Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
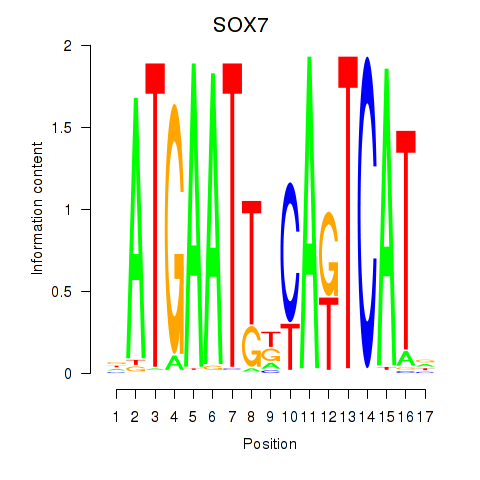
Results for SOX7
Z-value: 0.34
Transcription factors associated with SOX7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SOX7
|
ENSG00000171056.8 | SOX7 |
|
PINX1
|
ENSG00000258724.1 | PINX1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SOX7 | hg38_v1_chr8_-_10730498_10730525 | 0.13 | 5.0e-01 | Click! |
| PINX1 | hg38_v1_chr8_-_10839818_10839855 | 0.10 | 5.8e-01 | Click! |
Activity profile of SOX7 motif
Sorted Z-values of SOX7 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of SOX7
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_+_102995308 | 0.83 |
ENST00000612124.4
ENST00000374798.8 ENST00000487798.5 |
CYLC2
|
cylicin 2 |
| chr4_+_68447453 | 0.50 |
ENST00000305363.9
|
TMPRSS11E
|
transmembrane serine protease 11E |
| chr3_-_71305986 | 0.49 |
ENST00000647614.1
|
FOXP1
|
forkhead box P1 |
| chr12_-_56934403 | 0.38 |
ENST00000293502.2
|
SDR9C7
|
short chain dehydrogenase/reductase family 9C member 7 |
| chr7_+_134565098 | 0.25 |
ENST00000652743.1
|
AKR1B15
|
aldo-keto reductase family 1 member B15 |
| chr10_+_94683771 | 0.24 |
ENST00000339022.6
|
CYP2C18
|
cytochrome P450 family 2 subfamily C member 18 |
| chr17_-_35880350 | 0.24 |
ENST00000605140.6
ENST00000651122.1 ENST00000603197.6 |
CCL5
|
C-C motif chemokine ligand 5 |
| chr4_-_176269213 | 0.23 |
ENST00000296525.7
|
ASB5
|
ankyrin repeat and SOCS box containing 5 |
| chr16_+_33009175 | 0.22 |
ENST00000565407.2
|
IGHV3OR16-8
|
immunoglobulin heavy variable 3/OR16-8 (non-functional) |
| chr16_-_20352707 | 0.22 |
ENST00000396134.6
ENST00000573567.5 ENST00000570757.5 ENST00000396138.9 ENST00000571174.5 ENST00000576688.2 |
UMOD
|
uromodulin |
| chr14_-_106117159 | 0.22 |
ENST00000390601.3
|
IGHV3-11
|
immunoglobulin heavy variable 3-11 |
| chr16_-_33845229 | 0.21 |
ENST00000569103.2
|
IGHV3OR16-17
|
immunoglobulin heavy variable 3/OR16-17 (non-functional) |
| chr4_-_122456725 | 0.20 |
ENST00000226730.5
|
IL2
|
interleukin 2 |
| chr2_+_86907953 | 0.19 |
ENST00000409776.6
|
RGPD1
|
RANBP2 like and GRIP domain containing 1 |
| chrX_-_135781729 | 0.19 |
ENST00000617203.1
|
CT45A5
|
cancer/testis antigen family 45 member A5 |
| chr10_+_4963406 | 0.18 |
ENST00000380872.9
ENST00000442997.5 |
AKR1C1
|
aldo-keto reductase family 1 member C1 |
| chr4_-_47981535 | 0.16 |
ENST00000402813.9
|
CNGA1
|
cyclic nucleotide gated channel subunit alpha 1 |
| chrX_+_51710512 | 0.16 |
ENST00000602548.2
|
CENPVL1
|
centromere protein V like 1 |
| chr1_-_26067622 | 0.15 |
ENST00000374272.4
|
TRIM63
|
tripartite motif containing 63 |
| chr19_-_38426162 | 0.13 |
ENST00000587738.2
ENST00000586305.5 |
RASGRP4
|
RAS guanyl releasing protein 4 |
| chr2_-_89100352 | 0.12 |
ENST00000479981.1
|
IGKV1-16
|
immunoglobulin kappa variable 1-16 |
| chr6_+_116399395 | 0.12 |
ENST00000644499.1
|
ENSG00000285446.1
|
novel protein |
| chr1_+_100133135 | 0.12 |
ENST00000370143.5
ENST00000370141.7 |
TRMT13
|
tRNA methyltransferase 13 homolog |
| chr1_+_222928415 | 0.11 |
ENST00000284476.7
|
DISP1
|
dispatched RND transporter family member 1 |
| chr1_-_206003385 | 0.11 |
ENST00000617070.5
|
RAB7B
|
RAB7B, member RAS oncogene family |
| chr19_-_38426195 | 0.11 |
ENST00000615439.5
ENST00000614135.4 ENST00000622174.4 ENST00000587753.5 ENST00000454404.6 ENST00000617966.4 ENST00000618320.4 ENST00000293062.13 ENST00000433821.6 ENST00000426920.6 |
RASGRP4
|
RAS guanyl releasing protein 4 |
| chr1_-_206003442 | 0.10 |
ENST00000623893.1
|
RAB7B
|
RAB7B, member RAS oncogene family |
| chr6_+_143843316 | 0.10 |
ENST00000367576.6
|
LTV1
|
LTV1 ribosome biogenesis factor |
| chr16_+_28292485 | 0.10 |
ENST00000341901.5
|
SBK1
|
SH3 domain binding kinase 1 |
| chr6_+_150368892 | 0.09 |
ENST00000229447.9
ENST00000392256.6 |
IYD
|
iodotyrosine deiodinase |
| chr6_+_150368997 | 0.09 |
ENST00000392255.7
ENST00000500320.7 ENST00000344419.8 |
IYD
|
iodotyrosine deiodinase |
| chr19_-_5784599 | 0.09 |
ENST00000390672.2
ENST00000419421.3 |
PRR22
|
proline rich 22 |
| chr8_-_19602484 | 0.08 |
ENST00000454498.6
ENST00000520003.5 |
CSGALNACT1
|
chondroitin sulfate N-acetylgalactosaminyltransferase 1 |
| chr12_-_10420550 | 0.08 |
ENST00000381903.2
ENST00000396439.7 |
KLRC3
|
killer cell lectin like receptor C3 |
| chr3_-_71306012 | 0.08 |
ENST00000649431.1
ENST00000610810.5 |
FOXP1
|
forkhead box P1 |
| chr4_+_74308463 | 0.07 |
ENST00000413830.6
|
EPGN
|
epithelial mitogen |
| chr5_+_162067764 | 0.06 |
ENST00000639213.2
ENST00000414552.6 |
GABRG2
|
gamma-aminobutyric acid type A receptor subunit gamma2 |
| chr2_-_89222461 | 0.06 |
ENST00000482769.1
|
IGKV2-28
|
immunoglobulin kappa variable 2-28 |
| chr1_-_158426237 | 0.06 |
ENST00000641042.1
|
OR10K2
|
olfactory receptor family 10 subfamily K member 2 |
| chr11_+_35186820 | 0.05 |
ENST00000531110.6
ENST00000525685.6 |
CD44
|
CD44 molecule (Indian blood group) |
| chr12_+_63779894 | 0.05 |
ENST00000261234.11
|
RXYLT1
|
ribitol xylosyltransferase 1 |
| chr1_+_207325629 | 0.05 |
ENST00000618707.2
|
CD55
|
CD55 molecule (Cromer blood group) |
| chr5_+_162067458 | 0.05 |
ENST00000639975.1
ENST00000639111.2 ENST00000639683.1 |
GABRG2
|
gamma-aminobutyric acid type A receptor subunit gamma2 |
| chr10_+_24449426 | 0.04 |
ENST00000307544.10
|
KIAA1217
|
KIAA1217 |
| chr5_+_162067858 | 0.04 |
ENST00000361925.9
|
GABRG2
|
gamma-aminobutyric acid type A receptor subunit gamma2 |
| chr15_+_75347030 | 0.04 |
ENST00000566313.5
ENST00000355059.9 ENST00000568059.1 ENST00000568881.1 |
NEIL1
|
nei like DNA glycosylase 1 |
| chr6_-_111483700 | 0.03 |
ENST00000435970.5
ENST00000358835.7 |
REV3L
|
REV3 like, DNA directed polymerase zeta catalytic subunit |
| chr8_+_127409026 | 0.03 |
ENST00000465342.4
|
POU5F1B
|
POU class 5 homeobox 1B |
| chr8_-_86069662 | 0.03 |
ENST00000276616.3
|
PSKH2
|
protein serine kinase H2 |
| chr1_+_86993009 | 0.03 |
ENST00000370548.3
|
ENSG00000267561.2
|
novel protein |
| chr6_+_42563981 | 0.02 |
ENST00000372899.6
ENST00000372901.2 |
UBR2
|
ubiquitin protein ligase E3 component n-recognin 2 |
| chr8_+_12945667 | 0.02 |
ENST00000524591.7
|
TRMT9B
|
tRNA methyltransferase 9B (putative) |
| chrX_+_131083706 | 0.02 |
ENST00000370921.1
|
ARHGAP36
|
Rho GTPase activating protein 36 |
| chr8_+_28338640 | 0.02 |
ENST00000522209.1
|
PNOC
|
prepronociceptin |
| chr1_+_158931539 | 0.02 |
ENST00000368140.6
ENST00000368138.7 ENST00000392254.6 ENST00000392252.7 ENST00000368135.4 |
PYHIN1
|
pyrin and HIN domain family member 1 |
| chr11_+_56176618 | 0.02 |
ENST00000312298.1
|
OR5J2
|
olfactory receptor family 5 subfamily J member 2 |
| chr10_+_94402486 | 0.01 |
ENST00000225235.5
|
TBC1D12
|
TBC1 domain family member 12 |
| chr4_+_76074701 | 0.01 |
ENST00000355810.9
ENST00000349321.7 |
ART3
|
ADP-ribosyltransferase 3 (inactive) |
| chr6_-_85594063 | 0.01 |
ENST00000684150.1
|
SNX14
|
sorting nexin 14 |
| chr6_-_85594101 | 0.01 |
ENST00000682171.1
ENST00000505648.5 |
SNX14
|
sorting nexin 14 |
| chr11_+_7597182 | 0.01 |
ENST00000528883.5
|
PPFIBP2
|
PPFIA binding protein 2 |
| chr5_+_162067500 | 0.01 |
ENST00000639384.1
ENST00000640985.1 ENST00000638772.1 |
GABRG2
|
gamma-aminobutyric acid type A receptor subunit gamma2 |
| chr1_+_200027702 | 0.00 |
ENST00000367362.8
|
NR5A2
|
nuclear receptor subfamily 5 group A member 2 |
| chr19_-_2783241 | 0.00 |
ENST00000676943.1
ENST00000589251.5 ENST00000221566.7 ENST00000676984.1 |
SGTA
|
small glutamine rich tetratricopeptide repeat containing alpha |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0072023 | thick ascending limb development(GO:0072023) metanephric thick ascending limb development(GO:0072233) |
| 0.1 | 0.2 | GO:0033634 | positive regulation of cell-cell adhesion mediated by integrin(GO:0033634) |
| 0.1 | 0.2 | GO:0034164 | negative regulation of toll-like receptor 9 signaling pathway(GO:0034164) |
| 0.0 | 0.2 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.0 | 0.1 | GO:0007225 | patched ligand maturation(GO:0007225) signal maturation(GO:0035638) |
| 0.0 | 0.1 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) |
| 0.0 | 0.6 | GO:0072619 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.0 | 0.2 | GO:0009753 | response to jasmonic acid(GO:0009753) cellular response to jasmonic acid stimulus(GO:0071395) |
| 0.0 | 0.1 | GO:0050653 | chondroitin sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0050653) |
| 0.0 | 0.3 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.0 | 0.2 | GO:0006570 | tyrosine metabolic process(GO:0006570) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0033150 | cytoskeletal calyx(GO:0033150) |
| 0.0 | 0.1 | GO:0034448 | EGO complex(GO:0034448) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 0.0 | 0.2 | GO:0031726 | receptor signaling protein tyrosine kinase activator activity(GO:0030298) CCR1 chemokine receptor binding(GO:0031726) |
| 0.0 | 0.2 | GO:0004447 | iodide peroxidase activity(GO:0004447) |
| 0.0 | 0.2 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 0.0 | 0.3 | GO:0004303 | estradiol 17-beta-dehydrogenase activity(GO:0004303) |
| 0.0 | 0.4 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 0.0 | 0.1 | GO:0047237 | glucuronylgalactosylproteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047237) glucuronosyl-N-acetylgalactosaminyl-proteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047238) |
| 0.0 | 0.2 | GO:0005221 | intracellular cyclic nucleotide activated cation channel activity(GO:0005221) cyclic nucleotide-gated ion channel activity(GO:0043855) |
| 0.0 | 0.2 | GO:0019864 | IgG binding(GO:0019864) |
| 0.0 | 0.2 | GO:0008391 | arachidonic acid monooxygenase activity(GO:0008391) arachidonic acid epoxygenase activity(GO:0008392) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |