Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
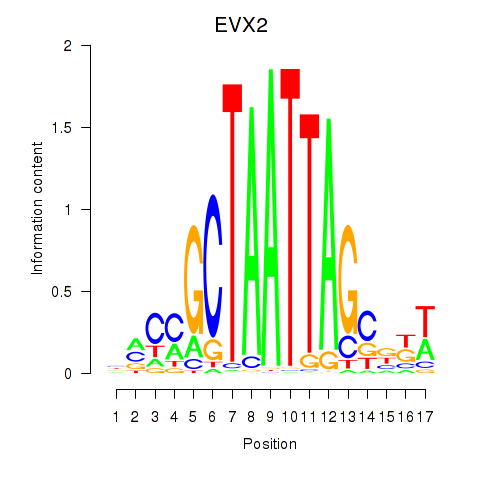
Results for EVX2
Z-value: 0.30
Transcription factors associated with EVX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
EVX2
|
ENSG00000174279.4 | EVX2 |
Activity profile of EVX2 motif
Sorted Z-values of EVX2 motif
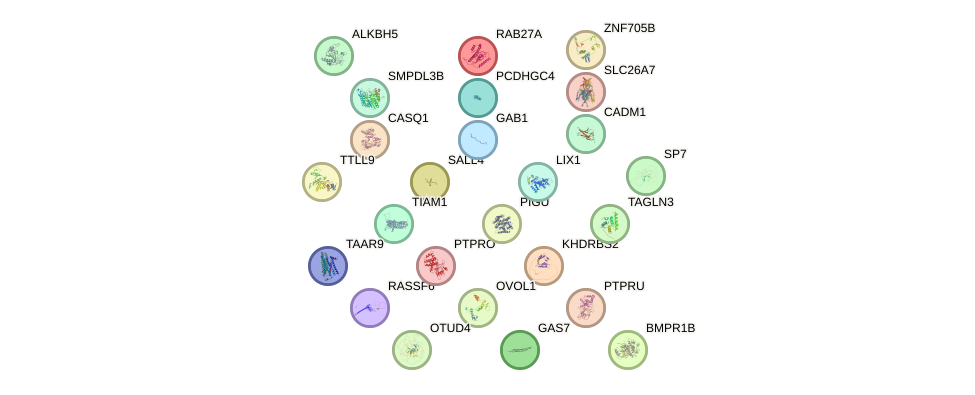
Network of associatons between targets according to the STRING database.
First level regulatory network of EVX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_-_55270280 | 0.48 |
ENST00000564609.5
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr15_-_55270383 | 0.45 |
ENST00000396307.6
|
RAB27A
|
RAB27A, member RAS oncogene family |
| chr15_-_55270874 | 0.44 |
ENST00000567380.5
ENST00000565972.5 ENST00000569493.5 |
RAB27A
|
RAB27A, member RAS oncogene family |
| chr4_+_143433491 | 0.42 |
ENST00000512843.1
|
GAB1
|
GRB2 associated binding protein 1 |
| chr3_+_111999326 | 0.37 |
ENST00000494932.1
|
TAGLN3
|
transgelin 3 |
| chr3_+_111998739 | 0.36 |
ENST00000393917.6
ENST00000273368.8 |
TAGLN3
|
transgelin 3 |
| chr3_+_111999189 | 0.35 |
ENST00000455401.6
|
TAGLN3
|
transgelin 3 |
| chr3_+_111998915 | 0.35 |
ENST00000478951.6
|
TAGLN3
|
transgelin 3 |
| chr6_-_62286161 | 0.32 |
ENST00000281156.5
|
KHDRBS2
|
KH RNA binding domain containing, signal transduction associated 2 |
| chr1_+_160190567 | 0.31 |
ENST00000368078.8
|
CASQ1
|
calsequestrin 1 |
| chr20_-_51802509 | 0.23 |
ENST00000371539.7
ENST00000217086.9 |
SALL4
|
spalt like transcription factor 4 |
| chr5_+_141484997 | 0.20 |
ENST00000617094.1
ENST00000306593.2 ENST00000610539.1 ENST00000618371.4 |
PCDHGC4
|
protocadherin gamma subfamily C, 4 |
| chr20_-_51802433 | 0.16 |
ENST00000395997.3
|
SALL4
|
spalt like transcription factor 4 |
| chr4_-_145180496 | 0.15 |
ENST00000447906.8
|
OTUD4
|
OTU deubiquitinase 4 |
| chr4_+_94974984 | 0.15 |
ENST00000672698.1
|
BMPR1B
|
bone morphogenetic protein receptor type 1B |
| chr1_+_27934980 | 0.15 |
ENST00000373894.8
|
SMPDL3B
|
sphingomyelin phosphodiesterase acid like 3B |
| chr12_-_53336342 | 0.15 |
ENST00000537210.2
ENST00000536324.4 |
SP7
|
Sp7 transcription factor |
| chr1_+_27935110 | 0.14 |
ENST00000549094.1
|
SMPDL3B
|
sphingomyelin phosphodiesterase acid like 3B |
| chr1_+_27935022 | 0.14 |
ENST00000411604.5
ENST00000373888.8 |
SMPDL3B
|
sphingomyelin phosphodiesterase acid like 3B |
| chr11_+_65787056 | 0.13 |
ENST00000335987.8
|
OVOL1
|
ovo like transcriptional repressor 1 |
| chr6_+_132538290 | 0.12 |
ENST00000434551.2
|
TAAR9
|
trace amine associated receptor 9 |
| chr20_+_31879798 | 0.09 |
ENST00000620043.4
|
TTLL9
|
tubulin tyrosine ligase like 9 |
| chr12_+_15546344 | 0.09 |
ENST00000674388.1
ENST00000542557.5 ENST00000445537.6 ENST00000544244.5 ENST00000442921.7 |
PTPRO
|
protein tyrosine phosphatase receptor type O |
| chr12_+_119668109 | 0.07 |
ENST00000229328.10
ENST00000630317.1 |
PRKAB1
|
protein kinase AMP-activated non-catalytic subunit beta 1 |
| chr8_+_7926337 | 0.07 |
ENST00000400120.3
|
ZNF705B
|
zinc finger protein 705B |
| chr8_+_91249307 | 0.05 |
ENST00000309536.6
ENST00000276609.8 |
SLC26A7
|
solute carrier family 26 member 7 |
| chr17_-_10026265 | 0.03 |
ENST00000437099.6
ENST00000396115.6 |
GAS7
|
growth arrest specific 7 |
| chr5_-_97142579 | 0.03 |
ENST00000274382.9
|
LIX1
|
limb and CNS expressed 1 |
| chr11_-_115504389 | 0.02 |
ENST00000545380.5
ENST00000452722.7 ENST00000331581.11 ENST00000537058.5 ENST00000536727.5 ENST00000542447.6 |
CADM1
|
cell adhesion molecule 1 |
| chr17_+_18183052 | 0.01 |
ENST00000541285.1
|
ALKBH5
|
alkB homolog 5, RNA demethylase |
| chr21_-_31160904 | 0.00 |
ENST00000636887.1
|
TIAM1
|
TIAM Rac1 associated GEF 1 |
| chr4_-_73620629 | 0.00 |
ENST00000342081.7
|
RASSF6
|
Ras association domain family member 6 |
| chr16_+_53099100 | 0.00 |
ENST00000565832.5
|
CHD9
|
chromodomain helicase DNA binding protein 9 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.4 | GO:1903435 | positive regulation of constitutive secretory pathway(GO:1903435) |
| 0.0 | 0.1 | GO:0044771 | meiotic cell cycle phase transition(GO:0044771) regulation of meiotic cell cycle phase transition(GO:1901993) negative regulation of meiotic cell cycle phase transition(GO:1901994) |
| 0.0 | 0.4 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 0.4 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.0 | 0.3 | GO:0014877 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 0.0 | 0.1 | GO:0036058 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.0 | 0.2 | GO:0001550 | ovarian cumulus expansion(GO:0001550) fused antrum stage(GO:0048165) negative regulation of chondrocyte proliferation(GO:1902731) |
| 0.0 | 0.4 | GO:0035728 | response to hepatocyte growth factor(GO:0035728) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | GO:0033093 | Weibel-Palade body(GO:0033093) |
| 0.1 | 0.3 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.0 | 0.2 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.4 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.0 | 0.4 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.1 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.2 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.4 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.4 | REACTOME SIGNALING BY CONSTITUTIVELY ACTIVE EGFR | Genes involved in Signaling by constitutively active EGFR |