Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
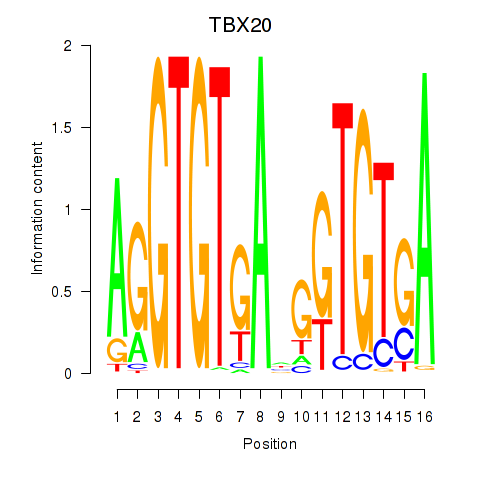
Results for TBX20
Z-value: 0.33
Transcription factors associated with TBX20
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
TBX20
|
ENSG00000164532.11 | TBX20 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| TBX20 | hg38_v1_chr7_-_35254074_35254106 | 0.11 | 5.8e-01 | Click! |
Activity profile of TBX20 motif
Sorted Z-values of TBX20 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of TBX20
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_+_57628507 | 0.79 |
ENST00000456916.5
ENST00000567154.5 ENST00000388813.9 ENST00000562558.6 ENST00000566271.6 ENST00000563374.5 ENST00000673126.2 ENST00000562631.7 ENST00000568234.5 ENST00000565770.5 ENST00000566164.5 |
ADGRG1
|
adhesion G protein-coupled receptor G1 |
| chr5_+_136059151 | 0.67 |
ENST00000503087.1
|
TGFBI
|
transforming growth factor beta induced |
| chr15_-_74212219 | 0.49 |
ENST00000449139.6
|
STRA6
|
signaling receptor and transporter of retinol STRA6 |
| chr15_-_74212256 | 0.46 |
ENST00000416286.7
|
STRA6
|
signaling receptor and transporter of retinol STRA6 |
| chr12_-_84912783 | 0.44 |
ENST00000680892.1
ENST00000266682.10 ENST00000680714.1 ENST00000552192.5 |
SLC6A15
|
solute carrier family 6 member 15 |
| chr12_-_121039204 | 0.41 |
ENST00000620239.5
|
OASL
|
2'-5'-oligoadenylate synthetase like |
| chr12_-_121039236 | 0.41 |
ENST00000257570.9
|
OASL
|
2'-5'-oligoadenylate synthetase like |
| chr6_-_32407123 | 0.40 |
ENST00000374993.4
ENST00000544175.2 ENST00000454136.7 ENST00000446536.2 |
BTNL2
|
butyrophilin like 2 |
| chr12_-_121039156 | 0.39 |
ENST00000339275.10
|
OASL
|
2'-5'-oligoadenylate synthetase like |
| chr20_+_45881218 | 0.35 |
ENST00000372523.1
|
ZSWIM1
|
zinc finger SWIM-type containing 1 |
| chr19_+_56404314 | 0.34 |
ENST00000333201.13
ENST00000391778.3 |
ZNF583
|
zinc finger protein 583 |
| chr12_-_121038967 | 0.34 |
ENST00000680620.1
ENST00000679655.1 ENST00000543677.2 |
OASL
|
2'-5'-oligoadenylate synthetase like |
| chr15_+_75724034 | 0.33 |
ENST00000332145.3
|
ODF3L1
|
outer dense fiber of sperm tails 3 like 1 |
| chr15_-_22160868 | 0.28 |
ENST00000604066.1
|
IGHV1OR15-1
|
immunoglobulin heavy variable 1/OR15-1 (non-functional) |
| chr12_+_15322257 | 0.27 |
ENST00000674316.1
|
PTPRO
|
protein tyrosine phosphatase receptor type O |
| chr1_-_11058839 | 0.26 |
ENST00000465788.1
|
SRM
|
spermidine synthase |
| chr3_+_187197486 | 0.26 |
ENST00000312295.5
|
RTP1
|
receptor transporter protein 1 |
| chrX_+_68693629 | 0.24 |
ENST00000374597.3
|
STARD8
|
StAR related lipid transfer domain containing 8 |
| chr11_+_15114912 | 0.22 |
ENST00000379556.8
|
INSC
|
INSC spindle orientation adaptor protein |
| chr3_+_40387175 | 0.22 |
ENST00000301825.8
|
ENTPD3
|
ectonucleoside triphosphate diphosphohydrolase 3 |
| chr19_-_40714641 | 0.22 |
ENST00000678467.1
|
COQ8B
|
coenzyme Q8B |
| chr12_+_15322529 | 0.21 |
ENST00000348962.7
|
PTPRO
|
protein tyrosine phosphatase receptor type O |
| chrX_-_101617921 | 0.21 |
ENST00000361910.9
ENST00000538627.5 ENST00000539247.5 |
ARMCX6
|
armadillo repeat containing X-linked 6 |
| chr15_-_21718245 | 0.20 |
ENST00000630556.1
|
ENSG00000281179.1
|
novel gene identicle to IGHV1OR15-1 |
| chr8_+_27325516 | 0.18 |
ENST00000346049.10
ENST00000420218.3 |
PTK2B
|
protein tyrosine kinase 2 beta |
| chr5_+_134525649 | 0.15 |
ENST00000282605.8
ENST00000681547.2 ENST00000361895.6 ENST00000402835.5 |
JADE2
|
jade family PHD finger 2 |
| chr16_+_57628684 | 0.14 |
ENST00000567397.5
ENST00000568979.5 ENST00000672974.1 |
ADGRG1
|
adhesion G protein-coupled receptor G1 |
| chr12_+_15322480 | 0.12 |
ENST00000674188.1
ENST00000281171.9 ENST00000543886.6 |
PTPRO
|
protein tyrosine phosphatase receptor type O |
| chrX_+_78945332 | 0.11 |
ENST00000544091.1
ENST00000171757.3 |
P2RY10
|
P2Y receptor family member 10 |
| chr19_-_39320839 | 0.11 |
ENST00000248668.5
|
LRFN1
|
leucine rich repeat and fibronectin type III domain containing 1 |
| chr3_-_15099122 | 0.10 |
ENST00000507357.1
ENST00000449050.5 ENST00000253699.7 ENST00000476527.6 |
RBSN
|
rabenosyn, RAB effector |
| chr9_+_107306459 | 0.09 |
ENST00000457811.1
|
RAD23B
|
RAD23 homolog B, nucleotide excision repair protein |
| chr11_-_64935882 | 0.09 |
ENST00000532246.1
ENST00000279168.7 |
GPHA2
|
glycoprotein hormone subunit alpha 2 |
| chrX_+_52369014 | 0.08 |
ENST00000286049.3
|
XAGE2
|
X antigen family member 2 |
| chr7_+_142424942 | 0.07 |
ENST00000426318.2
|
TRBV10-2
|
T cell receptor beta variable 10-2 |
| chr16_+_31033513 | 0.06 |
ENST00000313843.8
|
STX4
|
syntaxin 4 |
| chr7_-_33100886 | 0.06 |
ENST00000448915.1
|
RP9
|
RP9 pre-mRNA splicing factor |
| chr13_+_113105782 | 0.06 |
ENST00000541084.5
ENST00000346342.8 ENST00000375581.3 |
F7
|
coagulation factor VII |
| chr3_-_139678011 | 0.06 |
ENST00000646611.1
ENST00000645290.1 ENST00000647257.1 |
NMNAT3
|
nicotinamide nucleotide adenylyltransferase 3 |
| chrX_+_55089052 | 0.06 |
ENST00000374965.5
ENST00000449097.1 |
PAGE2
|
PAGE family member 2 |
| chr6_-_2841853 | 0.05 |
ENST00000380739.6
|
SERPINB1
|
serpin family B member 1 |
| chr11_+_6259806 | 0.05 |
ENST00000532715.5
ENST00000334619.7 ENST00000525014.1 ENST00000531712.5 ENST00000525462.1 |
CCKBR
|
cholecystokinin B receptor |
| chr2_+_176107272 | 0.04 |
ENST00000249504.7
|
HOXD11
|
homeobox D11 |
| chr3_-_131502946 | 0.04 |
ENST00000512877.1
ENST00000264995.8 ENST00000511168.5 ENST00000425847.6 |
MRPL3
|
mitochondrial ribosomal protein L3 |
| chr3_-_50359480 | 0.03 |
ENST00000266025.4
|
TMEM115
|
transmembrane protein 115 |
| chr15_-_84716153 | 0.03 |
ENST00000455959.7
|
SEC11A
|
SEC11 homolog A, signal peptidase complex subunit |
| chr15_-_84716384 | 0.03 |
ENST00000559729.5
|
SEC11A
|
SEC11 homolog A, signal peptidase complex subunit |
| chr1_-_158330957 | 0.03 |
ENST00000451207.5
|
CD1B
|
CD1b molecule |
| chr4_-_26490453 | 0.03 |
ENST00000295589.4
|
CCKAR
|
cholecystokinin A receptor |
| chr5_+_162067500 | 0.03 |
ENST00000639384.1
ENST00000640985.1 ENST00000638772.1 |
GABRG2
|
gamma-aminobutyric acid type A receptor subunit gamma2 |
| chr1_-_17054015 | 0.03 |
ENST00000375499.8
|
SDHB
|
succinate dehydrogenase complex iron sulfur subunit B |
| chr2_+_153871909 | 0.02 |
ENST00000392825.8
ENST00000434213.1 |
GALNT13
|
polypeptide N-acetylgalactosaminyltransferase 13 |
| chr2_+_60881515 | 0.02 |
ENST00000295025.12
|
REL
|
REL proto-oncogene, NF-kB subunit |
| chr15_-_84716099 | 0.02 |
ENST00000560266.5
|
SEC11A
|
SEC11 homolog A, signal peptidase complex subunit |
| chr11_-_14337074 | 0.02 |
ENST00000531421.5
|
RRAS2
|
RAS related 2 |
| chr2_+_60881553 | 0.02 |
ENST00000394479.4
|
REL
|
REL proto-oncogene, NF-kB subunit |
| chr15_-_84716063 | 0.02 |
ENST00000558217.5
ENST00000558196.1 ENST00000558134.5 |
SEC11A
|
SEC11 homolog A, signal peptidase complex subunit |
| chr19_+_7680822 | 0.01 |
ENST00000596148.3
ENST00000317378.5 ENST00000426877.2 |
TRAPPC5
|
trafficking protein particle complex 5 |
| chr6_-_106629472 | 0.01 |
ENST00000369063.8
|
RTN4IP1
|
reticulon 4 interacting protein 1 |
| chr6_-_136466858 | 0.01 |
ENST00000544465.5
|
MAP7
|
microtubule associated protein 7 |
| chr1_+_22637580 | 0.01 |
ENST00000402322.1
|
C1QA
|
complement C1q A chain |
| chr5_+_162067458 | 0.01 |
ENST00000639975.1
ENST00000639111.2 ENST00000639683.1 |
GABRG2
|
gamma-aminobutyric acid type A receptor subunit gamma2 |
| chr17_-_3696133 | 0.00 |
ENST00000225328.10
|
P2RX5
|
purinergic receptor P2X 5 |
| chr22_+_41560973 | 0.00 |
ENST00000306149.12
|
CSDC2
|
cold shock domain containing C2 |
| chr19_+_45668191 | 0.00 |
ENST00000652180.1
ENST00000590918.6 ENST00000263281.7 ENST00000304207.12 |
GIPR
|
gastric inhibitory polypeptide receptor |
| chr2_+_63840982 | 0.00 |
ENST00000480679.5
ENST00000613823.2 ENST00000677841.1 |
UGP2
|
UDP-glucose pyrophosphorylase 2 |
| chr2_+_85133376 | 0.00 |
ENST00000282111.4
|
TCF7L1
|
transcription factor 7 like 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | GO:0061143 | alveolar primary septum development(GO:0061143) |
| 0.1 | 0.6 | GO:0036058 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.1 | 0.2 | GO:0010752 | regulation of cGMP-mediated signaling(GO:0010752) regulation of B cell chemotaxis(GO:2000537) positive regulation of B cell chemotaxis(GO:2000538) |
| 0.1 | 0.9 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.0 | 0.4 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.0 | 0.3 | GO:0008295 | spermidine biosynthetic process(GO:0008295) |
| 0.0 | 0.2 | GO:0009134 | nucleoside diphosphate catabolic process(GO:0009134) |
| 0.0 | 0.1 | GO:0033594 | response to hydroxyisoflavone(GO:0033594) |
| 0.0 | 1.5 | GO:0045071 | negative regulation of viral genome replication(GO:0045071) |
| 0.0 | 0.1 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 0.0 | 0.1 | GO:0071942 | XPC complex(GO:0071942) |
| 0.0 | 0.2 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.1 | GO:0005787 | signal peptidase complex(GO:0005787) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.5 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.1 | 0.3 | GO:0004766 | spermidine synthase activity(GO:0004766) |
| 0.1 | 0.4 | GO:0005298 | proline:sodium symporter activity(GO:0005298) |
| 0.0 | 0.3 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.0 | 0.1 | GO:0031531 | thyrotropin-releasing hormone receptor binding(GO:0031531) |
| 0.0 | 0.9 | GO:0051183 | vitamin transporter activity(GO:0051183) |
| 0.0 | 0.6 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 0.2 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.0 | 1.6 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.2 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 1.5 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.4 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 0.3 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |