Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
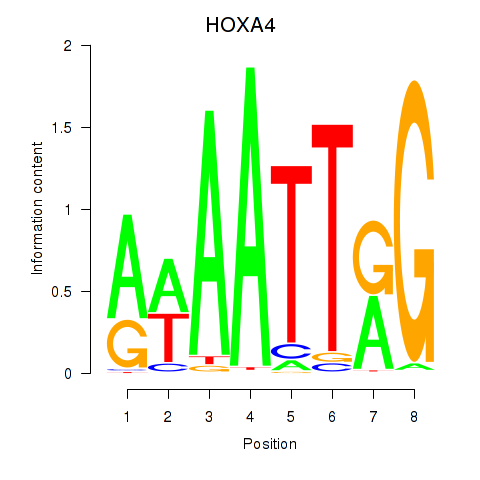
Results for HOXA4
Z-value: 0.53
Transcription factors associated with HOXA4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXA4
|
ENSG00000197576.14 | HOXA4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HOXA4 | hg38_v1_chr7_-_27130733_27130780 | 0.53 | 6.7e-03 | Click! |
Activity profile of HOXA4 motif
Sorted Z-values of HOXA4 motif
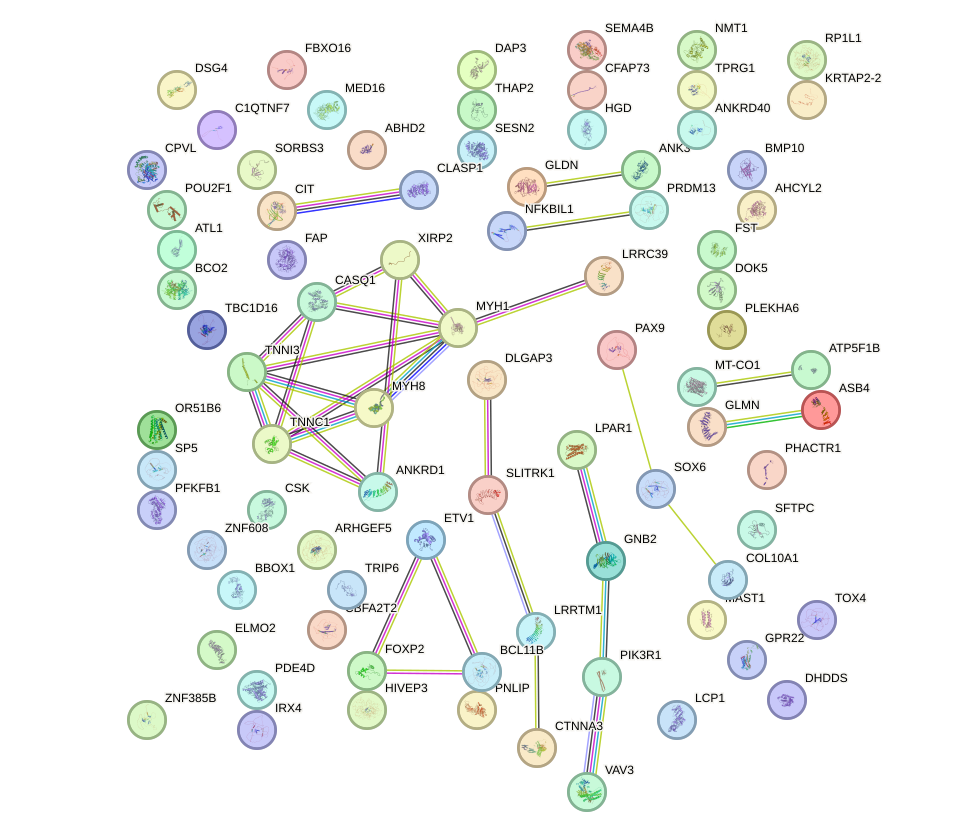
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXA4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_+_54475584 | 1.30 |
ENST00000262593.10
|
DOK5
|
docking protein 5 |
| chr20_+_54475647 | 1.22 |
ENST00000395939.5
|
DOK5
|
docking protein 5 |
| chr1_+_28259473 | 1.09 |
ENST00000253063.4
|
SESN2
|
sestrin 2 |
| chr5_-_59430600 | 0.98 |
ENST00000636120.1
|
PDE4D
|
phosphodiesterase 4D |
| chr19_+_12838437 | 0.78 |
ENST00000251472.9
|
MAST1
|
microtubule associated serine/threonine kinase 1 |
| chr5_-_124746630 | 0.76 |
ENST00000513986.2
|
ZNF608
|
zinc finger protein 608 |
| chr1_+_158461574 | 0.72 |
ENST00000641432.1
ENST00000641460.1 ENST00000641535.1 ENST00000641971.1 |
OR10K1
|
olfactory receptor family 10 subfamily K member 1 |
| chr15_+_74788542 | 0.58 |
ENST00000567571.5
|
CSK
|
C-terminal Src kinase |
| chr5_+_68290637 | 0.56 |
ENST00000336483.9
|
PIK3R1
|
phosphoinositide-3-kinase regulatory subunit 1 |
| chr5_+_53480619 | 0.55 |
ENST00000396947.7
ENST00000256759.8 |
FST
|
follistatin |
| chr10_-_90921079 | 0.54 |
ENST00000371697.4
|
ANKRD1
|
ankyrin repeat domain 1 |
| chr7_-_13986439 | 0.53 |
ENST00000443608.5
ENST00000438956.5 |
ETV1
|
ETS variant transcription factor 1 |
| chr6_+_12716760 | 0.51 |
ENST00000332995.12
|
PHACTR1
|
phosphatase and actin regulator 1 |
| chr2_-_68871382 | 0.50 |
ENST00000295379.2
|
BMP10
|
bone morphogenetic protein 10 |
| chr4_-_152352800 | 0.47 |
ENST00000393956.9
|
FBXW7
|
F-box and WD repeat domain containing 7 |
| chr2_+_167187364 | 0.45 |
ENST00000672671.1
|
XIRP2
|
xin actin binding repeat containing 2 |
| chr7_-_27130733 | 0.43 |
ENST00000428284.2
ENST00000360046.10 ENST00000610970.1 |
HOXA4
|
homeobox A4 |
| chr17_-_41055211 | 0.43 |
ENST00000542910.1
ENST00000398477.1 |
KRTAP2-2
|
keratin associated protein 2-2 |
| chr12_-_119803383 | 0.41 |
ENST00000392520.2
ENST00000678677.1 ENST00000679249.1 ENST00000676849.1 |
CIT
|
citron rho-interacting serine/threonine kinase |
| chr12_+_71664352 | 0.40 |
ENST00000547843.1
|
THAP2
|
THAP domain containing 2 |
| chr2_-_179562576 | 0.38 |
ENST00000336917.9
|
ZNF385B
|
zinc finger protein 385B |
| chr8_-_10655137 | 0.38 |
ENST00000382483.4
|
RP1L1
|
RP1 like 1 |
| chr17_-_10549694 | 0.37 |
ENST00000622564.4
|
MYH2
|
myosin heavy chain 2 |
| chr8_-_28490220 | 0.37 |
ENST00000517673.5
ENST00000380254.7 ENST00000518734.5 ENST00000346498.6 |
FBXO16
|
F-box protein 16 |
| chr17_-_10549612 | 0.36 |
ENST00000532183.6
ENST00000397183.6 ENST00000420805.1 |
MYH2
|
myosin heavy chain 2 |
| chr17_-_10549652 | 0.35 |
ENST00000245503.10
|
MYH2
|
myosin heavy chain 2 |
| chr18_+_31376777 | 0.34 |
ENST00000308128.9
ENST00000359747.4 |
DSG4
|
desmoglein 4 |
| chrM_+_5824 | 0.34 |
ENST00000361624.2
|
MT-CO1
|
mitochondrially encoded cytochrome c oxidase I |
| chr17_-_79950828 | 0.34 |
ENST00000572862.5
ENST00000573782.5 ENST00000574427.1 ENST00000570373.5 ENST00000340848.11 ENST00000576768.5 |
TBC1D16
|
TBC1 domain family member 16 |
| chr1_-_100178215 | 0.34 |
ENST00000370138.1
ENST00000370137.6 ENST00000342895.7 ENST00000620882.4 |
LRRC39
|
leucine rich repeat containing 39 |
| chr2_-_80304274 | 0.33 |
ENST00000409148.1
ENST00000415098.1 ENST00000452811.1 |
LRRTM1
|
leucine rich repeat transmembrane neuronal 1 |
| chr13_-_46142834 | 0.33 |
ENST00000674665.1
|
LCP1
|
lymphocyte cytosolic protein 1 |
| chr5_-_38557459 | 0.33 |
ENST00000511561.1
|
LIFR
|
LIF receptor subunit alpha |
| chr1_-_41918858 | 0.32 |
ENST00000372583.6
|
HIVEP3
|
HIVEP zinc finger 3 |
| chr11_-_16356538 | 0.31 |
ENST00000683767.1
|
SOX6
|
SRY-box transcription factor 6 |
| chr5_-_1882902 | 0.30 |
ENST00000231357.7
|
IRX4
|
iroquois homeobox 4 |
| chr6_+_12716801 | 0.30 |
ENST00000674595.1
|
PHACTR1
|
phosphatase and actin regulator 1 |
| chr1_+_167329044 | 0.30 |
ENST00000367862.9
|
POU2F1
|
POU class 2 homeobox 1 |
| chr2_+_170715317 | 0.30 |
ENST00000375281.4
|
SP5
|
Sp5 transcription factor |
| chr15_+_89088417 | 0.30 |
ENST00000569550.5
ENST00000565066.5 ENST00000565973.5 ENST00000352732.10 |
ABHD2
|
abhydrolase domain containing 2, acylglycerol lipase |
| chr8_-_94436926 | 0.29 |
ENST00000481490.3
|
FSBP
|
fibrinogen silencer binding protein |
| chr10_+_116545907 | 0.29 |
ENST00000369221.2
|
PNLIP
|
pancreatic lipase |
| chr1_-_41918666 | 0.28 |
ENST00000372584.5
|
HIVEP3
|
HIVEP zinc finger 3 |
| chr2_-_2324323 | 0.28 |
ENST00000648339.1
ENST00000647694.1 |
MYT1L
|
myelin transcription factor 1 like |
| chr17_+_44957907 | 0.28 |
ENST00000678938.1
|
NMT1
|
N-myristoyltransferase 1 |
| chr7_+_107470050 | 0.27 |
ENST00000304402.6
|
GPR22
|
G protein-coupled receptor 22 |
| chr6_+_12716554 | 0.27 |
ENST00000676159.1
|
PHACTR1
|
phosphatase and actin regulator 1 |
| chr20_-_46406582 | 0.27 |
ENST00000450812.5
ENST00000290246.11 ENST00000396391.5 |
ELMO2
|
engulfment and cell motility 2 |
| chr7_+_144355288 | 0.26 |
ENST00000498580.5
ENST00000056217.10 |
ARHGEF5
|
Rho guanine nucleotide exchange factor 5 |
| chr14_+_22515623 | 0.25 |
ENST00000390509.1
|
TRAJ28
|
T cell receptor alpha joining 28 |
| chr8_+_22567038 | 0.24 |
ENST00000523348.1
|
SORBS3
|
sorbin and SH3 domain containing 3 |
| chr15_+_51377247 | 0.24 |
ENST00000396399.6
|
GLDN
|
gliomedin |
| chr3_-_52452828 | 0.24 |
ENST00000496590.1
|
TNNC1
|
troponin C1, slow skeletal and cardiac type |
| chr6_+_99606833 | 0.24 |
ENST00000369215.5
|
PRDM13
|
PR/SET domain 13 |
| chr22_+_25742141 | 0.24 |
ENST00000536101.5
ENST00000335473.12 ENST00000407587.6 |
MYO18B
|
myosin XVIIIB |
| chr1_+_155689074 | 0.23 |
ENST00000343043.7
ENST00000421487.6 ENST00000535183.5 ENST00000368336.10 ENST00000465375.5 ENST00000470830.5 |
DAP3
|
death associated protein 3 |
| chr16_+_7332839 | 0.23 |
ENST00000355637.9
|
RBFOX1
|
RNA binding fox-1 homolog 1 |
| chr10_-_67696115 | 0.22 |
ENST00000433211.7
|
CTNNA3
|
catenin alpha 3 |
| chr4_+_15427998 | 0.22 |
ENST00000444304.3
|
C1QTNF7
|
C1q and TNF related 7 |
| chr2_-_162243375 | 0.22 |
ENST00000188790.9
ENST00000443424.5 |
FAP
|
fibroblast activation protein alpha |
| chr2_-_162242998 | 0.22 |
ENST00000627638.2
ENST00000447386.5 |
FAP
|
fibroblast activation protein alpha |
| chr16_+_7332744 | 0.21 |
ENST00000436368.6
ENST00000311745.9 ENST00000340209.8 ENST00000620507.4 |
RBFOX1
|
RNA binding fox-1 homolog 1 |
| chrX_-_85379659 | 0.21 |
ENST00000262753.9
|
POF1B
|
POF1B actin binding protein |
| chr11_+_27041313 | 0.21 |
ENST00000528583.5
|
BBOX1
|
gamma-butyrobetaine hydroxylase 1 |
| chr4_+_87832917 | 0.21 |
ENST00000395102.8
ENST00000497649.6 ENST00000540395.1 ENST00000560249.5 ENST00000511670.5 ENST00000361056.3 |
MEPE
|
matrix extracellular phosphoglycoprotein |
| chr3_-_120682215 | 0.21 |
ENST00000283871.10
|
HGD
|
homogentisate 1,2-dioxygenase |
| chr7_+_100867379 | 0.21 |
ENST00000200457.9
ENST00000619988.4 |
TRIP6
|
thyroid hormone receptor interactor 6 |
| chr19_-_893172 | 0.21 |
ENST00000325464.6
ENST00000312090.10 |
MED16
|
mediator complex subunit 16 |
| chr14_+_50560137 | 0.21 |
ENST00000358385.12
|
ATL1
|
atlastin GTPase 1 |
| chr14_+_36661852 | 0.21 |
ENST00000361487.7
|
PAX9
|
paired box 9 |
| chr2_-_80304700 | 0.20 |
ENST00000295057.4
|
LRRTM1
|
leucine rich repeat transmembrane neuronal 1 |
| chr1_+_160190567 | 0.20 |
ENST00000368078.8
|
CASQ1
|
calsequestrin 1 |
| chr10_-_104085847 | 0.20 |
ENST00000648076.2
|
COL17A1
|
collagen type XVII alpha 1 chain |
| chr7_+_114416286 | 0.20 |
ENST00000635534.1
|
FOXP2
|
forkhead box P2 |
| chr1_-_204359885 | 0.20 |
ENST00000414478.1
ENST00000272203.8 |
PLEKHA6
|
pleckstrin homology domain containing A6 |
| chr8_+_76681208 | 0.20 |
ENST00000651372.2
|
ZFHX4
|
zinc finger homeobox 4 |
| chr17_-_10421944 | 0.20 |
ENST00000403437.2
|
MYH8
|
myosin heavy chain 8 |
| chr3_+_188947199 | 0.19 |
ENST00000433971.5
|
TPRG1
|
tumor protein p63 regulated 1 |
| chrX_-_54998530 | 0.19 |
ENST00000545676.5
|
PFKFB1
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 |
| chr12_-_56643499 | 0.19 |
ENST00000551570.5
|
ATP5F1B
|
ATP synthase F1 subunit beta |
| chr15_+_90201301 | 0.19 |
ENST00000411539.6
|
SEMA4B
|
semaphorin 4B |
| chr11_+_5351508 | 0.19 |
ENST00000380219.1
|
OR51B6
|
olfactory receptor family 51 subfamily B member 6 |
| chr6_+_31546845 | 0.19 |
ENST00000376146.8
|
NFKBIL1
|
NFKB inhibitor like 1 |
| chr11_+_112175526 | 0.18 |
ENST00000532612.5
ENST00000438022.5 |
BCO2
|
beta-carotene oxygenase 2 |
| chr7_-_29195186 | 0.18 |
ENST00000449801.5
ENST00000409850.5 |
CPVL
|
carboxypeptidase vitellogenic like |
| chr14_+_39265278 | 0.18 |
ENST00000554392.5
ENST00000555716.5 ENST00000341749.7 ENST00000557038.5 |
MIA2
|
MIA SH3 domain ER export factor 2 |
| chr17_-_10518536 | 0.17 |
ENST00000226207.6
|
MYH1
|
myosin heavy chain 1 |
| chr7_+_95485898 | 0.17 |
ENST00000428113.5
|
ASB4
|
ankyrin repeat and SOCS box containing 4 |
| chr19_-_893200 | 0.17 |
ENST00000269814.8
ENST00000395808.7 |
MED16
|
mediator complex subunit 16 |
| chr7_+_100676112 | 0.17 |
ENST00000412215.5
ENST00000393924.1 |
GNB2
|
G protein subunit beta 2 |
| chr8_-_42501224 | 0.17 |
ENST00000520262.6
ENST00000517366.1 |
SLC20A2
|
solute carrier family 20 member 2 |
| chr7_+_95485934 | 0.17 |
ENST00000325885.6
|
ASB4
|
ankyrin repeat and SOCS box containing 4 |
| chr20_+_33562365 | 0.16 |
ENST00000346541.7
ENST00000397800.5 ENST00000492345.5 |
CBFA2T2
|
CBFA2/RUNX1 partner transcriptional co-repressor 2 |
| chr9_-_110999458 | 0.16 |
ENST00000374430.6
|
LPAR1
|
lysophosphatidic acid receptor 1 |
| chr10_-_60572599 | 0.15 |
ENST00000503366.5
|
ANK3
|
ankyrin 3 |
| chr17_-_50707855 | 0.15 |
ENST00000285243.7
|
ANKRD40
|
ankyrin repeat domain 40 |
| chr1_-_107688492 | 0.15 |
ENST00000415432.6
|
VAV3
|
vav guanine nucleotide exchange factor 3 |
| chrX_-_85379703 | 0.14 |
ENST00000373145.3
|
POF1B
|
POF1B actin binding protein |
| chr3_-_57199938 | 0.14 |
ENST00000473921.2
ENST00000295934.8 |
HESX1
|
HESX homeobox 1 |
| chr13_-_83882390 | 0.14 |
ENST00000377084.3
|
SLITRK1
|
SLIT and NTRK like family member 1 |
| chr6_-_116126120 | 0.13 |
ENST00000452729.1
ENST00000651968.1 ENST00000243222.8 |
COL10A1
|
collagen type X alpha 1 chain |
| chr7_+_129375643 | 0.13 |
ENST00000490911.5
|
AHCYL2
|
adenosylhomocysteinase like 2 |
| chr9_+_92964272 | 0.13 |
ENST00000468206.6
|
FGD3
|
FYVE, RhoGEF and PH domain containing 3 |
| chr6_-_55875583 | 0.13 |
ENST00000370830.4
|
BMP5
|
bone morphogenetic protein 5 |
| chr9_+_84669760 | 0.13 |
ENST00000304053.10
|
NTRK2
|
neurotrophic receptor tyrosine kinase 2 |
| chr11_-_59615673 | 0.12 |
ENST00000263847.6
|
OSBP
|
oxysterol binding protein |
| chr2_+_54456311 | 0.12 |
ENST00000615901.4
ENST00000356805.9 |
SPTBN1
|
spectrin beta, non-erythrocytic 1 |
| chr1_-_193106048 | 0.12 |
ENST00000367440.3
|
GLRX2
|
glutaredoxin 2 |
| chrX_-_123623155 | 0.12 |
ENST00000618150.4
|
THOC2
|
THO complex 2 |
| chr5_+_153490655 | 0.12 |
ENST00000518142.5
ENST00000285900.10 |
GRIA1
|
glutamate ionotropic receptor AMPA type subunit 1 |
| chr6_+_50818857 | 0.12 |
ENST00000393655.4
|
TFAP2B
|
transcription factor AP-2 beta |
| chr2_-_2324642 | 0.12 |
ENST00000650485.1
ENST00000649207.1 |
MYT1L
|
myelin transcription factor 1 like |
| chrX_+_41688967 | 0.12 |
ENST00000378142.9
|
GPR34
|
G protein-coupled receptor 34 |
| chr17_-_10469558 | 0.12 |
ENST00000255381.2
|
MYH4
|
myosin heavy chain 4 |
| chr11_+_112175501 | 0.11 |
ENST00000357685.11
ENST00000361053.8 |
BCO2
|
beta-carotene oxygenase 2 |
| chr12_+_53425070 | 0.11 |
ENST00000550839.1
|
AMHR2
|
anti-Mullerian hormone receptor type 2 |
| chr11_-_62545629 | 0.11 |
ENST00000528508.5
ENST00000533365.5 |
AHNAK
|
AHNAK nucleoprotein |
| chr14_-_37595224 | 0.11 |
ENST00000250448.5
|
FOXA1
|
forkhead box A1 |
| chr11_-_5343524 | 0.11 |
ENST00000300773.3
|
OR51B5
|
olfactory receptor family 51 subfamily B member 5 |
| chr9_-_110579880 | 0.11 |
ENST00000401783.6
ENST00000374461.1 |
SVEP1
|
sushi, von Willebrand factor type A, EGF and pentraxin domain containing 1 |
| chr9_+_84669707 | 0.11 |
ENST00000277120.8
ENST00000323115.10 |
NTRK2
|
neurotrophic receptor tyrosine kinase 2 |
| chr5_+_55853464 | 0.11 |
ENST00000652039.1
|
IL31RA
|
interleukin 31 receptor A |
| chr12_+_54053815 | 0.10 |
ENST00000430889.3
|
HOXC4
|
homeobox C4 |
| chr9_-_110579704 | 0.10 |
ENST00000374469.6
|
SVEP1
|
sushi, von Willebrand factor type A, EGF and pentraxin domain containing 1 |
| chr3_-_62374293 | 0.10 |
ENST00000486811.5
|
FEZF2
|
FEZ family zinc finger 2 |
| chr2_-_190062721 | 0.10 |
ENST00000260950.5
|
MSTN
|
myostatin |
| chr3_-_42702778 | 0.10 |
ENST00000457462.5
ENST00000441594.6 |
HHATL
|
hedgehog acyltransferase like |
| chr11_+_124183219 | 0.09 |
ENST00000641351.2
|
OR10D3
|
olfactory receptor family 10 subfamily D member 3 |
| chr12_+_106582996 | 0.09 |
ENST00000392842.6
|
RFX4
|
regulatory factor X4 |
| chr5_+_70025247 | 0.09 |
ENST00000380751.9
ENST00000380750.8 ENST00000503931.5 ENST00000506542.1 |
SERF1B
|
small EDRK-rich factor 1B |
| chr12_-_86256267 | 0.09 |
ENST00000620241.4
|
MGAT4C
|
MGAT4 family member C |
| chr18_-_23437927 | 0.09 |
ENST00000578520.5
ENST00000383233.8 |
TMEM241
|
transmembrane protein 241 |
| chr18_-_26548527 | 0.08 |
ENST00000580059.7
|
KCTD1
|
potassium channel tetramerization domain containing 1 |
| chr1_-_113871665 | 0.08 |
ENST00000528414.5
ENST00000460620.5 ENST00000359785.10 ENST00000420377.6 ENST00000525799.1 ENST00000538253.5 |
PTPN22
|
protein tyrosine phosphatase non-receptor type 22 |
| chr12_-_91004965 | 0.08 |
ENST00000261172.8
|
EPYC
|
epiphycan |
| chr10_+_11005301 | 0.08 |
ENST00000416382.6
ENST00000631460.1 ENST00000631816.1 |
CELF2
|
CUGBP Elav-like family member 2 |
| chr7_+_117611617 | 0.08 |
ENST00000468795.1
|
CFTR
|
CF transmembrane conductance regulator |
| chr6_-_116829037 | 0.07 |
ENST00000368549.7
ENST00000530250.1 ENST00000310357.8 |
GPRC6A
|
G protein-coupled receptor class C group 6 member A |
| chr22_+_22588155 | 0.07 |
ENST00000390302.3
|
IGLV2-33
|
immunoglobulin lambda variable 2-33 (non-functional) |
| chr11_+_124611420 | 0.07 |
ENST00000284288.3
|
PANX3
|
pannexin 3 |
| chr12_+_12785652 | 0.07 |
ENST00000356591.5
|
APOLD1
|
apolipoprotein L domain containing 1 |
| chr12_+_49750598 | 0.07 |
ENST00000552370.5
|
TMBIM6
|
transmembrane BAX inhibitor motif containing 6 |
| chr5_+_148203024 | 0.07 |
ENST00000325630.3
|
SPINK6
|
serine peptidase inhibitor Kazal type 6 |
| chr5_+_66958870 | 0.07 |
ENST00000405643.5
ENST00000407621.1 ENST00000432426.5 |
MAST4
|
microtubule associated serine/threonine kinase family member 4 |
| chrX_-_120311408 | 0.07 |
ENST00000309720.9
|
TMEM255A
|
transmembrane protein 255A |
| chrX_-_120311452 | 0.07 |
ENST00000371369.9
|
TMEM255A
|
transmembrane protein 255A |
| chrX_+_41689006 | 0.07 |
ENST00000378138.5
ENST00000620846.1 ENST00000649219.1 |
GPR34
|
G protein-coupled receptor 34 |
| chr7_-_13989658 | 0.06 |
ENST00000430479.6
ENST00000433547.1 ENST00000405192.6 |
ETV1
|
ETS variant transcription factor 1 |
| chr13_-_52848632 | 0.06 |
ENST00000377942.7
ENST00000338862.5 |
PCDH8
|
protocadherin 8 |
| chr14_-_80231052 | 0.06 |
ENST00000557010.5
|
DIO2
|
iodothyronine deiodinase 2 |
| chr6_-_31546552 | 0.06 |
ENST00000303892.10
ENST00000376151.4 |
ATP6V1G2
|
ATPase H+ transporting V1 subunit G2 |
| chr13_-_83882456 | 0.06 |
ENST00000674365.1
|
SLITRK1
|
SLIT and NTRK like family member 1 |
| chr7_-_13986498 | 0.06 |
ENST00000420159.6
ENST00000399357.7 ENST00000403527.5 |
ETV1
|
ETS variant transcription factor 1 |
| chr10_-_93702508 | 0.06 |
ENST00000359204.5
|
FRA10AC1
|
FRA10A associated CGG repeat 1 |
| chrX_-_120311533 | 0.05 |
ENST00000440464.5
ENST00000519908.1 |
TMEM255A
|
transmembrane protein 255A |
| chr10_+_113125536 | 0.05 |
ENST00000349937.7
|
TCF7L2
|
transcription factor 7 like 2 |
| chr5_+_67004618 | 0.05 |
ENST00000261569.11
ENST00000436277.5 |
MAST4
|
microtubule associated serine/threonine kinase family member 4 |
| chr11_+_121101243 | 0.05 |
ENST00000392793.6
ENST00000642222.1 |
TECTA
|
tectorin alpha |
| chr5_+_141430565 | 0.04 |
ENST00000613314.1
|
PCDHGA12
|
protocadherin gamma subfamily A, 12 |
| chr5_+_148202771 | 0.04 |
ENST00000514389.5
ENST00000621437.4 |
SPINK6
|
serine peptidase inhibitor Kazal type 6 |
| chr18_+_13612614 | 0.04 |
ENST00000586765.1
ENST00000677910.1 |
LDLRAD4
|
low density lipoprotein receptor class A domain containing 4 |
| chrX_-_111410420 | 0.04 |
ENST00000371993.7
ENST00000680476.1 |
DCX
|
doublecortin |
| chr2_+_113067970 | 0.04 |
ENST00000341010.6
|
IL1F10
|
interleukin 1 family member 10 |
| chr12_-_95073490 | 0.03 |
ENST00000330677.7
|
NR2C1
|
nuclear receptor subfamily 2 group C member 1 |
| chr5_-_146182591 | 0.03 |
ENST00000510191.5
ENST00000674277.1 ENST00000674447.1 ENST00000674270.1 ENST00000394434.7 ENST00000674290.1 ENST00000674398.1 ENST00000674174.1 |
LARS1
|
leucyl-tRNA synthetase 1 |
| chr17_-_48593748 | 0.03 |
ENST00000239151.6
|
HOXB5
|
homeobox B5 |
| chr2_+_53767772 | 0.03 |
ENST00000295304.5
|
CHAC2
|
ChaC glutathione specific gamma-glutamylcyclotransferase 2 |
| chr12_-_86256299 | 0.03 |
ENST00000552808.6
ENST00000547225.5 |
MGAT4C
|
MGAT4 family member C |
| chr5_-_146182475 | 0.02 |
ENST00000674158.1
ENST00000674191.1 ENST00000274562.13 |
LARS1
|
leucyl-tRNA synthetase 1 |
| chr11_-_13496018 | 0.02 |
ENST00000529816.1
|
PTH
|
parathyroid hormone |
| chr12_+_53454764 | 0.02 |
ENST00000439930.7
ENST00000548933.5 |
PCBP2
|
poly(rC) binding protein 2 |
| chr5_+_159916475 | 0.02 |
ENST00000306675.5
|
ADRA1B
|
adrenoceptor alpha 1B |
| chr10_-_73655984 | 0.02 |
ENST00000394810.3
|
SYNPO2L
|
synaptopodin 2 like |
| chr16_+_6483379 | 0.02 |
ENST00000552089.5
|
RBFOX1
|
RNA binding fox-1 homolog 1 |
| chr4_-_184217854 | 0.01 |
ENST00000296741.7
|
ENPP6
|
ectonucleotide pyrophosphatase/phosphodiesterase 6 |
| chr5_+_153491174 | 0.01 |
ENST00000521843.6
|
GRIA1
|
glutamate ionotropic receptor AMPA type subunit 1 |
| chr6_+_63211446 | 0.01 |
ENST00000370659.1
|
FKBP1C
|
FKBP prolyl isomerase family member 1C |
| chr7_-_13988863 | 0.01 |
ENST00000405358.8
|
ETV1
|
ETS variant transcription factor 1 |
| chr15_+_36702009 | 0.01 |
ENST00000562489.1
|
CDIN1
|
CDAN1 interacting nuclease 1 |
| chr3_+_122384167 | 0.01 |
ENST00000232125.9
ENST00000477892.5 ENST00000469967.1 |
FAM162A
|
family with sequence similarity 162 member A |
| chr1_+_26021768 | 0.01 |
ENST00000374280.4
|
EXTL1
|
exostosin like glycosyltransferase 1 |
| chr10_-_119178791 | 0.00 |
ENST00000298510.4
|
PRDX3
|
peroxiredoxin 3 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.1 | 2.5 | GO:0051386 | regulation of neurotrophin TRK receptor signaling pathway(GO:0051386) |
| 0.1 | 0.5 | GO:2000639 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.1 | 0.4 | GO:0097325 | melanocyte proliferation(GO:0097325) |
| 0.1 | 0.3 | GO:0048560 | establishment of anatomical structure orientation(GO:0048560) |
| 0.1 | 0.3 | GO:0016122 | tetraterpenoid metabolic process(GO:0016108) carotenoid metabolic process(GO:0016116) carotene catabolic process(GO:0016121) xanthophyll metabolic process(GO:0016122) terpene catabolic process(GO:0046247) |
| 0.1 | 1.0 | GO:0086024 | adrenergic receptor signaling pathway involved in positive regulation of heart rate(GO:0086024) |
| 0.1 | 0.6 | GO:0010989 | negative regulation of low-density lipoprotein particle clearance(GO:0010989) |
| 0.1 | 0.2 | GO:0002086 | diaphragm contraction(GO:0002086) |
| 0.1 | 0.2 | GO:0099538 | synaptic signaling via neuropeptide(GO:0099538) trans-synaptic signaling by neuropeptide(GO:0099540) trans-synaptic signaling by neuropeptide, modulating synaptic transmission(GO:0099551) |
| 0.1 | 0.3 | GO:0018008 | N-terminal peptidyl-glycine N-myristoylation(GO:0018008) |
| 0.1 | 0.2 | GO:0021758 | caudate nucleus development(GO:0021757) putamen development(GO:0021758) |
| 0.1 | 0.3 | GO:0002408 | myeloid dendritic cell chemotaxis(GO:0002408) |
| 0.1 | 0.3 | GO:0021778 | oligodendrocyte cell fate specification(GO:0021778) oligodendrocyte cell fate commitment(GO:0021779) glial cell fate specification(GO:0021780) |
| 0.1 | 1.1 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.1 | 0.3 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 0.0 | 0.1 | GO:0071676 | negative regulation of mononuclear cell migration(GO:0071676) |
| 0.0 | 0.2 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 0.0 | 0.2 | GO:0033133 | positive regulation of glucokinase activity(GO:0033133) |
| 0.0 | 0.2 | GO:0072660 | maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.0 | 0.5 | GO:0007512 | adult heart development(GO:0007512) |
| 0.0 | 0.1 | GO:0060738 | epithelial-mesenchymal signaling involved in prostate gland development(GO:0060738) |
| 0.0 | 0.6 | GO:0051798 | positive regulation of hair follicle development(GO:0051798) |
| 0.0 | 0.2 | GO:1904566 | response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904565) cellular response to 1-oleoyl-sn-glycerol 3-phosphate(GO:1904566) |
| 0.0 | 0.1 | GO:0006500 | N-terminal protein palmitoylation(GO:0006500) |
| 0.0 | 0.1 | GO:1900042 | positive regulation of interleukin-2 secretion(GO:1900042) |
| 0.0 | 0.6 | GO:0051531 | NFAT protein import into nucleus(GO:0051531) |
| 0.0 | 0.7 | GO:0048935 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.0 | 0.2 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 0.1 | GO:1902161 | positive regulation of cyclic nucleotide-gated ion channel activity(GO:1902161) |
| 0.0 | 0.5 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.0 | 0.2 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.0 | 0.1 | GO:1902613 | regulation of anti-Mullerian hormone signaling pathway(GO:1902612) negative regulation of anti-Mullerian hormone signaling pathway(GO:1902613) anti-Mullerian hormone signaling pathway(GO:1990262) |
| 0.0 | 0.1 | GO:0015783 | GDP-fucose transport(GO:0015783) purine nucleotide-sugar transport(GO:0036079) |
| 0.0 | 0.1 | GO:1902725 | negative regulation of satellite cell differentiation(GO:1902725) |
| 0.0 | 0.1 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.0 | 0.5 | GO:0002091 | negative regulation of receptor internalization(GO:0002091) |
| 0.0 | 0.2 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.0 | 0.3 | GO:2001214 | positive regulation of vasculogenesis(GO:2001214) |
| 0.0 | 0.3 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.0 | 0.1 | GO:0070425 | negative regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070425) negative regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070433) positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 0.0 | 0.2 | GO:0014877 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 0.0 | 0.2 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.0 | 0.1 | GO:0099566 | regulation of postsynaptic cytosolic calcium ion concentration(GO:0099566) |
| 0.0 | 0.2 | GO:0044341 | sodium-dependent phosphate transport(GO:0044341) |
| 0.0 | 0.3 | GO:0061365 | positive regulation of triglyceride lipase activity(GO:0061365) |
| 0.0 | 0.3 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.3 | GO:0036342 | post-anal tail morphogenesis(GO:0036342) |
| 0.0 | 0.1 | GO:0031438 | negative regulation of mRNA cleavage(GO:0031438) negative regulation of mRNA endonucleolytic cleavage involved in unfolded protein response(GO:1904721) |
| 0.0 | 0.1 | GO:0021914 | negative regulation of smoothened signaling pathway involved in ventral spinal cord patterning(GO:0021914) |
| 0.0 | 0.1 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.0 | 0.3 | GO:0048240 | sperm capacitation(GO:0048240) |
| 0.0 | 0.2 | GO:0031665 | negative regulation of lipopolysaccharide-mediated signaling pathway(GO:0031665) |
| 0.0 | 0.1 | GO:0042262 | DNA protection(GO:0042262) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:0005826 | actomyosin contractile ring(GO:0005826) |
| 0.1 | 0.6 | GO:1990578 | perinuclear endoplasmic reticulum membrane(GO:1990578) |
| 0.1 | 1.1 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.1 | 0.5 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.1 | 0.2 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 0.0 | 0.2 | GO:1990584 | cardiac Troponin complex(GO:1990584) |
| 0.0 | 0.3 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.0 | 0.2 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.0 | 0.4 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.0 | 0.1 | GO:0044308 | axonal spine(GO:0044308) |
| 0.0 | 0.3 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.0 | 0.3 | GO:0045275 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.0 | 0.5 | GO:0032982 | myosin filament(GO:0032982) |
| 0.0 | 0.6 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 0.1 | GO:0032437 | cuticular plate(GO:0032437) |
| 0.0 | 0.2 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.0 | 0.1 | GO:0031205 | endoplasmic reticulum Sec complex(GO:0031205) |
| 0.0 | 1.0 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 0.2 | GO:0016461 | unconventional myosin complex(GO:0016461) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.1 | 1.1 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 0.3 | GO:0004923 | leukemia inhibitory factor receptor activity(GO:0004923) |
| 0.1 | 0.2 | GO:0060175 | brain-derived neurotrophic factor-activated receptor activity(GO:0060175) |
| 0.1 | 0.4 | GO:0010861 | thyroid hormone receptor activator activity(GO:0010861) thyroid hormone receptor coactivator activity(GO:0030375) |
| 0.1 | 0.3 | GO:0019107 | glycylpeptide N-tetradecanoyltransferase activity(GO:0004379) myristoyltransferase activity(GO:0019107) |
| 0.1 | 0.2 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.1 | 0.5 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.1 | 0.6 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.0 | 0.2 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.0 | 2.5 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.0 | 0.2 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.0 | 0.2 | GO:0017018 | myosin phosphatase activity(GO:0017018) |
| 0.0 | 0.1 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 0.0 | 0.2 | GO:0015319 | sodium:inorganic phosphate symporter activity(GO:0015319) |
| 0.0 | 0.5 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.0 | 1.2 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.0 | 0.2 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 0.6 | GO:0048185 | activin binding(GO:0048185) |
| 0.0 | 0.1 | GO:0005260 | channel-conductance-controlling ATPase activity(GO:0005260) |
| 0.0 | 0.2 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.0 | 0.2 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.0 | 0.6 | GO:0034236 | protein kinase A catalytic subunit binding(GO:0034236) |
| 0.0 | 1.1 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.1 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.1 | GO:0005457 | GDP-fucose transmembrane transporter activity(GO:0005457) purine nucleotide-sugar transmembrane transporter activity(GO:0036080) |
| 0.0 | 0.6 | GO:0031698 | beta-2 adrenergic receptor binding(GO:0031698) |
| 0.0 | 0.2 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.0 | 0.1 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.0 | 0.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.1 | GO:0030613 | oxidoreductase activity, acting on phosphorus or arsenic in donors(GO:0030613) oxidoreductase activity, acting on phosphorus or arsenic in donors, disulfide as acceptor(GO:0030614) |
| 0.0 | 0.3 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.0 | 0.1 | GO:0055077 | gap junction hemi-channel activity(GO:0055077) |
| 0.0 | 0.1 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration(GO:0099583) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.0 | 0.4 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.5 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.0 | 0.1 | GO:0004800 | thyroxine 5'-deiodinase activity(GO:0004800) |
| 0.0 | 0.2 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.1 | GO:0060698 | endoribonuclease inhibitor activity(GO:0060698) |
| 0.0 | 0.3 | GO:0016676 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.9 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.0 | 1.1 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.0 | 0.7 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.0 | 1.2 | ST T CELL SIGNAL TRANSDUCTION | T Cell Signal Transduction |
| 0.0 | 0.1 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.6 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.0 | 1.0 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |