Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
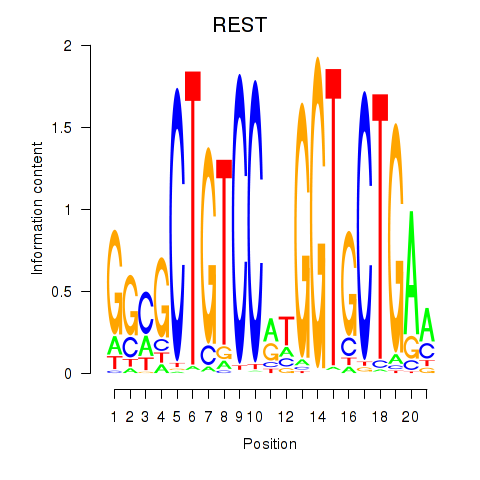
Results for REST
Z-value: 0.86
Transcription factors associated with REST
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
REST
|
ENSG00000084093.19 | REST |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| REST | hg38_v1_chr4_+_56908094_56908134 | -0.15 | 4.3e-01 | Click! |
Activity profile of REST motif
Sorted Z-values of REST motif
Network of associatons between targets according to the STRING database.
First level regulatory network of REST
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_+_63696643 | 1.85 |
ENST00000369996.3
|
TNFRSF6B
|
TNF receptor superfamily member 6b |
| chr5_+_33936386 | 1.82 |
ENST00000330120.5
|
RXFP3
|
relaxin family peptide receptor 3 |
| chr8_-_90082871 | 1.71 |
ENST00000265431.7
|
CALB1
|
calbindin 1 |
| chr9_+_132582581 | 1.60 |
ENST00000263610.7
|
BARHL1
|
BarH like homeobox 1 |
| chr19_-_51342130 | 1.50 |
ENST00000335624.5
|
VSIG10L
|
V-set and immunoglobulin domain containing 10 like |
| chr4_+_8580387 | 1.38 |
ENST00000382487.5
|
GPR78
|
G protein-coupled receptor 78 |
| chr6_+_35805347 | 1.18 |
ENST00000360215.3
|
LHFPL5
|
LHFPL tetraspan subfamily member 5 |
| chr12_+_71439976 | 1.10 |
ENST00000536515.5
ENST00000540815.2 |
LGR5
|
leucine rich repeat containing G protein-coupled receptor 5 |
| chr7_+_155070317 | 1.03 |
ENST00000287907.3
|
HTR5A
|
5-hydroxytryptamine receptor 5A |
| chr20_+_31968141 | 0.99 |
ENST00000562532.3
|
XKR7
|
XK related 7 |
| chr5_-_151924846 | 0.93 |
ENST00000274576.9
|
GLRA1
|
glycine receptor alpha 1 |
| chr5_-_151924824 | 0.90 |
ENST00000455880.2
|
GLRA1
|
glycine receptor alpha 1 |
| chrX_+_7147675 | 0.81 |
ENST00000674429.1
|
STS
|
steroid sulfatase |
| chr10_-_77638369 | 0.81 |
ENST00000372443.6
|
KCNMA1
|
potassium calcium-activated channel subfamily M alpha 1 |
| chr14_-_106627685 | 0.81 |
ENST00000390629.3
|
IGHV4-59
|
immunoglobulin heavy variable 4-59 |
| chr17_+_32486975 | 0.80 |
ENST00000313401.4
|
CDK5R1
|
cyclin dependent kinase 5 regulatory subunit 1 |
| chr3_-_42701513 | 0.78 |
ENST00000310417.9
|
HHATL
|
hedgehog acyltransferase like |
| chr12_-_29783798 | 0.78 |
ENST00000552618.5
ENST00000551659.5 ENST00000539277.6 |
TMTC1
|
transmembrane O-mannosyltransferase targeting cadherins 1 |
| chr15_-_21742799 | 0.74 |
ENST00000622410.2
|
ENSG00000278263.2
|
novel protein, identical to IGHV4-4 |
| chr11_+_62613236 | 0.73 |
ENST00000278833.4
|
ROM1
|
retinal outer segment membrane protein 1 |
| chr2_+_114442616 | 0.68 |
ENST00000410059.6
|
DPP10
|
dipeptidyl peptidase like 10 |
| chrX_+_7147237 | 0.67 |
ENST00000666110.2
|
STS
|
steroid sulfatase |
| chr14_+_92923143 | 0.65 |
ENST00000216492.10
ENST00000334654.4 |
CHGA
|
chromogranin A |
| chr4_+_157220691 | 0.65 |
ENST00000509417.5
ENST00000645636.1 ENST00000296526.12 ENST00000264426.14 |
GRIA2
|
glutamate ionotropic receptor AMPA type subunit 2 |
| chr15_-_72320149 | 0.62 |
ENST00000287202.10
|
CELF6
|
CUGBP Elav-like family member 6 |
| chr9_-_71446835 | 0.60 |
ENST00000357533.6
|
TRPM3
|
transient receptor potential cation channel subfamily M member 3 |
| chr15_-_31870651 | 0.59 |
ENST00000307050.6
ENST00000560598.2 |
OTUD7A
|
OTU deubiquitinase 7A |
| chr3_+_9703826 | 0.59 |
ENST00000613455.4
ENST00000383831.7 ENST00000383832.8 |
CPNE9
|
copine family member 9 |
| chr10_+_97849824 | 0.58 |
ENST00000370602.6
|
GOLGA7B
|
golgin A7 family member B |
| chr20_+_46029206 | 0.57 |
ENST00000243964.7
|
SLC12A5
|
solute carrier family 12 member 5 |
| chr6_+_109978297 | 0.56 |
ENST00000275169.5
|
GPR6
|
G protein-coupled receptor 6 |
| chr17_-_3398410 | 0.56 |
ENST00000322608.2
|
OR1E1
|
olfactory receptor family 1 subfamily E member 1 |
| chr6_+_109978256 | 0.54 |
ENST00000414000.3
|
GPR6
|
G protein-coupled receptor 6 |
| chr15_-_82709823 | 0.52 |
ENST00000666973.1
ENST00000664460.1 ENST00000669930.1 |
AP3B2
|
adaptor related protein complex 3 subunit beta 2 |
| chr20_+_46029165 | 0.52 |
ENST00000616201.4
ENST00000616202.4 ENST00000616933.4 ENST00000626937.2 |
SLC12A5
|
solute carrier family 12 member 5 |
| chr4_+_157220654 | 0.52 |
ENST00000393815.6
|
GRIA2
|
glutamate ionotropic receptor AMPA type subunit 2 |
| chr11_+_125903320 | 0.49 |
ENST00000525943.1
|
DDX25
|
DEAD-box helicase 25 |
| chr15_-_82709775 | 0.49 |
ENST00000535348.5
|
AP3B2
|
adaptor related protein complex 3 subunit beta 2 |
| chr12_+_113358542 | 0.48 |
ENST00000545182.6
ENST00000280800.5 |
PLBD2
|
phospholipase B domain containing 2 |
| chr2_-_51032390 | 0.48 |
ENST00000404971.5
|
NRXN1
|
neurexin 1 |
| chr2_-_51032536 | 0.47 |
ENST00000626899.1
ENST00000406316.6 |
NRXN1
|
neurexin 1 |
| chr7_-_101165114 | 0.44 |
ENST00000445482.2
|
VGF
|
VGF nerve growth factor inducible |
| chr3_-_48662877 | 0.44 |
ENST00000164024.5
|
CELSR3
|
cadherin EGF LAG seven-pass G-type receptor 3 |
| chr19_-_3971051 | 0.43 |
ENST00000545797.7
ENST00000596311.5 |
DAPK3
|
death associated protein kinase 3 |
| chr1_-_101996919 | 0.42 |
ENST00000370103.9
|
OLFM3
|
olfactomedin 3 |
| chr14_+_93430853 | 0.41 |
ENST00000553484.5
|
UNC79
|
unc-79 homolog, NALCN channel complex subunit |
| chr4_-_174829212 | 0.40 |
ENST00000340217.5
ENST00000274093.8 |
GLRA3
|
glycine receptor alpha 3 |
| chr5_+_175871570 | 0.38 |
ENST00000512824.5
ENST00000393745.8 |
CPLX2
|
complexin 2 |
| chr6_+_152750789 | 0.38 |
ENST00000367244.8
ENST00000367243.7 |
VIP
|
vasoactive intestinal peptide |
| chr15_-_82709859 | 0.38 |
ENST00000542200.2
ENST00000535359.6 ENST00000668990.2 ENST00000652847.1 |
AP3B2
|
adaptor related protein complex 3 subunit beta 2 |
| chr9_-_101594995 | 0.37 |
ENST00000636434.1
|
PPP3R2
|
protein phosphatase 3 regulatory subunit B, beta |
| chrX_-_47619850 | 0.37 |
ENST00000295987.13
ENST00000340666.5 |
SYN1
|
synapsin I |
| chr12_+_118981531 | 0.35 |
ENST00000267260.5
|
SRRM4
|
serine/arginine repetitive matrix 4 |
| chr2_-_51032151 | 0.35 |
ENST00000628515.2
|
NRXN1
|
neurexin 1 |
| chr1_-_241357085 | 0.35 |
ENST00000366564.5
|
RGS7
|
regulator of G protein signaling 7 |
| chr10_-_101031095 | 0.35 |
ENST00000645349.1
ENST00000619208.6 ENST00000644782.1 ENST00000370215.7 ENST00000646029.1 |
PDZD7
|
PDZ domain containing 7 |
| chr1_+_209938169 | 0.34 |
ENST00000367019.5
ENST00000537238.5 ENST00000637265.1 |
SYT14
|
synaptotagmin 14 |
| chr1_-_241357171 | 0.34 |
ENST00000440928.6
|
RGS7
|
regulator of G protein signaling 7 |
| chr9_-_101594918 | 0.34 |
ENST00000374806.2
|
PPP3R2
|
protein phosphatase 3 regulatory subunit B, beta |
| chr16_+_31459950 | 0.33 |
ENST00000564900.1
|
ARMC5
|
armadillo repeat containing 5 |
| chr9_+_137139139 | 0.32 |
ENST00000371561.8
|
GRIN1
|
glutamate ionotropic receptor NMDA type subunit 1 |
| chr1_+_209938207 | 0.32 |
ENST00000472886.5
|
SYT14
|
synaptotagmin 14 |
| chr6_+_25652272 | 0.32 |
ENST00000334979.6
|
SCGN
|
secretagogin, EF-hand calcium binding protein |
| chr15_-_82709886 | 0.30 |
ENST00000666055.1
ENST00000261722.8 ENST00000535513.2 |
AP3B2
|
adaptor related protein complex 3 subunit beta 2 |
| chr12_+_112791738 | 0.30 |
ENST00000389385.9
|
RPH3A
|
rabphilin 3A |
| chr9_+_137139481 | 0.30 |
ENST00000371546.8
ENST00000371550.8 ENST00000371553.7 ENST00000371555.8 ENST00000371559.8 ENST00000371560.4 |
GRIN1
|
glutamate ionotropic receptor NMDA type subunit 1 |
| chr7_-_101165558 | 0.30 |
ENST00000611537.1
ENST00000249330.3 |
VGF
|
VGF nerve growth factor inducible |
| chr3_-_114179052 | 0.30 |
ENST00000383673.5
ENST00000295881.9 |
DRD3
|
dopamine receptor D3 |
| chr10_-_49762276 | 0.28 |
ENST00000374103.9
|
OGDHL
|
oxoglutarate dehydrogenase L |
| chr14_-_106639589 | 0.27 |
ENST00000390630.3
|
IGHV4-61
|
immunoglobulin heavy variable 4-61 |
| chr2_-_51032091 | 0.27 |
ENST00000637511.1
ENST00000405472.7 ENST00000405581.3 ENST00000401669.7 |
NRXN1
|
neurexin 1 |
| chr1_+_154567820 | 0.26 |
ENST00000637900.1
|
CHRNB2
|
cholinergic receptor nicotinic beta 2 subunit |
| chr9_-_136205122 | 0.26 |
ENST00000371748.10
|
LHX3
|
LIM homeobox 3 |
| chr20_-_63653413 | 0.25 |
ENST00000370053.3
|
STMN3
|
stathmin 3 |
| chr1_+_65309517 | 0.25 |
ENST00000371069.5
|
DNAJC6
|
DnaJ heat shock protein family (Hsp40) member C6 |
| chr17_+_58148384 | 0.24 |
ENST00000268912.6
ENST00000641449.1 |
OR4D1
|
olfactory receptor family 4 subfamily D member 1 |
| chr14_-_74426102 | 0.24 |
ENST00000554953.1
ENST00000331628.8 |
SYNDIG1L
|
synapse differentiation inducing 1 like |
| chr10_-_44385043 | 0.23 |
ENST00000374426.6
ENST00000395794.2 ENST00000374429.6 ENST00000395793.7 ENST00000343575.10 ENST00000395795.5 |
CXCL12
|
C-X-C motif chemokine ligand 12 |
| chrX_-_153971810 | 0.23 |
ENST00000310441.12
|
HCFC1
|
host cell factor C1 |
| chr20_+_37383648 | 0.23 |
ENST00000373567.6
|
SRC
|
SRC proto-oncogene, non-receptor tyrosine kinase |
| chrX_-_153875847 | 0.23 |
ENST00000361699.8
ENST00000361981.7 |
L1CAM
|
L1 cell adhesion molecule |
| chrX_-_49200174 | 0.22 |
ENST00000472598.5
ENST00000263233.9 ENST00000479808.5 |
SYP
|
synaptophysin |
| chr15_-_26773737 | 0.21 |
ENST00000299267.8
|
GABRB3
|
gamma-aminobutyric acid type A receptor subunit beta3 |
| chr12_+_112791933 | 0.20 |
ENST00000551052.5
ENST00000415485.7 |
RPH3A
|
rabphilin 3A |
| chr17_+_44308573 | 0.20 |
ENST00000590941.5
ENST00000225441.11 ENST00000426726.8 |
RUNDC3A
|
RUN domain containing 3A |
| chr8_-_56445941 | 0.20 |
ENST00000517415.1
ENST00000314922.3 |
PENK
|
proenkephalin |
| chr5_+_64505981 | 0.20 |
ENST00000334025.3
|
RGS7BP
|
regulator of G protein signaling 7 binding protein |
| chr10_+_26216766 | 0.20 |
ENST00000376261.8
|
GAD2
|
glutamate decarboxylase 2 |
| chr14_-_94293071 | 0.19 |
ENST00000554723.5
|
SERPINA10
|
serpin family A member 10 |
| chr8_-_144336451 | 0.19 |
ENST00000569446.3
|
SCRT1
|
scratch family transcriptional repressor 1 |
| chr14_-_94293024 | 0.19 |
ENST00000393096.5
|
SERPINA10
|
serpin family A member 10 |
| chr19_+_40530464 | 0.18 |
ENST00000392023.1
|
SPTBN4
|
spectrin beta, non-erythrocytic 4 |
| chr1_+_154567770 | 0.18 |
ENST00000368476.4
|
CHRNB2
|
cholinergic receptor nicotinic beta 2 subunit |
| chr2_-_27308445 | 0.18 |
ENST00000296099.2
|
UCN
|
urocortin |
| chr14_-_94293260 | 0.18 |
ENST00000261994.9
|
SERPINA10
|
serpin family A member 10 |
| chr11_-_6655788 | 0.18 |
ENST00000299441.5
|
DCHS1
|
dachsous cadherin-related 1 |
| chr19_-_45423481 | 0.17 |
ENST00000589381.5
|
ERCC1
|
ERCC excision repair 1, endonuclease non-catalytic subunit |
| chr5_-_176610104 | 0.17 |
ENST00000303991.5
|
GPRIN1
|
G protein regulated inducer of neurite outgrowth 1 |
| chr2_+_225399684 | 0.16 |
ENST00000636099.1
|
NYAP2
|
neuronal tyrosine-phosphorylated phosphoinositide-3-kinase adaptor 2 |
| chr2_+_130756291 | 0.16 |
ENST00000458606.5
ENST00000423981.1 |
AMER3
|
APC membrane recruitment protein 3 |
| chr10_-_49762335 | 0.16 |
ENST00000419399.4
ENST00000432695.2 |
OGDHL
|
oxoglutarate dehydrogenase L |
| chr9_+_114329869 | 0.16 |
ENST00000431067.4
|
ORM2
|
orosomucoid 2 |
| chr6_+_138404206 | 0.16 |
ENST00000607197.6
ENST00000367697.7 |
HEBP2
|
heme binding protein 2 |
| chr17_-_3433841 | 0.16 |
ENST00000248384.1
|
OR1E2
|
olfactory receptor family 1 subfamily E member 2 |
| chr20_-_62220267 | 0.16 |
ENST00000317393.10
ENST00000611492.1 ENST00000340177.10 |
HRH3
|
histamine receptor H3 |
| chr6_+_25652201 | 0.15 |
ENST00000612225.4
ENST00000377961.3 |
SCGN
|
secretagogin, EF-hand calcium binding protein |
| chr19_-_45423501 | 0.15 |
ENST00000591636.5
ENST00000013807.9 ENST00000592023.5 |
ERCC1
|
ERCC excision repair 1, endonuclease non-catalytic subunit |
| chr14_-_51095091 | 0.14 |
ENST00000684578.1
|
TRIM9
|
tripartite motif containing 9 |
| chr1_-_151716782 | 0.14 |
ENST00000420342.1
ENST00000290583.9 |
CELF3
|
CUGBP Elav-like family member 3 |
| chr3_+_75906666 | 0.14 |
ENST00000487694.7
ENST00000602589.5 |
ROBO2
|
roundabout guidance receptor 2 |
| chr1_+_212432891 | 0.14 |
ENST00000366988.5
|
NENF
|
neudesin neurotrophic factor |
| chr17_-_29006145 | 0.13 |
ENST00000360295.13
|
SEZ6
|
seizure related 6 homolog |
| chr22_+_22711689 | 0.12 |
ENST00000390308.2
|
IGLV3-21
|
immunoglobulin lambda variable 3-21 |
| chr17_-_29005913 | 0.11 |
ENST00000442608.7
ENST00000317338.17 ENST00000335960.10 |
SEZ6
|
seizure related 6 homolog |
| chr11_+_637244 | 0.11 |
ENST00000176183.6
|
DRD4
|
dopamine receptor D4 |
| chr9_-_128275935 | 0.11 |
ENST00000610329.4
|
GOLGA2
|
golgin A2 |
| chr5_-_136365476 | 0.11 |
ENST00000378459.7
ENST00000502753.4 ENST00000513104.6 ENST00000352189.8 |
TRPC7
|
transient receptor potential cation channel subfamily C member 7 |
| chr8_+_67952028 | 0.11 |
ENST00000288368.5
|
PREX2
|
phosphatidylinositol-3,4,5-trisphosphate dependent Rac exchange factor 2 |
| chr11_-_66347560 | 0.11 |
ENST00000311181.5
|
B4GAT1
|
beta-1,4-glucuronyltransferase 1 |
| chr12_-_12684490 | 0.10 |
ENST00000540510.1
|
GPR19
|
G protein-coupled receptor 19 |
| chr3_+_120908072 | 0.09 |
ENST00000273666.10
ENST00000471454.6 ENST00000472879.5 ENST00000497029.5 ENST00000492541.5 |
STXBP5L
|
syntaxin binding protein 5 like |
| chr12_+_71439789 | 0.09 |
ENST00000266674.10
|
LGR5
|
leucine rich repeat containing G protein-coupled receptor 5 |
| chr10_+_26216665 | 0.09 |
ENST00000259271.7
|
GAD2
|
glutamate decarboxylase 2 |
| chr9_-_128275987 | 0.09 |
ENST00000490628.2
ENST00000421699.7 ENST00000611957.4 ENST00000450617.6 |
GOLGA2
|
golgin A2 |
| chr6_+_154039480 | 0.09 |
ENST00000229768.9
ENST00000419506.6 ENST00000524163.5 ENST00000414028.6 ENST00000435918.6 ENST00000337049.8 |
OPRM1
|
opioid receptor mu 1 |
| chr10_+_59177161 | 0.07 |
ENST00000373878.3
|
PHYHIPL
|
phytanoyl-CoA 2-hydroxylase interacting protein like |
| chr3_-_114178698 | 0.07 |
ENST00000467632.5
|
DRD3
|
dopamine receptor D3 |
| chr3_+_194136138 | 0.06 |
ENST00000232424.4
|
HES1
|
hes family bHLH transcription factor 1 |
| chr16_+_4958289 | 0.06 |
ENST00000251170.12
|
SEC14L5
|
SEC14 like lipid binding 5 |
| chr20_-_45515612 | 0.06 |
ENST00000217428.7
|
SPINT3
|
serine peptidase inhibitor, Kunitz type 3 |
| chr14_-_51095066 | 0.06 |
ENST00000360392.4
|
TRIM9
|
tripartite motif containing 9 |
| chr3_-_132035004 | 0.06 |
ENST00000429747.6
|
CPNE4
|
copine 4 |
| chr8_+_6708626 | 0.05 |
ENST00000285518.11
|
AGPAT5
|
1-acylglycerol-3-phosphate O-acyltransferase 5 |
| chr9_-_121050264 | 0.04 |
ENST00000223642.3
|
C5
|
complement C5 |
| chrX_+_153972729 | 0.04 |
ENST00000369982.5
|
TMEM187
|
transmembrane protein 187 |
| chr1_+_32886456 | 0.04 |
ENST00000373467.4
|
HPCA
|
hippocalcin |
| chr19_-_45423891 | 0.03 |
ENST00000300853.8
|
ERCC1
|
ERCC excision repair 1, endonuclease non-catalytic subunit |
| chr6_+_154039333 | 0.03 |
ENST00000428397.6
|
OPRM1
|
opioid receptor mu 1 |
| chr5_+_71719267 | 0.03 |
ENST00000296777.5
|
CARTPT
|
CART prepropeptide |
| chr6_+_154039416 | 0.02 |
ENST00000452687.6
|
OPRM1
|
opioid receptor mu 1 |
| chr19_-_45423839 | 0.00 |
ENST00000340192.11
|
ERCC1
|
ERCC excision repair 1, endonuclease non-catalytic subunit |
| chr11_-_125903190 | 0.00 |
ENST00000227474.8
ENST00000613398.4 ENST00000534158.5 ENST00000529801.1 |
PUS3
|
pseudouridine synthase 3 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | GO:0021586 | pons maturation(GO:0021586) |
| 0.3 | 1.8 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.3 | 0.8 | GO:0006500 | N-terminal protein palmitoylation(GO:0006500) |
| 0.2 | 0.7 | GO:1901899 | positive regulation of relaxation of cardiac muscle(GO:1901899) |
| 0.2 | 1.7 | GO:0035502 | metanephric part of ureteric bud development(GO:0035502) |
| 0.2 | 1.0 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.2 | 1.1 | GO:0030644 | cellular chloride ion homeostasis(GO:0030644) |
| 0.1 | 1.6 | GO:0097116 | gephyrin clustering involved in postsynaptic density assembly(GO:0097116) |
| 0.1 | 0.6 | GO:0071947 | protein deubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0071947) |
| 0.1 | 0.6 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.1 | 0.4 | GO:0050883 | negative regulation of sodium:proton antiporter activity(GO:0032416) musculoskeletal movement, spinal reflex action(GO:0050883) |
| 0.1 | 1.2 | GO:2001013 | epithelial cell proliferation involved in renal tubule morphogenesis(GO:2001013) |
| 0.1 | 1.5 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.1 | 0.4 | GO:0036515 | serotonergic neuron axon guidance(GO:0036515) |
| 0.1 | 0.3 | GO:0021526 | medial motor column neuron differentiation(GO:0021526) |
| 0.1 | 0.7 | GO:0033603 | positive regulation of dopamine secretion(GO:0033603) |
| 0.1 | 0.2 | GO:0019046 | release from viral latency(GO:0019046) |
| 0.1 | 0.8 | GO:0034465 | response to carbon monoxide(GO:0034465) |
| 0.1 | 0.6 | GO:0018231 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.1 | 0.4 | GO:2000987 | positive regulation of fear response(GO:1903367) positive regulation of behavioral fear response(GO:2000987) |
| 0.1 | 0.2 | GO:0003192 | mitral valve formation(GO:0003192) |
| 0.1 | 1.7 | GO:0099517 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.1 | 0.2 | GO:0035544 | negative regulation of SNARE complex assembly(GO:0035544) |
| 0.1 | 1.1 | GO:0003376 | sphingosine-1-phosphate signaling pathway(GO:0003376) |
| 0.0 | 0.3 | GO:0006540 | glutamate decarboxylation to succinate(GO:0006540) |
| 0.0 | 0.3 | GO:0000720 | pyrimidine dimer repair by nucleotide-excision repair(GO:0000720) |
| 0.0 | 0.2 | GO:0071393 | cellular response to progesterone stimulus(GO:0071393) |
| 0.0 | 1.8 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.0 | 1.5 | GO:0006706 | steroid catabolic process(GO:0006706) |
| 0.0 | 0.1 | GO:0050925 | negative regulation of negative chemotaxis(GO:0050925) |
| 0.0 | 0.2 | GO:0060012 | synaptic transmission, glycinergic(GO:0060012) |
| 0.0 | 0.1 | GO:0060164 | negative regulation of auditory receptor cell differentiation(GO:0045608) regulation of timing of neuron differentiation(GO:0060164) negative regulation of pro-B cell differentiation(GO:2000974) |
| 0.0 | 0.4 | GO:0070459 | prolactin secretion(GO:0070459) |
| 0.0 | 0.2 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.0 | 0.7 | GO:0060219 | camera-type eye photoreceptor cell differentiation(GO:0060219) |
| 0.0 | 0.1 | GO:0051944 | positive regulation of dopamine uptake involved in synaptic transmission(GO:0051586) positive regulation of catecholamine uptake involved in synaptic transmission(GO:0051944) |
| 0.0 | 0.2 | GO:0014063 | negative regulation of serotonin secretion(GO:0014063) |
| 0.0 | 0.0 | GO:0070093 | negative regulation of glucagon secretion(GO:0070093) |
| 0.0 | 0.2 | GO:0090166 | Golgi disassembly(GO:0090166) |
| 0.0 | 1.2 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.0 | 0.4 | GO:2001241 | positive regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001241) |
| 0.0 | 0.7 | GO:0002021 | response to dietary excess(GO:0002021) |
| 0.0 | 0.2 | GO:0060080 | inhibitory postsynaptic potential(GO:0060080) |
| 0.0 | 0.6 | GO:0016048 | detection of temperature stimulus(GO:0016048) |
| 0.0 | 0.4 | GO:0097091 | synaptic vesicle clustering(GO:0097091) |
| 0.0 | 0.2 | GO:0038003 | opioid receptor signaling pathway(GO:0038003) |
| 0.0 | 0.1 | GO:0030311 | poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.0 | 0.3 | GO:0048149 | behavioral response to ethanol(GO:0048149) |
| 0.0 | 0.3 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.0 | 0.2 | GO:0010940 | positive regulation of necrotic cell death(GO:0010940) |
| 0.0 | 0.0 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 0.0 | 0.2 | GO:0031282 | regulation of guanylate cyclase activity(GO:0031282) |
| 0.0 | 1.4 | GO:0007189 | adenylate cyclase-activating G-protein coupled receptor signaling pathway(GO:0007189) |
| 0.0 | 0.1 | GO:0032096 | negative regulation of response to food(GO:0032096) negative regulation of appetite(GO:0032099) |
| 0.0 | 1.1 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.0 | 0.2 | GO:0010459 | negative regulation of heart rate(GO:0010459) |
| 0.0 | 0.5 | GO:1903861 | positive regulation of dendrite extension(GO:1903861) |
| 0.0 | 0.2 | GO:0090036 | regulation of protein kinase C signaling(GO:0090036) |
| 0.0 | 1.6 | GO:0030901 | midbrain development(GO:0030901) |
| 0.0 | 1.9 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) |
| 0.0 | 0.2 | GO:1904469 | positive regulation of tumor necrosis factor secretion(GO:1904469) |
| 0.0 | 0.4 | GO:0042462 | eye photoreceptor cell development(GO:0042462) |
| 0.0 | 0.7 | GO:0090004 | positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.0 | 0.1 | GO:0016191 | synaptic vesicle uncoating(GO:0016191) |
| 0.0 | 0.4 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) |
| 0.2 | 0.8 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.1 | 1.7 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.1 | 2.1 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.1 | 0.3 | GO:0002139 | stereocilia coupling link(GO:0002139) |
| 0.1 | 1.2 | GO:0032426 | stereocilium tip(GO:0032426) |
| 0.0 | 0.3 | GO:0070554 | synaptobrevin 2-SNAP-25-syntaxin-3-complexin complex(GO:0070554) |
| 0.0 | 0.4 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.0 | 0.6 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.0 | 0.6 | GO:0002178 | palmitoyltransferase complex(GO:0002178) |
| 0.0 | 0.7 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.0 | 0.3 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.0 | 0.7 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.0 | 0.4 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.0 | 0.5 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 0.2 | GO:0043196 | varicosity(GO:0043196) |
| 0.0 | 2.2 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 1.3 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 0.2 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.0 | 0.4 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 1.6 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 0.2 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 1.2 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.2 | GO:0016934 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.4 | 1.5 | GO:0004773 | steryl-sulfatase activity(GO:0004773) |
| 0.3 | 1.8 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.3 | 1.7 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.3 | 0.8 | GO:0016534 | cyclin-dependent protein kinase 5 activator activity(GO:0016534) |
| 0.1 | 1.2 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.1 | 0.6 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.1 | 1.0 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.1 | 0.8 | GO:0060072 | large conductance calcium-activated potassium channel activity(GO:0060072) |
| 0.1 | 1.1 | GO:0022820 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 0.1 | 0.4 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.1 | 1.1 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.1 | 0.5 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.1 | 1.6 | GO:0033130 | acetylcholine receptor binding(GO:0033130) neuroligin family protein binding(GO:0097109) |
| 0.1 | 1.9 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.2 | GO:0051431 | histone deacetylase regulator activity(GO:0035033) corticotropin-releasing hormone receptor 2 binding(GO:0051431) |
| 0.0 | 0.5 | GO:0008430 | selenium binding(GO:0008430) |
| 0.0 | 0.2 | GO:0061676 | importin-alpha family protein binding(GO:0061676) |
| 0.0 | 0.7 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.3 | GO:1990599 | 3' overhang single-stranded DNA endodeoxyribonuclease activity(GO:1990599) |
| 0.0 | 1.3 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.3 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 0.0 | 0.4 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 0.2 | GO:0022851 | GABA-gated chloride ion channel activity(GO:0022851) |
| 0.0 | 0.7 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.2 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.2 | GO:0030250 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
| 0.0 | 0.2 | GO:0043995 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
| 0.0 | 0.6 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.1 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.0 | 0.6 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.2 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.0 | 0.2 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.4 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.0 | 0.1 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.0 | 0.1 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.8 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 1.2 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.2 | ST STAT3 PATHWAY | STAT3 Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.5 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.1 | 1.0 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 2.4 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.0 | 1.8 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.0 | 0.8 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.0 | 0.4 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 0.4 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.4 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.0 | 0.2 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.0 | 0.4 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.0 | 0.5 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 0.1 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.0 | 0.3 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.0 | 2.9 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |