Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
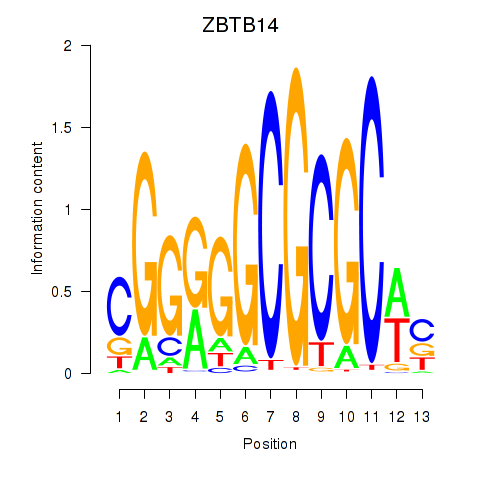
Results for ZBTB14
Z-value: 1.74
Transcription factors associated with ZBTB14
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ZBTB14
|
ENSG00000198081.11 | ZBTB14 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ZBTB14 | hg38_v1_chr18_-_5295679_5295737, hg38_v1_chr18_-_5296002_5296046 | 0.44 | 1.5e-02 | Click! |
Activity profile of ZBTB14 motif
Sorted Z-values of ZBTB14 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ZBTB14
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_-_48409292 | 10.24 |
ENST00000589194.5
ENST00000591279.5 ENST00000590855.5 ENST00000587107.5 ENST00000588970.5 ENST00000586525.5 ENST00000592387.5 ENST00000590800.6 |
ZBTB7C
|
zinc finger and BTB domain containing 7C |
| chr1_+_111346590 | 8.87 |
ENST00000369737.4
ENST00000369738.9 |
PIFO
|
primary cilia formation |
| chr2_-_27890348 | 8.54 |
ENST00000302188.8
|
RBKS
|
ribokinase |
| chr12_-_112013123 | 7.33 |
ENST00000550831.7
ENST00000549537.6 ENST00000355445.7 ENST00000552374.7 |
TMEM116
|
transmembrane protein 116 |
| chr9_+_68705414 | 7.32 |
ENST00000541509.5
|
PIP5K1B
|
phosphatidylinositol-4-phosphate 5-kinase type 1 beta |
| chr4_-_7042931 | 7.07 |
ENST00000310085.6
|
CCDC96
|
coiled-coil domain containing 96 |
| chr2_-_99141517 | 6.58 |
ENST00000355053.8
|
TSGA10
|
testis specific 10 |
| chr20_-_63831214 | 6.29 |
ENST00000302995.2
ENST00000245663.9 |
ZBTB46
|
zinc finger and BTB domain containing 46 |
| chr1_-_39901996 | 5.58 |
ENST00000397332.2
|
MYCL
|
MYCL proto-oncogene, bHLH transcription factor |
| chr22_-_31345770 | 5.54 |
ENST00000215919.3
|
PATZ1
|
POZ/BTB and AT hook containing zinc finger 1 |
| chr2_-_99141169 | 5.46 |
ENST00000674128.1
|
TSGA10
|
testis specific 10 |
| chr9_-_134068518 | 5.31 |
ENST00000371834.6
|
BRD3
|
bromodomain containing 3 |
| chr9_+_68705230 | 5.04 |
ENST00000265382.8
|
PIP5K1B
|
phosphatidylinositol-4-phosphate 5-kinase type 1 beta |
| chr6_+_18155399 | 4.98 |
ENST00000650836.2
ENST00000449850.2 ENST00000297792.9 |
KDM1B
|
lysine demethylase 1B |
| chr2_-_99141414 | 4.82 |
ENST00000393482.7
|
TSGA10
|
testis specific 10 |
| chr22_-_31346143 | 4.69 |
ENST00000405309.7
ENST00000351933.8 |
PATZ1
|
POZ/BTB and AT hook containing zinc finger 1 |
| chrX_-_8732116 | 4.66 |
ENST00000262648.8
|
ANOS1
|
anosmin 1 |
| chr1_-_223364059 | 4.45 |
ENST00000343846.7
ENST00000484758.6 ENST00000344029.6 ENST00000366878.9 ENST00000494793.6 ENST00000681285.1 ENST00000680429.1 ENST00000681669.1 ENST00000681305.1 |
SUSD4
|
sushi domain containing 4 |
| chr17_+_57256514 | 4.22 |
ENST00000284073.7
ENST00000674964.1 |
MSI2
|
musashi RNA binding protein 2 |
| chr11_+_72080803 | 3.99 |
ENST00000423494.6
ENST00000539587.6 ENST00000536917.2 ENST00000538478.5 ENST00000324866.11 ENST00000643715.1 ENST00000439209.5 |
LRTOMT
ENSG00000284922.2
|
leucine rich transmembrane and O-methyltransferase domain containing leucine rich transmembrane and O-methyltransferase domain containing |
| chr2_+_219229783 | 3.86 |
ENST00000453432.5
ENST00000409849.5 ENST00000323348.10 ENST00000416565.1 ENST00000410034.7 ENST00000447157.5 |
ANKZF1
|
ankyrin repeat and zinc finger peptidyl tRNA hydrolase 1 |
| chr5_+_77210667 | 3.82 |
ENST00000264917.10
|
PDE8B
|
phosphodiesterase 8B |
| chr19_+_708903 | 3.76 |
ENST00000338448.10
ENST00000264560.11 |
PALM
|
paralemmin |
| chr20_-_40689228 | 3.73 |
ENST00000373313.3
|
MAFB
|
MAF bZIP transcription factor B |
| chr6_+_135181268 | 3.72 |
ENST00000341911.10
ENST00000442647.7 ENST00000618728.4 ENST00000316528.12 ENST00000616088.4 |
MYB
|
MYB proto-oncogene, transcription factor |
| chr5_+_77210881 | 3.71 |
ENST00000340978.7
ENST00000346042.7 ENST00000342343.8 ENST00000333194.8 |
PDE8B
|
phosphodiesterase 8B |
| chr20_+_9069076 | 3.70 |
ENST00000378473.9
|
PLCB4
|
phospholipase C beta 4 |
| chr9_-_4300049 | 3.69 |
ENST00000381971.8
|
GLIS3
|
GLIS family zinc finger 3 |
| chr2_-_86337617 | 3.67 |
ENST00000538924.7
ENST00000535845.6 |
REEP1
|
receptor accessory protein 1 |
| chr6_+_135181323 | 3.65 |
ENST00000367814.8
|
MYB
|
MYB proto-oncogene, transcription factor |
| chr2_-_86337654 | 3.65 |
ENST00000165698.9
|
REEP1
|
receptor accessory protein 1 |
| chr19_-_6110463 | 3.63 |
ENST00000587181.1
ENST00000587321.1 ENST00000586806.1 ENST00000303657.10 ENST00000589742.5 ENST00000592546.5 |
RFX2
|
regulatory factor X2 |
| chr11_-_6419394 | 3.61 |
ENST00000311051.7
|
APBB1
|
amyloid beta precursor protein binding family B member 1 |
| chr8_+_98064522 | 3.57 |
ENST00000545282.1
|
ERICH5
|
glutamate rich 5 |
| chr13_-_36131286 | 3.52 |
ENST00000255448.8
ENST00000379892.4 |
DCLK1
|
doublecortin like kinase 1 |
| chr3_+_14402576 | 3.52 |
ENST00000613060.4
|
SLC6A6
|
solute carrier family 6 member 6 |
| chr1_-_39901861 | 3.52 |
ENST00000372816.3
ENST00000372815.1 |
MYCL
|
MYCL proto-oncogene, bHLH transcription factor |
| chr5_+_76403266 | 3.47 |
ENST00000274364.11
|
IQGAP2
|
IQ motif containing GTPase activating protein 2 |
| chr14_+_73569266 | 3.46 |
ENST00000613168.1
|
ACOT2
|
acyl-CoA thioesterase 2 |
| chr14_+_73569115 | 3.42 |
ENST00000622407.4
ENST00000238651.10 |
ACOT2
|
acyl-CoA thioesterase 2 |
| chr11_-_6419051 | 3.38 |
ENST00000299402.10
ENST00000532020.2 ENST00000609360.6 ENST00000389906.6 |
APBB1
|
amyloid beta precursor protein binding family B member 1 |
| chr6_+_31897775 | 3.38 |
ENST00000469372.5
ENST00000497706.5 |
C2
|
complement C2 |
| chr2_-_206765274 | 3.35 |
ENST00000454776.6
ENST00000449792.5 ENST00000374412.8 |
MDH1B
|
malate dehydrogenase 1B |
| chr15_-_51622738 | 3.34 |
ENST00000560891.6
|
DMXL2
|
Dmx like 2 |
| chr11_+_45885625 | 3.31 |
ENST00000241014.6
|
MAPK8IP1
|
mitogen-activated protein kinase 8 interacting protein 1 |
| chr7_+_121873089 | 3.30 |
ENST00000651065.1
|
PTPRZ1
|
protein tyrosine phosphatase receptor type Z1 |
| chr22_-_31346317 | 3.16 |
ENST00000266269.10
|
PATZ1
|
POZ/BTB and AT hook containing zinc finger 1 |
| chr15_-_51622613 | 3.12 |
ENST00000543779.6
ENST00000449909.7 |
DMXL2
|
Dmx like 2 |
| chr15_-_51622798 | 3.09 |
ENST00000251076.9
|
DMXL2
|
Dmx like 2 |
| chr21_+_41316747 | 3.09 |
ENST00000357985.7
ENST00000398647.7 ENST00000398652.7 |
FAM3B
|
FAM3 metabolism regulating signaling molecule B |
| chr4_-_148442342 | 3.04 |
ENST00000358102.8
|
NR3C2
|
nuclear receptor subfamily 3 group C member 2 |
| chr17_+_38705482 | 3.02 |
ENST00000620609.4
|
MLLT6
|
MLLT6, PHD finger containing |
| chr4_-_148442508 | 3.01 |
ENST00000625323.2
|
NR3C2
|
nuclear receptor subfamily 3 group C member 2 |
| chr3_+_72888031 | 2.99 |
ENST00000389617.9
|
GXYLT2
|
glucoside xylosyltransferase 2 |
| chr12_+_50057548 | 2.96 |
ENST00000228468.8
ENST00000447966.7 |
ASIC1
|
acid sensing ion channel subunit 1 |
| chr8_+_98064559 | 2.95 |
ENST00000318528.8
|
ERICH5
|
glutamate rich 5 |
| chr10_+_68827521 | 2.95 |
ENST00000298596.11
ENST00000399169.8 ENST00000399162.2 ENST00000399165.8 ENST00000642869.1 |
STOX1
|
storkhead box 1 |
| chr9_+_36136703 | 2.85 |
ENST00000377960.9
ENST00000377959.5 |
GLIPR2
|
GLI pathogenesis related 2 |
| chr10_+_93993897 | 2.84 |
ENST00000371380.8
|
PLCE1
|
phospholipase C epsilon 1 |
| chr3_+_14402592 | 2.79 |
ENST00000622186.5
ENST00000621751.4 |
SLC6A6
|
solute carrier family 6 member 6 |
| chr22_+_39349925 | 2.77 |
ENST00000318801.8
ENST00000216155.11 ENST00000406293.7 ENST00000328933.10 |
SYNGR1
|
synaptogyrin 1 |
| chr14_+_73537135 | 2.72 |
ENST00000311148.9
|
ACOT1
|
acyl-CoA thioesterase 1 |
| chr11_+_72080595 | 2.70 |
ENST00000647530.1
ENST00000539271.6 ENST00000642510.1 |
LRTOMT
|
leucine rich transmembrane and O-methyltransferase domain containing |
| chr9_+_36136752 | 2.70 |
ENST00000619700.1
|
GLIPR2
|
GLI pathogenesis related 2 |
| chr3_+_49412203 | 2.68 |
ENST00000273590.3
|
TCTA
|
T cell leukemia translocation altered |
| chr3_+_113948004 | 2.66 |
ENST00000638807.2
|
ZDHHC23
|
zinc finger DHHC-type palmitoyltransferase 23 |
| chr3_+_113947901 | 2.65 |
ENST00000330212.7
ENST00000498275.5 |
ZDHHC23
|
zinc finger DHHC-type palmitoyltransferase 23 |
| chr9_+_36136416 | 2.63 |
ENST00000396613.7
|
GLIPR2
|
GLI pathogenesis related 2 |
| chr22_-_50326935 | 2.63 |
ENST00000413817.8
|
DENND6B
|
DENN domain containing 6B |
| chr19_-_821929 | 2.62 |
ENST00000359894.6
ENST00000520876.8 ENST00000519502.2 |
PLPPR3
|
phospholipid phosphatase related 3 |
| chr17_+_74737211 | 2.61 |
ENST00000392612.7
ENST00000392610.5 |
RAB37
|
RAB37, member RAS oncogene family |
| chr14_+_73537346 | 2.60 |
ENST00000557556.1
|
ACOT1
|
acyl-CoA thioesterase 1 |
| chr8_+_17497108 | 2.56 |
ENST00000470360.5
|
SLC7A2
|
solute carrier family 7 member 2 |
| chr17_+_74737238 | 2.55 |
ENST00000392613.10
ENST00000613645.1 |
RAB37
|
RAB37, member RAS oncogene family |
| chr11_+_72080313 | 2.53 |
ENST00000307198.11
ENST00000538413.6 ENST00000642648.1 ENST00000289488.7 |
ENSG00000284922.2
LRTOMT
|
leucine rich transmembrane and O-methyltransferase domain containing leucine rich transmembrane and O-methyltransferase domain containing |
| chr19_+_40799796 | 2.48 |
ENST00000596517.1
|
EGLN2
|
egl-9 family hypoxia inducible factor 2 |
| chr14_-_64972143 | 2.46 |
ENST00000267512.9
|
RAB15
|
RAB15, member RAS oncogene family |
| chr13_+_23579346 | 2.45 |
ENST00000382258.8
ENST00000382263.3 |
TNFRSF19
|
TNF receptor superfamily member 19 |
| chr22_-_38794111 | 2.42 |
ENST00000406622.5
ENST00000216068.9 ENST00000406199.3 |
SUN2
DNAL4
|
Sad1 and UNC84 domain containing 2 dynein axonemal light chain 4 |
| chr8_+_74984496 | 2.41 |
ENST00000262207.9
|
CRISPLD1
|
cysteine rich secretory protein LCCL domain containing 1 |
| chr12_+_19129689 | 2.41 |
ENST00000429027.7
ENST00000540972.5 |
PLEKHA5
|
pleckstrin homology domain containing A5 |
| chr7_+_121873152 | 2.40 |
ENST00000650826.1
ENST00000650728.1 ENST00000393386.7 ENST00000651390.1 ENST00000651842.1 ENST00000650681.1 |
PTPRZ1
|
protein tyrosine phosphatase receptor type Z1 |
| chr11_-_118152775 | 2.39 |
ENST00000324727.9
|
SCN4B
|
sodium voltage-gated channel beta subunit 4 |
| chr16_-_1611985 | 2.38 |
ENST00000426508.7
|
IFT140
|
intraflagellar transport 140 |
| chr20_-_25058115 | 2.37 |
ENST00000323482.9
|
ACSS1
|
acyl-CoA synthetase short chain family member 1 |
| chr20_-_25058168 | 2.37 |
ENST00000432802.6
|
ACSS1
|
acyl-CoA synthetase short chain family member 1 |
| chr6_-_89412219 | 2.34 |
ENST00000369415.9
|
RRAGD
|
Ras related GTP binding D |
| chr1_+_210328894 | 2.32 |
ENST00000261458.8
ENST00000537898.5 ENST00000545154.5 |
HHAT
|
hedgehog acyltransferase |
| chr6_-_89412069 | 2.30 |
ENST00000359203.3
|
RRAGD
|
Ras related GTP binding D |
| chrY_+_12904860 | 2.21 |
ENST00000336079.8
|
DDX3Y
|
DEAD-box helicase 3 Y-linked |
| chr1_-_217137748 | 2.18 |
ENST00000493603.5
ENST00000366940.6 |
ESRRG
|
estrogen related receptor gamma |
| chr7_+_116953238 | 2.18 |
ENST00000393446.6
|
ST7
|
suppression of tumorigenicity 7 |
| chr8_+_17497078 | 2.17 |
ENST00000494857.6
ENST00000522656.5 |
SLC7A2
|
solute carrier family 7 member 2 |
| chr10_+_12349685 | 2.15 |
ENST00000378845.5
|
CAMK1D
|
calcium/calmodulin dependent protein kinase ID |
| chr12_+_123973123 | 2.15 |
ENST00000539644.5
|
ZNF664
|
zinc finger protein 664 |
| chr12_-_22544409 | 2.13 |
ENST00000536386.5
ENST00000396028.6 ENST00000545552.5 ENST00000446597.6 ENST00000333957.8 |
C2CD5
|
C2 calcium dependent domain containing 5 |
| chr3_-_9952337 | 2.11 |
ENST00000411976.2
ENST00000412055.6 |
PRRT3
|
proline rich transmembrane protein 3 |
| chr5_-_102296260 | 2.07 |
ENST00000310954.7
|
SLCO4C1
|
solute carrier organic anion transporter family member 4C1 |
| chr14_-_89619118 | 2.06 |
ENST00000345097.8
ENST00000555855.5 ENST00000555353.5 |
FOXN3
|
forkhead box N3 |
| chr6_-_39229465 | 2.05 |
ENST00000359534.4
|
KCNK5
|
potassium two pore domain channel subfamily K member 5 |
| chr19_+_14433284 | 2.03 |
ENST00000242783.11
|
PKN1
|
protein kinase N1 |
| chr11_-_119364166 | 2.03 |
ENST00000525735.1
|
USP2
|
ubiquitin specific peptidase 2 |
| chr9_-_89498038 | 2.03 |
ENST00000339861.8
ENST00000422704.7 ENST00000455551.6 |
SEMA4D
|
semaphorin 4D |
| chr6_-_118935146 | 2.03 |
ENST00000619706.5
ENST00000316316.10 |
MCM9
|
minichromosome maintenance 9 homologous recombination repair factor |
| chr19_+_40799707 | 2.02 |
ENST00000594380.1
ENST00000593397.1 ENST00000601733.1 ENST00000593525.1 |
EGLN2
|
egl-9 family hypoxia inducible factor 2 |
| chr15_+_74995520 | 1.99 |
ENST00000562327.5
ENST00000568018.5 ENST00000425597.8 ENST00000562212.5 ENST00000567920.5 ENST00000566872.5 ENST00000361900.10 |
SCAMP5
|
secretory carrier membrane protein 5 |
| chr7_-_120857124 | 1.99 |
ENST00000441017.5
ENST00000424710.5 ENST00000433758.5 |
TSPAN12
|
tetraspanin 12 |
| chr2_+_11155372 | 1.98 |
ENST00000441908.6
ENST00000295083.8 |
SLC66A3
|
solute carrier family 66 member 3 |
| chr1_+_59814939 | 1.98 |
ENST00000371208.5
|
HOOK1
|
hook microtubule tethering protein 1 |
| chr2_-_228181612 | 1.96 |
ENST00000344657.5
|
SPHKAP
|
SPHK1 interactor, AKAP domain containing |
| chr4_+_185396834 | 1.95 |
ENST00000335174.6
|
ANKRD37
|
ankyrin repeat domain 37 |
| chr7_+_121873317 | 1.95 |
ENST00000651863.1
ENST00000652298.1 ENST00000449182.1 |
PTPRZ1
|
protein tyrosine phosphatase receptor type Z1 |
| chr20_+_36461460 | 1.94 |
ENST00000482872.5
ENST00000495241.5 |
DLGAP4
|
DLG associated protein 4 |
| chr17_+_55266216 | 1.93 |
ENST00000573945.5
|
HLF
|
HLF transcription factor, PAR bZIP family member |
| chr2_-_228181669 | 1.92 |
ENST00000392056.8
|
SPHKAP
|
SPHK1 interactor, AKAP domain containing |
| chr9_+_17579059 | 1.92 |
ENST00000380607.5
|
SH3GL2
|
SH3 domain containing GRB2 like 2, endophilin A1 |
| chr19_+_40799501 | 1.89 |
ENST00000406058.6
ENST00000593726.5 |
EGLN2
|
egl-9 family hypoxia inducible factor 2 |
| chr2_+_24491239 | 1.89 |
ENST00000348332.8
|
NCOA1
|
nuclear receptor coactivator 1 |
| chr9_+_137241277 | 1.88 |
ENST00000340384.5
|
TUBB4B
|
tubulin beta 4B class IVb |
| chr19_-_461007 | 1.88 |
ENST00000264554.11
|
SHC2
|
SHC adaptor protein 2 |
| chr14_+_99793329 | 1.88 |
ENST00000334192.8
|
EML1
|
EMAP like 1 |
| chr3_-_36993103 | 1.88 |
ENST00000322716.8
|
EPM2AIP1
|
EPM2A interacting protein 1 |
| chr19_+_32405758 | 1.88 |
ENST00000392250.7
|
DPY19L3
|
dpy-19 like C-mannosyltransferase 3 |
| chr6_+_84033717 | 1.87 |
ENST00000257776.5
|
MRAP2
|
melanocortin 2 receptor accessory protein 2 |
| chr9_-_134068012 | 1.86 |
ENST00000303407.12
|
BRD3
|
bromodomain containing 3 |
| chr19_+_40799425 | 1.85 |
ENST00000593972.1
|
EGLN2
|
egl-9 family hypoxia inducible factor 2 |
| chr12_-_124567464 | 1.85 |
ENST00000458234.5
|
NCOR2
|
nuclear receptor corepressor 2 |
| chr3_-_133895577 | 1.80 |
ENST00000543906.5
|
RAB6B
|
RAB6B, member RAS oncogene family |
| chr10_+_12349533 | 1.80 |
ENST00000619168.5
|
CAMK1D
|
calcium/calmodulin dependent protein kinase ID |
| chr3_-_115147237 | 1.79 |
ENST00000357258.8
|
ZBTB20
|
zinc finger and BTB domain containing 20 |
| chr2_+_231708511 | 1.77 |
ENST00000341369.11
ENST00000409115.8 ENST00000409683.5 |
PTMA
|
prothymosin alpha |
| chr12_+_112013348 | 1.75 |
ENST00000455836.1
|
ERP29
|
endoplasmic reticulum protein 29 |
| chr17_+_12789457 | 1.75 |
ENST00000379672.10
ENST00000340825.7 |
ARHGAP44
|
Rho GTPase activating protein 44 |
| chr6_+_24494939 | 1.75 |
ENST00000348925.2
ENST00000357578.8 |
ALDH5A1
|
aldehyde dehydrogenase 5 family member A1 |
| chr19_+_32405789 | 1.74 |
ENST00000586987.5
|
DPY19L3
|
dpy-19 like C-mannosyltransferase 3 |
| chr4_+_74933108 | 1.74 |
ENST00000307428.7
|
PARM1
|
prostate androgen-regulated mucin-like protein 1 |
| chr3_-_115147277 | 1.74 |
ENST00000675478.1
|
ZBTB20
|
zinc finger and BTB domain containing 20 |
| chr19_+_32406076 | 1.73 |
ENST00000342179.9
ENST00000586427.1 |
DPY19L3
|
dpy-19 like C-mannosyltransferase 3 |
| chr18_+_3449413 | 1.73 |
ENST00000549253.5
|
TGIF1
|
TGFB induced factor homeobox 1 |
| chr7_+_17299234 | 1.73 |
ENST00000637807.1
|
ENSG00000283321.1
|
novel protein |
| chr1_+_236142526 | 1.72 |
ENST00000366592.8
|
GPR137B
|
G protein-coupled receptor 137B |
| chr1_+_174159501 | 1.72 |
ENST00000251507.8
ENST00000681986.1 |
RABGAP1L
|
RAB GTPase activating protein 1 like |
| chr4_+_41360759 | 1.71 |
ENST00000508501.5
ENST00000512946.5 ENST00000313860.11 ENST00000512632.5 ENST00000512820.5 |
LIMCH1
|
LIM and calponin homology domains 1 |
| chr4_+_74933095 | 1.69 |
ENST00000513238.5
|
PARM1
|
prostate androgen-regulated mucin-like protein 1 |
| chr2_+_229922482 | 1.69 |
ENST00000283946.8
ENST00000409992.1 |
FBXO36
|
F-box protein 36 |
| chr17_-_5500997 | 1.69 |
ENST00000568641.2
|
ENSG00000286190.2
|
novel protein |
| chr17_+_74432079 | 1.69 |
ENST00000392627.7
ENST00000392628.7 ENST00000582473.2 |
GPRC5C
|
G protein-coupled receptor class C group 5 member C |
| chr12_+_112013418 | 1.65 |
ENST00000261735.4
ENST00000552052.1 |
ERP29
|
endoplasmic reticulum protein 29 |
| chr10_-_102419934 | 1.64 |
ENST00000406432.5
|
PSD
|
pleckstrin and Sec7 domain containing |
| chr2_+_108786738 | 1.63 |
ENST00000412964.6
ENST00000295124.9 |
CCDC138
|
coiled-coil domain containing 138 |
| chr17_+_82023268 | 1.62 |
ENST00000306688.8
|
LRRC45
|
leucine rich repeat containing 45 |
| chr12_+_122021850 | 1.61 |
ENST00000261822.5
|
BCL7A
|
BAF chromatin remodeling complex subunit BCL7A |
| chr3_-_133895453 | 1.59 |
ENST00000486858.5
ENST00000477759.5 |
RAB6B
|
RAB6B, member RAS oncogene family |
| chr10_+_92848461 | 1.58 |
ENST00000443748.6
ENST00000371543.5 ENST00000260762.10 |
EXOC6
|
exocyst complex component 6 |
| chr19_-_4581755 | 1.58 |
ENST00000676793.1
|
SEMA6B
|
semaphorin 6B |
| chr6_+_125749623 | 1.58 |
ENST00000368364.4
|
HEY2
|
hes related family bHLH transcription factor with YRPW motif 2 |
| chr6_+_135181361 | 1.57 |
ENST00000527615.5
ENST00000420123.6 ENST00000525369.5 ENST00000528774.5 ENST00000533624.5 ENST00000534044.5 ENST00000534121.5 |
MYB
|
MYB proto-oncogene, transcription factor |
| chr18_+_13218769 | 1.56 |
ENST00000677055.1
ENST00000399848.7 |
LDLRAD4
|
low density lipoprotein receptor class A domain containing 4 |
| chr19_+_40799155 | 1.56 |
ENST00000303961.9
|
EGLN2
|
egl-9 family hypoxia inducible factor 2 |
| chr19_+_12995554 | 1.56 |
ENST00000397661.6
|
NFIX
|
nuclear factor I X |
| chr16_+_54930827 | 1.56 |
ENST00000394636.9
|
IRX5
|
iroquois homeobox 5 |
| chr3_+_119703001 | 1.54 |
ENST00000273390.9
ENST00000463700.1 ENST00000648112.1 |
CFAP91
ENSG00000285585.1
|
cilia and flagella associated protein 91 novel protein |
| chr4_-_2418608 | 1.54 |
ENST00000503000.1
|
ZFYVE28
|
zinc finger FYVE-type containing 28 |
| chr9_+_976648 | 1.54 |
ENST00000190165.3
|
DMRT3
|
doublesex and mab-3 related transcription factor 3 |
| chr4_+_128809791 | 1.54 |
ENST00000452328.6
ENST00000504089.5 |
JADE1
|
jade family PHD finger 1 |
| chr7_+_158856551 | 1.53 |
ENST00000407559.8
|
DYNC2I1
|
dynein 2 intermediate chain 1 |
| chr3_-_133895867 | 1.52 |
ENST00000285208.9
|
RAB6B
|
RAB6B, member RAS oncogene family |
| chr10_-_102419693 | 1.52 |
ENST00000611678.4
|
PSD
|
pleckstrin and Sec7 domain containing |
| chr1_-_45206594 | 1.51 |
ENST00000359600.6
|
ZSWIM5
|
zinc finger SWIM-type containing 5 |
| chr1_-_48776811 | 1.51 |
ENST00000371833.4
|
BEND5
|
BEN domain containing 5 |
| chr10_+_102419189 | 1.51 |
ENST00000432590.5
|
FBXL15
|
F-box and leucine rich repeat protein 15 |
| chr14_-_23301474 | 1.50 |
ENST00000561437.1
ENST00000559942.5 ENST00000560913.1 ENST00000559314.5 ENST00000558058.5 |
PPP1R3E
|
protein phosphatase 1 regulatory subunit 3E |
| chr16_+_58025745 | 1.50 |
ENST00000219271.4
|
MMP15
|
matrix metallopeptidase 15 |
| chr1_+_113390495 | 1.49 |
ENST00000307546.14
|
MAGI3
|
membrane associated guanylate kinase, WW and PDZ domain containing 3 |
| chr19_-_33064872 | 1.49 |
ENST00000254260.8
|
RHPN2
|
rhophilin Rho GTPase binding protein 2 |
| chr17_+_37489882 | 1.49 |
ENST00000617516.5
|
DUSP14
|
dual specificity phosphatase 14 |
| chr18_+_50967991 | 1.48 |
ENST00000588577.5
|
ELAC1
|
elaC ribonuclease Z 1 |
| chr2_-_60553618 | 1.47 |
ENST00000643716.1
ENST00000359629.10 ENST00000642384.2 ENST00000335712.11 |
BCL11A
|
BAF chromatin remodeling complex subunit BCL11A |
| chr4_+_128809684 | 1.47 |
ENST00000226319.11
ENST00000511647.5 |
JADE1
|
jade family PHD finger 1 |
| chr9_+_6757940 | 1.46 |
ENST00000381309.8
|
KDM4C
|
lysine demethylase 4C |
| chr8_+_28494190 | 1.46 |
ENST00000537916.2
ENST00000240093.8 ENST00000523546.1 |
FZD3
|
frizzled class receptor 3 |
| chr5_+_177446445 | 1.45 |
ENST00000507881.5
|
PRR7
|
proline rich 7, synaptic |
| chr7_+_29563820 | 1.44 |
ENST00000319694.3
|
PRR15
|
proline rich 15 |
| chr1_-_236281951 | 1.43 |
ENST00000354619.10
|
ERO1B
|
endoplasmic reticulum oxidoreductase 1 beta |
| chr13_-_95301319 | 1.43 |
ENST00000646439.1
ENST00000645532.1 |
ABCC4
|
ATP binding cassette subfamily C member 4 |
| chr5_-_38556625 | 1.43 |
ENST00000506990.5
ENST00000453190.7 |
LIFR
|
LIF receptor subunit alpha |
| chr11_-_73142308 | 1.38 |
ENST00000409418.9
|
FCHSD2
|
FCH and double SH3 domains 2 |
| chr19_-_18606779 | 1.38 |
ENST00000684169.1
ENST00000392386.8 |
CRLF1
|
cytokine receptor like factor 1 |
| chr5_-_180072086 | 1.37 |
ENST00000261947.4
|
RNF130
|
ring finger protein 130 |
| chr1_-_53945661 | 1.37 |
ENST00000194214.10
|
HSPB11
|
heat shock protein family B (small) member 11 |
| chr12_+_123973197 | 1.37 |
ENST00000392404.7
ENST00000337815.9 ENST00000538932.6 ENST00000618862.2 ENST00000389727.8 |
ZNF664
ENSG00000274874.2
RFLNA
|
zinc finger protein 664 novel protein refilin A |
| chr11_-_66568524 | 1.36 |
ENST00000679160.1
ENST00000678305.1 ENST00000310325.10 ENST00000677896.1 ENST00000677587.1 ENST00000679347.1 ENST00000677005.1 ENST00000678872.1 ENST00000679024.1 ENST00000678471.1 ENST00000524994.6 |
CTSF
|
cathepsin F |
| chr22_+_41381923 | 1.36 |
ENST00000266304.9
|
TEF
|
TEF transcription factor, PAR bZIP family member |
| chr20_+_50731571 | 1.35 |
ENST00000371610.7
|
PARD6B
|
par-6 family cell polarity regulator beta |
| chr19_+_35138778 | 1.35 |
ENST00000351325.9
ENST00000586871.5 ENST00000592174.1 |
FXYD1
ENSG00000221857.7
|
FXYD domain containing ion transport regulator 1 novel transcript |
| chr6_-_83065770 | 1.34 |
ENST00000369747.8
|
UBE3D
|
ubiquitin protein ligase E3D |
| chr6_+_167826848 | 1.34 |
ENST00000683244.1
ENST00000400825.8 |
AFDN
|
afadin, adherens junction formation factor |
| chr5_-_139904460 | 1.34 |
ENST00000340391.8
|
NRG2
|
neuregulin 2 |
| chr11_-_64844620 | 1.33 |
ENST00000342711.6
|
CDC42BPG
|
CDC42 binding protein kinase gamma |
| chr14_+_73644875 | 1.33 |
ENST00000554113.5
ENST00000553645.7 ENST00000555631.6 ENST00000311089.7 ENST00000555919.7 ENST00000554339.5 ENST00000554871.5 |
DNAL1
|
dynein axonemal light chain 1 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 8.9 | GO:1904897 | regulation of hepatic stellate cell proliferation(GO:1904897) positive regulation of hepatic stellate cell proliferation(GO:1904899) hepatic stellate cell proliferation(GO:1990922) |
| 2.8 | 8.5 | GO:0019303 | D-ribose catabolic process(GO:0019303) |
| 2.3 | 7.0 | GO:0050760 | negative regulation of thymidylate synthase biosynthetic process(GO:0050760) |
| 1.6 | 6.5 | GO:0006083 | acetate metabolic process(GO:0006083) |
| 1.6 | 4.7 | GO:1903400 | L-arginine transmembrane transport(GO:1903400) |
| 1.3 | 10.7 | GO:0045607 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 1.0 | 4.0 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.9 | 3.7 | GO:0021571 | rhombomere 5 development(GO:0021571) central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.9 | 2.6 | GO:1904017 | cellular response to Thyroglobulin triiodothyronine(GO:1904017) |
| 0.9 | 0.9 | GO:0035574 | histone H4-K20 demethylation(GO:0035574) |
| 0.8 | 9.8 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.8 | 3.9 | GO:0071630 | nucleus-associated proteasomal ubiquitin-dependent protein catabolic process(GO:0071630) |
| 0.8 | 5.4 | GO:0018103 | protein C-linked glycosylation(GO:0018103) peptidyl-tryptophan modification(GO:0018211) protein C-linked glycosylation via tryptophan(GO:0018317) protein C-linked glycosylation via 2'-alpha-mannosyl-L-tryptophan(GO:0018406) |
| 0.7 | 2.2 | GO:0034414 | tRNA 3'-trailer cleavage, endonucleolytic(GO:0034414) tRNA 3'-trailer cleavage(GO:0042779) |
| 0.7 | 2.2 | GO:1904049 | negative regulation of spontaneous neurotransmitter secretion(GO:1904049) |
| 0.6 | 9.1 | GO:1903025 | regulation of RNA polymerase II regulatory region sequence-specific DNA binding(GO:1903025) |
| 0.6 | 1.8 | GO:0072365 | regulation of cellular ketone metabolic process by negative regulation of transcription from RNA polymerase II promoter(GO:0072365) |
| 0.6 | 3.0 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 0.6 | 2.3 | GO:0061187 | regulation of chromatin silencing at rDNA(GO:0061187) negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 0.6 | 9.5 | GO:0070070 | proton-transporting V-type ATPase complex assembly(GO:0070070) vacuolar proton-transporting V-type ATPase complex assembly(GO:0070072) |
| 0.6 | 9.0 | GO:0070444 | oligodendrocyte progenitor proliferation(GO:0070444) regulation of oligodendrocyte progenitor proliferation(GO:0070445) |
| 0.6 | 1.7 | GO:0060392 | negative regulation of SMAD protein import into nucleus(GO:0060392) |
| 0.6 | 5.5 | GO:0045650 | negative regulation of macrophage differentiation(GO:0045650) |
| 0.5 | 2.2 | GO:0090156 | cellular sphingolipid homeostasis(GO:0090156) |
| 0.5 | 3.8 | GO:0060160 | negative regulation of dopamine receptor signaling pathway(GO:0060160) |
| 0.5 | 1.6 | GO:1900220 | semaphorin-plexin signaling pathway involved in bone trabecula morphogenesis(GO:1900220) |
| 0.5 | 1.6 | GO:0051228 | mitotic spindle disassembly(GO:0051228) spindle disassembly(GO:0051230) |
| 0.5 | 4.6 | GO:0071233 | cellular response to leucine(GO:0071233) cellular response to leucine starvation(GO:1990253) |
| 0.5 | 2.6 | GO:0072674 | multinuclear osteoclast differentiation(GO:0072674) osteoclast fusion(GO:0072675) |
| 0.5 | 2.0 | GO:0035407 | histone H3-T11 phosphorylation(GO:0035407) |
| 0.5 | 2.9 | GO:0071500 | cellular response to nitrosative stress(GO:0071500) |
| 0.5 | 9.2 | GO:0042424 | catechol-containing compound catabolic process(GO:0019614) catecholamine catabolic process(GO:0042424) |
| 0.5 | 1.9 | GO:2000173 | negative regulation of dendrite extension(GO:1903860) regulation of neuron remodeling(GO:1904799) negative regulation of neuron remodeling(GO:1904800) negative regulation of branching morphogenesis of a nerve(GO:2000173) |
| 0.5 | 1.8 | GO:0015910 | peroxisomal long-chain fatty acid import(GO:0015910) |
| 0.5 | 1.4 | GO:0061182 | negative regulation of chondrocyte development(GO:0061182) |
| 0.4 | 1.3 | GO:0031453 | regulation of heterochromatin assembly(GO:0031445) positive regulation of heterochromatin assembly(GO:0031453) |
| 0.4 | 0.9 | GO:2000836 | androgen secretion(GO:0035935) regulation of androgen secretion(GO:2000834) positive regulation of androgen secretion(GO:2000836) |
| 0.4 | 1.8 | GO:0098886 | modification of dendritic spine(GO:0098886) |
| 0.4 | 3.9 | GO:0060267 | positive regulation of respiratory burst(GO:0060267) |
| 0.4 | 0.9 | GO:0061502 | early endosome to recycling endosome transport(GO:0061502) |
| 0.4 | 1.3 | GO:1990926 | short-term synaptic potentiation(GO:1990926) calcium ion regulated lysosome exocytosis(GO:1990927) |
| 0.4 | 2.8 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.4 | 3.2 | GO:0031022 | nuclear migration along microfilament(GO:0031022) |
| 0.4 | 1.9 | GO:0043988 | histone H3-S28 phosphorylation(GO:0043988) |
| 0.4 | 5.0 | GO:0034720 | histone H3-K4 demethylation(GO:0034720) |
| 0.3 | 1.0 | GO:1902823 | negative regulation of late endosome to lysosome transport(GO:1902823) regulation of protein processing involved in protein targeting to mitochondrion(GO:1903216) negative regulation of protein processing involved in protein targeting to mitochondrion(GO:1903217) |
| 0.3 | 1.0 | GO:0006679 | glucosylceramide biosynthetic process(GO:0006679) |
| 0.3 | 1.6 | GO:0051138 | positive regulation of NK T cell differentiation(GO:0051138) |
| 0.3 | 0.9 | GO:2000374 | negative regulation of hydrogen peroxide catabolic process(GO:2000296) regulation of oxygen metabolic process(GO:2000374) |
| 0.3 | 1.2 | GO:0003186 | tricuspid valve morphogenesis(GO:0003186) |
| 0.3 | 0.9 | GO:0035494 | SNARE complex disassembly(GO:0035494) |
| 0.3 | 1.2 | GO:1900224 | positive regulation of nodal signaling pathway involved in determination of lateral mesoderm left/right asymmetry(GO:1900224) |
| 0.3 | 0.9 | GO:0070904 | L-ascorbic acid transport(GO:0015882) molecular hydrogen transport(GO:0015993) transepithelial L-ascorbic acid transport(GO:0070904) |
| 0.3 | 1.5 | GO:1904693 | midbrain morphogenesis(GO:1904693) |
| 0.3 | 1.4 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 0.3 | 3.4 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.3 | 1.4 | GO:0030070 | insulin processing(GO:0030070) |
| 0.3 | 3.3 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.3 | 1.4 | GO:2000672 | negative regulation of motor neuron apoptotic process(GO:2000672) |
| 0.3 | 1.9 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 0.3 | 6.7 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.3 | 0.8 | GO:0030910 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) positive regulation of mesenchymal cell proliferation involved in ureter development(GO:2000729) |
| 0.3 | 1.6 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.3 | 0.8 | GO:0051596 | methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 0.2 | 1.7 | GO:1901189 | arterial endothelial cell fate commitment(GO:0060844) positive regulation of ephrin receptor signaling pathway(GO:1901189) positive regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901297) positive regulation of canonical Wnt signaling pathway involved in heart development(GO:1905068) |
| 0.2 | 3.7 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.2 | 1.0 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.2 | 2.1 | GO:0098582 | innate vocalization behavior(GO:0098582) positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.2 | 2.1 | GO:0031340 | positive regulation of vesicle fusion(GO:0031340) |
| 0.2 | 0.9 | GO:0043060 | meiotic metaphase I plate congression(GO:0043060) meiotic spindle midzone assembly(GO:0051257) meiotic metaphase plate congression(GO:0051311) |
| 0.2 | 0.2 | GO:0035026 | leading edge cell differentiation(GO:0035026) |
| 0.2 | 0.7 | GO:0071676 | negative regulation of mononuclear cell migration(GO:0071676) |
| 0.2 | 1.4 | GO:0010734 | protein glutathionylation(GO:0010731) regulation of protein glutathionylation(GO:0010732) negative regulation of protein glutathionylation(GO:0010734) |
| 0.2 | 3.1 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.2 | 0.6 | GO:0009386 | translational attenuation(GO:0009386) |
| 0.2 | 1.4 | GO:0031860 | regulation of DNA-dependent DNA replication initiation(GO:0030174) telomeric 3' overhang formation(GO:0031860) |
| 0.2 | 0.6 | GO:0031508 | pericentric heterochromatin assembly(GO:0031508) |
| 0.2 | 3.6 | GO:0001675 | acrosome assembly(GO:0001675) |
| 0.2 | 1.2 | GO:1901341 | activation of store-operated calcium channel activity(GO:0032237) positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.2 | 0.2 | GO:0019336 | phenol-containing compound catabolic process(GO:0019336) |
| 0.2 | 0.6 | GO:0043490 | malate-aspartate shuttle(GO:0043490) |
| 0.2 | 1.1 | GO:0010746 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) |
| 0.2 | 0.9 | GO:0000379 | tRNA-type intron splice site recognition and cleavage(GO:0000379) |
| 0.2 | 3.6 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.2 | 0.5 | GO:0001544 | initiation of primordial ovarian follicle growth(GO:0001544) |
| 0.2 | 0.7 | GO:1902568 | positive regulation of eosinophil degranulation(GO:0043311) positive regulation of eosinophil activation(GO:1902568) |
| 0.2 | 0.9 | GO:0016095 | polyprenol catabolic process(GO:0016095) |
| 0.2 | 0.5 | GO:0036245 | cellular response to menadione(GO:0036245) |
| 0.2 | 1.2 | GO:2000467 | positive regulation of glycogen (starch) synthase activity(GO:2000467) |
| 0.2 | 0.7 | GO:0072061 | inner medullary collecting duct development(GO:0072061) |
| 0.2 | 0.7 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.2 | 0.5 | GO:0016340 | calcium-dependent cell-matrix adhesion(GO:0016340) |
| 0.2 | 1.2 | GO:0050893 | sensory processing(GO:0050893) |
| 0.2 | 6.6 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.2 | 0.7 | GO:1901377 | mycotoxin catabolic process(GO:0043387) aflatoxin catabolic process(GO:0046223) organic heteropentacyclic compound catabolic process(GO:1901377) regulation of glutathione biosynthetic process(GO:1903786) positive regulation of glutathione biosynthetic process(GO:1903788) |
| 0.2 | 0.5 | GO:2000176 | regulation of pro-T cell differentiation(GO:2000174) positive regulation of pro-T cell differentiation(GO:2000176) |
| 0.2 | 0.5 | GO:0031443 | fast-twitch skeletal muscle fiber contraction(GO:0031443) |
| 0.2 | 0.6 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.2 | 0.6 | GO:2000820 | pathogenesis(GO:0009405) negative regulation of transcription from RNA polymerase II promoter involved in smooth muscle cell differentiation(GO:2000820) |
| 0.2 | 0.8 | GO:0006850 | mitochondrial pyruvate transport(GO:0006850) mitochondrial pyruvate transmembrane transport(GO:1902361) |
| 0.2 | 0.6 | GO:0071816 | tail-anchored membrane protein insertion into ER membrane(GO:0071816) |
| 0.1 | 2.7 | GO:0032310 | prostaglandin secretion(GO:0032310) |
| 0.1 | 0.3 | GO:0002071 | glandular epithelial cell maturation(GO:0002071) |
| 0.1 | 0.4 | GO:1903296 | regulation of glutamate secretion, neurotransmission(GO:1903294) positive regulation of glutamate secretion, neurotransmission(GO:1903296) |
| 0.1 | 3.5 | GO:0070493 | thrombin receptor signaling pathway(GO:0070493) |
| 0.1 | 1.0 | GO:0032380 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.1 | 0.3 | GO:0003221 | right ventricular cardiac muscle tissue morphogenesis(GO:0003221) |
| 0.1 | 0.4 | GO:0021817 | nucleokinesis involved in cell motility in cerebral cortex radial glia guided migration(GO:0021817) nuclear migration along microtubule(GO:0030473) |
| 0.1 | 1.1 | GO:0010637 | negative regulation of mitochondrial fusion(GO:0010637) |
| 0.1 | 0.4 | GO:1902303 | regulation of heart rate by hormone(GO:0003064) negative regulation of potassium ion export(GO:1902303) |
| 0.1 | 1.3 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.1 | 0.8 | GO:1903385 | regulation of homophilic cell adhesion(GO:1903385) |
| 0.1 | 0.3 | GO:0035880 | embryonic nail plate morphogenesis(GO:0035880) |
| 0.1 | 2.8 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.1 | 1.1 | GO:0097026 | dendritic cell dendrite assembly(GO:0097026) |
| 0.1 | 1.9 | GO:1990416 | cellular response to brain-derived neurotrophic factor stimulus(GO:1990416) |
| 0.1 | 1.4 | GO:0060136 | embryonic process involved in female pregnancy(GO:0060136) |
| 0.1 | 1.4 | GO:0042985 | negative regulation of amyloid precursor protein biosynthetic process(GO:0042985) |
| 0.1 | 0.4 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 0.1 | 0.9 | GO:0006498 | N-terminal protein lipidation(GO:0006498) N-terminal protein myristoylation(GO:0006499) |
| 0.1 | 0.7 | GO:0044240 | multicellular organism lipid catabolic process(GO:0044240) |
| 0.1 | 1.2 | GO:0060897 | neural plate anterior/posterior regionalization(GO:0021999) neural plate regionalization(GO:0060897) |
| 0.1 | 1.0 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.1 | 1.7 | GO:0051386 | regulation of neurotrophin TRK receptor signaling pathway(GO:0051386) |
| 0.1 | 0.5 | GO:0019605 | butyrate metabolic process(GO:0019605) |
| 0.1 | 0.5 | GO:0035513 | oxidative RNA demethylation(GO:0035513) oxidative single-stranded RNA demethylation(GO:0035553) |
| 0.1 | 5.9 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.1 | 7.9 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.1 | 0.3 | GO:0061428 | negative regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061428) |
| 0.1 | 7.5 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.1 | 1.7 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.1 | 1.0 | GO:0090625 | mRNA cleavage involved in gene silencing by siRNA(GO:0090625) |
| 0.1 | 0.2 | GO:0060024 | rhythmic synaptic transmission(GO:0060024) |
| 0.1 | 1.0 | GO:0040031 | snRNA modification(GO:0040031) |
| 0.1 | 0.5 | GO:0010700 | negative regulation of norepinephrine secretion(GO:0010700) |
| 0.1 | 0.8 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 0.1 | 0.9 | GO:0060591 | chondroblast differentiation(GO:0060591) |
| 0.1 | 1.5 | GO:0060573 | ventral spinal cord interneuron specification(GO:0021521) cell fate specification involved in pattern specification(GO:0060573) |
| 0.1 | 1.0 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.1 | 0.5 | GO:0009305 | protein biotinylation(GO:0009305) histone biotinylation(GO:0071110) |
| 0.1 | 0.8 | GO:0072757 | cellular response to camptothecin(GO:0072757) |
| 0.1 | 1.0 | GO:0000710 | meiotic mismatch repair(GO:0000710) |
| 0.1 | 0.7 | GO:0040034 | regulation of development, heterochronic(GO:0040034) regulation of timing of cell differentiation(GO:0048505) |
| 0.1 | 0.7 | GO:0045627 | positive regulation of T-helper 1 cell differentiation(GO:0045627) negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.1 | 0.5 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 0.1 | 0.6 | GO:0030950 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) |
| 0.1 | 1.3 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.1 | 0.6 | GO:0070086 | ubiquitin-dependent endocytosis(GO:0070086) |
| 0.1 | 0.4 | GO:0042091 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.1 | 1.7 | GO:0045475 | locomotor rhythm(GO:0045475) |
| 0.1 | 0.2 | GO:0034036 | purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.1 | 0.4 | GO:2000847 | negative regulation of steroid hormone secretion(GO:2000832) negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 0.1 | 0.2 | GO:1900127 | positive regulation of hyaluronan biosynthetic process(GO:1900127) |
| 0.1 | 0.7 | GO:0045079 | negative regulation of chemokine biosynthetic process(GO:0045079) |
| 0.1 | 0.9 | GO:0032782 | bile acid secretion(GO:0032782) |
| 0.1 | 0.7 | GO:0051549 | positive regulation of keratinocyte migration(GO:0051549) |
| 0.1 | 4.5 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.1 | 1.9 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.1 | 0.5 | GO:0051176 | positive regulation of sulfur metabolic process(GO:0051176) |
| 0.1 | 0.9 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.1 | 0.5 | GO:0097056 | selenocysteinyl-tRNA(Sec) biosynthetic process(GO:0097056) |
| 0.1 | 3.2 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 2.5 | GO:0086027 | AV node cell action potential(GO:0086016) AV node cell to bundle of His cell signaling(GO:0086027) |
| 0.1 | 1.2 | GO:0006655 | phosphatidylglycerol biosynthetic process(GO:0006655) |
| 0.1 | 7.2 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 0.1 | 0.4 | GO:0003025 | regulation of systemic arterial blood pressure by baroreceptor feedback(GO:0003025) |
| 0.1 | 0.3 | GO:0044837 | assembly of actomyosin apparatus involved in cytokinesis(GO:0000912) actomyosin contractile ring assembly(GO:0000915) actomyosin contractile ring organization(GO:0044837) |
| 0.1 | 2.1 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.1 | 0.9 | GO:0097267 | omega-hydroxylase P450 pathway(GO:0097267) |
| 0.1 | 0.4 | GO:0009138 | pyrimidine nucleoside diphosphate metabolic process(GO:0009138) |
| 0.1 | 0.3 | GO:0034970 | histone H3-R2 methylation(GO:0034970) |
| 0.1 | 5.6 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.1 | 1.0 | GO:0060059 | skeletal muscle satellite cell differentiation(GO:0014816) embryonic retina morphogenesis in camera-type eye(GO:0060059) |
| 0.1 | 0.5 | GO:1904688 | regulation of cap-independent translational initiation(GO:1903677) positive regulation of cap-independent translational initiation(GO:1903679) regulation of cytoplasmic translational initiation(GO:1904688) positive regulation of cytoplasmic translational initiation(GO:1904690) |
| 0.1 | 0.4 | GO:1904217 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.1 | 1.4 | GO:0031441 | negative regulation of mRNA 3'-end processing(GO:0031441) |
| 0.1 | 13.5 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.1 | 0.7 | GO:0070544 | histone H3-K36 demethylation(GO:0070544) |
| 0.1 | 0.6 | GO:0097327 | response to antineoplastic agent(GO:0097327) |
| 0.1 | 0.9 | GO:0015824 | proline transport(GO:0015824) |
| 0.1 | 0.9 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.1 | 0.5 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) nuclear retention of pre-mRNA at the site of transcription(GO:0071033) |
| 0.1 | 0.2 | GO:0036302 | atrioventricular canal development(GO:0036302) |
| 0.1 | 0.7 | GO:0034244 | negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.1 | 0.7 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.1 | 1.1 | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway(GO:0071880) |
| 0.1 | 0.4 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.1 | 2.3 | GO:0060325 | face morphogenesis(GO:0060325) |
| 0.1 | 0.3 | GO:0002143 | tRNA wobble position uridine thiolation(GO:0002143) |
| 0.1 | 0.4 | GO:0035865 | cellular response to potassium ion(GO:0035865) |
| 0.1 | 0.4 | GO:0018076 | N-terminal peptidyl-lysine acetylation(GO:0018076) |
| 0.1 | 1.2 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) |
| 0.1 | 0.4 | GO:0009048 | dosage compensation(GO:0007549) dosage compensation by inactivation of X chromosome(GO:0009048) |
| 0.1 | 0.4 | GO:0015853 | adenine transport(GO:0015853) |
| 0.1 | 0.2 | GO:0034402 | recruitment of 3'-end processing factors to RNA polymerase II holoenzyme complex(GO:0034402) |
| 0.1 | 0.1 | GO:1904562 | phosphatidylinositol 5-phosphate metabolic process(GO:1904562) |
| 0.1 | 0.3 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.1 | 1.0 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 0.1 | 0.3 | GO:0021966 | corticospinal neuron axon guidance(GO:0021966) |
| 0.1 | 0.5 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.1 | 0.5 | GO:1901315 | negative regulation of histone ubiquitination(GO:0033183) histone H2A K63-linked ubiquitination(GO:0070535) negative regulation of protein K63-linked ubiquitination(GO:1900045) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.1 | 1.3 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.1 | 0.3 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 0.1 | 0.5 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.1 | 0.8 | GO:0007042 | lysosomal lumen acidification(GO:0007042) |
| 0.1 | 0.8 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.1 | 1.3 | GO:0070884 | regulation of calcineurin-NFAT signaling cascade(GO:0070884) |
| 0.1 | 0.6 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.1 | 0.4 | GO:0071105 | response to interleukin-11(GO:0071105) |
| 0.1 | 0.2 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 0.1 | 0.7 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.1 | 0.4 | GO:0098706 | ferric iron import into cell(GO:0097461) ferric iron import across plasma membrane(GO:0098706) |
| 0.1 | 0.8 | GO:1990118 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 1.1 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.1 | 0.3 | GO:0034499 | late endosome to Golgi transport(GO:0034499) |
| 0.1 | 1.0 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) regulation of parathyroid hormone secretion(GO:2000828) |
| 0.1 | 0.2 | GO:0002528 | regulation of vascular permeability involved in acute inflammatory response(GO:0002528) |
| 0.1 | 0.5 | GO:0071376 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.1 | 1.8 | GO:0035066 | positive regulation of histone acetylation(GO:0035066) positive regulation of peptidyl-lysine acetylation(GO:2000758) |
| 0.1 | 0.2 | GO:0075071 | autophagy of host cells involved in interaction with symbiont(GO:0075044) autophagy involved in symbiotic interaction(GO:0075071) |
| 0.1 | 0.5 | GO:1902255 | positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator(GO:1902255) |
| 0.1 | 0.7 | GO:0060539 | diaphragm development(GO:0060539) |
| 0.1 | 0.1 | GO:0044340 | canonical Wnt signaling pathway involved in regulation of cell proliferation(GO:0044340) |
| 0.1 | 0.5 | GO:0000480 | endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000480) |
| 0.1 | 1.3 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.1 | 1.0 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.1 | 9.7 | GO:0046546 | male gonad development(GO:0008584) development of primary male sexual characteristics(GO:0046546) |
| 0.1 | 0.2 | GO:0070092 | glucagon secretion(GO:0070091) regulation of glucagon secretion(GO:0070092) |
| 0.1 | 0.4 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.1 | 2.6 | GO:0051973 | positive regulation of telomerase activity(GO:0051973) |
| 0.1 | 1.8 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.1 | 0.2 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.1 | 0.4 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.1 | 2.2 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.1 | 0.2 | GO:0046726 | positive regulation by virus of viral protein levels in host cell(GO:0046726) |
| 0.1 | 0.1 | GO:0061056 | sclerotome development(GO:0061056) |
| 0.1 | 0.3 | GO:0007056 | spindle assembly involved in female meiosis(GO:0007056) |
| 0.1 | 0.7 | GO:0045974 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.1 | 0.8 | GO:0035268 | protein mannosylation(GO:0035268) |
| 0.0 | 2.6 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.0 | 0.4 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.0 | 1.5 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 0.5 | GO:0007028 | cytoplasm organization(GO:0007028) |
| 0.0 | 0.5 | GO:0003356 | regulation of cilium beat frequency(GO:0003356) |
| 0.0 | 0.6 | GO:0043568 | positive regulation of insulin-like growth factor receptor signaling pathway(GO:0043568) |
| 0.0 | 0.5 | GO:0001767 | establishment of lymphocyte polarity(GO:0001767) |
| 0.0 | 0.3 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.0 | 0.8 | GO:0002115 | store-operated calcium entry(GO:0002115) |
| 0.0 | 0.6 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.0 | 1.3 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 1.1 | GO:0010745 | negative regulation of macrophage derived foam cell differentiation(GO:0010745) |
| 0.0 | 0.1 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 0.0 | 0.3 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.0 | 0.8 | GO:0098828 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.0 | 2.3 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.0 | 1.9 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.0 | 0.7 | GO:0031293 | membrane protein intracellular domain proteolysis(GO:0031293) |
| 0.0 | 0.2 | GO:1903756 | regulation of transcription from RNA polymerase II promoter by histone modification(GO:1903756) negative regulation of transcription from RNA polymerase II promoter by histone modification(GO:1903758) |
| 0.0 | 0.6 | GO:2000680 | rubidium ion transport(GO:0035826) regulation of rubidium ion transport(GO:2000680) |
| 0.0 | 0.9 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 0.3 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.0 | 1.3 | GO:0010954 | positive regulation of protein processing(GO:0010954) |
| 0.0 | 0.6 | GO:0006684 | sphingomyelin metabolic process(GO:0006684) |
| 0.0 | 0.2 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.0 | 1.7 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.0 | 4.5 | GO:0030449 | regulation of complement activation(GO:0030449) |
| 0.0 | 0.4 | GO:0097502 | mannosylation(GO:0097502) |
| 0.0 | 0.6 | GO:0008228 | opsonization(GO:0008228) |
| 0.0 | 2.6 | GO:0001942 | hair follicle development(GO:0001942) |
| 0.0 | 0.2 | GO:0045541 | negative regulation of cholesterol biosynthetic process(GO:0045541) negative regulation of cholesterol metabolic process(GO:0090206) |
| 0.0 | 1.2 | GO:2000144 | positive regulation of DNA-templated transcription, initiation(GO:2000144) |
| 0.0 | 0.1 | GO:0051582 | positive regulation of neurotransmitter uptake(GO:0051582) positive regulation of dopamine uptake involved in synaptic transmission(GO:0051586) positive regulation of catecholamine uptake involved in synaptic transmission(GO:0051944) |
| 0.0 | 0.6 | GO:0034243 | regulation of transcription elongation from RNA polymerase II promoter(GO:0034243) |
| 0.0 | 3.3 | GO:0006112 | energy reserve metabolic process(GO:0006112) |
| 0.0 | 1.1 | GO:0071549 | cellular response to dexamethasone stimulus(GO:0071549) |
| 0.0 | 0.3 | GO:0061709 | pexophagy(GO:0030242) reticulophagy(GO:0061709) |
| 0.0 | 0.3 | GO:0050966 | detection of mechanical stimulus involved in sensory perception of pain(GO:0050966) |
| 0.0 | 0.2 | GO:1902037 | negative regulation of hematopoietic stem cell differentiation(GO:1902037) |
| 0.0 | 0.9 | GO:0007141 | male meiosis I(GO:0007141) |
| 0.0 | 0.7 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.0 | 0.5 | GO:0031643 | positive regulation of myelination(GO:0031643) |
| 0.0 | 0.1 | GO:0071051 | polyadenylation-dependent snoRNA 3'-end processing(GO:0071051) |
| 0.0 | 0.3 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.0 | 1.3 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.1 | GO:0071139 | resolution of recombination intermediates(GO:0071139) resolution of mitotic recombination intermediates(GO:0071140) |
| 0.0 | 1.5 | GO:0046710 | GDP metabolic process(GO:0046710) |
| 0.0 | 0.1 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.0 | 1.6 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.0 | 0.9 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.0 | 0.3 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.0 | 0.3 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.0 | 0.5 | GO:0099624 | atrial cardiac muscle cell membrane repolarization(GO:0099624) |
| 0.0 | 0.2 | GO:1900121 | negative regulation of receptor binding(GO:1900121) |
| 0.0 | 0.7 | GO:0036499 | PERK-mediated unfolded protein response(GO:0036499) |
| 0.0 | 1.2 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.0 | 0.6 | GO:0035411 | catenin import into nucleus(GO:0035411) |
| 0.0 | 0.3 | GO:0043586 | tongue development(GO:0043586) |
| 0.0 | 0.3 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 0.0 | 0.7 | GO:0045956 | positive regulation of calcium ion-dependent exocytosis(GO:0045956) |
| 0.0 | 0.1 | GO:0045113 | regulation of integrin biosynthetic process(GO:0045113) |
| 0.0 | 0.5 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.0 | 1.1 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 2.5 | GO:0008543 | fibroblast growth factor receptor signaling pathway(GO:0008543) |
| 0.0 | 0.2 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.0 | 0.3 | GO:0070862 | negative regulation of protein exit from endoplasmic reticulum(GO:0070862) negative regulation of retrograde protein transport, ER to cytosol(GO:1904153) |
| 0.0 | 0.2 | GO:0070863 | positive regulation of protein exit from endoplasmic reticulum(GO:0070863) |
| 0.0 | 2.3 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 1.4 | GO:0043647 | inositol phosphate metabolic process(GO:0043647) |
| 0.0 | 0.6 | GO:0042133 | neurotransmitter metabolic process(GO:0042133) |
| 0.0 | 0.6 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.6 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.0 | 0.2 | GO:1900119 | positive regulation of execution phase of apoptosis(GO:1900119) |
| 0.0 | 0.2 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.0 | 0.4 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.0 | 0.9 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 0.4 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.0 | 1.1 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.0 | 0.1 | GO:0034552 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) respiratory chain complex II assembly(GO:0034552) mitochondrial respiratory chain complex II assembly(GO:0034553) mitochondrial respiratory chain complex II biogenesis(GO:0097032) |
| 0.0 | 0.1 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.0 | 0.3 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.0 | 0.4 | GO:0010863 | positive regulation of phospholipase C activity(GO:0010863) |
| 0.0 | 0.1 | GO:0015742 | alpha-ketoglutarate transport(GO:0015742) |
| 0.0 | 0.4 | GO:0045599 | negative regulation of fat cell differentiation(GO:0045599) |
| 0.0 | 0.2 | GO:1904424 | regulation of GTP binding(GO:1904424) |
| 0.0 | 1.1 | GO:0046849 | bone remodeling(GO:0046849) |
| 0.0 | 0.4 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.0 | 0.3 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.3 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.0 | 0.2 | GO:0097354 | protein prenylation(GO:0018342) prenylation(GO:0097354) |
| 0.0 | 0.1 | GO:0051967 | negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.0 | 0.1 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.0 | 0.1 | GO:2000580 | regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.0 | 0.3 | GO:0032743 | positive regulation of interleukin-2 production(GO:0032743) |
| 0.0 | 0.2 | GO:0070932 | histone H3 deacetylation(GO:0070932) |
| 0.0 | 0.1 | GO:0000722 | telomere maintenance via recombination(GO:0000722) |
| 0.0 | 0.2 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.0 | 1.2 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.0 | 0.2 | GO:0016576 | histone dephosphorylation(GO:0016576) |
| 0.0 | 0.2 | GO:0010832 | negative regulation of myotube differentiation(GO:0010832) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 9.5 | GO:0043291 | RAVE complex(GO:0043291) |
| 1.8 | 9.0 | GO:0072534 | perineuronal net(GO:0072534) |
| 1.0 | 9.2 | GO:1990761 | growth cone lamellipodium(GO:1990761) |
| 0.8 | 4.6 | GO:1990131 | Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.6 | 12.4 | GO:0031254 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.5 | 1.4 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.4 | 2.2 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.4 | 1.5 | GO:0097362 | MCM8-MCM9 complex(GO:0097362) |
| 0.4 | 3.3 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.3 | 1.0 | GO:0097454 | Schwann cell microvillus(GO:0097454) |
| 0.3 | 1.0 | GO:0044753 | amphisome(GO:0044753) |
| 0.3 | 3.9 | GO:0098799 | outer mitochondrial membrane protein complex(GO:0098799) |
| 0.3 | 5.8 | GO:0000786 | nucleosome(GO:0000786) |
| 0.2 | 1.2 | GO:0032444 | activin responsive factor complex(GO:0032444) |
| 0.2 | 0.9 | GO:0005715 | late recombination nodule(GO:0005715) |
| 0.2 | 0.7 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.2 | 0.4 | GO:0097636 | intrinsic component of autophagosome membrane(GO:0097636) integral component of autophagosome membrane(GO:0097637) |
| 0.2 | 1.8 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.2 | 0.9 | GO:0097123 | cyclin A1-CDK2 complex(GO:0097123) |
| 0.2 | 7.8 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.2 | 2.4 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.2 | 1.9 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.2 | 3.7 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.2 | 1.0 | GO:0032301 | MutSalpha complex(GO:0032301) |
| 0.2 | 1.7 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.2 | 0.9 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.2 | 1.2 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.2 | 3.8 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.2 | 3.3 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.2 | 7.6 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.2 | 0.9 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.2 | 0.6 | GO:0071818 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.2 | 0.3 | GO:0000805 | X chromosome(GO:0000805) |
| 0.1 | 2.6 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.1 | 1.3 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.1 | 2.6 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.1 | 2.8 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.1 | 8.9 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.1 | 1.1 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.1 | 1.4 | GO:0044233 | ER-mitochondrion membrane contact site(GO:0044233) |
| 0.1 | 0.9 | GO:0097450 | astrocyte end-foot(GO:0097450) |
| 0.1 | 2.7 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.1 | 3.2 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 1.0 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 0.1 | 0.6 | GO:0070695 | FHF complex(GO:0070695) |
| 0.1 | 0.9 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.1 | 1.6 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.1 | 0.7 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.1 | 17.2 | GO:0031514 | motile cilium(GO:0031514) |
| 0.1 | 1.1 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.1 | 0.4 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.1 | 0.4 | GO:0045293 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 0.1 | 0.7 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.1 | 0.2 | GO:0097232 | lamellar body membrane(GO:0097232) alveolar lamellar body membrane(GO:0097233) |
| 0.1 | 0.3 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.1 | 0.9 | GO:0032009 | early phagosome(GO:0032009) |
| 0.1 | 0.4 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.1 | 1.0 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.1 | 0.8 | GO:0005664 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.1 | 1.6 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.1 | 1.4 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.1 | 0.6 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.1 | 0.9 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.1 | 0.6 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.1 | 5.0 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.1 | 0.2 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.1 | 1.0 | GO:0070578 | RISC-loading complex(GO:0070578) |
| 0.1 | 0.2 | GO:0002945 | cyclin K-CDK13 complex(GO:0002945) |
| 0.0 | 0.4 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.0 | 0.5 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 3.2 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.0 | 0.8 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.0 | 0.5 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.0 | 0.7 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.2 | GO:0016589 | NURF complex(GO:0016589) |
| 0.0 | 5.2 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.0 | 3.1 | GO:0005844 | polysome(GO:0005844) |
| 0.0 | 2.8 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 0.9 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 0.7 | GO:0048500 | signal recognition particle(GO:0048500) |
| 0.0 | 0.8 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 1.2 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 2.1 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.3 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.5 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.6 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.0 | 0.5 | GO:0071203 | WASH complex(GO:0071203) |
| 0.0 | 0.6 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.0 | 5.3 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 0.5 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 0.3 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.0 | 0.8 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.0 | 0.2 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.0 | 0.3 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 0.4 | GO:0046930 | pore complex(GO:0046930) |
| 0.0 | 0.3 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.0 | 1.6 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 1.5 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 1.7 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.5 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.1 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.1 | GO:0089717 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.0 | 0.8 | GO:0031304 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.4 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 0.8 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 0.5 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.0 | 0.2 | GO:0005720 | nuclear heterochromatin(GO:0005720) |
| 0.0 | 0.1 | GO:0035061 | interchromatin granule(GO:0035061) |
| 0.0 | 0.2 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 0.4 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 0.3 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.4 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.0 | 0.1 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.0 | 0.5 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 0.2 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.0 | 0.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 9.7 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 0.4 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.0 | 0.4 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 0.1 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.0 | 0.4 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.0 | 0.2 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.0 | 0.1 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.0 | 0.3 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.0 | 1.3 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.0 | 0.2 | GO:0008091 | spectrin(GO:0008091) |
| 0.0 | 1.9 | GO:0005923 | bicellular tight junction(GO:0005923) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.1 | 12.4 | GO:0052810 | 1-phosphatidylinositol-5-kinase activity(GO:0052810) |
| 1.6 | 4.7 | GO:0005292 | high-affinity basic amino acid transmembrane transporter activity(GO:0005287) high-affinity arginine transmembrane transporter activity(GO:0005289) high-affinity lysine transmembrane transporter activity(GO:0005292) |
| 1.2 | 7.3 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 1.2 | 4.7 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 1.1 | 9.8 | GO:0019826 | oxygen sensor activity(GO:0019826) |
| 1.0 | 9.2 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.9 | 2.6 | GO:0033142 | progesterone receptor binding(GO:0033142) |
| 0.8 | 12.4 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.8 | 3.8 | GO:0031750 | D3 dopamine receptor binding(GO:0031750) |
| 0.7 | 2.2 | GO:0042781 | 3'-tRNA processing endoribonuclease activity(GO:0042781) |
| 0.7 | 5.0 | GO:0034648 | histone demethylase activity (H3-dimethyl-K4 specific)(GO:0034648) |
| 0.6 | 1.9 | GO:0031783 | corticotropin hormone receptor binding(GO:0031780) type 5 melanocortin receptor binding(GO:0031783) |
| 0.6 | 3.0 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.6 | 3.3 | GO:0030060 | L-malate dehydrogenase activity(GO:0030060) |
| 0.5 | 2.7 | GO:0016404 | 15-hydroxyprostaglandin dehydrogenase (NAD+) activity(GO:0016404) |
| 0.5 | 2.0 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.4 | 2.8 | GO:0005127 | ciliary neurotrophic factor receptor binding(GO:0005127) |
| 0.4 | 1.6 | GO:0031826 | type 2A serotonin receptor binding(GO:0031826) |
| 0.4 | 1.6 | GO:0035939 | microsatellite binding(GO:0035939) |
| 0.4 | 4.7 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.3 | 1.4 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.3 | 1.0 | GO:1904713 | beta-catenin destruction complex binding(GO:1904713) |
| 0.3 | 10.0 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.3 | 2.7 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.3 | 0.9 | GO:0070890 | L-ascorbate:sodium symporter activity(GO:0008520) sodium-dependent multivitamin transmembrane transporter activity(GO:0008523) L-ascorbic acid transporter activity(GO:0015229) sodium-dependent L-ascorbate transmembrane transporter activity(GO:0070890) |
| 0.3 | 0.6 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) thiol oxidase activity(GO:0016972) |
| 0.3 | 0.9 | GO:0035575 | histone demethylase activity (H4-K20 specific)(GO:0035575) |
| 0.3 | 1.1 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.3 | 0.8 | GO:0004345 | glucose-6-phosphate dehydrogenase activity(GO:0004345) |
| 0.3 | 1.8 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.3 | 1.6 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.3 | 0.8 | GO:0004416 | hydroxyacylglutathione hydrolase activity(GO:0004416) |
| 0.2 | 0.7 | GO:0004605 | phosphatidate cytidylyltransferase activity(GO:0004605) |
| 0.2 | 6.6 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.2 | 2.2 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.2 | 3.3 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.2 | 2.3 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
| 0.2 | 1.1 | GO:0033857 | diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.2 | 0.7 | GO:0047389 | glycerophosphocholine phosphodiesterase activity(GO:0047389) |
| 0.2 | 1.1 | GO:0004803 | transposase activity(GO:0004803) |
| 0.2 | 2.9 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.2 | 9.5 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.2 | 1.0 | GO:0008321 | Ral guanyl-nucleotide exchange factor activity(GO:0008321) |
| 0.2 | 1.0 | GO:0032143 | single thymine insertion binding(GO:0032143) |
| 0.2 | 3.4 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.2 | 3.2 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.2 | 6.7 | GO:0005328 | neurotransmitter:sodium symporter activity(GO:0005328) |
| 0.2 | 0.6 | GO:0050405 | [hydroxymethylglutaryl-CoA reductase (NADPH)] kinase activity(GO:0047322) [acetyl-CoA carboxylase] kinase activity(GO:0050405) |
| 0.2 | 5.8 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.2 | 0.9 | GO:0004882 | androgen receptor activity(GO:0004882) |
| 0.2 | 1.6 | GO:0042285 | UDP-xylosyltransferase activity(GO:0035252) xylosyltransferase activity(GO:0042285) |
| 0.2 | 4.2 | GO:0043495 | protein anchor(GO:0043495) |
| 0.2 | 4.9 | GO:0016864 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.2 | 0.5 | GO:0050146 | nucleoside phosphotransferase activity(GO:0050146) |
| 0.2 | 6.9 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.2 | 0.7 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.2 | 1.0 | GO:0098821 | BMP receptor activity(GO:0098821) |
| 0.2 | 7.2 | GO:0019200 | carbohydrate kinase activity(GO:0019200) |
| 0.2 | 2.8 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.2 | 2.6 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.2 | 2.2 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.2 | 0.6 | GO:0032440 | 2-alkenal reductase [NAD(P)] activity(GO:0032440) |
| 0.2 | 7.5 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.2 | 4.2 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 1.0 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.1 | 1.3 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.1 | 0.4 | GO:0016422 | mRNA (2'-O-methyladenosine-N6-)-methyltransferase activity(GO:0016422) |
| 0.1 | 0.9 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.1 | 0.9 | GO:0032137 | guanine/thymine mispair binding(GO:0032137) |
| 0.1 | 0.4 | GO:0070336 | flap-structured DNA binding(GO:0070336) |
| 0.1 | 0.5 | GO:0035515 | oxidative RNA demethylase activity(GO:0035515) |
| 0.1 | 0.4 | GO:0102008 | cytosolic dipeptidase activity(GO:0102008) |
| 0.1 | 0.6 | GO:0008160 | protein tyrosine phosphatase activator activity(GO:0008160) |
| 0.1 | 4.6 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.1 | 0.7 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.1 | 6.2 | GO:0016409 | palmitoyltransferase activity(GO:0016409) |
| 0.1 | 0.5 | GO:0004079 | biotin-[acetyl-CoA-carboxylase] ligase activity(GO:0004077) biotin-[methylcrotonoyl-CoA-carboxylase] ligase activity(GO:0004078) biotin-[methylmalonyl-CoA-carboxytransferase] ligase activity(GO:0004079) biotin-[propionyl-CoA-carboxylase (ATP-hydrolyzing)] ligase activity(GO:0004080) biotin-protein ligase activity(GO:0018271) |
| 0.1 | 0.3 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.1 | 3.1 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 1.1 | GO:0001727 | lipid kinase activity(GO:0001727) |
| 0.1 | 1.4 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.1 | 1.7 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.1 | 0.9 | GO:0000213 | tRNA-intron endonuclease activity(GO:0000213) |
| 0.1 | 1.2 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.1 | 0.6 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.1 | 0.4 | GO:0003865 | 3-oxo-5-alpha-steroid 4-dehydrogenase activity(GO:0003865) steroid dehydrogenase activity, acting on the CH-CH group of donors(GO:0033765) cholestenone 5-alpha-reductase activity(GO:0047751) |
| 0.1 | 1.3 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.1 | 2.1 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.1 | 3.4 | GO:0016505 | peptidase activator activity involved in apoptotic process(GO:0016505) |
| 0.1 | 0.3 | GO:0086059 | voltage-gated calcium channel activity involved SA node cell action potential(GO:0086059) |
| 0.1 | 0.3 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.1 | 4.4 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 2.5 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.1 | 1.2 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.1 | 0.5 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.1 | 3.2 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 1.0 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.1 | 0.4 | GO:0047273 | galactosylgalactosylglucosylceramide beta-D-acetylgalactosaminyltransferase activity(GO:0047273) |
| 0.1 | 0.5 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.1 | 1.8 | GO:0005112 | Notch binding(GO:0005112) |
| 0.1 | 1.2 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 0.4 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.1 | 0.1 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.1 | 1.7 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.1 | 0.9 | GO:0046870 | cadmium ion binding(GO:0046870) |
| 0.1 | 0.4 | GO:0052851 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.1 | 2.9 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.1 | 1.4 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.1 | 0.1 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.1 | 0.7 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.1 | 0.7 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.1 | 0.8 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.1 | 15.4 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.1 | 1.6 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.1 | 0.7 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.1 | 0.5 | GO:0050833 | pyruvate transmembrane transporter activity(GO:0050833) |
| 0.1 | 0.5 | GO:0030911 | TPR domain binding(GO:0030911) |
| 0.1 | 0.2 | GO:0051765 | inositol tetrakisphosphate kinase activity(GO:0051765) |
| 0.1 | 2.1 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 1.9 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.1 | 3.0 | GO:0098811 | transcriptional activator activity, RNA polymerase II transcription factor binding(GO:0001190) transcriptional repressor activity, RNA polymerase II activating transcription factor binding(GO:0098811) |
| 0.1 | 5.0 | GO:0061733 | peptide-lysine-N-acetyltransferase activity(GO:0061733) |
| 0.1 | 0.9 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.1 | 0.6 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.1 | 2.4 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 0.6 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.0 | 1.4 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.0 | 0.3 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.0 | 0.3 | GO:0001225 | RNA polymerase II transcription coactivator binding(GO:0001225) |
| 0.0 | 1.0 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 0.0 | 0.9 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.5 | GO:0031685 | G-protein coupled adenosine receptor activity(GO:0001609) adenosine receptor binding(GO:0031685) |
| 0.0 | 0.6 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.5 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.0 | 0.2 | GO:0035241 | protein-arginine omega-N monomethyltransferase activity(GO:0035241) |
| 0.0 | 0.4 | GO:0010385 | double-stranded methylated DNA binding(GO:0010385) |
| 0.0 | 0.7 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.0 | 1.5 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 3.4 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.0 | 0.1 | GO:0060001 | minus-end directed microfilament motor activity(GO:0060001) |
| 0.0 | 0.4 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.0 | 1.1 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 4.9 | GO:0005496 | steroid binding(GO:0005496) |
| 0.0 | 0.2 | GO:0004800 | thyroxine 5'-deiodinase activity(GO:0004800) |
| 0.0 | 1.2 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 3.6 | GO:0000987 | core promoter proximal region sequence-specific DNA binding(GO:0000987) |
| 0.0 | 18.9 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.7 | GO:0016857 | racemase and epimerase activity, acting on carbohydrates and derivatives(GO:0016857) |
| 0.0 | 2.2 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.0 | 0.9 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.0 | 1.0 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 0.9 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 1.8 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.7 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 1.0 | GO:0031748 | D1 dopamine receptor binding(GO:0031748) |
| 0.0 | 0.6 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.0 | 0.3 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.0 | 0.3 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.0 | 0.9 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 1.5 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.0 | 1.0 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.0 | 0.2 | GO:0008318 | protein prenyltransferase activity(GO:0008318) |
| 0.0 | 0.2 | GO:0050659 | N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase activity(GO:0050659) |
| 0.0 | 0.2 | GO:0010858 | calcium-dependent protein kinase regulator activity(GO:0010858) |
| 0.0 | 0.4 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.0 | 0.9 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.0 | 0.1 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 0.2 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 1.2 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.4 | GO:0004935 | adrenergic receptor activity(GO:0004935) |
| 0.0 | 0.1 | GO:0016509 | long-chain-3-hydroxyacyl-CoA dehydrogenase activity(GO:0016509) |
| 0.0 | 0.5 | GO:0016594 | glycine binding(GO:0016594) |
| 0.0 | 2.3 | GO:0042626 | ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 0.5 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.0 | 1.9 | GO:0004004 | ATP-dependent RNA helicase activity(GO:0004004) RNA-dependent ATPase activity(GO:0008186) |
| 0.0 | 0.9 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.6 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.2 | GO:0015288 | porin activity(GO:0015288) |
| 0.0 | 0.9 | GO:0043175 | RNA polymerase core enzyme binding(GO:0043175) |
| 0.0 | 0.4 | GO:0009881 | photoreceptor activity(GO:0009881) |
| 0.0 | 0.2 | GO:0030267 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.0 | 0.4 | GO:0001055 | RNA polymerase II activity(GO:0001055) |
| 0.0 | 0.1 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.0 | 0.9 | GO:0001221 | transcription cofactor binding(GO:0001221) |
| 0.0 | 0.3 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.0 | 4.0 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.5 | GO:0008198 | ferrous iron binding(GO:0008198) |
| 0.0 | 0.1 | GO:0000982 | transcription factor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0000982) |
| 0.0 | 0.2 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.1 | GO:0044547 | DNA topoisomerase binding(GO:0044547) |
| 0.0 | 0.5 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 1.4 | GO:0051287 | NAD binding(GO:0051287) |
| 0.0 | 1.3 | GO:0042805 | actinin binding(GO:0042805) |
| 0.0 | 0.3 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.0 | 0.1 | GO:0072590 | N-acetyl-L-aspartate-L-glutamate ligase activity(GO:0072590) |
| 0.0 | 0.9 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.1 | GO:0015140 | malate transmembrane transporter activity(GO:0015140) |
| 0.0 | 0.6 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.1 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.0 | 0.3 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.4 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.1 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.0 | 0.3 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.0 | 0.4 | GO:0016875 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.0 | 0.1 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.0 | 0.4 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 1.2 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 0.1 | GO:0003696 | satellite DNA binding(GO:0003696) |
| 0.0 | 0.1 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 2.3 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 4.4 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.1 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.0 | 0.1 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.0 | 0.7 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 0.5 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.2 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 12.6 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.3 | 9.3 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.2 | 0.7 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.2 | 16.4 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.1 | 5.3 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.1 | 1.7 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 1.5 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.1 | 13.2 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.1 | 10.0 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.1 | 3.6 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.1 | 2.4 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.1 | 0.9 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 2.3 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 0.9 | ST ADRENERGIC | Adrenergic Pathway |
| 0.1 | 1.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 1.2 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 2.1 | SIG IL4RECEPTOR IN B LYPHOCYTES | Genes related to IL4 rceptor signaling in B lymphocytes |
| 0.0 | 0.4 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.0 | 2.2 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.0 | 2.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.3 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 0.6 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.0 | 0.4 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.0 | 1.4 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 0.8 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 0.7 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.0 | 0.5 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 0.7 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 0.6 | ST G ALPHA I PATHWAY | G alpha i Pathway |
| 0.0 | 0.6 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 0.3 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 1.0 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 0.2 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.0 | 0.2 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 0.2 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 0.2 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 0.4 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.0 | 0.2 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 1.2 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.5 | PID INSULIN PATHWAY | Insulin Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.4 | 4.7 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.2 | 9.6 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.2 | 7.2 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.2 | 10.6 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.2 | 4.3 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.2 | 3.4 | REACTOME INITIAL TRIGGERING OF COMPLEMENT | Genes involved in Initial triggering of complement |
| 0.2 | 5.3 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.1 | 3.1 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.1 | 1.9 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 3.1 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 1.3 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.1 | 3.1 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.1 | 2.1 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 2.7 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.1 | 2.1 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.1 | 1.4 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.1 | 0.7 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.1 | 2.6 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.1 | 2.3 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.1 | 1.6 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 1.9 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.1 | 0.9 | REACTOME NRIF SIGNALS CELL DEATH FROM THE NUCLEUS | Genes involved in NRIF signals cell death from the nucleus |
| 0.1 | 1.2 | REACTOME HIGHLY CALCIUM PERMEABLE POSTSYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Highly calcium permeable postsynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.9 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.0 | 0.4 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.6 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.0 | 0.7 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.0 | 0.1 | REACTOME SIGNALING BY EGFR IN CANCER | Genes involved in Signaling by EGFR in Cancer |
| 0.0 | 1.7 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 0.5 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.0 | 4.5 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.8 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 1.0 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 1.3 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 2.2 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.2 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.0 | 0.9 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.0 | 1.1 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 1.1 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 0.4 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.0 | 0.7 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 0.5 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 1.8 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 4.7 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 0.7 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.9 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.0 | 0.1 | REACTOME VIF MEDIATED DEGRADATION OF APOBEC3G | Genes involved in Vif-mediated degradation of APOBEC3G |
| 0.0 | 0.8 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.4 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.0 | 0.4 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.0 | 0.4 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.0 | 0.2 | REACTOME ACTIVATION OF CHAPERONES BY ATF6 ALPHA | Genes involved in Activation of Chaperones by ATF6-alpha |
| 0.0 | 0.6 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.0 | 1.5 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 1.0 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.2 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.0 | 0.6 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.0 | 2.4 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.1 | REACTOME GAP JUNCTION DEGRADATION | Genes involved in Gap junction degradation |
| 0.0 | 0.2 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.4 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.2 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 0.3 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 0.6 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 0.9 | REACTOME SCFSKP2 MEDIATED DEGRADATION OF P27 P21 | Genes involved in SCF(Skp2)-mediated degradation of p27/p21 |
| 0.0 | 0.2 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |