Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
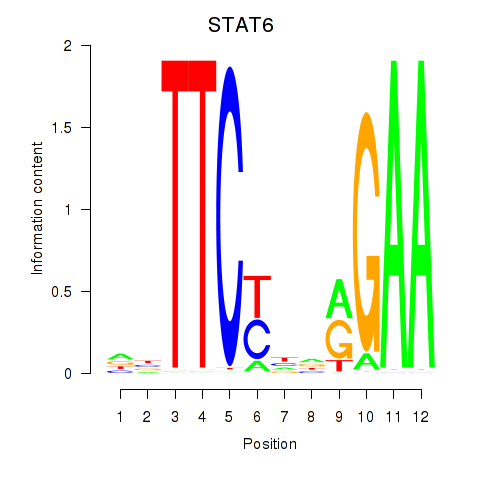
Results for STAT6
Z-value: 0.85
Transcription factors associated with STAT6
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
STAT6
|
ENSG00000166888.12 | STAT6 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| STAT6 | hg38_v1_chr12_-_57111338_57111433 | 0.11 | 5.7e-01 | Click! |
Activity profile of STAT6 motif
Sorted Z-values of STAT6 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of STAT6
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_205422050 | 10.05 |
ENST00000367153.9
|
LEMD1
|
LEM domain containing 1 |
| chrX_+_136169624 | 3.23 |
ENST00000394153.6
|
FHL1
|
four and a half LIM domains 1 |
| chrX_+_136169833 | 3.17 |
ENST00000628032.2
|
FHL1
|
four and a half LIM domains 1 |
| chrX_+_136169664 | 2.93 |
ENST00000456445.5
|
FHL1
|
four and a half LIM domains 1 |
| chr10_+_100347225 | 2.69 |
ENST00000370355.3
|
SCD
|
stearoyl-CoA desaturase |
| chr2_+_113127588 | 2.66 |
ENST00000409930.4
|
IL1RN
|
interleukin 1 receptor antagonist |
| chrX_+_136169891 | 2.25 |
ENST00000449474.5
|
FHL1
|
four and a half LIM domains 1 |
| chr17_-_17836997 | 2.23 |
ENST00000395757.6
|
SREBF1
|
sterol regulatory element binding transcription factor 1 |
| chr11_-_128587551 | 2.09 |
ENST00000392668.8
|
ETS1
|
ETS proto-oncogene 1, transcription factor |
| chr1_-_160646958 | 1.86 |
ENST00000538290.2
|
SLAMF1
|
signaling lymphocytic activation molecule family member 1 |
| chr11_+_70078291 | 1.77 |
ENST00000355303.9
|
ANO1
|
anoctamin 1 |
| chr4_+_54100161 | 1.75 |
ENST00000326902.7
ENST00000503800.1 |
GSX2
|
GS homeobox 2 |
| chr3_+_50269140 | 1.62 |
ENST00000616701.5
ENST00000433753.4 ENST00000611067.4 |
SEMA3B
|
semaphorin 3B |
| chr17_-_17836973 | 1.59 |
ENST00000261646.11
ENST00000355815.8 |
SREBF1
|
sterol regulatory element binding transcription factor 1 |
| chr9_+_36572854 | 1.59 |
ENST00000543751.5
ENST00000536860.5 ENST00000541717.4 ENST00000536329.5 ENST00000536987.5 ENST00000298048.7 ENST00000495529.5 ENST00000545008.5 |
MELK
|
maternal embryonic leucine zipper kinase |
| chr13_+_48653921 | 1.53 |
ENST00000682523.1
|
CYSLTR2
|
cysteinyl leukotriene receptor 2 |
| chr12_-_27971970 | 1.50 |
ENST00000395872.5
ENST00000201015.8 |
PTHLH
|
parathyroid hormone like hormone |
| chr11_-_68213577 | 1.45 |
ENST00000402789.5
ENST00000402185.6 ENST00000458496.1 |
KMT5B
|
lysine methyltransferase 5B |
| chr2_-_189179754 | 1.41 |
ENST00000374866.9
ENST00000618828.1 |
COL5A2
|
collagen type V alpha 2 chain |
| chr1_-_8026283 | 1.39 |
ENST00000474874.5
ENST00000469499.5 ENST00000377482.10 |
ERRFI1
|
ERBB receptor feedback inhibitor 1 |
| chrX_-_107717054 | 1.37 |
ENST00000503515.1
ENST00000372397.6 |
TSC22D3
|
TSC22 domain family member 3 |
| chr5_-_151686908 | 1.35 |
ENST00000231061.9
|
SPARC
|
secreted protein acidic and cysteine rich |
| chr14_-_68979314 | 1.27 |
ENST00000684713.1
ENST00000683198.1 ENST00000684598.1 ENST00000682331.1 ENST00000682291.1 ENST00000683342.1 |
ACTN1
|
actinin alpha 1 |
| chr14_-_68979436 | 1.26 |
ENST00000193403.10
|
ACTN1
|
actinin alpha 1 |
| chr8_+_49072335 | 1.26 |
ENST00000399653.8
ENST00000522267.6 ENST00000303202.8 |
PPDPFL
|
pancreatic progenitor cell differentiation and proliferation factor like |
| chr12_+_8992029 | 1.26 |
ENST00000543895.1
|
KLRG1
|
killer cell lectin like receptor G1 |
| chr22_+_44069043 | 1.24 |
ENST00000404989.1
|
PARVB
|
parvin beta |
| chr11_+_65314853 | 1.21 |
ENST00000279249.3
|
CDC42EP2
|
CDC42 effector protein 2 |
| chr2_+_190927649 | 1.19 |
ENST00000409428.5
ENST00000409215.5 |
GLS
|
glutaminase |
| chr15_-_29822028 | 1.18 |
ENST00000545208.6
|
TJP1
|
tight junction protein 1 |
| chr15_-_29822077 | 1.10 |
ENST00000677774.1
|
TJP1
|
tight junction protein 1 |
| chr15_+_66453418 | 1.04 |
ENST00000566326.1
|
MAP2K1
|
mitogen-activated protein kinase kinase 1 |
| chr1_-_75932392 | 1.04 |
ENST00000284142.7
|
ASB17
|
ankyrin repeat and SOCS box containing 17 |
| chr17_-_41350824 | 1.02 |
ENST00000007735.4
|
KRT33A
|
keratin 33A |
| chr10_+_74176741 | 0.97 |
ENST00000673027.1
ENST00000372734.5 ENST00000541550.6 |
ADK
|
adenosine kinase |
| chr8_+_11809135 | 0.97 |
ENST00000528643.5
ENST00000525777.5 |
FDFT1
|
farnesyl-diphosphate farnesyltransferase 1 |
| chr16_+_30064142 | 0.93 |
ENST00000562168.5
ENST00000569545.5 |
ALDOA
|
aldolase, fructose-bisphosphate A |
| chr1_-_160647037 | 0.92 |
ENST00000302035.11
|
SLAMF1
|
signaling lymphocytic activation molecule family member 1 |
| chr15_-_29822418 | 0.91 |
ENST00000614355.5
ENST00000495972.6 ENST00000346128.10 |
TJP1
|
tight junction protein 1 |
| chr1_-_120100688 | 0.91 |
ENST00000652264.1
|
NOTCH2
|
notch receptor 2 |
| chr4_+_72031902 | 0.90 |
ENST00000344413.6
ENST00000308744.12 |
NPFFR2
|
neuropeptide FF receptor 2 |
| chr1_+_156282917 | 0.88 |
ENST00000295694.9
ENST00000357501.6 |
TMEM79
|
transmembrane protein 79 |
| chr3_-_73624840 | 0.86 |
ENST00000308537.4
ENST00000263666.9 |
PDZRN3
|
PDZ domain containing ring finger 3 |
| chr20_+_59676661 | 0.84 |
ENST00000355648.8
|
PHACTR3
|
phosphatase and actin regulator 3 |
| chr1_+_150272772 | 0.84 |
ENST00000369098.3
ENST00000369099.8 |
C1orf54
|
chromosome 1 open reading frame 54 |
| chr6_-_116829037 | 0.83 |
ENST00000368549.7
ENST00000530250.1 ENST00000310357.8 |
GPRC6A
|
G protein-coupled receptor class C group 6 member A |
| chr2_+_68734773 | 0.82 |
ENST00000409202.8
|
ARHGAP25
|
Rho GTPase activating protein 25 |
| chr4_+_6693870 | 0.82 |
ENST00000296370.4
|
S100P
|
S100 calcium binding protein P |
| chr1_-_40862354 | 0.81 |
ENST00000372638.4
|
CITED4
|
Cbp/p300 interacting transactivator with Glu/Asp rich carboxy-terminal domain 4 |
| chr16_+_30064274 | 0.79 |
ENST00000563060.6
|
ALDOA
|
aldolase, fructose-bisphosphate A |
| chr1_-_160647287 | 0.79 |
ENST00000235739.6
|
SLAMF1
|
signaling lymphocytic activation molecule family member 1 |
| chr3_-_10108231 | 0.78 |
ENST00000524279.1
ENST00000450660.3 |
FANCD2OS
|
FANCD2 opposite strand |
| chr3_-_98517096 | 0.78 |
ENST00000513873.1
|
CLDND1
|
claudin domain containing 1 |
| chr19_+_2785481 | 0.75 |
ENST00000307741.11
ENST00000585338.1 |
THOP1
|
thimet oligopeptidase 1 |
| chr11_-_60952559 | 0.74 |
ENST00000538739.2
|
SLC15A3
|
solute carrier family 15 member 3 |
| chr16_+_30064462 | 0.72 |
ENST00000412304.6
|
ALDOA
|
aldolase, fructose-bisphosphate A |
| chr3_+_189789734 | 0.71 |
ENST00000437221.5
ENST00000392463.6 ENST00000392461.7 ENST00000449992.5 ENST00000456148.1 |
TP63
|
tumor protein p63 |
| chr6_-_27912396 | 0.70 |
ENST00000303324.4
|
OR2B2
|
olfactory receptor family 2 subfamily B member 2 |
| chr3_+_159069252 | 0.70 |
ENST00000640015.1
ENST00000476809.7 ENST00000485419.7 |
IQCJ-SCHIP1
|
IQCJ-SCHIP1 readthrough |
| chr2_+_201451711 | 0.69 |
ENST00000194530.8
ENST00000392249.6 |
STRADB
|
STE20 related adaptor beta |
| chr2_-_133568393 | 0.68 |
ENST00000317721.10
ENST00000405974.7 ENST00000409261.6 ENST00000409213.5 |
NCKAP5
|
NCK associated protein 5 |
| chr3_+_159273235 | 0.67 |
ENST00000638749.1
|
IQCJ-SCHIP1
|
IQCJ-SCHIP1 readthrough |
| chr6_+_30557287 | 0.66 |
ENST00000376560.8
|
PRR3
|
proline rich 3 |
| chr12_-_54385727 | 0.66 |
ENST00000551109.5
ENST00000546970.5 |
ZNF385A
|
zinc finger protein 385A |
| chr11_-_60952067 | 0.65 |
ENST00000681275.1
|
SLC15A3
|
solute carrier family 15 member 3 |
| chr8_-_42768602 | 0.63 |
ENST00000534622.5
|
CHRNA6
|
cholinergic receptor nicotinic alpha 6 subunit |
| chr6_+_30557274 | 0.63 |
ENST00000376557.3
|
PRR3
|
proline rich 3 |
| chr8_-_42768781 | 0.63 |
ENST00000276410.7
|
CHRNA6
|
cholinergic receptor nicotinic alpha 6 subunit |
| chr1_+_213989691 | 0.62 |
ENST00000607425.1
|
PROX1
|
prospero homeobox 1 |
| chr10_+_96304392 | 0.60 |
ENST00000630152.1
|
DNTT
|
DNA nucleotidylexotransferase |
| chr10_+_96304425 | 0.60 |
ENST00000371174.5
|
DNTT
|
DNA nucleotidylexotransferase |
| chr3_+_189789643 | 0.59 |
ENST00000354600.10
|
TP63
|
tumor protein p63 |
| chr19_-_54100792 | 0.59 |
ENST00000391761.5
ENST00000356532.7 ENST00000616447.4 ENST00000359649.8 ENST00000358375.8 ENST00000391760.1 ENST00000351806.8 |
OSCAR
|
osteoclast associated Ig-like receptor |
| chr9_-_128724088 | 0.57 |
ENST00000406904.2
ENST00000452105.5 ENST00000372667.9 ENST00000372663.9 |
ZDHHC12
|
zinc finger DHHC-type palmitoyltransferase 12 |
| chr5_+_127649018 | 0.55 |
ENST00000379445.7
|
CTXN3
|
cortexin 3 |
| chr18_-_26865732 | 0.54 |
ENST00000672188.1
|
AQP4
|
aquaporin 4 |
| chr18_-_26865689 | 0.54 |
ENST00000675739.1
ENST00000383168.9 ENST00000672981.2 ENST00000578776.1 |
AQP4
|
aquaporin 4 |
| chr6_-_41201128 | 0.54 |
ENST00000483722.2
|
TREML2
|
triggering receptor expressed on myeloid cells like 2 |
| chr11_-_60952134 | 0.53 |
ENST00000679573.1
ENST00000681882.1 ENST00000681951.1 ENST00000227880.8 |
SLC15A3
|
solute carrier family 15 member 3 |
| chr2_+_131011683 | 0.51 |
ENST00000355771.7
|
ARHGEF4
|
Rho guanine nucleotide exchange factor 4 |
| chr6_-_41286665 | 0.51 |
ENST00000589614.5
ENST00000244709.9 ENST00000334475.10 ENST00000591620.1 |
TREM1
|
triggering receptor expressed on myeloid cells 1 |
| chr7_+_129375643 | 0.51 |
ENST00000490911.5
|
AHCYL2
|
adenosylhomocysteinase like 2 |
| chr3_+_35641421 | 0.51 |
ENST00000449196.5
|
ARPP21
|
cAMP regulated phosphoprotein 21 |
| chr1_+_159437845 | 0.50 |
ENST00000642080.1
|
OR10J1
|
olfactory receptor family 10 subfamily J member 1 |
| chr10_+_12068945 | 0.49 |
ENST00000263035.9
|
DHTKD1
|
dehydrogenase E1 and transketolase domain containing 1 |
| chr9_+_87497222 | 0.49 |
ENST00000358077.9
|
DAPK1
|
death associated protein kinase 1 |
| chr11_+_112961402 | 0.47 |
ENST00000613217.4
ENST00000316851.12 ENST00000620046.4 ENST00000531044.5 ENST00000529356.5 |
NCAM1
|
neural cell adhesion molecule 1 |
| chr12_+_43758936 | 0.47 |
ENST00000440781.6
ENST00000431837.5 ENST00000550616.5 ENST00000613694.5 ENST00000551736.5 |
IRAK4
|
interleukin 1 receptor associated kinase 4 |
| chr12_-_70637405 | 0.46 |
ENST00000548122.2
ENST00000551525.5 ENST00000550358.5 ENST00000334414.11 |
PTPRB
|
protein tyrosine phosphatase receptor type B |
| chr15_-_23687290 | 0.45 |
ENST00000649030.2
|
NDN
|
necdin, MAGE family member |
| chr9_+_87497675 | 0.45 |
ENST00000472284.5
ENST00000469640.6 |
DAPK1
|
death associated protein kinase 1 |
| chr4_-_120066777 | 0.44 |
ENST00000296509.11
|
MAD2L1
|
mitotic arrest deficient 2 like 1 |
| chr1_+_26432803 | 0.43 |
ENST00000430232.5
|
DHDDS
|
dehydrodolichyl diphosphate synthase subunit |
| chr2_-_203535253 | 0.43 |
ENST00000457812.5
ENST00000319170.10 ENST00000630330.2 ENST00000308091.8 ENST00000453034.5 ENST00000420371.2 |
RAPH1
|
Ras association (RalGDS/AF-6) and pleckstrin homology domains 1 |
| chr15_+_91100194 | 0.40 |
ENST00000394232.6
|
SV2B
|
synaptic vesicle glycoprotein 2B |
| chrX_-_139832235 | 0.40 |
ENST00000327569.7
ENST00000361648.6 |
ATP11C
|
ATPase phospholipid transporting 11C |
| chr11_-_59212869 | 0.40 |
ENST00000361050.4
|
MPEG1
|
macrophage expressed 1 |
| chr7_+_32979445 | 0.38 |
ENST00000418354.5
ENST00000490776.3 |
FKBP9
|
FKBP prolyl isomerase 9 |
| chr13_-_74133892 | 0.38 |
ENST00000377669.7
|
KLF12
|
Kruppel like factor 12 |
| chr3_+_141262614 | 0.38 |
ENST00000504264.5
|
PXYLP1
|
2-phosphoxylose phosphatase 1 |
| chr17_+_7481332 | 0.38 |
ENST00000412468.3
|
SLC35G6
|
solute carrier family 35 member G6 |
| chr3_-_57199938 | 0.38 |
ENST00000473921.2
ENST00000295934.8 |
HESX1
|
HESX homeobox 1 |
| chr9_+_137742957 | 0.36 |
ENST00000637318.1
ENST00000478940.1 |
EHMT1
|
euchromatic histone lysine methyltransferase 1 |
| chr11_+_65333834 | 0.36 |
ENST00000528416.6
ENST00000415073.6 ENST00000252268.8 |
DPF2
|
double PHD fingers 2 |
| chr7_-_77199808 | 0.35 |
ENST00000248598.6
|
FGL2
|
fibrinogen like 2 |
| chr4_-_115113822 | 0.35 |
ENST00000613194.4
|
NDST4
|
N-deacetylase and N-sulfotransferase 4 |
| chr9_+_87497852 | 0.34 |
ENST00000408954.8
|
DAPK1
|
death associated protein kinase 1 |
| chr1_-_67833448 | 0.34 |
ENST00000370982.4
|
GNG12
|
G protein subunit gamma 12 |
| chr16_-_29745951 | 0.34 |
ENST00000329410.4
|
C16orf54
|
chromosome 16 open reading frame 54 |
| chr11_+_112961247 | 0.33 |
ENST00000621518.4
ENST00000618266.4 ENST00000615112.4 ENST00000615285.4 ENST00000619839.4 ENST00000401611.6 ENST00000621128.4 |
NCAM1
|
neural cell adhesion molecule 1 |
| chrX_-_101291325 | 0.32 |
ENST00000356784.2
|
TAF7L
|
TATA-box binding protein associated factor 7 like |
| chr14_-_37172553 | 0.32 |
ENST00000331299.6
|
SLC25A21
|
solute carrier family 25 member 21 |
| chr12_-_52703522 | 0.31 |
ENST00000341809.8
|
KRT77
|
keratin 77 |
| chr12_+_56041893 | 0.31 |
ENST00000552361.1
ENST00000646449.2 |
RPS26
|
ribosomal protein S26 |
| chr7_+_112480853 | 0.30 |
ENST00000439068.6
ENST00000312849.4 |
LSMEM1
|
leucine rich single-pass membrane protein 1 |
| chr1_-_108192818 | 0.30 |
ENST00000370041.4
|
SLC25A24
|
solute carrier family 25 member 24 |
| chr2_+_233778330 | 0.30 |
ENST00000389758.3
|
MROH2A
|
maestro heat like repeat family member 2A |
| chr8_+_10095704 | 0.30 |
ENST00000382490.9
|
MSRA
|
methionine sulfoxide reductase A |
| chr17_+_36211055 | 0.29 |
ENST00000617405.5
ENST00000617416.4 ENST00000613173.4 ENST00000620732.4 ENST00000620098.4 ENST00000620576.4 ENST00000620055.4 ENST00000610565.4 ENST00000620250.1 |
CCL4L2
|
C-C motif chemokine ligand 4 like 2 |
| chr6_+_116529013 | 0.29 |
ENST00000628083.1
ENST00000368597.6 ENST00000452373.5 ENST00000405399.5 |
CALHM4
|
calcium homeostasis modulator family member 4 |
| chr6_-_11232658 | 0.29 |
ENST00000379433.5
ENST00000379446.10 ENST00000620854.4 |
NEDD9
|
neural precursor cell expressed, developmentally down-regulated 9 |
| chr1_+_42153399 | 0.28 |
ENST00000372581.2
|
GUCA2B
|
guanylate cyclase activator 2B |
| chr2_+_196639686 | 0.28 |
ENST00000389175.9
|
CCDC150
|
coiled-coil domain containing 150 |
| chr7_-_75772172 | 0.28 |
ENST00000005180.9
|
CCL26
|
C-C motif chemokine ligand 26 |
| chr20_+_43945677 | 0.27 |
ENST00000358131.5
|
TOX2
|
TOX high mobility group box family member 2 |
| chr13_-_51845169 | 0.27 |
ENST00000627246.3
ENST00000629372.3 |
TMEM272
|
transmembrane protein 272 |
| chr2_+_167248638 | 0.27 |
ENST00000295237.10
|
XIRP2
|
xin actin binding repeat containing 2 |
| chr8_+_10095551 | 0.26 |
ENST00000522907.5
ENST00000528246.5 |
MSRA
|
methionine sulfoxide reductase A |
| chr16_-_67660694 | 0.26 |
ENST00000219251.13
ENST00000620338.4 |
ACD
|
ACD shelterin complex subunit and telomerase recruitment factor |
| chr12_-_124495252 | 0.26 |
ENST00000405201.5
|
NCOR2
|
nuclear receptor corepressor 2 |
| chr9_-_70869076 | 0.26 |
ENST00000677594.1
|
TRPM3
|
transient receptor potential cation channel subfamily M member 3 |
| chrX_-_50643649 | 0.26 |
ENST00000460112.3
|
SHROOM4
|
shroom family member 4 |
| chr17_+_36210924 | 0.25 |
ENST00000615418.4
|
CCL4L2
|
C-C motif chemokine ligand 4 like 2 |
| chr15_+_84981981 | 0.25 |
ENST00000339708.9
|
PDE8A
|
phosphodiesterase 8A |
| chrX_+_149929573 | 0.25 |
ENST00000457775.3
|
HSFX4
|
heat shock transcription factor family, X-linked member 4 |
| chr14_+_38207893 | 0.25 |
ENST00000267377.3
|
SSTR1
|
somatostatin receptor 1 |
| chr5_+_150671588 | 0.25 |
ENST00000523553.1
|
MYOZ3
|
myozenin 3 |
| chr1_-_153312919 | 0.25 |
ENST00000683862.1
|
PGLYRP3
|
peptidoglycan recognition protein 3 |
| chr12_-_32755876 | 0.24 |
ENST00000324868.13
|
YARS2
|
tyrosyl-tRNA synthetase 2 |
| chr8_-_81447428 | 0.24 |
ENST00000256103.3
ENST00000519260.1 |
PMP2
|
peripheral myelin protein 2 |
| chrX_+_102937272 | 0.24 |
ENST00000218249.7
|
RAB40AL
|
RAB40A like |
| chr5_-_88823763 | 0.23 |
ENST00000635898.1
ENST00000626391.2 ENST00000628656.2 |
MEF2C
|
myocyte enhancer factor 2C |
| chr18_+_35581734 | 0.23 |
ENST00000591924.5
ENST00000269195.6 |
GALNT1
|
polypeptide N-acetylgalactosaminyltransferase 1 |
| chr19_+_14941489 | 0.23 |
ENST00000248072.3
|
OR7C2
|
olfactory receptor family 7 subfamily C member 2 |
| chr14_-_77423517 | 0.22 |
ENST00000555603.1
|
NOXRED1
|
NADP dependent oxidoreductase domain containing 1 |
| chr15_+_84981834 | 0.22 |
ENST00000394553.6
|
PDE8A
|
phosphodiesterase 8A |
| chr12_+_51888217 | 0.22 |
ENST00000340970.8
|
ANKRD33
|
ankyrin repeat domain 33 |
| chr6_+_83512501 | 0.22 |
ENST00000369700.4
|
PRSS35
|
serine protease 35 |
| chr14_+_22052503 | 0.21 |
ENST00000390449.3
|
TRAV21
|
T cell receptor alpha variable 21 |
| chr6_-_32853813 | 0.21 |
ENST00000643049.2
|
TAP1
|
transporter 1, ATP binding cassette subfamily B member |
| chr12_+_51888083 | 0.21 |
ENST00000301190.11
|
ANKRD33
|
ankyrin repeat domain 33 |
| chr12_-_81758665 | 0.21 |
ENST00000549325.5
ENST00000550584.6 |
PPFIA2
|
PTPRF interacting protein alpha 2 |
| chr6_+_28080565 | 0.20 |
ENST00000377325.2
ENST00000683778.1 |
ZNF165
|
zinc finger protein 165 |
| chr15_+_67548992 | 0.20 |
ENST00000354498.9
|
MAP2K5
|
mitogen-activated protein kinase kinase 5 |
| chr20_-_56525925 | 0.20 |
ENST00000243913.8
|
GCNT7
|
glucosaminyl (N-acetyl) transferase family member 7 |
| chr7_-_139716980 | 0.20 |
ENST00000342645.7
|
HIPK2
|
homeodomain interacting protein kinase 2 |
| chr6_-_32853618 | 0.20 |
ENST00000354258.5
|
TAP1
|
transporter 1, ATP binding cassette subfamily B member |
| chr5_+_7396099 | 0.20 |
ENST00000338316.9
|
ADCY2
|
adenylate cyclase 2 |
| chr1_+_77918128 | 0.20 |
ENST00000342754.5
|
NEXN
|
nexilin F-actin binding protein |
| chr2_-_202870761 | 0.19 |
ENST00000420558.5
ENST00000418208.5 |
ICA1L
|
islet cell autoantigen 1 like |
| chr5_+_147878703 | 0.19 |
ENST00000296694.5
|
SCGB3A2
|
secretoglobin family 3A member 2 |
| chr17_-_64230727 | 0.19 |
ENST00000583097.5
ENST00000615733.4 |
TEX2
|
testis expressed 2 |
| chr3_+_121593363 | 0.18 |
ENST00000338040.6
|
FBXO40
|
F-box protein 40 |
| chr9_-_132079856 | 0.18 |
ENST00000651555.1
ENST00000651950.1 ENST00000357028.6 ENST00000474263.1 ENST00000292035.10 |
MED27
|
mediator complex subunit 27 |
| chr1_+_192636121 | 0.17 |
ENST00000543215.5
ENST00000391995.7 |
RGS13
|
regulator of G protein signaling 13 |
| chr12_-_6342066 | 0.17 |
ENST00000162749.7
ENST00000440083.6 |
TNFRSF1A
|
TNF receptor superfamily member 1A |
| chr7_-_123199960 | 0.17 |
ENST00000194130.7
|
SLC13A1
|
solute carrier family 13 member 1 |
| chr12_+_6384144 | 0.16 |
ENST00000228918.9
ENST00000536876.5 ENST00000543190.5 |
LTBR
|
lymphotoxin beta receptor |
| chrX_-_135764444 | 0.16 |
ENST00000597510.6
|
CT45A3
|
cancer/testis antigen family 45 member A3 |
| chr8_-_116874746 | 0.15 |
ENST00000297338.7
|
RAD21
|
RAD21 cohesin complex component |
| chr6_+_31571957 | 0.15 |
ENST00000454783.5
|
LTA
|
lymphotoxin alpha |
| chr11_-_790062 | 0.15 |
ENST00000330106.5
|
CEND1
|
cell cycle exit and neuronal differentiation 1 |
| chr19_+_13118235 | 0.15 |
ENST00000292431.5
|
NACC1
|
nucleus accumbens associated 1 |
| chr2_-_229921316 | 0.13 |
ENST00000428959.5
ENST00000675423.1 |
TRIP12
|
thyroid hormone receptor interactor 12 |
| chr2_+_27217361 | 0.12 |
ENST00000264705.9
ENST00000403525.5 |
CAD
|
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase |
| chr14_+_21918161 | 0.11 |
ENST00000390439.2
|
TRAV13-2
|
T cell receptor alpha variable 13-2 |
| chr5_+_139273752 | 0.11 |
ENST00000509990.5
ENST00000506147.5 ENST00000512107.5 |
MATR3
|
matrin 3 |
| chr12_-_81759307 | 0.11 |
ENST00000547623.5
ENST00000549396.6 |
PPFIA2
|
PTPRF interacting protein alpha 2 |
| chr4_+_70472189 | 0.10 |
ENST00000304887.6
|
MUC7
|
mucin 7, secreted |
| chr14_-_24609660 | 0.10 |
ENST00000557220.6
ENST00000216338.9 ENST00000382548.4 |
GZMH
|
granzyme H |
| chrX_-_73214793 | 0.10 |
ENST00000373517.4
|
NAP1L2
|
nucleosome assembly protein 1 like 2 |
| chr17_-_3298360 | 0.09 |
ENST00000323404.2
|
OR3A1
|
olfactory receptor family 3 subfamily A member 1 |
| chr22_-_29061831 | 0.09 |
ENST00000216071.5
|
C22orf31
|
chromosome 22 open reading frame 31 |
| chr3_-_185938006 | 0.09 |
ENST00000342294.4
ENST00000453386.7 ENST00000382191.4 |
TRA2B
|
transformer 2 beta homolog |
| chrX_-_140784366 | 0.09 |
ENST00000674533.1
|
CDR1
|
cerebellar degeneration related protein 1 |
| chr10_-_59362460 | 0.08 |
ENST00000422313.6
ENST00000435852.6 ENST00000614220.4 ENST00000618804.5 ENST00000621119.4 |
FAM13C
|
family with sequence similarity 13 member C |
| chr12_-_81758641 | 0.08 |
ENST00000552948.5
ENST00000548586.5 |
PPFIA2
|
PTPRF interacting protein alpha 2 |
| chr8_+_103298836 | 0.08 |
ENST00000523739.5
ENST00000358755.5 |
FZD6
|
frizzled class receptor 6 |
| chr1_-_173603041 | 0.08 |
ENST00000367714.4
|
SLC9C2
|
solute carrier family 9 member C2 (putative) |
| chr1_-_17045219 | 0.08 |
ENST00000491274.5
|
SDHB
|
succinate dehydrogenase complex iron sulfur subunit B |
| chr1_-_115841116 | 0.07 |
ENST00000320238.3
|
NHLH2
|
nescient helix-loop-helix 2 |
| chr3_+_141402322 | 0.07 |
ENST00000510338.5
ENST00000504673.5 |
ZBTB38
|
zinc finger and BTB domain containing 38 |
| chr10_-_95693893 | 0.07 |
ENST00000371209.5
ENST00000680144.1 ENST00000430368.6 ENST00000371217.10 |
TCTN3
|
tectonic family member 3 |
| chr17_+_7558296 | 0.07 |
ENST00000438470.5
ENST00000436057.5 |
TNFSF13
|
TNF superfamily member 13 |
| chr20_-_23086316 | 0.07 |
ENST00000246006.5
|
CD93
|
CD93 molecule |
| chr11_+_63888515 | 0.07 |
ENST00000509502.6
ENST00000512060.1 |
MARK2
|
microtubule affinity regulating kinase 2 |
| chr1_+_202462730 | 0.07 |
ENST00000290419.9
ENST00000491336.5 |
PPP1R12B
|
protein phosphatase 1 regulatory subunit 12B |
| chr15_+_91099943 | 0.07 |
ENST00000545111.6
|
SV2B
|
synaptic vesicle glycoprotein 2B |
| chr12_+_26011713 | 0.06 |
ENST00000542004.5
|
RASSF8
|
Ras association domain family member 8 |
| chr17_-_35880350 | 0.06 |
ENST00000605140.6
ENST00000651122.1 ENST00000603197.6 |
CCL5
|
C-C motif chemokine ligand 5 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.6 | GO:0035744 | T-helper 1 cell cytokine production(GO:0035744) |
| 0.6 | 3.2 | GO:1901350 | cell-cell signaling involved in cell-cell junction organization(GO:1901350) |
| 0.5 | 1.4 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.3 | 2.7 | GO:2000660 | negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 0.3 | 3.8 | GO:0045542 | positive regulation of cholesterol biosynthetic process(GO:0045542) |
| 0.3 | 1.1 | GO:0007499 | ectoderm and mesoderm interaction(GO:0007499) |
| 0.2 | 1.0 | GO:0006175 | adenosine salvage(GO:0006169) dATP biosynthetic process(GO:0006175) |
| 0.2 | 1.8 | GO:0015705 | iodide transport(GO:0015705) |
| 0.2 | 0.9 | GO:0042335 | cuticle development(GO:0042335) |
| 0.2 | 1.8 | GO:0021798 | forebrain dorsal/ventral pattern formation(GO:0021798) |
| 0.2 | 0.6 | GO:0002194 | hepatocyte cell migration(GO:0002194) otic placode formation(GO:0043049) branching involved in pancreas morphogenesis(GO:0061114) acinar cell differentiation(GO:0090425) positive regulation of forebrain neuron differentiation(GO:2000979) |
| 0.2 | 1.2 | GO:0006543 | glutamine catabolic process(GO:0006543) |
| 0.2 | 1.4 | GO:0033591 | response to L-ascorbic acid(GO:0033591) |
| 0.2 | 1.5 | GO:0061737 | leukotriene signaling pathway(GO:0061737) |
| 0.2 | 2.3 | GO:0030578 | PML body organization(GO:0030578) |
| 0.2 | 0.7 | GO:1902164 | mRNA localization resulting in posttranscriptional regulation of gene expression(GO:0010609) platelet alpha granule organization(GO:0070889) regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902162) positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:1902164) |
| 0.2 | 1.0 | GO:0008204 | ergosterol biosynthetic process(GO:0006696) ergosterol metabolic process(GO:0008204) |
| 0.2 | 1.4 | GO:0070236 | negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.1 | 1.0 | GO:0090170 | regulation of Golgi inheritance(GO:0090170) |
| 0.1 | 0.4 | GO:0016094 | polyprenol biosynthetic process(GO:0016094) |
| 0.1 | 1.4 | GO:1903243 | negative regulation of cardiac muscle adaptation(GO:0010616) skin morphogenesis(GO:0043589) negative regulation of cardiac muscle hypertrophy in response to stress(GO:1903243) |
| 0.1 | 0.4 | GO:0046967 | cytosol to ER transport(GO:0046967) |
| 0.1 | 2.4 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.1 | 1.2 | GO:0071963 | establishment or maintenance of cell polarity regulating cell shape(GO:0071963) |
| 0.1 | 11.3 | GO:0043268 | positive regulation of potassium ion transport(GO:0043268) |
| 0.1 | 2.5 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 0.3 | GO:0072365 | regulation of cellular ketone metabolic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072365) |
| 0.1 | 0.5 | GO:0070945 | neutrophil mediated killing of gram-negative bacterium(GO:0070945) |
| 0.1 | 0.2 | GO:0032824 | natural killer cell differentiation involved in immune response(GO:0002325) negative regulation of natural killer cell differentiation(GO:0032824) regulation of natural killer cell differentiation involved in immune response(GO:0032826) negative regulation of natural killer cell differentiation involved in immune response(GO:0032827) |
| 0.1 | 0.5 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.1 | 0.6 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.1 | 1.2 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.1 | 0.2 | GO:0002876 | positive regulation of chronic inflammatory response to antigenic stimulus(GO:0002876) |
| 0.1 | 0.3 | GO:1904744 | positive regulation of telomeric DNA binding(GO:1904744) |
| 0.1 | 0.3 | GO:0015742 | alpha-ketoglutarate transport(GO:0015742) |
| 0.1 | 0.2 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.1 | 0.9 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.1 | 0.4 | GO:0002329 | pre-B cell differentiation(GO:0002329) |
| 0.0 | 0.2 | GO:0038170 | somatostatin receptor signaling pathway(GO:0038169) somatostatin signaling pathway(GO:0038170) |
| 0.0 | 0.2 | GO:2000342 | negative regulation of interleukin-8 biosynthetic process(GO:0045415) negative regulation of chemokine (C-X-C motif) ligand 2 production(GO:2000342) |
| 0.0 | 0.6 | GO:0030091 | protein repair(GO:0030091) |
| 0.0 | 1.5 | GO:0002076 | osteoblast development(GO:0002076) |
| 0.0 | 1.3 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.0 | 2.7 | GO:0046949 | fatty-acyl-CoA biosynthetic process(GO:0046949) |
| 0.0 | 0.4 | GO:0090267 | positive regulation of mitotic cell cycle spindle assembly checkpoint(GO:0090267) |
| 0.0 | 0.1 | GO:0044205 | 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.0 | 0.2 | GO:0070127 | tRNA aminoacylation for mitochondrial protein translation(GO:0070127) |
| 0.0 | 1.1 | GO:0006833 | water transport(GO:0006833) |
| 0.0 | 0.1 | GO:0070100 | negative regulation of chemokine-mediated signaling pathway(GO:0070100) |
| 0.0 | 0.2 | GO:0021699 | cerebellum maturation(GO:0021590) cerebellar cortex maturation(GO:0021699) |
| 0.0 | 0.2 | GO:0003172 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.0 | 0.4 | GO:0030916 | otic vesicle formation(GO:0030916) |
| 0.0 | 0.4 | GO:0060992 | response to fungicide(GO:0060992) |
| 0.0 | 0.3 | GO:0030210 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.0 | 1.3 | GO:0014046 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.0 | 0.1 | GO:0035880 | embryonic nail plate morphogenesis(GO:0035880) |
| 0.0 | 0.2 | GO:0006561 | proline biosynthetic process(GO:0006561) |
| 0.0 | 1.2 | GO:0033198 | response to ATP(GO:0033198) |
| 0.0 | 1.6 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 0.3 | GO:0015866 | ADP transport(GO:0015866) |
| 0.0 | 0.6 | GO:2000479 | regulation of cAMP-dependent protein kinase activity(GO:2000479) |
| 0.0 | 0.5 | GO:1903206 | negative regulation of hydrogen peroxide-induced cell death(GO:1903206) |
| 0.0 | 0.2 | GO:1904321 | response to forskolin(GO:1904321) cellular response to forskolin(GO:1904322) |
| 0.0 | 1.6 | GO:0008631 | intrinsic apoptotic signaling pathway in response to oxidative stress(GO:0008631) |
| 0.0 | 0.5 | GO:0070498 | interleukin-1-mediated signaling pathway(GO:0070498) |
| 0.0 | 0.1 | GO:1901314 | negative regulation of histone ubiquitination(GO:0033183) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) |
| 0.0 | 0.1 | GO:0001757 | somite specification(GO:0001757) |
| 0.0 | 0.2 | GO:2000304 | positive regulation of sphingolipid biosynthetic process(GO:0090154) positive regulation of ceramide biosynthetic process(GO:2000304) |
| 0.0 | 0.3 | GO:0031284 | positive regulation of guanylate cyclase activity(GO:0031284) |
| 0.0 | 0.1 | GO:0071442 | positive regulation of histone H3-K14 acetylation(GO:0071442) |
| 0.0 | 0.7 | GO:2001240 | negative regulation of signal transduction in absence of ligand(GO:1901099) negative regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001240) |
| 0.0 | 0.4 | GO:2000678 | negative regulation of transcription regulatory region DNA binding(GO:2000678) |
| 0.0 | 0.6 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 1.3 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.0 | 0.3 | GO:0010818 | T cell chemotaxis(GO:0010818) |
| 0.0 | 0.2 | GO:1902358 | sulfate transmembrane transport(GO:1902358) |
| 0.0 | 0.1 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.0 | 0.5 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 0.3 | GO:0016048 | detection of temperature stimulus(GO:0016048) |
| 0.0 | 0.4 | GO:0007413 | axonal fasciculation(GO:0007413) |
| 0.0 | 0.2 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.0 | 1.8 | GO:0015992 | proton transport(GO:0015992) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 2.5 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.1 | 2.9 | GO:0005921 | gap junction(GO:0005921) |
| 0.1 | 0.4 | GO:0042825 | TAP complex(GO:0042825) |
| 0.1 | 1.5 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 0.5 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.0 | 3.8 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
| 0.0 | 1.3 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 2.9 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 0.1 | GO:0097679 | other organism cytoplasm(GO:0097679) |
| 0.0 | 0.8 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.0 | 0.2 | GO:0000798 | nuclear cohesin complex(GO:0000798) |
| 0.0 | 0.4 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.0 | 1.2 | GO:0000791 | euchromatin(GO:0000791) |
| 0.0 | 3.6 | GO:0045335 | phagocytic vesicle(GO:0045335) |
| 0.0 | 0.3 | GO:0070187 | telosome(GO:0070187) |
| 0.0 | 0.6 | GO:0000780 | condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.0 | 13.8 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 0.1 | GO:0045257 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.0 | 0.7 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.2 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 1.1 | GO:0034707 | chloride channel complex(GO:0034707) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 2.7 | GO:0005150 | interleukin-1, Type I receptor binding(GO:0005150) |
| 0.8 | 3.8 | GO:0032810 | sterol response element binding(GO:0032810) |
| 0.7 | 2.7 | GO:0016215 | stearoyl-CoA 9-desaturase activity(GO:0004768) acyl-CoA desaturase activity(GO:0016215) |
| 0.4 | 1.8 | GO:0015111 | iodide transmembrane transporter activity(GO:0015111) |
| 0.3 | 1.9 | GO:0015333 | peptide:proton symporter activity(GO:0015333) proton-dependent peptide secondary active transmembrane transporter activity(GO:0022897) |
| 0.2 | 1.0 | GO:0004001 | adenosine kinase activity(GO:0004001) |
| 0.2 | 2.4 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.2 | 1.5 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.2 | 1.2 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.2 | 1.2 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.2 | 1.0 | GO:0004310 | farnesyl-diphosphate farnesyltransferase activity(GO:0004310) squalene synthase activity(GO:0051996) |
| 0.2 | 1.4 | GO:0043426 | MRF binding(GO:0043426) |
| 0.1 | 0.4 | GO:0002094 | polyprenyltransferase activity(GO:0002094) |
| 0.1 | 0.6 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.1 | 0.9 | GO:0038049 | transcription factor activity, ligand-activated RNA polymerase II transcription factor binding(GO:0038049) |
| 0.1 | 2.5 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.1 | 0.5 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.1 | 0.5 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.1 | 1.1 | GO:0097371 | MDM2/MDM4 family protein binding(GO:0097371) |
| 0.1 | 1.6 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.1 | 1.5 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.1 | 1.1 | GO:0015250 | water channel activity(GO:0015250) |
| 0.1 | 0.6 | GO:0042799 | histone methyltransferase activity (H4-K20 specific)(GO:0042799) |
| 0.1 | 0.6 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 0.3 | GO:0031728 | CCR3 chemokine receptor binding(GO:0031728) |
| 0.1 | 0.4 | GO:0015440 | peptide-transporting ATPase activity(GO:0015440) MHC class Ib protein binding(GO:0023029) TAP2 binding(GO:0046979) |
| 0.0 | 0.2 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.0 | 0.3 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.0 | 1.3 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.0 | 0.2 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.0 | 2.4 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 0.8 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.0 | 10.2 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 0.4 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.0 | 0.3 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.0 | 0.9 | GO:0031628 | opioid receptor binding(GO:0031628) |
| 0.0 | 1.2 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 1.3 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 1.2 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 0.2 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.0 | 4.2 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.0 | 0.1 | GO:0004087 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.0 | 0.2 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 0.0 | 0.3 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.0 | 0.5 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.1 | GO:0060002 | microfilament motor activity(GO:0000146) plus-end directed microfilament motor activity(GO:0060002) |
| 0.0 | 0.3 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ADP transmembrane transporter activity(GO:0015217) |
| 0.0 | 0.8 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.4 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.0 | 0.1 | GO:0051538 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) 3 iron, 4 sulfur cluster binding(GO:0051538) |
| 0.0 | 0.2 | GO:0046790 | virion binding(GO:0046790) |
| 0.0 | 1.6 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.6 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.4 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 1.4 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.1 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.0 | 0.5 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 0.5 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.2 | GO:0008271 | secondary active sulfate transmembrane transporter activity(GO:0008271) |
| 0.0 | 1.4 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.2 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.0 | 0.5 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.8 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 0.1 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.0 | 3.7 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 0.2 | GO:0004383 | guanylate cyclase activity(GO:0004383) adenylate cyclase binding(GO:0008179) |
| 0.0 | 0.3 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 0.2 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.3 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 0.4 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.0 | GO:0008745 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.4 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.1 | 2.9 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 5.7 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 1.5 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 3.1 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 1.3 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 1.2 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 1.4 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 1.7 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 2.4 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.3 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.0 | 1.2 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.5 | PID MET PATHWAY | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) |
| 0.0 | 1.1 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.8 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 3.8 | REACTOME RORA ACTIVATES CIRCADIAN EXPRESSION | Genes involved in RORA Activates Circadian Expression |
| 0.1 | 3.2 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 1.0 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.1 | 1.3 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 1.1 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.1 | 1.5 | REACTOME EICOSANOID LIGAND BINDING RECEPTORS | Genes involved in Eicosanoid ligand-binding receptors |
| 0.1 | 1.3 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.1 | 0.9 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.1 | 3.1 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 2.4 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 1.2 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 1.0 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 0.8 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.0 | 0.3 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.7 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.0 | 1.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 1.4 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.4 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 1.8 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.2 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.0 | 0.4 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.0 | 0.2 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.0 | 0.5 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |