Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
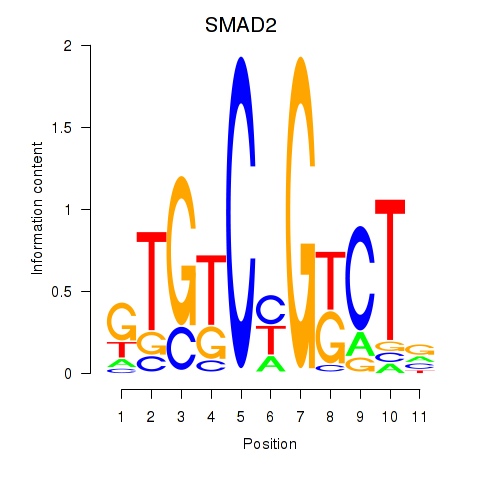
Results for SMAD2
Z-value: 0.53
Transcription factors associated with SMAD2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SMAD2
|
ENSG00000175387.16 | SMAD2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SMAD2 | hg38_v1_chr18_-_47931107_47931146 | -0.54 | 2.0e-03 | Click! |
Activity profile of SMAD2 motif
Sorted Z-values of SMAD2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SMAD2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chrX_-_48470243 | 1.87 |
ENST00000429543.2
ENST00000620913.5 |
SLC38A5
|
solute carrier family 38 member 5 |
| chrX_-_48470163 | 1.84 |
ENST00000595796.5
|
SLC38A5
|
solute carrier family 38 member 5 |
| chr5_-_151686908 | 1.42 |
ENST00000231061.9
|
SPARC
|
secreted protein acidic and cysteine rich |
| chr6_-_30687200 | 1.37 |
ENST00000399199.7
|
PPP1R18
|
protein phosphatase 1 regulatory subunit 18 |
| chr12_-_119804298 | 1.35 |
ENST00000678652.1
ENST00000678494.1 |
CIT
|
citron rho-interacting serine/threonine kinase |
| chr12_+_53050179 | 1.33 |
ENST00000546602.5
ENST00000552570.5 ENST00000549700.5 |
TNS2
|
tensin 2 |
| chrX_-_107775740 | 1.30 |
ENST00000372383.9
|
TSC22D3
|
TSC22 domain family member 3 |
| chr19_-_43781249 | 1.30 |
ENST00000615047.4
|
KCNN4
|
potassium calcium-activated channel subfamily N member 4 |
| chr12_-_119803383 | 1.23 |
ENST00000392520.2
ENST00000678677.1 ENST00000679249.1 ENST00000676849.1 |
CIT
|
citron rho-interacting serine/threonine kinase |
| chrX_-_107775951 | 1.21 |
ENST00000315660.8
ENST00000372384.6 ENST00000502650.1 ENST00000506724.1 |
TSC22D3
|
TSC22 domain family member 3 |
| chr11_-_107858777 | 1.21 |
ENST00000525815.6
|
SLC35F2
|
solute carrier family 35 member F2 |
| chr9_+_122375286 | 1.03 |
ENST00000373698.7
|
PTGS1
|
prostaglandin-endoperoxide synthase 1 |
| chr17_-_7394240 | 1.00 |
ENST00000576362.5
ENST00000571078.5 |
PLSCR3
|
phospholipid scramblase 3 |
| chr17_-_7394514 | 0.95 |
ENST00000571802.1
ENST00000619711.5 ENST00000576201.5 ENST00000573213.1 ENST00000324822.15 |
PLSCR3
|
phospholipid scramblase 3 |
| chr16_+_55479188 | 0.93 |
ENST00000219070.9
|
MMP2
|
matrix metallopeptidase 2 |
| chr9_+_125747345 | 0.92 |
ENST00000342287.9
ENST00000373489.10 ENST00000373487.8 |
PBX3
|
PBX homeobox 3 |
| chr1_-_149936816 | 0.87 |
ENST00000439741.4
|
MTMR11
|
myotubularin related protein 11 |
| chr9_+_34990250 | 0.83 |
ENST00000454002.6
ENST00000545841.5 |
DNAJB5
|
DnaJ heat shock protein family (Hsp40) member B5 |
| chr2_-_227164194 | 0.83 |
ENST00000396625.5
|
COL4A4
|
collagen type IV alpha 4 chain |
| chr16_+_28878480 | 0.81 |
ENST00000395503.9
|
ATP2A1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr1_-_149936324 | 0.79 |
ENST00000369140.7
|
MTMR11
|
myotubularin related protein 11 |
| chr7_+_129126518 | 0.79 |
ENST00000467614.2
|
ENSG00000230626.4
|
novel protein similar to mitogen-activated protein kinase kinase 2 MAP2K2 |
| chr17_-_78925376 | 0.78 |
ENST00000262768.11
|
TIMP2
|
TIMP metallopeptidase inhibitor 2 |
| chr4_+_37244735 | 0.75 |
ENST00000309447.6
|
NWD2
|
NACHT and WD repeat domain containing 2 |
| chr20_+_44715360 | 0.73 |
ENST00000190983.5
|
CCN5
|
cellular communication network factor 5 |
| chr16_+_28878382 | 0.72 |
ENST00000357084.7
|
ATP2A1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr20_+_44714853 | 0.71 |
ENST00000372865.4
|
CCN5
|
cellular communication network factor 5 |
| chr17_+_4950147 | 0.69 |
ENST00000522301.5
|
ENO3
|
enolase 3 |
| chr12_+_53050014 | 0.69 |
ENST00000314250.11
|
TNS2
|
tensin 2 |
| chr22_+_37560472 | 0.65 |
ENST00000430687.1
ENST00000249014.5 |
CDC42EP1
|
CDC42 effector protein 1 |
| chr22_-_37427433 | 0.65 |
ENST00000452946.1
ENST00000402918.7 |
ELFN2
ELFN2
|
extracellular leucine rich repeat and fibronectin type III domain containing 2 extracellular leucine rich repeat and fibronectin type III domain containing 2 |
| chr12_-_52814106 | 0.63 |
ENST00000551956.2
|
KRT4
|
keratin 4 |
| chr10_+_71212524 | 0.58 |
ENST00000335350.10
|
UNC5B
|
unc-5 netrin receptor B |
| chr19_-_14114156 | 0.58 |
ENST00000589994.6
|
PRKACA
|
protein kinase cAMP-activated catalytic subunit alpha |
| chr7_-_102616692 | 0.58 |
ENST00000521076.5
ENST00000462172.5 ENST00000522801.5 ENST00000262940.12 ENST00000449970.6 |
RASA4
|
RAS p21 protein activator 4 |
| chr12_-_6631632 | 0.58 |
ENST00000431922.1
|
LPAR5
|
lysophosphatidic acid receptor 5 |
| chr10_-_14330879 | 0.58 |
ENST00000357447.7
|
FRMD4A
|
FERM domain containing 4A |
| chr7_-_75994574 | 0.58 |
ENST00000439537.5
ENST00000493111.7 ENST00000417509.5 ENST00000485200.1 |
TMEM120A
|
transmembrane protein 120A |
| chr21_-_5973383 | 0.57 |
ENST00000464664.3
|
ENSG00000274559.3
|
novel histone H2B family protein |
| chr22_-_43891995 | 0.54 |
ENST00000438734.1
ENST00000597664.6 ENST00000216177.8 ENST00000593866.5 ENST00000381198.6 |
PNPLA5
|
patatin like phospholipase domain containing 5 |
| chr3_+_50279080 | 0.53 |
ENST00000316436.4
|
LSMEM2
|
leucine rich single-pass membrane protein 2 |
| chr1_+_37474572 | 0.51 |
ENST00000373087.7
|
ZC3H12A
|
zinc finger CCCH-type containing 12A |
| chr5_-_64768619 | 0.51 |
ENST00000513458.9
|
SREK1IP1
|
SREK1 interacting protein 1 |
| chr7_-_102517755 | 0.51 |
ENST00000306682.6
ENST00000465829.6 ENST00000541662.5 |
RASA4B
|
RAS p21 protein activator 4B |
| chr4_-_41214450 | 0.50 |
ENST00000513140.5
|
APBB2
|
amyloid beta precursor protein binding family B member 2 |
| chr19_-_41353904 | 0.50 |
ENST00000221930.6
|
TGFB1
|
transforming growth factor beta 1 |
| chr5_+_71455636 | 0.47 |
ENST00000358731.9
ENST00000380675.3 |
BDP1
|
B double prime 1, subunit of RNA polymerase III transcription initiation factor IIIB |
| chr19_-_49362376 | 0.47 |
ENST00000601519.5
ENST00000593945.6 ENST00000539846.5 ENST00000596757.1 ENST00000311227.6 |
TEAD2
|
TEA domain transcription factor 2 |
| chr16_+_2009870 | 0.47 |
ENST00000567649.1
|
NPW
|
neuropeptide W |
| chr20_+_37346128 | 0.46 |
ENST00000373578.7
|
SRC
|
SRC proto-oncogene, non-receptor tyrosine kinase |
| chr11_-_101583985 | 0.46 |
ENST00000344327.8
|
TRPC6
|
transient receptor potential cation channel subfamily C member 6 |
| chr22_+_37639660 | 0.45 |
ENST00000649765.2
ENST00000451997.6 |
SH3BP1
ENSG00000285304.1
|
SH3 domain binding protein 1 novel protein |
| chr4_-_41214602 | 0.44 |
ENST00000508676.5
ENST00000506352.5 ENST00000295974.12 |
APBB2
|
amyloid beta precursor protein binding family B member 2 |
| chr11_-_117295485 | 0.41 |
ENST00000680971.1
|
BACE1
|
beta-secretase 1 |
| chr13_-_26760741 | 0.41 |
ENST00000405846.5
|
GPR12
|
G protein-coupled receptor 12 |
| chr21_-_34887148 | 0.40 |
ENST00000399240.5
|
RUNX1
|
RUNX family transcription factor 1 |
| chr4_-_41214535 | 0.39 |
ENST00000508593.6
|
APBB2
|
amyloid beta precursor protein binding family B member 2 |
| chr1_+_2476315 | 0.39 |
ENST00000419816.6
|
PLCH2
|
phospholipase C eta 2 |
| chr1_+_2476284 | 0.39 |
ENST00000378486.8
|
PLCH2
|
phospholipase C eta 2 |
| chr6_+_34514853 | 0.38 |
ENST00000538621.2
|
PACSIN1
|
protein kinase C and casein kinase substrate in neurons 1 |
| chr14_-_106038355 | 0.37 |
ENST00000390597.3
|
IGHV2-5
|
immunoglobulin heavy variable 2-5 |
| chr3_+_130850585 | 0.37 |
ENST00000505330.5
ENST00000504381.5 ENST00000507488.6 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr14_-_104978360 | 0.37 |
ENST00000333244.6
|
AHNAK2
|
AHNAK nucleoprotein 2 |
| chr21_-_37267511 | 0.37 |
ENST00000398998.1
|
VPS26C
|
VPS26 endosomal protein sorting factor C |
| chr17_+_3723889 | 0.36 |
ENST00000325418.5
|
HASPIN
|
histone H3 associated protein kinase |
| chr12_-_8612850 | 0.35 |
ENST00000229335.11
ENST00000537228.5 |
AICDA
|
activation induced cytidine deaminase |
| chr6_+_43132472 | 0.35 |
ENST00000489707.5
|
PTK7
|
protein tyrosine kinase 7 (inactive) |
| chrX_-_47629845 | 0.34 |
ENST00000469388.1
ENST00000396992.8 ENST00000377005.6 |
CFP
|
complement factor properdin |
| chr14_-_56810448 | 0.34 |
ENST00000339475.10
ENST00000555006.5 ENST00000672264.2 ENST00000554559.5 ENST00000555804.1 |
OTX2
|
orthodenticle homeobox 2 |
| chr16_-_2009797 | 0.34 |
ENST00000563630.6
ENST00000562103.2 |
ZNF598
|
zinc finger protein 598, E3 ubiquitin ligase |
| chr22_-_36028773 | 0.34 |
ENST00000438146.7
|
RBFOX2
|
RNA binding fox-1 homolog 2 |
| chr1_-_44017296 | 0.32 |
ENST00000357730.6
ENST00000360584.6 ENST00000528803.1 |
SLC6A9
|
solute carrier family 6 member 9 |
| chr4_-_102828048 | 0.32 |
ENST00000508249.1
|
UBE2D3
|
ubiquitin conjugating enzyme E2 D3 |
| chr2_-_20225433 | 0.32 |
ENST00000381150.5
|
SDC1
|
syndecan 1 |
| chr16_-_4937064 | 0.31 |
ENST00000590782.6
ENST00000345988.7 |
PPL
|
periplakin |
| chr5_+_74766981 | 0.31 |
ENST00000296802.9
|
NSA2
|
NSA2 ribosome biogenesis factor |
| chrX_-_135098695 | 0.31 |
ENST00000433425.4
|
SMIM10L2B
|
small integral membrane protein 10 like 2B |
| chr7_-_14841267 | 0.31 |
ENST00000406247.7
ENST00000399322.7 |
DGKB
|
diacylglycerol kinase beta |
| chr12_+_100794769 | 0.31 |
ENST00000392977.8
ENST00000546991.1 ENST00000392979.7 |
ANO4
|
anoctamin 4 |
| chr22_+_50089879 | 0.30 |
ENST00000545383.5
ENST00000262794.10 |
MOV10L1
|
Mov10 like RISC complex RNA helicase 1 |
| chr11_-_107719657 | 0.30 |
ENST00000525934.1
ENST00000531293.1 |
SLN
|
sarcolipin |
| chr9_+_122519141 | 0.30 |
ENST00000340750.1
|
OR1J4
|
olfactory receptor family 1 subfamily J member 4 |
| chr3_-_195876635 | 0.30 |
ENST00000672669.1
ENST00000672886.1 ENST00000672098.1 ENST00000671767.1 ENST00000672548.1 |
TNK2
|
tyrosine kinase non receptor 2 |
| chr18_+_23135452 | 0.29 |
ENST00000580153.5
ENST00000256925.12 |
CABLES1
|
Cdk5 and Abl enzyme substrate 1 |
| chr20_+_32010429 | 0.29 |
ENST00000452892.3
ENST00000262659.12 |
CCM2L
|
CCM2 like scaffold protein |
| chr4_-_39977836 | 0.29 |
ENST00000303538.13
ENST00000503396.5 |
PDS5A
|
PDS5 cohesin associated factor A |
| chr1_+_15456727 | 0.29 |
ENST00000359621.5
|
CELA2A
|
chymotrypsin like elastase 2A |
| chr11_-_18791768 | 0.29 |
ENST00000358540.7
|
PTPN5
|
protein tyrosine phosphatase non-receptor type 5 |
| chr2_-_219309484 | 0.28 |
ENST00000409251.7
ENST00000451506.5 ENST00000446182.5 |
PTPRN
|
protein tyrosine phosphatase receptor type N |
| chr11_-_3837858 | 0.28 |
ENST00000396979.1
|
RHOG
|
ras homolog family member G |
| chr4_-_102828022 | 0.28 |
ENST00000502690.5
|
UBE2D3
|
ubiquitin conjugating enzyme E2 D3 |
| chr5_+_33440947 | 0.27 |
ENST00000455217.6
ENST00000265112.8 |
TARS1
|
threonyl-tRNA synthetase 1 |
| chr17_-_42609356 | 0.27 |
ENST00000309428.10
|
RETREG3
|
reticulophagy regulator family member 3 |
| chr4_-_113761068 | 0.27 |
ENST00000342666.9
ENST00000514328.5 ENST00000515496.5 ENST00000508738.5 ENST00000379773.6 |
CAMK2D
|
calcium/calmodulin dependent protein kinase II delta |
| chr1_+_36306359 | 0.27 |
ENST00000453908.8
|
SH3D21
|
SH3 domain containing 21 |
| chr15_-_82262660 | 0.26 |
ENST00000557844.1
ENST00000359445.7 ENST00000268206.12 |
EFL1
|
elongation factor like GTPase 1 |
| chr20_-_23086316 | 0.26 |
ENST00000246006.5
|
CD93
|
CD93 molecule |
| chrX_+_38561530 | 0.26 |
ENST00000378482.7
ENST00000286824.6 |
TSPAN7
|
tetraspanin 7 |
| chr20_+_1895365 | 0.25 |
ENST00000358771.5
|
SIRPA
|
signal regulatory protein alpha |
| chr10_+_103555124 | 0.25 |
ENST00000437579.1
|
NEURL1
|
neuralized E3 ubiquitin protein ligase 1 |
| chr10_-_73433550 | 0.24 |
ENST00000299432.7
|
MSS51
|
MSS51 mitochondrial translational activator |
| chr5_+_74767234 | 0.24 |
ENST00000610426.5
|
NSA2
|
NSA2 ribosome biogenesis factor |
| chrX_+_21940693 | 0.24 |
ENST00000404933.7
ENST00000379404.5 |
SMS
|
spermine synthase |
| chr15_+_45587366 | 0.24 |
ENST00000220531.9
|
BLOC1S6
|
biogenesis of lysosomal organelles complex 1 subunit 6 |
| chr15_+_45587580 | 0.24 |
ENST00000566801.5
ENST00000565323.6 ENST00000568816.5 |
BLOC1S6
|
biogenesis of lysosomal organelles complex 1 subunit 6 |
| chr9_+_122264603 | 0.23 |
ENST00000297908.7
|
MRRF
|
mitochondrial ribosome recycling factor |
| chr16_+_2148603 | 0.23 |
ENST00000210187.11
|
RAB26
|
RAB26, member RAS oncogene family |
| chr12_-_21941300 | 0.23 |
ENST00000684084.1
|
ABCC9
|
ATP binding cassette subfamily C member 9 |
| chr3_+_111674654 | 0.23 |
ENST00000636933.1
ENST00000393934.7 ENST00000477665.2 |
PLCXD2
|
phosphatidylinositol specific phospholipase C X domain containing 2 |
| chr11_-_18791563 | 0.23 |
ENST00000396168.1
|
PTPN5
|
protein tyrosine phosphatase non-receptor type 5 |
| chr4_-_103019634 | 0.22 |
ENST00000510559.1
ENST00000296422.12 ENST00000394789.7 |
SLC9B1
|
solute carrier family 9 member B1 |
| chr16_+_24729641 | 0.22 |
ENST00000395799.8
|
TNRC6A
|
trinucleotide repeat containing adaptor 6A |
| chr19_-_14496088 | 0.22 |
ENST00000393033.9
ENST00000345425.6 ENST00000586027.5 ENST00000591349.5 ENST00000587210.1 |
GIPC1
|
GIPC PDZ domain containing family member 1 |
| chrX_+_153764233 | 0.22 |
ENST00000361971.10
|
PLXNB3
|
plexin B3 |
| chr11_+_75562056 | 0.21 |
ENST00000533603.5
|
SERPINH1
|
serpin family H member 1 |
| chrX_+_153764178 | 0.21 |
ENST00000538966.5
|
PLXNB3
|
plexin B3 |
| chr7_-_92836555 | 0.21 |
ENST00000424848.3
|
CDK6
|
cyclin dependent kinase 6 |
| chr2_-_219309350 | 0.21 |
ENST00000295718.7
|
PTPRN
|
protein tyrosine phosphatase receptor type N |
| chr17_-_3668640 | 0.21 |
ENST00000611779.4
|
TAX1BP3
|
Tax1 binding protein 3 |
| chr9_-_133131109 | 0.20 |
ENST00000372062.8
|
RALGDS
|
ral guanine nucleotide dissociation stimulator |
| chr9_-_134917872 | 0.20 |
ENST00000616356.4
ENST00000371806.4 |
FCN1
|
ficolin 1 |
| chr5_-_151141631 | 0.20 |
ENST00000523714.5
ENST00000521749.5 |
ANXA6
|
annexin A6 |
| chr4_-_163613505 | 0.20 |
ENST00000339875.9
|
MARCHF1
|
membrane associated ring-CH-type finger 1 |
| chr11_+_66857056 | 0.19 |
ENST00000309602.5
ENST00000393952.3 |
LRFN4
|
leucine rich repeat and fibronectin type III domain containing 4 |
| chr15_+_68277724 | 0.19 |
ENST00000306917.5
|
FEM1B
|
fem-1 homolog B |
| chr6_+_41042557 | 0.19 |
ENST00000373158.6
ENST00000470917.1 |
TSPO2
|
translocator protein 2 |
| chr1_-_44031352 | 0.19 |
ENST00000372306.7
ENST00000475075.6 |
SLC6A9
|
solute carrier family 6 member 9 |
| chr1_-_1307861 | 0.19 |
ENST00000354700.10
|
ACAP3
|
ArfGAP with coiled-coil, ankyrin repeat and PH domains 3 |
| chr19_+_11355386 | 0.19 |
ENST00000251473.9
ENST00000591329.5 ENST00000586380.5 |
PLPPR2
|
phospholipid phosphatase related 2 |
| chr1_-_149812359 | 0.18 |
ENST00000369167.2
ENST00000545683.1 |
H2BC18
|
H2B clustered histone 18 |
| chr17_+_42609641 | 0.18 |
ENST00000681413.1
ENST00000251413.8 ENST00000591509.5 |
TUBG1
|
tubulin gamma 1 |
| chr22_-_38455199 | 0.18 |
ENST00000303592.3
|
KCNJ4
|
potassium inwardly rectifying channel subfamily J member 4 |
| chr1_-_206003442 | 0.18 |
ENST00000623893.1
|
RAB7B
|
RAB7B, member RAS oncogene family |
| chr9_-_98708856 | 0.18 |
ENST00000259455.4
|
GABBR2
|
gamma-aminobutyric acid type B receptor subunit 2 |
| chr9_-_28670285 | 0.18 |
ENST00000379992.6
ENST00000308675.5 ENST00000613945.3 |
LINGO2
|
leucine rich repeat and Ig domain containing 2 |
| chr10_+_18261080 | 0.18 |
ENST00000377319.8
ENST00000377331.8 |
CACNB2
|
calcium voltage-gated channel auxiliary subunit beta 2 |
| chr9_+_33524241 | 0.18 |
ENST00000290943.10
|
ANKRD18B
|
ankyrin repeat domain 18B |
| chr12_-_57016517 | 0.18 |
ENST00000441881.5
ENST00000458521.7 |
TAC3
|
tachykinin precursor 3 |
| chr4_-_102828159 | 0.18 |
ENST00000394803.9
|
UBE2D3
|
ubiquitin conjugating enzyme E2 D3 |
| chr2_-_17800195 | 0.18 |
ENST00000402989.5
ENST00000428868.1 |
SMC6
|
structural maintenance of chromosomes 6 |
| chr6_-_143450662 | 0.17 |
ENST00000237283.9
|
ADAT2
|
adenosine deaminase tRNA specific 2 |
| chr4_-_102827723 | 0.17 |
ENST00000349311.12
|
UBE2D3
|
ubiquitin conjugating enzyme E2 D3 |
| chr14_-_56810380 | 0.17 |
ENST00000672125.1
ENST00000673481.1 |
OTX2
|
orthodenticle homeobox 2 |
| chr18_+_3448456 | 0.17 |
ENST00000549780.5
|
TGIF1
|
TGFB induced factor homeobox 1 |
| chr4_-_113761441 | 0.17 |
ENST00000394524.7
|
CAMK2D
|
calcium/calmodulin dependent protein kinase II delta |
| chr4_-_184217854 | 0.17 |
ENST00000296741.7
|
ENPP6
|
ectonucleotide pyrophosphatase/phosphodiesterase 6 |
| chr19_+_47130782 | 0.17 |
ENST00000597808.5
ENST00000413379.7 ENST00000600706.5 ENST00000270225.12 ENST00000598840.5 ENST00000600753.1 ENST00000392776.3 |
SAE1
|
SUMO1 activating enzyme subunit 1 |
| chr8_-_94895195 | 0.17 |
ENST00000308108.9
ENST00000396133.7 |
CCNE2
|
cyclin E2 |
| chr5_+_134526176 | 0.16 |
ENST00000681820.1
ENST00000512386.6 ENST00000612830.2 |
JADE2
|
jade family PHD finger 2 |
| chr19_+_35775515 | 0.16 |
ENST00000378944.9
|
ARHGAP33
|
Rho GTPase activating protein 33 |
| chr9_+_15422704 | 0.16 |
ENST00000380821.7
ENST00000610884.4 ENST00000421710.5 |
SNAPC3
|
small nuclear RNA activating complex polypeptide 3 |
| chrX_-_30577759 | 0.15 |
ENST00000378962.4
|
TASL
|
TLR adaptor interacting with endolysosomal SLC15A4 |
| chr1_-_206003385 | 0.15 |
ENST00000617070.5
|
RAB7B
|
RAB7B, member RAS oncogene family |
| chr9_+_137251261 | 0.15 |
ENST00000620243.4
ENST00000388931.7 ENST00000412566.5 ENST00000611378.4 |
STPG3
|
sperm-tail PG-rich repeat containing 3 |
| chr4_+_92303946 | 0.15 |
ENST00000282020.9
|
GRID2
|
glutamate ionotropic receptor delta type subunit 2 |
| chr9_-_14722725 | 0.15 |
ENST00000380911.4
|
CER1
|
cerberus 1, DAN family BMP antagonist |
| chr12_+_109900518 | 0.15 |
ENST00000312777.9
ENST00000536408.2 |
TCHP
|
trichoplein keratin filament binding |
| chr17_-_3668557 | 0.15 |
ENST00000225525.4
|
TAX1BP3
|
Tax1 binding protein 3 |
| chr4_-_113761724 | 0.15 |
ENST00000511664.6
|
CAMK2D
|
calcium/calmodulin dependent protein kinase II delta |
| chr15_-_73368951 | 0.15 |
ENST00000261917.4
|
HCN4
|
hyperpolarization activated cyclic nucleotide gated potassium channel 4 |
| chr3_+_40505992 | 0.15 |
ENST00000420891.5
ENST00000314529.10 ENST00000418905.1 |
ZNF620
|
zinc finger protein 620 |
| chr13_-_96053370 | 0.15 |
ENST00000376712.4
ENST00000397618.7 ENST00000376747.8 ENST00000376714.7 ENST00000638479.1 ENST00000621375.5 |
UGGT2
|
UDP-glucose glycoprotein glucosyltransferase 2 |
| chr4_-_55592225 | 0.15 |
ENST00000295645.9
|
PDCL2
|
phosducin like 2 |
| chr5_+_64768921 | 0.14 |
ENST00000381070.8
ENST00000508024.1 |
CWC27
|
CWC27 spliceosome associated cyclophilin |
| chr3_-_165196369 | 0.14 |
ENST00000475390.2
|
SLITRK3
|
SLIT and NTRK like family member 3 |
| chr18_-_3013114 | 0.14 |
ENST00000677752.1
|
LPIN2
|
lipin 2 |
| chr19_-_14496144 | 0.14 |
ENST00000393028.5
|
GIPC1
|
GIPC PDZ domain containing family member 1 |
| chr1_-_160070150 | 0.13 |
ENST00000644903.1
|
KCNJ10
|
potassium inwardly rectifying channel subfamily J member 10 |
| chr12_-_95996302 | 0.13 |
ENST00000261208.8
ENST00000538703.5 ENST00000541929.5 |
HAL
|
histidine ammonia-lyase |
| chr11_+_75240761 | 0.13 |
ENST00000562197.3
|
TPBGL
|
trophoblast glycoprotein like |
| chr14_+_80955577 | 0.13 |
ENST00000642209.1
ENST00000298171.7 |
TSHR
|
thyroid stimulating hormone receptor |
| chr19_-_45785659 | 0.12 |
ENST00000537879.1
ENST00000596586.5 ENST00000595946.1 |
DMWD
ENSG00000268434.5
|
DM1 locus, WD repeat containing novel protein |
| chr19_+_48469354 | 0.12 |
ENST00000452733.7
ENST00000641098.1 |
CYTH2
|
cytohesin 2 |
| chr6_+_101393699 | 0.12 |
ENST00000369134.9
ENST00000684068.1 ENST00000683903.1 ENST00000681975.1 |
GRIK2
|
glutamate ionotropic receptor kainate type subunit 2 |
| chr17_-_41424583 | 0.12 |
ENST00000225550.4
|
KRT37
|
keratin 37 |
| chr3_-_71130963 | 0.11 |
ENST00000649695.2
|
FOXP1
|
forkhead box P1 |
| chr20_+_58651785 | 0.11 |
ENST00000358029.8
|
STX16
|
syntaxin 16 |
| chr22_+_24495242 | 0.10 |
ENST00000382760.2
ENST00000326010.10 |
UPB1
|
beta-ureidopropionase 1 |
| chr1_+_27726005 | 0.10 |
ENST00000530324.5
ENST00000234549.11 ENST00000373949.5 ENST00000010299.10 |
FAM76A
|
family with sequence similarity 76 member A |
| chr17_+_75633420 | 0.09 |
ENST00000375215.3
|
SMIM5
|
small integral membrane protein 5 |
| chr7_+_69967464 | 0.09 |
ENST00000664521.1
|
AUTS2
|
activator of transcription and developmental regulator AUTS2 |
| chr17_-_42185452 | 0.09 |
ENST00000293330.1
|
HCRT
|
hypocretin neuropeptide precursor |
| chr17_-_7207245 | 0.09 |
ENST00000649971.1
|
DLG4
|
discs large MAGUK scaffold protein 4 |
| chr17_+_68205453 | 0.09 |
ENST00000674770.2
|
AMZ2
|
archaelysin family metallopeptidase 2 |
| chr5_+_134525649 | 0.09 |
ENST00000282605.8
ENST00000681547.2 ENST00000361895.6 ENST00000402835.5 |
JADE2
|
jade family PHD finger 2 |
| chr1_-_151146611 | 0.08 |
ENST00000341697.7
ENST00000368914.8 |
SEMA6C
|
semaphorin 6C |
| chr3_+_52495330 | 0.08 |
ENST00000321725.10
|
STAB1
|
stabilin 1 |
| chr5_+_134526100 | 0.08 |
ENST00000395003.5
|
JADE2
|
jade family PHD finger 2 |
| chr4_-_5888400 | 0.08 |
ENST00000397890.6
|
CRMP1
|
collapsin response mediator protein 1 |
| chr6_-_27873525 | 0.08 |
ENST00000618305.2
|
H4C13
|
H4 clustered histone 13 |
| chr1_+_2050387 | 0.07 |
ENST00000378567.8
|
PRKCZ
|
protein kinase C zeta |
| chr1_+_99646025 | 0.07 |
ENST00000263174.9
ENST00000605497.5 ENST00000615664.1 |
PALMD
|
palmdelphin |
| chr12_+_6927946 | 0.07 |
ENST00000396684.3
|
ATN1
|
atrophin 1 |
| chr15_+_75206398 | 0.07 |
ENST00000565074.1
|
C15orf39
|
chromosome 15 open reading frame 39 |
| chr7_+_130381092 | 0.07 |
ENST00000484324.1
|
CPA1
|
carboxypeptidase A1 |
| chr1_-_160070102 | 0.07 |
ENST00000638728.1
ENST00000637644.1 |
KCNJ10
|
potassium inwardly rectifying channel subfamily J member 10 |
| chr12_+_103930600 | 0.06 |
ENST00000680316.1
ENST00000679861.1 |
HSP90B1
|
heat shock protein 90 beta family member 1 |
| chr3_-_71130557 | 0.06 |
ENST00000497355.7
|
FOXP1
|
forkhead box P1 |
| chr7_+_155458129 | 0.06 |
ENST00000297375.4
|
EN2
|
engrailed homeobox 2 |
| chr5_+_175871570 | 0.06 |
ENST00000512824.5
ENST00000393745.8 |
CPLX2
|
complexin 2 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0014724 | regulation of twitch skeletal muscle contraction(GO:0014724) fast-twitch skeletal muscle fiber contraction(GO:0031443) relaxation of skeletal muscle(GO:0090076) |
| 0.3 | 4.2 | GO:0015816 | glycine transport(GO:0015816) |
| 0.3 | 2.5 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.2 | 1.4 | GO:0033591 | response to L-ascorbic acid(GO:0033591) |
| 0.2 | 0.5 | GO:0070898 | RNA polymerase III transcriptional preinitiation complex assembly(GO:0070898) |
| 0.2 | 0.9 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.1 | 0.8 | GO:0032487 | regulation of Rap protein signal transduction(GO:0032487) |
| 0.1 | 0.5 | GO:0052251 | induction by symbiont of host defense response(GO:0044416) induction of host immune response by virus(GO:0046730) active induction of host immune response by virus(GO:0046732) modulation by symbiont of host defense response(GO:0052031) induction by organism of defense response of other organism involved in symbiotic interaction(GO:0052251) modulation by organism of defense response of other organism involved in symbiotic interaction(GO:0052255) positive regulation by symbiont of host defense response(GO:0052509) positive regulation by organism of defense response of other organism involved in symbiotic interaction(GO:0052510) modulation by organism of immune response of other organism involved in symbiotic interaction(GO:0052552) modulation by symbiont of host immune response(GO:0052553) modulation by virus of host immune response(GO:0075528) |
| 0.1 | 0.4 | GO:0090310 | negative regulation of methylation-dependent chromatin silencing(GO:0090310) |
| 0.1 | 0.9 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.1 | 0.3 | GO:0072344 | rescue of stalled ribosome(GO:0072344) |
| 0.1 | 0.5 | GO:0010587 | miRNA catabolic process(GO:0010587) negative regulation by host of viral genome replication(GO:0044828) response to ionomycin(GO:1904636) cellular response to ionomycin(GO:1904637) |
| 0.1 | 0.3 | GO:1904692 | positive regulation of type B pancreatic cell proliferation(GO:1904692) |
| 0.1 | 0.3 | GO:1901876 | regulation of calcium ion binding(GO:1901876) negative regulation of calcium ion binding(GO:1901877) |
| 0.1 | 0.3 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 0.1 | 2.3 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.1 | 0.5 | GO:0071393 | cellular response to progesterone stimulus(GO:0071393) |
| 0.1 | 0.3 | GO:0034164 | negative regulation of toll-like receptor 9 signaling pathway(GO:0034164) |
| 0.1 | 0.3 | GO:0048627 | myoblast development(GO:0048627) |
| 0.1 | 0.2 | GO:0002752 | cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 0.1 | 0.6 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.1 | 0.4 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.1 | 2.0 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.1 | 1.3 | GO:0050862 | positive regulation of T cell receptor signaling pathway(GO:0050862) |
| 0.1 | 0.8 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.1 | 1.0 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.0 | 2.9 | GO:0032467 | positive regulation of cytokinesis(GO:0032467) |
| 0.0 | 0.2 | GO:0060743 | epithelial cell maturation involved in prostate gland development(GO:0060743) |
| 0.0 | 0.6 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.0 | 0.7 | GO:0098909 | regulation of cardiac muscle cell action potential involved in regulation of contraction(GO:0098909) |
| 0.0 | 0.2 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.0 | 0.4 | GO:0010593 | negative regulation of lamellipodium assembly(GO:0010593) |
| 0.0 | 0.2 | GO:1904627 | response to phorbol 13-acetate 12-myristate(GO:1904627) cellular response to phorbol 13-acetate 12-myristate(GO:1904628) |
| 0.0 | 0.1 | GO:0019483 | beta-alanine biosynthetic process(GO:0019483) |
| 0.0 | 0.5 | GO:2000543 | positive regulation of gastrulation(GO:2000543) |
| 0.0 | 0.3 | GO:0010724 | regulation of definitive erythrocyte differentiation(GO:0010724) |
| 0.0 | 0.6 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.0 | 0.2 | GO:0048170 | positive regulation of long-term neuronal synaptic plasticity(GO:0048170) |
| 0.0 | 0.5 | GO:0016081 | synaptic vesicle docking(GO:0016081) |
| 0.0 | 0.2 | GO:0008215 | spermine metabolic process(GO:0008215) |
| 0.0 | 0.1 | GO:0008616 | queuosine biosynthetic process(GO:0008616) queuosine metabolic process(GO:0046116) |
| 0.0 | 0.1 | GO:0060738 | epithelial-mesenchymal signaling involved in prostate gland development(GO:0060738) |
| 0.0 | 0.5 | GO:0048368 | lateral mesoderm development(GO:0048368) |
| 0.0 | 0.3 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.0 | 0.1 | GO:0019556 | histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 0.0 | 0.5 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.0 | 0.2 | GO:1900454 | positive regulation of long term synaptic depression(GO:1900454) |
| 0.0 | 0.4 | GO:0030854 | positive regulation of granulocyte differentiation(GO:0030854) |
| 0.0 | 0.3 | GO:0042256 | mature ribosome assembly(GO:0042256) |
| 0.0 | 0.3 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.0 | 0.6 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.0 | 0.4 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.0 | 0.2 | GO:0003433 | chondrocyte development involved in endochondral bone morphogenesis(GO:0003433) |
| 0.0 | 0.1 | GO:0031247 | actin rod assembly(GO:0031247) |
| 0.0 | 0.2 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.0 | 0.1 | GO:1904529 | regulation of actin filament binding(GO:1904529) regulation of actin binding(GO:1904616) |
| 0.0 | 0.2 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.0 | 0.2 | GO:0060075 | regulation of resting membrane potential(GO:0060075) |
| 0.0 | 0.3 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.0 | 0.6 | GO:0000470 | maturation of LSU-rRNA(GO:0000470) |
| 0.0 | 0.2 | GO:0007217 | tachykinin receptor signaling pathway(GO:0007217) |
| 0.0 | 1.0 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 0.3 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.0 | 0.2 | GO:0002097 | tRNA wobble base modification(GO:0002097) |
| 0.0 | 0.1 | GO:0006011 | UDP-glucose metabolic process(GO:0006011) |
| 0.0 | 0.1 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.0 | 0.4 | GO:2000009 | negative regulation of protein localization to cell surface(GO:2000009) |
| 0.0 | 0.2 | GO:0019695 | choline metabolic process(GO:0019695) |
| 0.0 | 1.1 | GO:0046580 | negative regulation of Ras protein signal transduction(GO:0046580) negative regulation of small GTPase mediated signal transduction(GO:0051058) |
| 0.0 | 0.1 | GO:2001181 | positive regulation of interleukin-10 secretion(GO:2001181) |
| 0.0 | 0.7 | GO:0043403 | skeletal muscle tissue regeneration(GO:0043403) |
| 0.0 | 0.2 | GO:0045974 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.0 | 0.4 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.0 | 0.3 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 0.2 | GO:0000722 | telomere maintenance via recombination(GO:0000722) |
| 0.0 | 0.2 | GO:0000212 | meiotic spindle organization(GO:0000212) |
| 0.0 | 0.5 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 0.1 | GO:2000620 | innate vocalization behavior(GO:0098582) positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.0 | 0.2 | GO:0072619 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0070931 | Golgi-associated vesicle lumen(GO:0070931) |
| 0.1 | 0.5 | GO:0000126 | transcription factor TFIIIB complex(GO:0000126) |
| 0.1 | 0.5 | GO:0071149 | TEAD-2-YAP complex(GO:0071149) |
| 0.1 | 0.8 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.1 | 0.7 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.1 | 1.5 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.1 | 0.5 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.1 | 0.2 | GO:0097135 | cyclin E2-CDK2 complex(GO:0097135) |
| 0.1 | 1.5 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 0.1 | GO:0098855 | HCN channel complex(GO:0098855) |
| 0.0 | 0.2 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.0 | 0.3 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 0.0 | 0.2 | GO:0031510 | SUMO activating enzyme complex(GO:0031510) |
| 0.0 | 1.1 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.5 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.0 | 0.2 | GO:0035061 | interchromatin granule(GO:0035061) |
| 0.0 | 0.3 | GO:0071546 | pi-body(GO:0071546) |
| 0.0 | 0.2 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.0 | 0.2 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.0 | 0.6 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 0.2 | GO:0005827 | polar microtubule(GO:0005827) |
| 0.0 | 0.2 | GO:0045179 | apical cortex(GO:0045179) |
| 0.0 | 0.5 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.0 | 0.6 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.2 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.0 | 1.1 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 1.9 | GO:0031985 | Golgi cisterna(GO:0031985) |
| 0.0 | 0.9 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.4 | GO:0030137 | COPI-coated vesicle(GO:0030137) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 4.2 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.3 | 2.5 | GO:0043426 | MRF binding(GO:0043426) |
| 0.2 | 1.0 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.1 | 1.3 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.1 | 0.5 | GO:0001156 | TFIIIB-type transcription factor activity(GO:0001026) TFIIIC-class transcription factor binding(GO:0001156) |
| 0.1 | 0.4 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.1 | 2.3 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.1 | 0.5 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.1 | 0.7 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.1 | 0.4 | GO:0051022 | GDP-dissociation inhibitor binding(GO:0051021) Rho GDP-dissociation inhibitor binding(GO:0051022) |
| 0.1 | 1.9 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.1 | 0.2 | GO:0008281 | sulfonylurea receptor activity(GO:0008281) |
| 0.1 | 0.2 | GO:0098770 | FBXO family protein binding(GO:0098770) |
| 0.0 | 0.5 | GO:0071253 | connexin binding(GO:0071253) |
| 0.0 | 0.6 | GO:0070679 | inositol 1,4,5 trisphosphate binding(GO:0070679) |
| 0.0 | 0.4 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.0 | 0.3 | GO:1904929 | coreceptor activity involved in Wnt signaling pathway, planar cell polarity pathway(GO:1904929) |
| 0.0 | 0.2 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.0 | 0.6 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.0 | 0.8 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.1 | GO:0005222 | intracellular cAMP activated cation channel activity(GO:0005222) |
| 0.0 | 0.3 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.0 | 0.2 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.0 | 0.2 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 2.6 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 0.6 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.0 | 0.6 | GO:0098641 | GTP-Rho binding(GO:0017049) cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.5 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 0.2 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.0 | 0.5 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.0 | 0.2 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.0 | 0.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.0 | 0.1 | GO:0008479 | queuine tRNA-ribosyltransferase activity(GO:0008479) |
| 0.0 | 0.5 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.0 | 0.8 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.0 | 0.5 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.2 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.0 | 0.2 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 0.1 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.0 | 0.3 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.8 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 1.3 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.9 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 0.1 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.0 | 1.3 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.2 | GO:0015385 | sodium:proton antiporter activity(GO:0015385) |
| 0.0 | 0.2 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.0 | 0.3 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.0 | 0.1 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.0 | 0.2 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.0 | 0.4 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 0.2 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.5 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 2.5 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 1.0 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 1.1 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.0 | 0.6 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.0 | 0.6 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.6 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.0 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.1 | 3.7 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 1.7 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 0.6 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 0.5 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 0.5 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.0 | 1.1 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 0.6 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.0 | 1.3 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 0.4 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.6 | REACTOME RAS ACTIVATION UOPN CA2 INFUX THROUGH NMDA RECEPTOR | Genes involved in Ras activation uopn Ca2+ infux through NMDA receptor |
| 0.0 | 0.8 | REACTOME INWARDLY RECTIFYING K CHANNELS | Genes involved in Inwardly rectifying K+ channels |
| 0.0 | 0.3 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 1.0 | REACTOME EXTRACELLULAR MATRIX ORGANIZATION | Genes involved in Extracellular matrix organization |
| 0.0 | 0.2 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.5 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 1.0 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.0 | 0.2 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 1.2 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.0 | 1.4 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |