Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
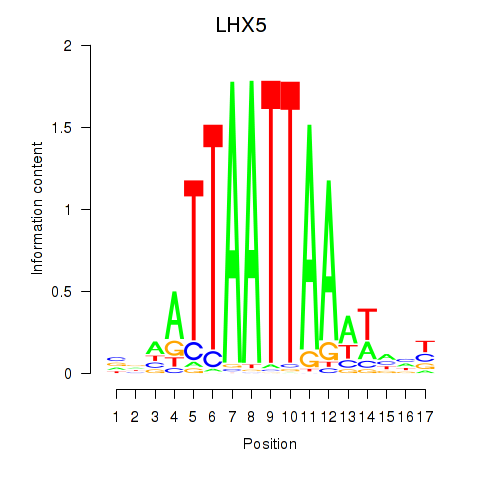
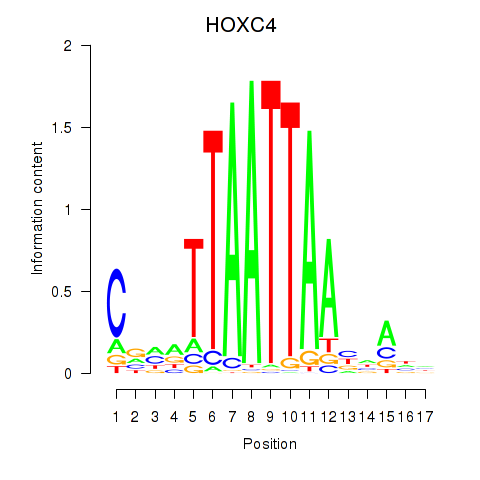
Results for OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4
Z-value: 0.68
Transcription factors associated with OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
OTP
|
ENSG00000171540.8 | OTP |
|
PHOX2B
|
ENSG00000109132.7 | PHOX2B |
|
LHX1
|
ENSG00000273706.5 | LHX1 |
|
LMX1A
|
ENSG00000162761.14 | LMX1A |
|
LHX5
|
ENSG00000089116.4 | LHX5 |
|
HOXC4
|
ENSG00000198353.8 | HOXC4 |
|
HOXC4
|
ENSG00000198353.8 | HOXC4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| LHX5 | hg38_v1_chr12_-_113471851_113471876 | 0.39 | 3.1e-02 | Click! |
| PHOX2B | hg38_v1_chr4_-_41748713_41748731 | 0.27 | 1.5e-01 | Click! |
| LMX1A | hg38_v1_chr1_-_165356212_165356241, hg38_v1_chr1_-_165356703_165356721 | 0.26 | 1.7e-01 | Click! |
| HOXC4 | hg38_v1_chr12_+_54053815_54053875 | 0.12 | 5.2e-01 | Click! |
| OTP | hg38_v1_chr5_-_77638713_77638713 | -0.07 | 7.0e-01 | Click! |
Activity profile of OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4 motif
Sorted Z-values of OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of OTP_PHOX2B_LHX1_LMX1A_LHX5_HOXC4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_+_130018565 | 3.03 |
ENST00000361794.7
ENST00000526087.5 ENST00000533560.5 |
L3MBTL3
|
L3MBTL histone methyl-lysine binding protein 3 |
| chr9_+_72577939 | 2.72 |
ENST00000645773.1
|
TMC1
|
transmembrane channel like 1 |
| chr15_+_41286011 | 1.89 |
ENST00000661438.1
|
ENSG00000285920.2
|
novel protein |
| chr11_-_122116215 | 1.89 |
ENST00000560104.2
|
BLID
|
BH3-like motif containing, cell death inducer |
| chrX_+_43656289 | 1.85 |
ENST00000338702.4
|
MAOA
|
monoamine oxidase A |
| chr3_-_142029108 | 1.70 |
ENST00000497579.5
|
TFDP2
|
transcription factor Dp-2 |
| chr9_+_72577369 | 1.42 |
ENST00000651183.1
|
TMC1
|
transmembrane channel like 1 |
| chr9_+_122371036 | 1.37 |
ENST00000619306.5
ENST00000426608.6 ENST00000223423.8 |
PTGS1
|
prostaglandin-endoperoxide synthase 1 |
| chr9_+_72577788 | 1.35 |
ENST00000645208.2
|
TMC1
|
transmembrane channel like 1 |
| chr17_+_59155726 | 1.18 |
ENST00000578777.5
ENST00000577457.1 ENST00000582995.5 ENST00000262293.9 ENST00000614081.1 |
PRR11
|
proline rich 11 |
| chr11_-_107858777 | 1.16 |
ENST00000525815.6
|
SLC35F2
|
solute carrier family 35 member F2 |
| chr6_-_32190170 | 1.12 |
ENST00000375050.6
|
PBX2
|
PBX homeobox 2 |
| chr7_+_134843884 | 1.03 |
ENST00000445569.6
|
CALD1
|
caldesmon 1 |
| chr11_-_5301946 | 1.02 |
ENST00000380224.2
|
OR51B4
|
olfactory receptor family 51 subfamily B member 4 |
| chr4_+_168497066 | 0.97 |
ENST00000261509.10
|
PALLD
|
palladin, cytoskeletal associated protein |
| chr3_-_165078480 | 0.95 |
ENST00000264382.8
|
SI
|
sucrase-isomaltase |
| chr5_-_126595237 | 0.94 |
ENST00000637206.1
ENST00000553117.5 |
ALDH7A1
|
aldehyde dehydrogenase 7 family member A1 |
| chr6_+_113857333 | 0.91 |
ENST00000612661.2
|
MARCKS
|
myristoylated alanine rich protein kinase C substrate |
| chr15_+_21579912 | 0.87 |
ENST00000628444.1
|
LINC02203
|
long intergenic non-protein coding RNA 2203 |
| chr9_+_122371014 | 0.86 |
ENST00000362012.7
|
PTGS1
|
prostaglandin-endoperoxide synthase 1 |
| chr4_+_168497044 | 0.86 |
ENST00000505667.6
|
PALLD
|
palladin, cytoskeletal associated protein |
| chr12_+_26011713 | 0.84 |
ENST00000542004.5
|
RASSF8
|
Ras association domain family member 8 |
| chr9_+_122370523 | 0.82 |
ENST00000643810.1
ENST00000540753.6 |
PTGS1
|
prostaglandin-endoperoxide synthase 1 |
| chr12_-_10826358 | 0.80 |
ENST00000240619.2
|
TAS2R10
|
taste 2 receptor member 10 |
| chr2_+_90038848 | 0.75 |
ENST00000390270.2
|
IGKV3D-20
|
immunoglobulin kappa variable 3D-20 |
| chr12_+_26195313 | 0.70 |
ENST00000422622.3
|
SSPN
|
sarcospan |
| chr15_+_22015233 | 0.70 |
ENST00000639059.1
ENST00000640156.1 |
ENSG00000285472.1
ENSG00000284500.3
|
novel protein novel transcript |
| chr4_-_39032343 | 0.69 |
ENST00000381938.4
|
TMEM156
|
transmembrane protein 156 |
| chr14_+_30577752 | 0.66 |
ENST00000547532.5
ENST00000555429.1 |
G2E3
|
G2/M-phase specific E3 ubiquitin protein ligase |
| chrX_-_18672101 | 0.66 |
ENST00000379984.4
|
RS1
|
retinoschisin 1 |
| chr7_-_87713287 | 0.65 |
ENST00000416177.1
ENST00000265724.8 ENST00000543898.5 |
ABCB1
|
ATP binding cassette subfamily B member 1 |
| chr12_+_26195647 | 0.65 |
ENST00000535504.1
|
SSPN
|
sarcospan |
| chr14_-_67412112 | 0.64 |
ENST00000216446.9
|
PLEK2
|
pleckstrin 2 |
| chr12_+_26195543 | 0.60 |
ENST00000242729.7
|
SSPN
|
sarcospan |
| chr5_+_151020438 | 0.59 |
ENST00000622181.4
ENST00000614343.4 ENST00000388825.9 ENST00000521650.5 ENST00000517973.1 |
GPX3
|
glutathione peroxidase 3 |
| chr17_-_9791586 | 0.59 |
ENST00000571134.2
|
DHRS7C
|
dehydrogenase/reductase 7C |
| chr10_-_32935511 | 0.57 |
ENST00000423113.5
|
ITGB1
|
integrin subunit beta 1 |
| chr14_+_103385450 | 0.57 |
ENST00000416682.6
|
MARK3
|
microtubule affinity regulating kinase 3 |
| chr7_-_111392915 | 0.54 |
ENST00000450877.5
|
IMMP2L
|
inner mitochondrial membrane peptidase subunit 2 |
| chr7_+_107583919 | 0.54 |
ENST00000491150.5
|
BCAP29
|
B cell receptor associated protein 29 |
| chr1_-_113871665 | 0.53 |
ENST00000528414.5
ENST00000460620.5 ENST00000359785.10 ENST00000420377.6 ENST00000525799.1 ENST00000538253.5 |
PTPN22
|
protein tyrosine phosphatase non-receptor type 22 |
| chr10_-_104085847 | 0.52 |
ENST00000648076.2
|
COL17A1
|
collagen type XVII alpha 1 chain |
| chrX_-_101617921 | 0.52 |
ENST00000361910.9
ENST00000538627.5 ENST00000539247.5 |
ARMCX6
|
armadillo repeat containing X-linked 6 |
| chr4_+_182448995 | 0.51 |
ENST00000510504.1
|
TENM3
|
teneurin transmembrane protein 3 |
| chr14_+_34993240 | 0.50 |
ENST00000677647.1
|
SRP54
|
signal recognition particle 54 |
| chr2_+_90172802 | 0.50 |
ENST00000390277.3
|
IGKV3D-11
|
immunoglobulin kappa variable 3D-11 |
| chr2_-_89027700 | 0.48 |
ENST00000483158.1
|
IGKV3-11
|
immunoglobulin kappa variable 3-11 |
| chr2_+_102337148 | 0.48 |
ENST00000311734.6
ENST00000409584.5 |
IL1RL1
|
interleukin 1 receptor like 1 |
| chr1_-_201171545 | 0.47 |
ENST00000367333.6
|
TMEM9
|
transmembrane protein 9 |
| chr20_-_35147285 | 0.47 |
ENST00000374491.3
ENST00000374492.8 |
EDEM2
|
ER degradation enhancing alpha-mannosidase like protein 2 |
| chr9_+_121567057 | 0.46 |
ENST00000394340.7
ENST00000436835.5 ENST00000259371.6 |
DAB2IP
|
DAB2 interacting protein |
| chr3_-_20012250 | 0.45 |
ENST00000389050.5
|
PP2D1
|
protein phosphatase 2C like domain containing 1 |
| chrX_+_154304923 | 0.44 |
ENST00000426989.5
ENST00000426203.5 ENST00000369912.2 |
TKTL1
|
transketolase like 1 |
| chr9_-_74952904 | 0.43 |
ENST00000376854.6
|
C9orf40
|
chromosome 9 open reading frame 40 |
| chr12_-_11134644 | 0.42 |
ENST00000539585.1
|
TAS2R30
|
taste 2 receptor member 30 |
| chr3_-_185821092 | 0.42 |
ENST00000421047.3
|
IGF2BP2
|
insulin like growth factor 2 mRNA binding protein 2 |
| chr20_-_57690624 | 0.41 |
ENST00000414037.5
|
PMEPA1
|
prostate transmembrane protein, androgen induced 1 |
| chr1_+_117001744 | 0.41 |
ENST00000256652.8
ENST00000682167.1 ENST00000369470.1 |
CD101
|
CD101 molecule |
| chr2_+_186694007 | 0.40 |
ENST00000304698.10
|
FAM171B
|
family with sequence similarity 171 member B |
| chr12_-_9869345 | 0.40 |
ENST00000228438.3
|
CLEC2B
|
C-type lectin domain family 2 member B |
| chr1_-_157700738 | 0.40 |
ENST00000368186.9
ENST00000496769.1 ENST00000368184.8 |
FCRL3
|
Fc receptor like 3 |
| chr17_+_41966787 | 0.39 |
ENST00000393892.8
ENST00000587679.1 |
CNP
|
2',3'-cyclic nucleotide 3' phosphodiesterase |
| chr7_-_44541262 | 0.39 |
ENST00000289547.8
ENST00000546276.5 ENST00000423141.1 |
NPC1L1
|
NPC1 like intracellular cholesterol transporter 1 |
| chr17_+_76467597 | 0.38 |
ENST00000392492.8
|
AANAT
|
aralkylamine N-acetyltransferase |
| chr14_+_103385374 | 0.38 |
ENST00000678179.1
ENST00000676938.1 ENST00000678619.1 ENST00000440884.7 ENST00000560417.6 ENST00000679330.1 ENST00000556744.2 ENST00000676897.1 ENST00000677560.1 ENST00000561314.6 ENST00000677829.1 ENST00000677133.1 ENST00000676645.1 ENST00000678175.1 ENST00000429436.7 ENST00000677360.1 ENST00000678237.1 ENST00000677347.1 ENST00000677432.1 |
MARK3
|
microtubule affinity regulating kinase 3 |
| chr12_-_21910853 | 0.38 |
ENST00000544039.5
|
ABCC9
|
ATP binding cassette subfamily C member 9 |
| chr12_+_12357418 | 0.37 |
ENST00000298571.6
|
BORCS5
|
BLOC-1 related complex subunit 5 |
| chr17_+_41966814 | 0.37 |
ENST00000393888.1
ENST00000441615.2 |
CNP
|
2',3'-cyclic nucleotide 3' phosphodiesterase |
| chr2_+_101839815 | 0.37 |
ENST00000421882.5
|
MAP4K4
|
mitogen-activated protein kinase kinase kinase kinase 4 |
| chr17_-_74776323 | 0.37 |
ENST00000582870.5
ENST00000581136.5 ENST00000579218.5 ENST00000583476.5 ENST00000580301.5 ENST00000583757.5 ENST00000357814.8 ENST00000582524.5 |
NAT9
|
N-acetyltransferase 9 (putative) |
| chr3_-_33645433 | 0.36 |
ENST00000635664.1
ENST00000485378.6 ENST00000313350.10 ENST00000487200.5 |
CLASP2
|
cytoplasmic linker associated protein 2 |
| chr6_-_111793871 | 0.36 |
ENST00000368667.6
|
FYN
|
FYN proto-oncogene, Src family tyrosine kinase |
| chrX_+_108045050 | 0.36 |
ENST00000458383.1
ENST00000217957.10 |
VSIG1
|
V-set and immunoglobulin domain containing 1 |
| chr17_-_445939 | 0.34 |
ENST00000329099.4
|
RFLNB
|
refilin B |
| chr17_+_58755821 | 0.34 |
ENST00000308249.4
|
PPM1E
|
protein phosphatase, Mg2+/Mn2+ dependent 1E |
| chr3_+_12287899 | 0.34 |
ENST00000643888.2
|
PPARG
|
peroxisome proliferator activated receptor gamma |
| chr13_-_61415508 | 0.34 |
ENST00000409204.4
|
PCDH20
|
protocadherin 20 |
| chr6_+_29111560 | 0.34 |
ENST00000377169.2
|
OR2J3
|
olfactory receptor family 2 subfamily J member 3 |
| chr2_-_25168571 | 0.32 |
ENST00000264708.7
ENST00000449220.1 ENST00000395826.7 |
POMC
|
proopiomelanocortin |
| chr20_+_45416551 | 0.32 |
ENST00000639292.1
|
PIGT
|
phosphatidylinositol glycan anchor biosynthesis class T |
| chrX_+_108044967 | 0.32 |
ENST00000415430.7
|
VSIG1
|
V-set and immunoglobulin domain containing 1 |
| chr12_-_95116967 | 0.32 |
ENST00000551521.5
|
FGD6
|
FYVE, RhoGEF and PH domain containing 6 |
| chr3_+_12287859 | 0.31 |
ENST00000309576.11
ENST00000397015.7 |
PPARG
|
peroxisome proliferator activated receptor gamma |
| chr2_-_20823048 | 0.31 |
ENST00000402479.6
ENST00000237822.8 ENST00000626491.2 ENST00000432947.1 ENST00000403006.6 ENST00000419825.2 ENST00000381090.7 ENST00000412261.5 ENST00000619656.4 ENST00000541941.5 ENST00000440866.6 ENST00000435420.6 |
LDAH
|
lipid droplet associated hydrolase |
| chr10_+_52128343 | 0.31 |
ENST00000672084.1
|
PRKG1
|
protein kinase cGMP-dependent 1 |
| chr2_+_90234809 | 0.31 |
ENST00000443397.5
|
IGKV3D-7
|
immunoglobulin kappa variable 3D-7 |
| chr19_-_35812838 | 0.29 |
ENST00000653904.2
|
PRODH2
|
proline dehydrogenase 2 |
| chr4_+_118888918 | 0.29 |
ENST00000434046.6
|
SYNPO2
|
synaptopodin 2 |
| chr20_+_58907981 | 0.28 |
ENST00000656419.1
|
GNAS
|
GNAS complex locus |
| chr4_+_94207596 | 0.28 |
ENST00000359052.8
|
SMARCAD1
|
SWI/SNF-related, matrix-associated actin-dependent regulator of chromatin, subfamily a, containing DEAD/H box 1 |
| chr11_-_117316230 | 0.28 |
ENST00000313005.11
ENST00000528053.5 |
BACE1
|
beta-secretase 1 |
| chr2_+_181986015 | 0.28 |
ENST00000409702.1
|
PPP1R1C
|
protein phosphatase 1 regulatory inhibitor subunit 1C |
| chr1_+_157993273 | 0.27 |
ENST00000360089.8
ENST00000368173.7 |
KIRREL1
|
kirre like nephrin family adhesion molecule 1 |
| chr12_-_56741535 | 0.27 |
ENST00000647707.1
|
ENSG00000285625.1
|
novel protein |
| chr10_-_93482194 | 0.27 |
ENST00000358334.9
ENST00000371488.3 |
MYOF
|
myoferlin |
| chr10_-_93482326 | 0.27 |
ENST00000359263.9
|
MYOF
|
myoferlin |
| chr12_-_54259531 | 0.26 |
ENST00000550411.5
ENST00000439541.6 |
CBX5
|
chromobox 5 |
| chr1_-_197146620 | 0.26 |
ENST00000367409.9
ENST00000680265.1 |
ASPM
|
assembly factor for spindle microtubules |
| chr17_-_66229380 | 0.26 |
ENST00000205948.11
|
APOH
|
apolipoprotein H |
| chrX_+_80420466 | 0.26 |
ENST00000308293.5
|
TENT5D
|
terminal nucleotidyltransferase 5D |
| chr5_+_127649018 | 0.26 |
ENST00000379445.7
|
CTXN3
|
cortexin 3 |
| chr13_-_35476682 | 0.26 |
ENST00000379919.6
|
MAB21L1
|
mab-21 like 1 |
| chr9_-_28670285 | 0.25 |
ENST00000379992.6
ENST00000308675.5 ENST00000613945.3 |
LINGO2
|
leucine rich repeat and Ig domain containing 2 |
| chr15_-_19988117 | 0.25 |
ENST00000558565.2
|
IGHV3OR15-7
|
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr12_-_118359639 | 0.25 |
ENST00000541786.5
ENST00000419821.6 ENST00000541878.5 |
TAOK3
|
TAO kinase 3 |
| chr4_-_39977836 | 0.25 |
ENST00000303538.13
ENST00000503396.5 |
PDS5A
|
PDS5 cohesin associated factor A |
| chr18_+_63587297 | 0.24 |
ENST00000269489.9
|
SERPINB13
|
serpin family B member 13 |
| chr6_+_47781982 | 0.24 |
ENST00000489301.6
ENST00000638973.1 ENST00000371211.6 ENST00000393699.2 |
OPN5
|
opsin 5 |
| chr2_-_89085787 | 0.24 |
ENST00000390252.2
|
IGKV3-15
|
immunoglobulin kappa variable 3-15 |
| chr18_+_63587336 | 0.23 |
ENST00000344731.10
|
SERPINB13
|
serpin family B member 13 |
| chr12_-_111685720 | 0.23 |
ENST00000327551.6
|
BRAP
|
BRCA1 associated protein |
| chr4_-_119322128 | 0.22 |
ENST00000274024.4
|
FABP2
|
fatty acid binding protein 2 |
| chr16_-_66550005 | 0.22 |
ENST00000527284.6
|
TK2
|
thymidine kinase 2 |
| chr16_-_66549839 | 0.21 |
ENST00000527800.6
ENST00000677555.1 ENST00000563369.6 |
TK2
|
thymidine kinase 2 |
| chr18_-_36798482 | 0.21 |
ENST00000590258.2
|
TPGS2
|
tubulin polyglutamylase complex subunit 2 |
| chr2_+_181985846 | 0.21 |
ENST00000682840.1
ENST00000409137.7 ENST00000280295.7 |
PPP1R1C
|
protein phosphatase 1 regulatory inhibitor subunit 1C |
| chr18_+_23689439 | 0.21 |
ENST00000313654.14
|
LAMA3
|
laminin subunit alpha 3 |
| chr18_-_49492305 | 0.21 |
ENST00000615479.4
ENST00000583637.5 ENST00000618613.5 ENST00000615760.4 ENST00000578528.1 ENST00000578532.5 ENST00000580387.5 ENST00000579248.5 ENST00000580261.6 ENST00000581373.5 ENST00000618619.4 ENST00000617346.4 ENST00000583036.5 ENST00000332968.11 |
RPL17
RPL17-C18orf32
|
ribosomal protein L17 RPL17-C18orf32 readthrough |
| chr11_+_94768331 | 0.21 |
ENST00000317829.12
ENST00000433060.3 |
AMOTL1
|
angiomotin like 1 |
| chr19_+_53962925 | 0.21 |
ENST00000270458.4
|
CACNG8
|
calcium voltage-gated channel auxiliary subunit gamma 8 |
| chr19_-_51417791 | 0.20 |
ENST00000353836.9
|
SIGLEC10
|
sialic acid binding Ig like lectin 10 |
| chr5_+_154941063 | 0.20 |
ENST00000523037.6
ENST00000265229.12 ENST00000439747.7 ENST00000522038.5 |
MRPL22
|
mitochondrial ribosomal protein L22 |
| chr15_+_64387828 | 0.20 |
ENST00000261884.8
|
TRIP4
|
thyroid hormone receptor interactor 4 |
| chr3_+_111999326 | 0.20 |
ENST00000494932.1
|
TAGLN3
|
transgelin 3 |
| chr4_+_70226116 | 0.20 |
ENST00000317987.6
|
FDCSP
|
follicular dendritic cell secreted protein |
| chr12_+_8157034 | 0.20 |
ENST00000396570.7
|
ZNF705A
|
zinc finger protein 705A |
| chr14_-_106211453 | 0.19 |
ENST00000390606.3
|
IGHV3-20
|
immunoglobulin heavy variable 3-20 |
| chr2_-_25168903 | 0.19 |
ENST00000405623.5
|
POMC
|
proopiomelanocortin |
| chr2_-_25168690 | 0.19 |
ENST00000380794.5
|
POMC
|
proopiomelanocortin |
| chr3_+_44799187 | 0.19 |
ENST00000425755.5
|
KIF15
|
kinesin family member 15 |
| chr11_+_60056587 | 0.19 |
ENST00000395032.6
ENST00000358152.6 |
MS4A3
|
membrane spanning 4-domains A3 |
| chr16_+_11965234 | 0.19 |
ENST00000562385.1
|
TNFRSF17
|
TNF receptor superfamily member 17 |
| chr16_-_66550091 | 0.19 |
ENST00000564917.5
ENST00000677420.1 |
TK2
|
thymidine kinase 2 |
| chr1_-_211134135 | 0.19 |
ENST00000638983.1
ENST00000271751.10 ENST00000639952.1 |
ENSG00000284299.1
KCNH1
|
novel protein potassium voltage-gated channel subfamily H member 1 |
| chr2_+_68734773 | 0.18 |
ENST00000409202.8
|
ARHGAP25
|
Rho GTPase activating protein 25 |
| chr18_+_23689609 | 0.18 |
ENST00000399516.7
|
LAMA3
|
laminin subunit alpha 3 |
| chr2_-_162152404 | 0.18 |
ENST00000375497.3
|
GCG
|
glucagon |
| chr11_+_60056653 | 0.18 |
ENST00000278865.8
|
MS4A3
|
membrane spanning 4-domains A3 |
| chr10_+_18260715 | 0.18 |
ENST00000615785.4
ENST00000617363.4 ENST00000396576.6 |
CACNB2
|
calcium voltage-gated channel auxiliary subunit beta 2 |
| chr15_+_94355956 | 0.17 |
ENST00000557742.1
|
MCTP2
|
multiple C2 and transmembrane domain containing 2 |
| chr9_+_34652167 | 0.17 |
ENST00000441545.7
ENST00000553620.5 |
IL11RA
|
interleukin 11 receptor subunit alpha |
| chr7_+_116222804 | 0.17 |
ENST00000393481.6
|
TES
|
testin LIM domain protein |
| chr19_-_51417700 | 0.17 |
ENST00000529627.1
ENST00000439889.6 |
SIGLEC10
|
sialic acid binding Ig like lectin 10 |
| chr8_+_12104389 | 0.16 |
ENST00000400085.7
|
ZNF705D
|
zinc finger protein 705D |
| chr6_-_82247697 | 0.16 |
ENST00000306270.12
ENST00000610980.4 |
IBTK
|
inhibitor of Bruton tyrosine kinase |
| chr19_-_43883964 | 0.16 |
ENST00000587539.2
|
ZNF404
|
zinc finger protein 404 |
| chr3_+_45886501 | 0.16 |
ENST00000395963.2
|
CCR9
|
C-C motif chemokine receptor 9 |
| chr7_-_22822829 | 0.16 |
ENST00000358435.9
ENST00000621567.1 |
TOMM7
|
translocase of outer mitochondrial membrane 7 |
| chr9_-_26947222 | 0.16 |
ENST00000520884.5
ENST00000397292.8 |
PLAA
|
phospholipase A2 activating protein |
| chr9_-_111036207 | 0.16 |
ENST00000541779.5
|
LPAR1
|
lysophosphatidic acid receptor 1 |
| chr7_-_22822779 | 0.16 |
ENST00000372879.8
|
TOMM7
|
translocase of outer mitochondrial membrane 7 |
| chr1_-_109075944 | 0.16 |
ENST00000338366.6
|
TAF13
|
TATA-box binding protein associated factor 13 |
| chrX_-_72239022 | 0.16 |
ENST00000373657.2
ENST00000334463.4 |
ERCC6L
|
ERCC excision repair 6 like, spindle assembly checkpoint helicase |
| chr3_+_111998739 | 0.16 |
ENST00000393917.6
ENST00000273368.8 |
TAGLN3
|
transgelin 3 |
| chr8_-_85341705 | 0.15 |
ENST00000517618.5
|
CA1
|
carbonic anhydrase 1 |
| chr14_+_71586261 | 0.15 |
ENST00000358550.6
|
SIPA1L1
|
signal induced proliferation associated 1 like 1 |
| chr20_-_7940444 | 0.15 |
ENST00000378789.4
|
HAO1
|
hydroxyacid oxidase 1 |
| chr6_+_26087417 | 0.15 |
ENST00000357618.10
ENST00000309234.10 |
HFE
|
homeostatic iron regulator |
| chr8_-_48921419 | 0.14 |
ENST00000020945.4
|
SNAI2
|
snail family transcriptional repressor 2 |
| chr1_-_158554405 | 0.14 |
ENST00000641282.1
ENST00000641622.1 |
OR6Y1
|
olfactory receptor family 6 subfamily Y member 1 |
| chr1_-_247760556 | 0.14 |
ENST00000641256.1
|
OR1C1
|
olfactory receptor family 1 subfamily C member 1 |
| chr3_+_130850585 | 0.14 |
ENST00000505330.5
ENST00000504381.5 ENST00000507488.6 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr4_-_76023489 | 0.14 |
ENST00000306602.3
|
CXCL10
|
C-X-C motif chemokine ligand 10 |
| chr7_-_122098831 | 0.14 |
ENST00000681213.1
ENST00000679419.1 |
AASS
|
aminoadipate-semialdehyde synthase |
| chr10_-_49188380 | 0.14 |
ENST00000374153.7
ENST00000374148.1 ENST00000374151.7 |
TMEM273
|
transmembrane protein 273 |
| chr3_+_111999189 | 0.14 |
ENST00000455401.6
|
TAGLN3
|
transgelin 3 |
| chr18_+_57352541 | 0.14 |
ENST00000324000.4
|
ST8SIA3
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 |
| chr9_+_69145463 | 0.14 |
ENST00000636438.1
|
TJP2
|
tight junction protein 2 |
| chr5_+_90474879 | 0.14 |
ENST00000504930.5
ENST00000514483.5 |
POLR3G
|
RNA polymerase III subunit G |
| chr5_+_67004618 | 0.14 |
ENST00000261569.11
ENST00000436277.5 |
MAST4
|
microtubule associated serine/threonine kinase family member 4 |
| chr4_-_151227881 | 0.13 |
ENST00000652233.1
ENST00000514152.5 |
SH3D19
|
SH3 domain containing 19 |
| chr5_+_90474848 | 0.13 |
ENST00000651687.1
|
POLR3G
|
RNA polymerase III subunit G |
| chr11_-_89808575 | 0.13 |
ENST00000329758.5
|
TRIM49
|
tripartite motif containing 49 |
| chr4_+_87650277 | 0.13 |
ENST00000339673.11
ENST00000282479.8 |
DMP1
|
dentin matrix acidic phosphoprotein 1 |
| chr19_-_3772211 | 0.13 |
ENST00000555978.5
ENST00000555633.3 |
RAX2
|
retina and anterior neural fold homeobox 2 |
| chr11_-_66002123 | 0.13 |
ENST00000532707.5
ENST00000526451.5 ENST00000312234.6 ENST00000533544.6 ENST00000530462.5 ENST00000525767.5 ENST00000529964.5 ENST00000527249.5 |
EIF1AD
|
eukaryotic translation initiation factor 1A domain containing |
| chr15_+_92900338 | 0.12 |
ENST00000625990.3
|
CHD2
|
chromodomain helicase DNA binding protein 2 |
| chr8_-_85341659 | 0.12 |
ENST00000522389.5
|
CA1
|
carbonic anhydrase 1 |
| chr6_-_155455830 | 0.12 |
ENST00000159060.3
|
NOX3
|
NADPH oxidase 3 |
| chr3_-_151316795 | 0.12 |
ENST00000260843.5
|
GPR87
|
G protein-coupled receptor 87 |
| chr7_-_150323489 | 0.12 |
ENST00000683684.1
ENST00000478393.5 |
ACTR3C
|
actin related protein 3C |
| chr4_+_154563003 | 0.12 |
ENST00000302068.9
ENST00000509493.1 |
FGB
|
fibrinogen beta chain |
| chr1_+_10450004 | 0.12 |
ENST00000377049.4
|
CORT
|
cortistatin |
| chr8_-_61689768 | 0.12 |
ENST00000517847.6
ENST00000389204.8 ENST00000517661.5 ENST00000517903.5 ENST00000522603.5 ENST00000541428.5 ENST00000522349.5 ENST00000522835.5 ENST00000518306.5 |
ASPH
|
aspartate beta-hydroxylase |
| chr10_-_73655984 | 0.11 |
ENST00000394810.3
|
SYNPO2L
|
synaptopodin 2 like |
| chr10_+_120851341 | 0.11 |
ENST00000263461.11
|
WDR11
|
WD repeat domain 11 |
| chrX_-_101407893 | 0.11 |
ENST00000676156.1
ENST00000675592.1 ENST00000674634.2 ENST00000649178.1 ENST00000218516.4 |
GLA
|
galactosidase alpha |
| chr13_-_37598750 | 0.11 |
ENST00000379743.8
ENST00000379742.4 ENST00000379749.8 ENST00000379747.9 ENST00000541179.5 ENST00000541481.5 |
POSTN
|
periostin |
| chr2_-_74392025 | 0.11 |
ENST00000440727.1
ENST00000409240.5 |
DCTN1
|
dynactin subunit 1 |
| chr6_+_124983356 | 0.10 |
ENST00000519799.5
ENST00000368414.6 ENST00000359704.2 |
RNF217
|
ring finger protein 217 |
| chr20_+_59019397 | 0.10 |
ENST00000217133.2
|
TUBB1
|
tubulin beta 1 class VI |
| chr3_+_111998915 | 0.10 |
ENST00000478951.6
|
TAGLN3
|
transgelin 3 |
| chr7_-_87475839 | 0.10 |
ENST00000359206.8
|
ABCB4
|
ATP binding cassette subfamily B member 4 |
| chr2_+_165239388 | 0.10 |
ENST00000424833.5
ENST00000375437.7 ENST00000631182.3 |
SCN2A
|
sodium voltage-gated channel alpha subunit 2 |
| chr21_-_14210948 | 0.10 |
ENST00000681601.1
|
LIPI
|
lipase I |
| chr11_-_102705737 | 0.10 |
ENST00000260229.5
|
MMP27
|
matrix metallopeptidase 27 |
| chrM_-_14669 | 0.10 |
ENST00000361681.2
|
MT-ND6
|
mitochondrially encoded NADH:ubiquinone oxidoreductase core subunit 6 |
| chr9_-_128724088 | 0.10 |
ENST00000406904.2
ENST00000452105.5 ENST00000372667.9 ENST00000372663.9 |
ZDHHC12
|
zinc finger DHHC-type palmitoyltransferase 12 |
| chr1_-_111488795 | 0.10 |
ENST00000472933.2
|
TMIGD3
|
transmembrane and immunoglobulin domain containing 3 |
| chr10_+_18400562 | 0.10 |
ENST00000377315.5
ENST00000650685.1 |
CACNB2
|
calcium voltage-gated channel auxiliary subunit beta 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 5.5 | GO:0060005 | vestibular reflex(GO:0060005) |
| 0.2 | 0.9 | GO:0031456 | glycine betaine biosynthetic process from choline(GO:0019285) glycine betaine metabolic process(GO:0031455) glycine betaine biosynthetic process(GO:0031456) |
| 0.2 | 3.0 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.2 | 0.5 | GO:0061300 | cerebellum vasculature development(GO:0061300) |
| 0.2 | 0.8 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.2 | 0.5 | GO:0036510 | trimming of terminal mannose on C branch(GO:0036510) |
| 0.1 | 3.1 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.1 | 1.0 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.1 | 0.4 | GO:0030187 | melatonin metabolic process(GO:0030186) melatonin biosynthetic process(GO:0030187) |
| 0.1 | 0.5 | GO:0006617 | SRP-dependent cotranslational protein targeting to membrane, signal sequence recognition(GO:0006617) |
| 0.1 | 1.9 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.1 | 0.3 | GO:0061182 | negative regulation of chondrocyte development(GO:0061182) |
| 0.1 | 0.5 | GO:1902523 | negative regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070425) negative regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070433) positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 0.1 | 0.7 | GO:2000852 | regulation of corticosterone secretion(GO:2000852) |
| 0.1 | 0.3 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.1 | 0.7 | GO:2000230 | negative regulation of pancreatic stellate cell proliferation(GO:2000230) |
| 0.1 | 0.6 | GO:0006982 | response to lipid hydroperoxide(GO:0006982) |
| 0.1 | 0.5 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.1 | 0.4 | GO:0006772 | thiamine metabolic process(GO:0006772) |
| 0.1 | 1.8 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.1 | 0.7 | GO:1901529 | positive regulation of anion channel activity(GO:1901529) |
| 0.1 | 0.6 | GO:0007161 | calcium-independent cell-matrix adhesion(GO:0007161) regulation of collagen catabolic process(GO:0010710) |
| 0.1 | 0.2 | GO:1900390 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) negative regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002590) regulation of T cell antigen processing and presentation(GO:0002625) positive regulation of iron ion transport(GO:0034758) positive regulation of iron ion transmembrane transport(GO:0034761) regulation of iron ion import(GO:1900390) regulation of ferrous iron import into cell(GO:1903989) positive regulation of ferrous iron import into cell(GO:1903991) regulation of ferrous iron binding(GO:1904432) positive regulation of ferrous iron binding(GO:1904434) regulation of transferrin receptor binding(GO:1904435) positive regulation of transferrin receptor binding(GO:1904437) regulation of ferrous iron import across plasma membrane(GO:1904438) positive regulation of ferrous iron import across plasma membrane(GO:1904440) response to iron ion starvation(GO:1990641) |
| 0.1 | 0.9 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.1 | 0.5 | GO:0036324 | vascular endothelial growth factor receptor-2 signaling pathway(GO:0036324) |
| 0.1 | 0.4 | GO:0046092 | deoxycytidine metabolic process(GO:0046092) |
| 0.1 | 0.5 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 0.1 | 0.3 | GO:0009786 | regulation of asymmetric cell division(GO:0009786) |
| 0.0 | 0.1 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 0.0 | 0.1 | GO:0061011 | hepatic duct development(GO:0061011) |
| 0.0 | 0.1 | GO:1901420 | negative regulation of response to alcohol(GO:1901420) |
| 0.0 | 0.1 | GO:0019878 | lysine biosynthetic process(GO:0009085) lysine biosynthetic process via aminoadipic acid(GO:0019878) |
| 0.0 | 0.1 | GO:0002428 | antigen processing and presentation of peptide antigen via MHC class Ib(GO:0002428) |
| 0.0 | 0.5 | GO:0097283 | keratinocyte apoptotic process(GO:0097283) regulation of keratinocyte apoptotic process(GO:1902172) |
| 0.0 | 0.1 | GO:0002305 | gamma-delta intraepithelial T cell differentiation(GO:0002304) CD8-positive, gamma-delta intraepithelial T cell differentiation(GO:0002305) |
| 0.0 | 0.3 | GO:0060087 | relaxation of vascular smooth muscle(GO:0060087) |
| 0.0 | 0.2 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 0.0 | 0.3 | GO:0045040 | protein import into mitochondrial outer membrane(GO:0045040) |
| 0.0 | 0.3 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.0 | 0.7 | GO:0009642 | response to light intensity(GO:0009642) |
| 0.0 | 0.1 | GO:2001025 | response to cyclosporin A(GO:1905237) positive regulation of response to drug(GO:2001025) |
| 0.0 | 0.9 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.7 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.0 | 0.3 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 0.2 | GO:1904565 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 0.0 | 1.2 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.0 | 0.9 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 0.4 | GO:1902951 | negative regulation of dendritic spine maintenance(GO:1902951) |
| 0.0 | 0.5 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.0 | 0.1 | GO:1990523 | bone regeneration(GO:1990523) |
| 0.0 | 0.3 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.0 | 0.4 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.0 | 0.4 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.0 | 0.5 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.1 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.0 | 0.1 | GO:0001868 | regulation of complement activation, lectin pathway(GO:0001868) negative regulation of complement activation, lectin pathway(GO:0001869) |
| 0.0 | 0.1 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.0 | 0.3 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.0 | 0.1 | GO:1900222 | regulation of cell growth by extracellular stimulus(GO:0001560) negative regulation of beta-amyloid clearance(GO:1900222) positive regulation of cardiac vascular smooth muscle cell differentiation(GO:2000724) |
| 0.0 | 0.3 | GO:0034392 | negative regulation of smooth muscle cell apoptotic process(GO:0034392) negative regulation of fibrinolysis(GO:0051918) |
| 0.0 | 1.1 | GO:0009954 | proximal/distal pattern formation(GO:0009954) |
| 0.0 | 0.1 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 0.0 | 0.1 | GO:0002769 | natural killer cell inhibitory signaling pathway(GO:0002769) |
| 0.0 | 1.1 | GO:1900739 | regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900739) positive regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway(GO:1900740) |
| 0.0 | 0.4 | GO:0098856 | intestinal cholesterol absorption(GO:0030299) intestinal lipid absorption(GO:0098856) |
| 0.0 | 0.3 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.0 | 0.3 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.0 | 0.2 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.0 | 0.1 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.0 | 0.1 | GO:0048631 | regulation of skeletal muscle tissue growth(GO:0048631) negative regulation of satellite cell differentiation(GO:1902725) |
| 0.0 | 0.0 | GO:2000296 | negative regulation of hydrogen peroxide catabolic process(GO:2000296) |
| 0.0 | 0.1 | GO:0048840 | otolith development(GO:0048840) |
| 0.0 | 0.8 | GO:0021762 | substantia nigra development(GO:0021762) |
| 0.0 | 0.1 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.0 | 0.1 | GO:0043950 | positive regulation of cAMP-mediated signaling(GO:0043950) |
| 0.0 | 0.2 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 0.1 | GO:0071847 | TNFSF11-mediated signaling pathway(GO:0071847) |
| 0.0 | 0.3 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.0 | 0.1 | GO:0048669 | microglia differentiation(GO:0014004) microglia development(GO:0014005) collateral sprouting in absence of injury(GO:0048669) smooth endoplasmic reticulum calcium ion homeostasis(GO:0051563) cellular response to norepinephrine stimulus(GO:0071874) |
| 0.0 | 0.1 | GO:0016139 | glycoside catabolic process(GO:0016139) |
| 0.0 | 0.1 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.0 | 1.5 | GO:0030449 | regulation of complement activation(GO:0030449) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 5.5 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.3 | 0.9 | GO:0001674 | female germ cell nucleus(GO:0001674) |
| 0.2 | 0.6 | GO:0034677 | integrin alpha7-beta1 complex(GO:0034677) |
| 0.1 | 0.5 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.1 | 0.5 | GO:1990032 | parallel fiber(GO:1990032) |
| 0.1 | 0.6 | GO:0005873 | plus-end kinesin complex(GO:0005873) |
| 0.1 | 0.3 | GO:0070931 | Golgi-associated vesicle lumen(GO:0070931) |
| 0.1 | 1.0 | GO:0030478 | actin cap(GO:0030478) |
| 0.1 | 0.2 | GO:1990357 | terminal web(GO:1990357) |
| 0.1 | 0.4 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.1 | 0.8 | GO:0035749 | myelin sheath adaxonal region(GO:0035749) |
| 0.1 | 0.3 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.0 | 0.4 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.0 | 1.9 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 0.5 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) |
| 0.0 | 1.8 | GO:0002102 | podosome(GO:0002102) |
| 0.0 | 0.3 | GO:0036449 | microtubule minus-end(GO:0036449) meiotic spindle(GO:0072687) |
| 0.0 | 0.1 | GO:0071821 | FANCM-MHF complex(GO:0071821) |
| 0.0 | 0.3 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.4 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.0 | 2.9 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 0.5 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.3 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.1 | GO:0071942 | XPC complex(GO:0071942) |
| 0.0 | 0.5 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.0 | 0.3 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.0 | 0.3 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.1 | GO:0042825 | TAP complex(GO:0042825) |
| 0.0 | 0.7 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.0 | 0.1 | GO:0000444 | MIS12/MIND type complex(GO:0000444) |
| 0.0 | 0.1 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 0.0 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.0 | 0.1 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 0.3 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 3.1 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.4 | 5.5 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.2 | 0.7 | GO:0090555 | phosphatidylethanolamine-translocating ATPase activity(GO:0090555) |
| 0.2 | 1.0 | GO:0004558 | alpha-1,4-glucosidase activity(GO:0004558) |
| 0.2 | 1.9 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.2 | 0.5 | GO:0002113 | interleukin-33 binding(GO:0002113) |
| 0.1 | 0.8 | GO:0070996 | type 1 melanocortin receptor binding(GO:0070996) |
| 0.1 | 0.4 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.1 | 0.4 | GO:0008281 | sulfonylurea receptor activity(GO:0008281) |
| 0.1 | 0.4 | GO:0004802 | transketolase activity(GO:0004802) |
| 0.1 | 0.4 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 0.1 | 0.6 | GO:0098639 | collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.1 | 0.3 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.1 | 0.5 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) |
| 0.1 | 2.1 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.1 | 0.5 | GO:0035662 | Toll-like receptor 4 binding(GO:0035662) |
| 0.1 | 0.5 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.1 | 0.3 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.1 | 0.7 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 0.4 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.1 | 1.2 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.1 | 0.3 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.1 | 0.5 | GO:0004571 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) |
| 0.0 | 0.6 | GO:0008430 | selenium binding(GO:0008430) |
| 0.0 | 0.1 | GO:0052853 | (S)-2-hydroxy-acid oxidase activity(GO:0003973) very-long-chain-(S)-2-hydroxy-acid oxidase activity(GO:0052852) long-chain-(S)-2-hydroxy-long-chain-acid oxidase activity(GO:0052853) medium-chain-(S)-2-hydroxy-acid oxidase activity(GO:0052854) |
| 0.0 | 0.2 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.0 | 0.4 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.0 | 0.3 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.0 | 0.7 | GO:0070492 | phosphatidylinositol-5-phosphate binding(GO:0010314) oligosaccharide binding(GO:0070492) |
| 0.0 | 1.0 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.0 | 0.5 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.0 | 0.9 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.0 | 0.1 | GO:0004597 | peptide-aspartate beta-dioxygenase activity(GO:0004597) |
| 0.0 | 0.1 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.0 | 0.3 | GO:0004064 | arylesterase activity(GO:0004064) |
| 0.0 | 0.1 | GO:0090554 | phosphatidylcholine-translocating ATPase activity(GO:0090554) |
| 0.0 | 0.3 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.0 | 0.6 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.0 | 0.3 | GO:0015266 | protein channel activity(GO:0015266) |
| 0.0 | 0.1 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.0 | 0.0 | GO:0003918 | DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 0.0 | 0.1 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.0 | 0.3 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.0 | 0.2 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.0 | 0.1 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.0 | 0.1 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) |
| 0.0 | 0.2 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.0 | 0.1 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.0 | 0.2 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.0 | 0.4 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.0 | 0.3 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 1.0 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 0.1 | GO:0016524 | latrotoxin receptor activity(GO:0016524) |
| 0.0 | 0.4 | GO:0051010 | dystroglycan binding(GO:0002162) microtubule plus-end binding(GO:0051010) |
| 0.0 | 0.1 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.0 | 0.3 | GO:0051430 | corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.0 | 0.3 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.0 | 0.1 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.1 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 0.4 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.0 | 1.1 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 0.6 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.2 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.1 | 1.9 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.1 | 0.9 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 0.9 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.0 | 0.7 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.0 | 0.7 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.0 | 3.1 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.0 | 0.5 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.0 | 0.2 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 1.0 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.4 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.0 | 0.3 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.4 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.0 | 0.2 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.3 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |