal like |
| chr1_-_24143112 | 0.52 |
ENST00000270800.2
|
IL22RA1
|
interleukin 22 receptor subunit alpha 1 |
| chr19_-_18940289 | 0.51 |
ENST00000542541.6
ENST00000433218.6 |
HOMER3
|
homer scaffold protein 3 |
| chr17_-_9791586 | 0.51 |
ENST00000571134.2
|
DHRS7C
|
dehydrogenase/reductase 7C |
| chr10_-_119536533 | 0.50 |
ENST00000392865.5
|
RGS10
|
regulator of G protein signaling 10 |
| chrX_+_71301742 | 0.49 |
ENST00000373829.8
ENST00000538820.1 |
ITGB1BP2
|
integrin subunit beta 1 binding protein 2 |
| chr17_-_676348 | 0.49 |
ENST00000681510.1
ENST00000679680.1 |
VPS53
|
VPS53 subunit of GARP complex |
| chr1_-_16980607 | 0.49 |
ENST00000375535.4
|
MFAP2
|
microfibril associated protein 2 |
| chr11_-_89063631 | 0.48 |
ENST00000455756.6
|
GRM5
|
glutamate metabotropic receptor 5 |
| chr17_-_7204502 | 0.48 |
ENST00000486626.8
ENST00000648263.1 |
DLG4
|
discs large MAGUK scaffold protein 4 |
| chr8_-_65789084 | 0.47 |
ENST00000379419.8
|
PDE7A
|
phosphodiesterase 7A |
| chr11_-_125592448 | 0.46 |
ENST00000648911.1
|
FEZ1
|
fasciculation and elongation protein zeta 1 |
| chr16_-_79599902 | 0.44 |
ENST00000569649.1
|
MAF
|
MAF bZIP transcription factor |
| chrX_+_150983299 | 0.44 |
ENST00000325307.12
|
HMGB3
|
high mobility group box 3 |
| chr22_+_31092447 | 0.43 |
ENST00000455608.5
|
SMTN
|
smoothelin |
| chr22_+_31093358 | 0.43 |
ENST00000404574.5
|
SMTN
|
smoothelin |
| chr11_-_14891643 | 0.42 |
ENST00000532378.5
|
CYP2R1
|
cytochrome P450 family 2 subfamily R member 1 |
| chr17_+_7438267 | 0.41 |
ENST00000575235.5
|
FGF11
|
fibroblast growth factor 11 |
| chr3_+_35680932 | 0.41 |
ENST00000396481.6
|
ARPP21
|
cAMP regulated phosphoprotein 21 |
| chr19_+_53867874 | 0.41 |
ENST00000448420.5
ENST00000439000.5 ENST00000391771.1 ENST00000391770.9 |
MYADM
|
myeloid associated differentiation marker |
| chr19_+_44777860 | 0.40 |
ENST00000341505.4
ENST00000647358.2 |
CBLC
|
Cbl proto-oncogene C |
| chr20_+_3796288 | 0.40 |
ENST00000439880.6
ENST00000245960.10 |
CDC25B
|
cell division cycle 25B |
| chr19_+_15728024 | 0.40 |
ENST00000305899.5
|
OR10H2
|
olfactory receptor family 10 subfamily H member 2 |
| chr17_+_76384601 | 0.38 |
ENST00000592299.6
ENST00000590959.5 ENST00000323374.8 |
SPHK1
|
sphingosine kinase 1 |
| chr2_+_68774782 | 0.37 |
ENST00000409030.7
ENST00000409220.5 |
ARHGAP25
|
Rho GTPase activating protein 25 |
| chr5_-_172006567 | 0.37 |
ENST00000517395.6
ENST00000265094.9 ENST00000393802.6 |
FBXW11
|
F-box and WD repeat domain containing 11 |
| chrX_+_70290077 | 0.37 |
ENST00000374403.4
|
KIF4A
|
kinesin family member 4A |
| chr5_-_16936231 | 0.36 |
ENST00000507288.1
ENST00000274203.13 ENST00000513610.6 |
MYO10
|
myosin X |
| chr1_-_150876697 | 0.36 |
ENST00000515192.5
|
ARNT
|
aryl hydrocarbon receptor nuclear translocator |
| chr9_+_133061981 | 0.35 |
ENST00000372080.8
|
CEL
|
carboxyl ester lipase |
| chr17_-_78903193 | 0.35 |
ENST00000322630.3
ENST00000586713.5 |
CEP295NL
|
CEP295 N-terminal like |
| chr8_+_69466617 | 0.35 |
ENST00000525061.5
ENST00000260128.8 ENST00000458141.6 |
SULF1
|
sulfatase 1 |
| chr3_+_42856021 | 0.35 |
ENST00000493193.1
|
ACKR2
|
atypical chemokine receptor 2 |
| chr4_+_8199363 | 0.34 |
ENST00000382521.7
ENST00000457650.7 |
SH3TC1
|
SH3 domain and tetratricopeptide repeats 1 |
| chr17_+_76385256 | 0.34 |
ENST00000392496.3
|
SPHK1
|
sphingosine kinase 1 |
| chr16_+_23835946 | 0.34 |
ENST00000321728.12
ENST00000643927.1 |
PRKCB
|
protein kinase C beta |
| chr4_+_8199239 | 0.34 |
ENST00000245105.8
|
SH3TC1
|
SH3 domain and tetratricopeptide repeats 1 |
| chr16_-_30091226 | 0.33 |
ENST00000279386.6
ENST00000627355.2 |
TBX6
|
T-box transcription factor 6 |
| chr11_+_125063295 | 0.33 |
ENST00000532000.5
ENST00000403796.7 ENST00000308074.4 |
SLC37A2
|
solute carrier family 37 member 2 |
| chr20_-_32536383 | 0.33 |
ENST00000375678.7
|
NOL4L
|
nucleolar protein 4 like |
| chr5_-_172006817 | 0.33 |
ENST00000296933.10
|
FBXW11
|
F-box and WD repeat domain containing 11 |
| chr7_-_4959172 | 0.33 |
ENST00000401401.8
ENST00000406755.5 ENST00000404774.7 ENST00000612910.1 |
MMD2
|
monocyte to macrophage differentiation associated 2 |
| chr20_+_44355692 | 0.32 |
ENST00000316673.8
ENST00000609795.5 ENST00000457232.5 ENST00000609262.5 |
HNF4A
|
hepatocyte nuclear factor 4 alpha |
| chr15_+_22094522 | 0.32 |
ENST00000328795.5
|
OR4N4
|
olfactory receptor family 4 subfamily N member 4 |
| chrX_+_47585212 | 0.32 |
ENST00000445623.1
|
TIMP1
|
TIMP metallopeptidase inhibitor 1 |
| chr11_-_14892217 | 0.31 |
ENST00000334636.10
|
CYP2R1
|
cytochrome P450 family 2 subfamily R member 1 |
| chr11_+_71538025 | 0.31 |
ENST00000398534.4
|
KRTAP5-8
|
keratin associated protein 5-8 |
| chr19_+_40717091 | 0.31 |
ENST00000263370.3
|
ITPKC
|
inositol-trisphosphate 3-kinase C |
| chr9_-_35689913 | 0.30 |
ENST00000329305.6
ENST00000645482.3 ENST00000647435.1 ENST00000378292.9 |
TPM2
|
tropomyosin 2 |
| chr9_+_128882119 | 0.30 |
ENST00000372600.9
ENST00000372599.7 |
LRRC8A
|
leucine rich repeat containing 8 VRAC subunit A |
| chr20_-_768424 | 0.29 |
ENST00000675066.1
|
SLC52A3
|
solute carrier family 52 member 3 |
| chr1_+_23743462 | 0.29 |
ENST00000609199.1
|
ELOA
|
elongin A |
| chr7_+_31687208 | 0.29 |
ENST00000409146.3
ENST00000342032.8 |
PPP1R17
|
protein phosphatase 1 regulatory subunit 17 |
| chr21_-_32813695 | 0.29 |
ENST00000479548.2
ENST00000490358.5 |
C21orf62
|
chromosome 21 open reading frame 62 |
| chr1_-_243163310 | 0.29 |
ENST00000492145.1
ENST00000490813.5 ENST00000464936.5 |
CEP170
|
centrosomal protein 170 |
| chr19_-_14560170 | 0.28 |
ENST00000679223.1
|
DNAJB1
|
DnaJ heat shock protein family (Hsp40) member B1 |
| chr17_-_41811930 | 0.28 |
ENST00000393928.6
|
P3H4
|
prolyl 3-hydroxylase family member 4 (inactive) |
| chr10_+_94402486 | 0.28 |
ENST00000225235.5
|
TBC1D12
|
TBC1 domain family member 12 |
| chrX_+_68693629 | 0.27 |
ENST00000374597.3
|
STARD8
|
StAR related lipid transfer domain containing 8 |
| chr17_-_76726753 | 0.27 |
ENST00000617192.4
|
JMJD6
|
jumonji domain containing 6, arginine demethylase and lysine hydroxylase |
| chr1_-_216423396 | 0.27 |
ENST00000366942.3
ENST00000674083.1 ENST00000307340.8 |
USH2A
|
usherin |
| chr10_-_101588126 | 0.27 |
ENST00000339310.7
ENST00000299206.8 ENST00000413344.5 ENST00000429502.1 ENST00000430045.1 ENST00000370172.5 ENST00000370162.8 ENST00000628479.2 |
POLL
|
DNA polymerase lambda |
| chr18_+_49562049 | 0.27 |
ENST00000261292.9
ENST00000427224.6 ENST00000580036.5 |
LIPG
|
lipase G, endothelial type |
| chr10_-_13707536 | 0.27 |
ENST00000632570.1
ENST00000477221.2 |
FRMD4A
|
FERM domain containing 4A |
| chr6_-_26250625 | 0.26 |
ENST00000618052.2
|
H3C7
|
H3 clustered histone 7 |
| chr9_-_128881922 | 0.26 |
ENST00000320665.10
ENST00000436267.7 ENST00000302586.8 ENST00000651925.1 |
KYAT1
ENSG00000286112.1
|
kynurenine aminotransferase 1 kynurenine aminotransferase 1 |
| chr15_+_75647877 | 0.26 |
ENST00000308527.6
|
SNX33
|
sorting nexin 33 |
| chr13_-_33205997 | 0.26 |
ENST00000399365.7
|
STARD13
|
StAR related lipid transfer domain containing 13 |
| chr11_+_76783349 | 0.26 |
ENST00000333090.5
|
TSKU
|
tsukushi, small leucine rich proteoglycan |
| chr6_-_56542802 | 0.25 |
ENST00000524186.1
|
DST
|
dystonin |
| chr10_+_76318330 | 0.25 |
ENST00000496424.2
|
LRMDA
|
leucine rich melanocyte differentiation associated |
| chr12_+_56021316 | 0.25 |
ENST00000547791.2
|
IKZF4
|
IKAROS family zinc finger 4 |
| chr12_+_12891554 | 0.24 |
ENST00000014914.6
|
GPRC5A
|
G protein-coupled receptor class C group 5 member A |
| chr11_+_118956289 | 0.24 |
ENST00000264031.3
|
UPK2
|
uroplakin 2 |
| chr15_+_67067780 | 0.24 |
ENST00000679624.1
|
SMAD3
|
SMAD family member 3 |
| chr5_-_36241798 | 0.24 |
ENST00000514504.5
|
NADK2
|
NAD kinase 2, mitochondrial |
| chr17_+_4833331 | 0.24 |
ENST00000355280.11
ENST00000347992.11 |
MINK1
|
misshapen like kinase 1 |
| chr13_+_32031706 | 0.24 |
ENST00000542859.6
|
FRY
|
FRY microtubule binding protein |
| chr3_+_191329020 | 0.24 |
ENST00000392456.4
|
CCDC50
|
coiled-coil domain containing 50 |
| chr11_+_57597563 | 0.24 |
ENST00000619430.2
ENST00000457869.1 ENST00000340687.10 ENST00000278407.9 ENST00000378323.8 ENST00000378324.6 ENST00000403558.1 |
SERPING1
|
serpin family G member 1 |
| chr11_-_63015804 | 0.23 |
ENST00000535878.5
ENST00000545207.5 |
SLC22A8
|
solute carrier family 22 member 8 |
| chr8_+_66493556 | 0.23 |
ENST00000305454.8
ENST00000522977.5 ENST00000480005.1 |
VXN
|
vexin |
| chr11_+_64237420 | 0.23 |
ENST00000541681.1
|
VEGFB
|
vascular endothelial growth factor B |
| chr5_+_149141817 | 0.23 |
ENST00000504238.5
|
ABLIM3
|
actin binding LIM protein family member 3 |
| chr17_+_51165815 | 0.23 |
ENST00000513177.5
|
NME2
|
NME/NM23 nucleoside diphosphate kinase 2 |
| chr7_-_76626127 | 0.23 |
ENST00000454397.1
|
POMZP3
|
POM121 and ZP3 fusion |
| chr1_-_93847150 | 0.23 |
ENST00000370244.5
|
BCAR3
|
BCAR3 adaptor protein, NSP family member |
| chr17_+_82235769 | 0.22 |
ENST00000619321.2
|
SLC16A3
|
solute carrier family 16 member 3 |
| chr1_+_50103903 | 0.22 |
ENST00000371827.5
|
ELAVL4
|
ELAV like RNA binding protein 4 |
| chr9_-_124507382 | 0.22 |
ENST00000373588.9
ENST00000620110.4 |
NR5A1
|
nuclear receptor subfamily 5 group A member 1 |
| chr1_-_150876571 | 0.22 |
ENST00000354396.6
ENST00000358595.10 ENST00000505755.5 |
ARNT
|
aryl hydrocarbon receptor nuclear translocator |
| chr2_+_232539683 | 0.22 |
ENST00000651502.1
|
CHRNG
|
cholinergic receptor nicotinic gamma subunit |
| chrX_-_46759055 | 0.22 |
ENST00000328306.4
ENST00000616978.5 |
SLC9A7
|
solute carrier family 9 member A7 |
| chr3_+_184230373 | 0.22 |
ENST00000426955.6
|
VWA5B2
|
von Willebrand factor A domain containing 5B2 |
| chr18_+_57352541 | 0.22 |
ENST00000324000.4
|
ST8SIA3
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 |
| chr7_+_143381907 | 0.22 |
ENST00000392910.6
|
ZYX
|
zyxin |
| chr2_+_232539720 | 0.21 |
ENST00000389492.3
|
CHRNG
|
cholinergic receptor nicotinic gamma subunit |
| chr22_+_23894375 | 0.21 |
ENST00000215754.8
|
MIF
|
macrophage migration inhibitory factor |
| chr7_+_143381561 | 0.21 |
ENST00000354434.8
|
ZYX
|
zyxin |
| chr11_+_57598184 | 0.21 |
ENST00000677625.1
ENST00000676670.1 |
SERPING1
|
serpin family G member 1 |
| chr7_-_155510158 | 0.21 |
ENST00000682997.1
|
CNPY1
|
canopy FGF signaling regulator 1 |
| chr12_+_48818763 | 0.21 |
ENST00000548279.5
ENST00000547230.5 |
CACNB3
|
calcium voltage-gated channel auxiliary subunit beta 3 |
| chr7_+_155298561 | 0.21 |
ENST00000476756.1
|
INSIG1
|
insulin induced gene 1 |
| chr2_-_29921580 | 0.20 |
ENST00000389048.8
|
ALK
|
ALK receptor tyrosine kinase |
| chr17_+_47209035 | 0.20 |
ENST00000572316.5
ENST00000354968.5 ENST00000576874.5 ENST00000536623.6 |
MYL4
|
myosin light chain 4 |
| chr9_-_101594995 | 0.20 |
ENST00000636434.1
|
PPP3R2
|
protein phosphatase 3 regulatory subunit B, beta |
| chr5_+_149141483 | 0.20 |
ENST00000326685.11
ENST00000309868.12 |
ABLIM3
|
actin binding LIM protein family member 3 |
| chr14_+_51847145 | 0.20 |
ENST00000615906.4
|
GNG2
|
G protein subunit gamma 2 |
| chr12_+_48818478 | 0.20 |
ENST00000547818.5
ENST00000301050.7 ENST00000547392.5 |
CACNB3
|
calcium voltage-gated channel auxiliary subunit beta 3 |
| chr19_-_2256406 | 0.20 |
ENST00000300961.10
|
JSRP1
|
junctional sarcoplasmic reticulum protein 1 |
| chr9_+_128882502 | 0.20 |
ENST00000259324.5
|
LRRC8A
|
leucine rich repeat containing 8 VRAC subunit A |
| chr19_-_45322867 | 0.20 |
ENST00000221476.4
|
CKM
|
creatine kinase, M-type |
| chr2_+_79512993 | 0.20 |
ENST00000496558.5
ENST00000451966.5 ENST00000402739.9 ENST00000629316.2 |
CTNNA2
|
catenin alpha 2 |
| chr5_+_157460173 | 0.20 |
ENST00000435489.7
ENST00000311946.8 |
NIPAL4
|
NIPA like domain containing 4 |
| chr7_+_134745460 | 0.19 |
ENST00000436461.6
|
CALD1
|
caldesmon 1 |
| chr5_+_149141573 | 0.19 |
ENST00000506113.5
|
ABLIM3
|
actin binding LIM protein family member 3 |
| chr9_-_101594918 | 0.19 |
ENST00000374806.2
|
PPP3R2
|
protein phosphatase 3 regulatory subunit B, beta |
| chrX_-_107775951 | 0.19 |
ENST00000315660.8
ENST00000372384.6 ENST00000502650.1 ENST00000506724.1 |
TSC22D3
|
TSC22 domain family member 3 |
| chr15_-_77696142 | 0.19 |
ENST00000561030.5
|
LINGO1
|
leucine rich repeat and Ig domain containing 1 |
| chr2_+_136765508 | 0.19 |
ENST00000409968.6
|
THSD7B
|
thrombospondin type 1 domain containing 7B |
| chr10_+_100999287 | 0.19 |
ENST00000370220.1
|
LZTS2
|
leucine zipper tumor suppressor 2 |
| chr12_-_66678934 | 0.19 |
ENST00000545666.5
ENST00000398016.7 ENST00000359742.9 ENST00000538211.5 |
GRIP1
|
glutamate receptor interacting protein 1 |
| chr19_-_42412347 | 0.19 |
ENST00000601189.1
ENST00000599211.1 |
LIPE
|
lipase E, hormone sensitive type |
| chr1_+_12464912 | 0.19 |
ENST00000543766.2
|
VPS13D
|
vacuolar protein sorting 13 homolog D |
| chr17_-_3398410 | 0.19 |
ENST00000322608.2
|
OR1E1
|
olfactory receptor family 1 subfamily E member 1 |
| chr14_+_92923143 | 0.19 |
ENST00000216492.10
ENST00000334654.4 |
CHGA
|
chromogranin A |
| chr22_-_35840218 | 0.19 |
ENST00000414461.6
ENST00000416721.6 ENST00000449924.6 ENST00000262829.11 ENST00000397305.3 |
RBFOX2
|
RNA binding fox-1 homolog 2 |
| chr12_+_4269771 | 0.19 |
ENST00000676411.1
|
CCND2
|
cyclin D2 |
| chr15_-_67254625 | 0.19 |
ENST00000261880.10
ENST00000561452.5 |
AAGAB
|
alpha and gamma adaptin binding protein |
| chr3_-_139539679 | 0.19 |
ENST00000483943.6
ENST00000672186.1 ENST00000232219.6 ENST00000617459.4 ENST00000492918.1 |
RBP1
|
retinol binding protein 1 |
| chr12_+_119668109 | 0.19 |
ENST00000229328.10
ENST00000630317.1 |
PRKAB1
|
protein kinase AMP-activated non-catalytic subunit beta 1 |
| chr12_+_112978504 | 0.18 |
ENST00000392583.7
|
OAS2
|
2'-5'-oligoadenylate synthetase 2 |
| chr17_-_41118369 | 0.18 |
ENST00000391413.4
|
KRTAP4-11
|
keratin associated protein 4-11 |
| chr1_+_155078829 | 0.18 |
ENST00000368408.4
|
EFNA3
|
ephrin A3 |
| chr22_-_37486357 | 0.18 |
ENST00000356998.8
ENST00000416983.7 ENST00000424765.2 |
MFNG
|
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr15_+_21651844 | 0.18 |
ENST00000623441.1
|
OR4N4C
|
olfactory receptor family 4 subfamily N member 4C |
| chr5_-_36241944 | 0.18 |
ENST00000511088.1
ENST00000282512.7 ENST00000506945.5 |
NADK2
|
NAD kinase 2, mitochondrial |
| chr6_-_144008364 | 0.18 |
ENST00000625622.2
|
PLAGL1
|
PLAG1 like zinc finger 1 |
| chr12_-_52321395 | 0.18 |
ENST00000293670.3
|
KRT83
|
keratin 83 |
| chr2_+_219460719 | 0.18 |
ENST00000396688.5
|
SPEG
|
striated muscle enriched protein kinase |
| chr7_-_100479142 | 0.18 |
ENST00000300181.7
ENST00000393991.5 |
TSC22D4
|
TSC22 domain family member 4 |
| chr17_+_47209338 | 0.18 |
ENST00000393450.5
|
MYL4
|
myosin light chain 4 |
| chr6_-_40587314 | 0.18 |
ENST00000338305.7
|
LRFN2
|
leucine rich repeat and fibronectin type III domain containing 2 |
| chr2_+_74002685 | 0.18 |
ENST00000305799.8
|
TET3
|
tet methylcytosine dioxygenase 3 |
| chr2_-_88947820 | 0.17 |
ENST00000496168.1
|
IGKV1-5
|
immunoglobulin kappa variable 1-5 |
| chr22_-_35840577 | 0.17 |
ENST00000405409.6
|
RBFOX2
|
RNA binding fox-1 homolog 2 |
| chr19_+_15949008 | 0.17 |
ENST00000322107.1
|
OR10H4
|
olfactory receptor family 10 subfamily H member 4 |
| chr12_+_93572664 | 0.17 |
ENST00000551556.2
|
SOCS2
|
suppressor of cytokine signaling 2 |
| chr2_-_181680490 | 0.17 |
ENST00000684145.1
ENST00000295108.4 ENST00000684079.1 ENST00000683430.1 |
CERKL
NEUROD1
|
ceramide kinase like neuronal differentiation 1 |
| chr9_+_122614738 | 0.17 |
ENST00000297913.3
|
OR1Q1
|
olfactory receptor family 1 subfamily Q member 1 |
| chr1_-_228406761 | 0.17 |
ENST00000366699.3
ENST00000284551.11 |
TRIM11
|
tripartite motif containing 11 |
| chr3_+_159852933 | 0.17 |
ENST00000482804.1
|
SCHIP1
|
schwannomin interacting protein 1 |
| chr6_-_144008396 | 0.17 |
ENST00000354765.6
ENST00000416623.5 ENST00000649211.1 |
PLAGL1
|
PLAG1 like zinc finger 1 |
| chr3_+_49803212 | 0.17 |
ENST00000333323.6
|
INKA1
|
inka box actin regulator 1 |
| chr1_-_231421146 | 0.17 |
ENST00000667629.1
ENST00000670301.1 ENST00000658954.1 |
EGLN1
|
egl-9 family hypoxia inducible factor 1 |
| chr12_-_76084612 | 0.17 |
ENST00000535020.6
ENST00000552342.5 |
NAP1L1
|
nucleosome assembly protein 1 like 1 |
| chr17_-_76726453 | 0.17 |
ENST00000585429.1
|
JMJD6
|
jumonji domain containing 6, arginine demethylase and lysine hydroxylase |
| chr6_-_144008118 | 0.17 |
ENST00000629195.2
ENST00000650125.1 ENST00000647880.1 ENST00000649307.1 ENST00000674357.1 ENST00000417959.4 ENST00000635591.1 |
PLAGL1
HYMAI
|
PLAG1 like zinc finger 1 hydatidiform mole associated and imprinted |
| chr3_+_46877705 | 0.17 |
ENST00000449590.6
|
PTH1R
|
parathyroid hormone 1 receptor |
| chr2_-_26346755 | 0.16 |
ENST00000651242.1
ENST00000433584.1 ENST00000333478.10 |
ADGRF3
|
adhesion G protein-coupled receptor F3 |
| chr8_-_22156741 | 0.16 |
ENST00000424267.6
|
LGI3
|
leucine rich repeat LGI family member 3 |
| chr11_+_5389377 | 0.16 |
ENST00000328611.5
|
OR51M1
|
olfactory receptor family 51 subfamily M member 1 |
| chr1_+_228149922 | 0.16 |
ENST00000366714.3
|
GJC2
|
gap junction protein gamma 2 |
| chr22_-_37484505 | 0.16 |
ENST00000442496.1
|
MFNG
|
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr15_+_88638947 | 0.16 |
ENST00000559876.2
|
ISG20
|
interferon stimulated exonuclease gene 20 |
| chr5_+_114056017 | 0.16 |
ENST00000512097.9
|
KCNN2
|
potassium calcium-activated channel subfamily N member 2 |
| chr17_-_3292600 | 0.16 |
ENST00000615105.1
|
OR3A1
|
olfactory receptor family 3 subfamily A member 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0046521 | sphingoid catabolic process(GO:0046521) |
| 0.2 | 0.7 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 0.2 | 0.6 | GO:0070079 | peptidyl-lysine hydroxylation to 5-hydroxy-L-lysine(GO:0018395) histone arginine demethylation(GO:0070077) histone H3-R2 demethylation(GO:0070078) histone H4-R3 demethylation(GO:0070079) |
| 0.2 | 0.6 | GO:1901420 | negative regulation of response to alcohol(GO:1901420) |
| 0.2 | 1.3 | GO:0048050 | post-embryonic eye morphogenesis(GO:0048050) |
| 0.2 | 0.6 | GO:0072717 | cellular response to actinomycin D(GO:0072717) |
| 0.1 | 0.4 | GO:0001868 | regulation of complement activation, lectin pathway(GO:0001868) negative regulation of complement activation, lectin pathway(GO:0001869) |
| 0.1 | 0.7 | GO:0036378 | calcitriol biosynthetic process from calciol(GO:0036378) |
| 0.1 | 0.6 | GO:0019074 | viral genome packaging(GO:0019072) viral RNA genome packaging(GO:0019074) |
| 0.1 | 0.6 | GO:0018125 | peptidyl-cysteine methylation(GO:0018125) |
| 0.1 | 0.6 | GO:0007206 | phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.1 | 0.4 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.1 | 1.6 | GO:0030202 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.1 | 0.3 | GO:0014846 | esophagus smooth muscle contraction(GO:0014846) |
| 0.1 | 0.7 | GO:0090170 | regulation of Golgi inheritance(GO:0090170) |
| 0.1 | 0.4 | GO:0018106 | peptidyl-histidine phosphorylation(GO:0018106) |
| 0.1 | 0.3 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.1 | 0.4 | GO:0044240 | multicellular organism lipid catabolic process(GO:0044240) |
| 0.1 | 0.6 | GO:0002329 | pre-B cell differentiation(GO:0002329) |
| 0.1 | 0.2 | GO:1901301 | regulation of cargo loading into COPII-coated vesicle(GO:1901301) |
| 0.1 | 0.3 | GO:0010982 | regulation of high-density lipoprotein particle clearance(GO:0010982) |
| 0.1 | 1.3 | GO:0045199 | maintenance of apical/basal cell polarity(GO:0035090) maintenance of epithelial cell apical/basal polarity(GO:0045199) |
| 0.1 | 0.3 | GO:0061073 | ciliary body morphogenesis(GO:0061073) |
| 0.1 | 0.5 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.1 | 0.2 | GO:1901899 | positive regulation of relaxation of cardiac muscle(GO:1901899) |
| 0.1 | 0.2 | GO:1904428 | negative regulation of tubulin deacetylation(GO:1904428) |
| 0.1 | 0.2 | GO:0044725 | chromatin reprogramming in the zygote(GO:0044725) |
| 0.1 | 0.6 | GO:0046886 | positive regulation of hormone biosynthetic process(GO:0046886) |
| 0.1 | 0.2 | GO:0007538 | primary sex determination(GO:0007538) |
| 0.1 | 0.3 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.1 | 0.1 | GO:0090134 | mesendoderm migration(GO:0090133) cell migration involved in mesendoderm migration(GO:0090134) |
| 0.1 | 1.2 | GO:0060712 | spongiotrophoblast layer development(GO:0060712) |
| 0.1 | 0.2 | GO:0002905 | mature B cell apoptotic process(GO:0002901) regulation of mature B cell apoptotic process(GO:0002905) negative regulation of mature B cell apoptotic process(GO:0002906) |
| 0.1 | 0.2 | GO:0014740 | negative regulation of muscle hyperplasia(GO:0014740) |
| 0.1 | 0.2 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.1 | GO:0010931 | macrophage tolerance induction(GO:0010931) regulation of macrophage tolerance induction(GO:0010932) positive regulation of macrophage tolerance induction(GO:0010933) |
| 0.0 | 0.1 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 0.0 | 0.1 | GO:0019364 | NADP catabolic process(GO:0006742) pyridine nucleotide catabolic process(GO:0019364) |
| 0.0 | 0.4 | GO:0098903 | regulation of membrane repolarization during action potential(GO:0098903) |
| 0.0 | 0.5 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.0 | 0.3 | GO:0006287 | base-excision repair, gap-filling(GO:0006287) |
| 0.0 | 0.3 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.0 | 0.3 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.0 | 0.3 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.0 | 0.2 | GO:0030806 | negative regulation of cyclic nucleotide catabolic process(GO:0030806) negative regulation of cAMP catabolic process(GO:0030821) |
| 0.0 | 1.2 | GO:0023019 | signal transduction involved in regulation of gene expression(GO:0023019) |
| 0.0 | 0.4 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.3 | GO:1902961 | positive regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902961) positive regulation of aspartic-type peptidase activity(GO:1905247) |
| 0.0 | 0.6 | GO:0021942 | radial glia guided migration of Purkinje cell(GO:0021942) |
| 0.0 | 0.2 | GO:0035803 | egg coat formation(GO:0035803) |
| 0.0 | 0.2 | GO:0061056 | sclerotome development(GO:0061056) |
| 0.0 | 0.3 | GO:0014859 | negative regulation of skeletal muscle cell proliferation(GO:0014859) negative regulation of skeletal muscle satellite cell proliferation(GO:1902723) |
| 0.0 | 0.3 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.0 | 0.1 | GO:0090096 | regulation of metanephric cap mesenchymal cell proliferation(GO:0090095) positive regulation of metanephric cap mesenchymal cell proliferation(GO:0090096) |
| 0.0 | 0.5 | GO:0097113 | AMPA glutamate receptor clustering(GO:0097113) glutamate receptor clustering(GO:0097688) |
| 0.0 | 0.8 | GO:0071481 | cellular response to X-ray(GO:0071481) |
| 0.0 | 0.1 | GO:0019836 | hemolysis by symbiont of host erythrocytes(GO:0019836) hemolysis in other organism(GO:0044179) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 0.0 | 0.5 | GO:1902902 | negative regulation of autophagosome assembly(GO:1902902) |
| 0.0 | 0.1 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) |
| 0.0 | 0.2 | GO:2000048 | negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 0.0 | 0.2 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.0 | 0.2 | GO:0071477 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 0.0 | 0.2 | GO:0030311 | poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.0 | 0.6 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.0 | 0.1 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) |
| 0.0 | 0.4 | GO:0030011 | maintenance of cell polarity(GO:0030011) |
| 0.0 | 0.2 | GO:0035879 | plasma membrane lactate transport(GO:0035879) |
| 0.0 | 1.1 | GO:0098926 | acetylcholine receptor signaling pathway(GO:0095500) postsynaptic signal transduction(GO:0098926) signal transduction involved in cellular response to ammonium ion(GO:1903831) response to acetylcholine(GO:1905144) cellular response to acetylcholine(GO:1905145) |
| 0.0 | 0.1 | GO:1904075 | regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.0 | 0.6 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.0 | 0.1 | GO:0038018 | Wnt receptor catabolic process(GO:0038018) |
| 0.0 | 0.1 | GO:0002302 | CD8-positive, alpha-beta T cell differentiation involved in immune response(GO:0002302) |
| 0.0 | 0.8 | GO:0050908 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.0 | 0.7 | GO:0042753 | positive regulation of circadian rhythm(GO:0042753) |
| 0.0 | 0.2 | GO:0060754 | positive regulation of mast cell chemotaxis(GO:0060754) |
| 0.0 | 0.1 | GO:0038156 | interleukin-5-mediated signaling pathway(GO:0038043) interleukin-3-mediated signaling pathway(GO:0038156) |
| 0.0 | 0.2 | GO:0097052 | L-kynurenine metabolic process(GO:0097052) |
| 0.0 | 0.1 | GO:0001808 | type IV hypersensitivity(GO:0001806) regulation of type IV hypersensitivity(GO:0001807) negative regulation of type IV hypersensitivity(GO:0001808) |
| 0.0 | 0.4 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.0 | 0.1 | GO:0019255 | UDP-glucuronate biosynthetic process(GO:0006065) glucose 1-phosphate metabolic process(GO:0019255) UDP-glucuronate metabolic process(GO:0046398) |
| 0.0 | 0.1 | GO:0039534 | negative regulation of MDA-5 signaling pathway(GO:0039534) |
| 0.0 | 0.1 | GO:0051594 | detection of carbohydrate stimulus(GO:0009730) detection of hexose stimulus(GO:0009732) detection of monosaccharide stimulus(GO:0034287) detection of glucose(GO:0051594) |
| 0.0 | 0.3 | GO:1902951 | negative regulation of dendritic spine maintenance(GO:1902951) |
| 0.0 | 0.3 | GO:0071313 | cellular response to caffeine(GO:0071313) |
| 0.0 | 0.3 | GO:0036089 | cleavage furrow formation(GO:0036089) |
| 0.0 | 0.1 | GO:0098528 | skeletal muscle fiber differentiation(GO:0098528) |
| 0.0 | 0.2 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) |
| 0.0 | 0.1 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.0 | 0.1 | GO:1903377 | negative regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903377) |
| 0.0 | 0.1 | GO:0021564 | vagus nerve development(GO:0021564) |
| 0.0 | 0.1 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.0 | 0.2 | GO:0035878 | nail development(GO:0035878) |
| 0.0 | 0.2 | GO:0098914 | membrane repolarization during atrial cardiac muscle cell action potential(GO:0098914) |
| 0.0 | 0.1 | GO:0043686 | co-translational protein modification(GO:0043686) |
| 0.0 | 0.1 | GO:0044501 | modulation of signal transduction in other organism(GO:0044501) modulation by symbiont of host signal transduction pathway(GO:0052027) modulation of signal transduction in other organism involved in symbiotic interaction(GO:0052250) modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade(GO:0085032) |
| 0.0 | 0.1 | GO:0032903 | viral protein processing(GO:0019082) regulation of nerve growth factor production(GO:0032903) negative regulation of nerve growth factor production(GO:0032904) dibasic protein processing(GO:0090472) |
| 0.0 | 0.1 | GO:0070460 | thyroid-stimulating hormone secretion(GO:0070460) regulation of thyroid-stimulating hormone secretion(GO:2000612) |
| 0.0 | 0.1 | GO:2000639 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.0 | 0.1 | GO:0021586 | olfactory learning(GO:0008355) pons maturation(GO:0021586) cellular alkene metabolic process(GO:0043449) |
| 0.0 | 0.1 | GO:0046707 | IDP metabolic process(GO:0046707) IDP catabolic process(GO:0046709) |
| 0.0 | 0.1 | GO:0070495 | regulation of thrombin receptor signaling pathway(GO:0070494) negative regulation of thrombin receptor signaling pathway(GO:0070495) |
| 0.0 | 0.2 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.0 | 0.1 | GO:1901350 | cell-cell signaling involved in cell-cell junction organization(GO:1901350) |
| 0.0 | 1.6 | GO:0043268 | positive regulation of potassium ion transport(GO:0043268) |
| 0.0 | 0.3 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 0.0 | 0.2 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.0 | 0.0 | GO:0034165 | positive regulation of toll-like receptor 9 signaling pathway(GO:0034165) |
| 0.0 | 0.2 | GO:1990118 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 0.1 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.2 | GO:0007144 | female meiosis I(GO:0007144) |
| 0.0 | 0.1 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.0 | 0.1 | GO:0007499 | ectoderm and mesoderm interaction(GO:0007499) |
| 0.0 | 0.1 | GO:0036324 | vascular endothelial growth factor receptor-2 signaling pathway(GO:0036324) |
| 0.0 | 0.1 | GO:0033058 | directional locomotion(GO:0033058) |
| 0.0 | 0.3 | GO:0007130 | synaptonemal complex assembly(GO:0007130) |
| 0.0 | 0.1 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.0 | 0.2 | GO:2001214 | positive regulation of vasculogenesis(GO:2001214) |
| 0.0 | 0.4 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.1 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.0 | 0.1 | GO:0060056 | mammary gland involution(GO:0060056) |
| 0.0 | 0.2 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.0 | 0.2 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.0 | 0.1 | GO:0031086 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.0 | 0.0 | GO:2000097 | regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 0.0 | 0.1 | GO:0030807 | positive regulation of cyclic nucleotide catabolic process(GO:0030807) positive regulation of cAMP catabolic process(GO:0030822) |
| 0.0 | 0.0 | GO:1903215 | negative regulation of protein targeting to mitochondrion(GO:1903215) |
| 0.0 | 0.1 | GO:1904628 | response to phorbol 13-acetate 12-myristate(GO:1904627) cellular response to phorbol 13-acetate 12-myristate(GO:1904628) |
| 0.0 | 0.4 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 0.0 | 0.1 | GO:0060437 | lung growth(GO:0060437) |
| 0.0 | 0.2 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 0.0 | 0.0 | GO:0007225 | patched ligand maturation(GO:0007225) signal maturation(GO:0035638) |
| 0.0 | 0.0 | GO:0036496 | regulation of translational initiation by eIF2 alpha dephosphorylation(GO:0036496) |
| 0.0 | 0.7 | GO:0097435 | fibril organization(GO:0097435) |
| 0.0 | 0.1 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.0 | 0.0 | GO:0021593 | rhombomere morphogenesis(GO:0021593) rhombomere 3 morphogenesis(GO:0021658) |
| 0.0 | 1.8 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 1.3 | GO:0032392 | DNA geometric change(GO:0032392) |
| 0.0 | 0.1 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0043512 | inhibin complex(GO:0043511) inhibin A complex(GO:0043512) |
| 0.1 | 0.8 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.1 | 0.3 | GO:0002139 | stereocilia coupling link(GO:0002139) periciliary membrane compartment(GO:1990075) |
| 0.1 | 0.5 | GO:1990745 | EARP complex(GO:1990745) |
| 0.1 | 1.3 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.1 | 0.3 | GO:0031673 | H zone(GO:0031673) |
| 0.1 | 0.2 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.0 | 0.2 | GO:0016035 | zeta DNA polymerase complex(GO:0016035) |
| 0.0 | 0.7 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.5 | GO:0098839 | postsynaptic density membrane(GO:0098839) |
| 0.0 | 0.2 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.1 | GO:0070381 | endosome to plasma membrane transport vesicle(GO:0070381) |
| 0.0 | 0.6 | GO:0042588 | zymogen granule(GO:0042588) |
| 0.0 | 0.3 | GO:0070449 | elongin complex(GO:0070449) |
| 0.0 | 0.1 | GO:1990032 | parallel fiber(GO:1990032) |
| 0.0 | 1.6 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 0.3 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.0 | 0.4 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.2 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.0 | 1.0 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 0.4 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 0.1 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.0 | 0.4 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.1 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 0.0 | 0.2 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.0 | 0.1 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.0 | 0.2 | GO:0030478 | actin cap(GO:0030478) |
| 0.0 | 0.2 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.0 | 0.5 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 0.1 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.0 | 0.1 | GO:0002178 | palmitoyltransferase complex(GO:0002178) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.6 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.2 | 0.6 | GO:0004874 | aryl hydrocarbon receptor activity(GO:0004874) |
| 0.2 | 0.6 | GO:0033746 | histone demethylase activity (H3-R2 specific)(GO:0033746) histone demethylase activity (H4-R3 specific)(GO:0033749) |
| 0.2 | 0.6 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.2 | 0.7 | GO:0030343 | vitamin D3 25-hydroxylase activity(GO:0030343) vitamin D 25-hydroxylase activity(GO:0070643) |
| 0.1 | 0.6 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.5 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.1 | 0.7 | GO:0008481 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.1 | 0.3 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 0.1 | 0.3 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.1 | 0.3 | GO:0004698 | calcium-dependent protein kinase C activity(GO:0004698) |
| 0.1 | 0.7 | GO:0070699 | type II activin receptor binding(GO:0070699) |
| 0.1 | 0.3 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.1 | 0.3 | GO:0061513 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.1 | 0.3 | GO:0047804 | cysteine-S-conjugate beta-lyase activity(GO:0047804) L-phenylalanine aminotransferase activity(GO:0070546) |
| 0.1 | 0.2 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.1 | 0.4 | GO:0032038 | myosin II heavy chain binding(GO:0032038) |
| 0.1 | 0.6 | GO:0099583 | neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration(GO:0099583) |
| 0.1 | 0.5 | GO:0031812 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 0.1 | 0.4 | GO:0004771 | sterol esterase activity(GO:0004771) |
| 0.0 | 0.6 | GO:0004075 | biotin carboxylase activity(GO:0004075) |
| 0.0 | 0.2 | GO:0004991 | parathyroid hormone receptor activity(GO:0004991) |
| 0.0 | 0.2 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.0 | 1.3 | GO:0000400 | four-way junction DNA binding(GO:0000400) |
| 0.0 | 0.2 | GO:0004167 | dopachrome isomerase activity(GO:0004167) |
| 0.0 | 0.4 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.0 | 0.2 | GO:0031962 | mineralocorticoid receptor binding(GO:0031962) |
| 0.0 | 0.3 | GO:0004465 | lipoprotein lipase activity(GO:0004465) |
| 0.0 | 1.1 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.3 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.0 | 0.2 | GO:0048101 | calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.0 | 0.1 | GO:0051748 | UTP:glucose-1-phosphate uridylyltransferase activity(GO:0003983) UTP-monosaccharide-1-phosphate uridylyltransferase activity(GO:0051748) |
| 0.0 | 0.2 | GO:0070579 | methylcytosine dioxygenase activity(GO:0070579) |
| 0.0 | 0.6 | GO:0035256 | G-protein coupled glutamate receptor binding(GO:0035256) |
| 0.0 | 0.3 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.0 | 0.2 | GO:0001010 | transcription factor activity, sequence-specific DNA binding transcription factor recruiting(GO:0001010) |
| 0.0 | 0.5 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.0 | 0.1 | GO:0004346 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.0 | 0.6 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.0 | 0.2 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.0 | 0.1 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.0 | 0.4 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.3 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 0.2 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.0 | 0.1 | GO:0004912 | interleukin-3 receptor activity(GO:0004912) interleukin-5 receptor activity(GO:0004914) |
| 0.0 | 0.7 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.0 | 0.2 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.0 | 0.3 | GO:0051575 | 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.0 | 0.7 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.0 | 0.6 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.0 | 0.2 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.0 | 0.3 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.0 | 0.2 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.0 | 0.1 | GO:0042284 | sphingolipid delta-4 desaturase activity(GO:0042284) |
| 0.0 | 0.1 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 0.0 | 0.6 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.0 | 0.1 | GO:0004090 | carbonyl reductase (NADPH) activity(GO:0004090) |
| 0.0 | 0.2 | GO:0008269 | JAK pathway signal transduction adaptor activity(GO:0008269) |
| 0.0 | 0.2 | GO:0043426 | MRF binding(GO:0043426) |
| 0.0 | 0.3 | GO:0004673 | protein histidine kinase activity(GO:0004673) |
| 0.0 | 0.1 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.0 | 0.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.0 | 0.2 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.2 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.1 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.0 | 0.2 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.2 | GO:1903763 | gap junction channel activity involved in cell communication by electrical coupling(GO:1903763) |
| 0.0 | 0.1 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.0 | 0.5 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.7 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 0.1 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.0 | 0.1 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 0.0 | 0.2 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.2 | GO:0022820 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 0.0 | 0.6 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.1 | GO:0004800 | thyroxine 5'-deiodinase activity(GO:0004800) |
| 0.0 | 0.2 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.0 | 0.2 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.0 | 0.1 | GO:0098770 | FBXO family protein binding(GO:0098770) |
| 0.0 | 0.1 | GO:0019144 | ADP-sugar diphosphatase activity(GO:0019144) |
| 0.0 | 0.1 | GO:0016524 | latrotoxin receptor activity(GO:0016524) |
| 0.0 | 0.9 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.0 | 0.1 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.0 | 0.1 | GO:0031432 | titin binding(GO:0031432) |
| 0.0 | 0.1 | GO:0035473 | lipase binding(GO:0035473) |
| 0.0 | 0.1 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.0 | 0.3 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.4 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 0.2 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.3 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 0.2 | GO:0019841 | retinol binding(GO:0019841) |
| 0.0 | 0.6 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 1.0 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.5 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.0 | 0.6 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.1 | GO:0016802 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.5 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.4 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 0.3 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 0.4 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 0.3 | GO:0005523 | tropomyosin binding(GO:0005523) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.1 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 0.4 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.0 | 0.7 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.0 | 2.0 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.2 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.0 | 0.5 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 0.2 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.0 | 0.7 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.2 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.0 | 0.9 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.0 | 0.5 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 0.6 | PID ECADHERIN KERATINOCYTE PATHWAY | E-cadherin signaling in keratinocytes |
| 0.0 | 0.6 | ST G ALPHA I PATHWAY | G alpha i Pathway |
| 0.0 | 0.5 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.7 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 2.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.3 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.2 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.1 | 0.7 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.0 | 1.6 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.4 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.0 | 0.9 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.4 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 0.7 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.0 | 0.4 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 1.3 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 0.2 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.5 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.4 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.7 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.9 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.4 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.1 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.0 | 0.6 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.6 | REACTOME REGULATION OF HYPOXIA INDUCIBLE FACTOR HIF BY OXYGEN | Genes involved in Regulation of Hypoxia-inducible Factor (HIF) by Oxygen |
| 0.0 | 0.5 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 1.9 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.0 | 0.4 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.1 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 0.1 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.0 | 0.2 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.5 | REACTOME SMAD2 SMAD3 SMAD4 HETEROTRIMER REGULATES TRANSCRIPTION | Genes involved in SMAD2/SMAD3:SMAD4 heterotrimer regulates transcription |
| 0.0 | 0.4 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |