Project
Inflammatory response time course, HUVEC (Wada, 2009)
Navigation
Downloads
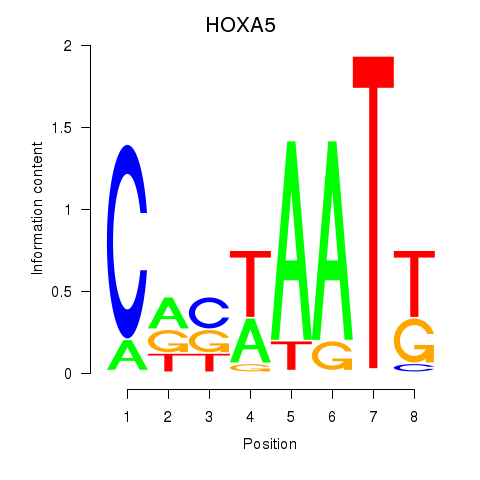
Results for HOXA5
Z-value: 0.37
Transcription factors associated with HOXA5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXA5
|
ENSG00000106004.5 | HOXA5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HOXA5 | hg38_v1_chr7_-_27143672_27143719 | -0.54 | 5.1e-03 | Click! |
Activity profile of HOXA5 motif
Sorted Z-values of HOXA5 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXA5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_73740541 | 0.59 |
ENST00000401931.1
ENST00000307407.8 |
CXCL8
|
C-X-C motif chemokine ligand 8 |
| chr7_-_22193824 | 0.57 |
ENST00000401957.6
|
RAPGEF5
|
Rap guanine nucleotide exchange factor 5 |
| chr7_-_22194709 | 0.57 |
ENST00000458533.5
|
RAPGEF5
|
Rap guanine nucleotide exchange factor 5 |
| chr3_+_190615308 | 0.56 |
ENST00000412080.1
|
IL1RAP
|
interleukin 1 receptor accessory protein |
| chr12_-_89526253 | 0.44 |
ENST00000547474.1
|
POC1B-GALNT4
|
POC1B-GALNT4 readthrough |
| chr22_-_41286168 | 0.43 |
ENST00000356244.8
|
RANGAP1
|
Ran GTPase activating protein 1 |
| chr6_-_154356735 | 0.42 |
ENST00000367220.8
ENST00000265198.8 ENST00000520261.1 |
IPCEF1
|
interaction protein for cytohesin exchange factors 1 |
| chr5_+_70025247 | 0.39 |
ENST00000380751.9
ENST00000380750.8 ENST00000503931.5 ENST00000506542.1 |
SERF1B
|
small EDRK-rich factor 1B |
| chr10_+_102226293 | 0.39 |
ENST00000370005.4
|
ELOVL3
|
ELOVL fatty acid elongase 3 |
| chr12_+_78036248 | 0.38 |
ENST00000644176.1
|
NAV3
|
neuron navigator 3 |
| chr11_+_20022550 | 0.37 |
ENST00000533917.5
|
NAV2
|
neuron navigator 2 |
| chr3_-_79019444 | 0.36 |
ENST00000618833.4
ENST00000436010.6 ENST00000618846.4 |
ROBO1
|
roundabout guidance receptor 1 |
| chr18_-_36129305 | 0.31 |
ENST00000269187.10
ENST00000590986.5 ENST00000440549.6 |
SLC39A6
|
solute carrier family 39 member 6 |
| chr6_+_125956696 | 0.31 |
ENST00000229633.7
|
HINT3
|
histidine triad nucleotide binding protein 3 |
| chr2_+_108377947 | 0.30 |
ENST00000272452.7
|
SULT1C4
|
sulfotransferase family 1C member 4 |
| chr5_+_67004618 | 0.30 |
ENST00000261569.11
ENST00000436277.5 |
MAST4
|
microtubule associated serine/threonine kinase family member 4 |
| chr13_-_40982880 | 0.22 |
ENST00000635415.1
|
ELF1
|
E74 like ETS transcription factor 1 |
| chr13_+_97222296 | 0.21 |
ENST00000343600.8
ENST00000376673.7 ENST00000679496.1 ENST00000345429.10 |
MBNL2
|
muscleblind like splicing regulator 2 |
| chr17_-_29761415 | 0.21 |
ENST00000649863.1
|
SSH2
|
slingshot protein phosphatase 2 |
| chr2_-_160062589 | 0.20 |
ENST00000392771.1
ENST00000283243.13 |
PLA2R1
|
phospholipase A2 receptor 1 |
| chr12_-_9607903 | 0.20 |
ENST00000229402.4
|
KLRB1
|
killer cell lectin like receptor B1 |
| chr3_+_12351493 | 0.18 |
ENST00000683699.1
|
PPARG
|
peroxisome proliferator activated receptor gamma |
| chr17_-_29761390 | 0.18 |
ENST00000324677.12
|
SSH2
|
slingshot protein phosphatase 2 |
| chr6_+_122779707 | 0.17 |
ENST00000368444.8
ENST00000356535.4 |
FABP7
|
fatty acid binding protein 7 |
| chr17_+_19533818 | 0.17 |
ENST00000436810.6
ENST00000270570.8 ENST00000575023.5 ENST00000395585.5 |
SLC47A1
|
solute carrier family 47 member 1 |
| chr17_-_51046868 | 0.17 |
ENST00000510283.5
ENST00000510855.1 |
SPAG9
|
sperm associated antigen 9 |
| chr2_+_113406368 | 0.16 |
ENST00000453673.3
|
IGKV1OR2-108
|
immunoglobulin kappa variable 1/OR2-108 (non-functional) |
| chr11_+_33039996 | 0.16 |
ENST00000432887.5
ENST00000528898.1 ENST00000531632.6 |
TCP11L1
|
t-complex 11 like 1 |
| chr19_-_1021114 | 0.16 |
ENST00000333175.9
ENST00000356663.8 |
TMEM259
|
transmembrane protein 259 |
| chr12_-_91180365 | 0.15 |
ENST00000547937.5
|
DCN
|
decorin |
| chr8_-_33567118 | 0.15 |
ENST00000256257.2
|
RNF122
|
ring finger protein 122 |
| chr3_-_191282383 | 0.14 |
ENST00000427544.6
|
UTS2B
|
urotensin 2B |
| chr7_-_93226449 | 0.14 |
ENST00000394468.7
ENST00000453812.2 |
HEPACAM2
|
HEPACAM family member 2 |
| chr10_-_50885619 | 0.14 |
ENST00000373997.8
|
A1CF
|
APOBEC1 complementation factor |
| chrX_-_66033664 | 0.13 |
ENST00000427538.5
|
VSIG4
|
V-set and immunoglobulin domain containing 4 |
| chr6_+_26365215 | 0.13 |
ENST00000527422.5
ENST00000356386.6 ENST00000396948.5 |
BTN3A2
|
butyrophilin subfamily 3 member A2 |
| chr13_+_30427950 | 0.13 |
ENST00000436446.1
|
UBE2L5
|
ubiquitin conjugating enzyme E2 L5 |
| chr8_-_134510182 | 0.13 |
ENST00000521673.5
|
ZFAT
|
zinc finger and AT-hook domain containing |
| chr16_-_72094371 | 0.13 |
ENST00000426362.6
|
TXNL4B
|
thioredoxin like 4B |
| chr11_+_66011994 | 0.13 |
ENST00000312134.3
|
CST6
|
cystatin E/M |
| chr6_+_26365176 | 0.12 |
ENST00000377708.7
|
BTN3A2
|
butyrophilin subfamily 3 member A2 |
| chr3_+_130850585 | 0.12 |
ENST00000505330.5
ENST00000504381.5 ENST00000507488.6 |
ATP2C1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr7_+_142384328 | 0.12 |
ENST00000390361.3
|
TRBV7-3
|
T cell receptor beta variable 7-3 |
| chr18_+_616711 | 0.12 |
ENST00000579494.1
|
CLUL1
|
clusterin like 1 |
| chr18_+_61333424 | 0.12 |
ENST00000262717.9
|
CDH20
|
cadherin 20 |
| chr18_+_616672 | 0.12 |
ENST00000338387.11
|
CLUL1
|
clusterin like 1 |
| chr3_+_98166696 | 0.12 |
ENST00000641450.1
|
OR5H15
|
olfactory receptor family 5 subfamily H member 15 |
| chr10_-_67665642 | 0.12 |
ENST00000682945.1
ENST00000330298.6 ENST00000682758.1 |
CTNNA3
|
catenin alpha 3 |
| chr4_+_183905266 | 0.11 |
ENST00000308497.9
|
STOX2
|
storkhead box 2 |
| chr8_+_54616078 | 0.11 |
ENST00000220676.2
|
RP1
|
RP1 axonemal microtubule associated |
| chr14_+_23556253 | 0.10 |
ENST00000556015.5
ENST00000554970.1 ENST00000288014.7 ENST00000554789.1 |
THTPA
|
thiamine triphosphatase |
| chr1_-_157841800 | 0.10 |
ENST00000368174.5
|
CD5L
|
CD5 molecule like |
| chr16_-_62036412 | 0.10 |
ENST00000577390.6
|
CDH8
|
cadherin 8 |
| chr1_-_170074568 | 0.10 |
ENST00000367767.5
ENST00000361580.7 ENST00000538366.5 |
KIFAP3
|
kinesin associated protein 3 |
| chr1_+_248038172 | 0.09 |
ENST00000366479.4
|
OR2L2
|
olfactory receptor family 2 subfamily L member 2 |
| chr16_-_66730216 | 0.09 |
ENST00000569320.5
|
DYNC1LI2
|
dynein cytoplasmic 1 light intermediate chain 2 |
| chr18_-_63644250 | 0.09 |
ENST00000341074.10
ENST00000436264.1 |
SERPINB4
|
serpin family B member 4 |
| chr19_+_10286944 | 0.09 |
ENST00000380770.5
|
ICAM4
|
intercellular adhesion molecule 4 (Landsteiner-Wiener blood group) |
| chr2_+_165239388 | 0.09 |
ENST00000424833.5
ENST00000375437.7 ENST00000631182.3 |
SCN2A
|
sodium voltage-gated channel alpha subunit 2 |
| chr6_-_17706852 | 0.09 |
ENST00000262077.3
|
NUP153
|
nucleoporin 153 |
| chr16_-_62036399 | 0.09 |
ENST00000584337.5
|
CDH8
|
cadherin 8 |
| chr18_+_13611429 | 0.09 |
ENST00000587757.5
|
LDLRAD4
|
low density lipoprotein receptor class A domain containing 4 |
| chr5_+_72848115 | 0.09 |
ENST00000679378.1
|
TNPO1
|
transportin 1 |
| chr19_-_54100792 | 0.09 |
ENST00000391761.5
ENST00000356532.7 ENST00000616447.4 ENST00000359649.8 ENST00000358375.8 ENST00000391760.1 ENST00000351806.8 |
OSCAR
|
osteoclast associated Ig-like receptor |
| chr10_+_84194621 | 0.09 |
ENST00000332904.7
|
CDHR1
|
cadherin related family member 1 |
| chr3_-_161372821 | 0.08 |
ENST00000617024.1
ENST00000359175.8 |
SPTSSB
|
serine palmitoyltransferase small subunit B |
| chr19_-_43754901 | 0.08 |
ENST00000270066.11
ENST00000601170.5 |
SMG9
|
SMG9 nonsense mediated mRNA decay factor |
| chr17_-_76737321 | 0.08 |
ENST00000359995.10
ENST00000508921.7 ENST00000583836.1 ENST00000358156.6 ENST00000392485.2 |
SRSF2
|
serine and arginine rich splicing factor 2 |
| chr11_-_72112068 | 0.08 |
ENST00000537644.5
ENST00000538919.5 ENST00000539395.1 ENST00000542531.5 |
ANAPC15
|
anaphase promoting complex subunit 15 |
| chr7_+_2354810 | 0.08 |
ENST00000360876.9
ENST00000413917.5 ENST00000397011.2 |
EIF3B
|
eukaryotic translation initiation factor 3 subunit B |
| chr19_-_39320839 | 0.08 |
ENST00000248668.5
|
LRFN1
|
leucine rich repeat and fibronectin type III domain containing 1 |
| chr19_-_51019699 | 0.08 |
ENST00000358789.8
|
KLK10
|
kallikrein related peptidase 10 |
| chr4_-_102825767 | 0.08 |
ENST00000505207.5
ENST00000502404.5 ENST00000507845.5 |
UBE2D3
|
ubiquitin conjugating enzyme E2 D3 |
| chr4_-_102825854 | 0.07 |
ENST00000350435.11
|
UBE2D3
|
ubiquitin conjugating enzyme E2 D3 |
| chr18_+_34710307 | 0.07 |
ENST00000679796.1
|
DTNA
|
dystrobrevin alpha |
| chr5_-_132543513 | 0.07 |
ENST00000231454.6
|
IL5
|
interleukin 5 |
| chr8_-_41309434 | 0.07 |
ENST00000220772.8
|
SFRP1
|
secreted frizzled related protein 1 |
| chr3_-_190449782 | 0.07 |
ENST00000354905.3
|
TMEM207
|
transmembrane protein 207 |
| chr2_+_90154073 | 0.07 |
ENST00000611391.1
|
IGKV1D-13
|
immunoglobulin kappa variable 1D-13 |
| chr2_+_165239432 | 0.07 |
ENST00000636071.2
ENST00000636985.2 |
SCN2A
|
sodium voltage-gated channel alpha subunit 2 |
| chr4_-_70666492 | 0.07 |
ENST00000254801.9
ENST00000391614.7 |
JCHAIN
|
joining chain of multimeric IgA and IgM |
| chr18_-_3874270 | 0.07 |
ENST00000400149.7
ENST00000400155.5 ENST00000400150.7 |
DLGAP1
|
DLG associated protein 1 |
| chr4_-_102825526 | 0.07 |
ENST00000504211.5
ENST00000508476.5 |
UBE2D3
|
ubiquitin conjugating enzyme E2 D3 |
| chr7_-_100119840 | 0.07 |
ENST00000437822.6
|
TAF6
|
TATA-box binding protein associated factor 6 |
| chr1_+_204870831 | 0.07 |
ENST00000404076.5
ENST00000539706.6 |
NFASC
|
neurofascin |
| chr14_+_77708076 | 0.06 |
ENST00000238688.9
ENST00000557342.6 ENST00000557623.5 ENST00000557431.5 ENST00000556831.5 ENST00000556375.5 ENST00000553981.1 |
SLIRP
|
SRA stem-loop interacting RNA binding protein |
| chr11_-_11353241 | 0.06 |
ENST00000528848.3
|
CSNK2A3
|
casein kinase 2 alpha 3 |
| chr10_+_99329349 | 0.06 |
ENST00000356713.5
|
CNNM1
|
cyclin and CBS domain divalent metal cation transport mediator 1 |
| chr17_-_28897724 | 0.06 |
ENST00000394908.9
|
FLOT2
|
flotillin 2 |
| chr1_-_152325232 | 0.06 |
ENST00000368799.2
|
FLG
|
filaggrin |
| chr6_+_131637296 | 0.06 |
ENST00000358229.6
ENST00000357639.8 |
ENPP3
|
ectonucleotide pyrophosphatase/phosphodiesterase 3 |
| chr1_+_241652275 | 0.06 |
ENST00000366552.6
ENST00000437684.7 |
WDR64
|
WD repeat domain 64 |
| chr3_+_178558700 | 0.06 |
ENST00000432997.5
ENST00000455865.5 |
KCNMB2
|
potassium calcium-activated channel subfamily M regulatory beta subunit 2 |
| chr2_-_232776555 | 0.06 |
ENST00000438786.1
ENST00000233826.4 ENST00000409779.1 |
KCNJ13
|
potassium inwardly rectifying channel subfamily J member 13 |
| chr16_-_21278282 | 0.06 |
ENST00000572914.2
|
CRYM
|
crystallin mu |
| chr11_-_115504389 | 0.06 |
ENST00000545380.5
ENST00000452722.7 ENST00000331581.11 ENST00000537058.5 ENST00000536727.5 ENST00000542447.6 |
CADM1
|
cell adhesion molecule 1 |
| chr8_+_91249307 | 0.05 |
ENST00000309536.6
ENST00000276609.8 |
SLC26A7
|
solute carrier family 26 member 7 |
| chr2_-_215138603 | 0.05 |
ENST00000272895.12
|
ABCA12
|
ATP binding cassette subfamily A member 12 |
| chr10_-_20897288 | 0.05 |
ENST00000377122.9
|
NEBL
|
nebulette |
| chrX_-_108736556 | 0.05 |
ENST00000372129.4
|
IRS4
|
insulin receptor substrate 4 |
| chr1_-_154178174 | 0.05 |
ENST00000302206.9
|
TPM3
|
tropomyosin 3 |
| chr10_-_27746060 | 0.05 |
ENST00000375790.9
|
MKX
|
mohawk homeobox |
| chr4_+_119027335 | 0.05 |
ENST00000627783.2
|
SYNPO2
|
synaptopodin 2 |
| chr4_+_73409340 | 0.05 |
ENST00000511370.1
|
ALB
|
albumin |
| chr1_+_154257071 | 0.05 |
ENST00000428595.1
|
UBAP2L
|
ubiquitin associated protein 2 like |
| chr3_+_172039556 | 0.05 |
ENST00000415807.7
ENST00000421757.5 |
FNDC3B
|
fibronectin type III domain containing 3B |
| chr10_-_27745810 | 0.05 |
ENST00000419761.6
|
MKX
|
mohawk homeobox |
| chr3_-_62373538 | 0.05 |
ENST00000283268.8
|
FEZF2
|
FEZ family zinc finger 2 |
| chr1_+_177170916 | 0.04 |
ENST00000361539.5
|
BRINP2
|
BMP/retinoic acid inducible neural specific 2 |
| chr7_+_143935233 | 0.04 |
ENST00000408955.3
|
OR2F2
|
olfactory receptor family 2 subfamily F member 2 |
| chr2_-_79087986 | 0.04 |
ENST00000305089.8
|
REG1B
|
regenerating family member 1 beta |
| chr12_-_21775581 | 0.04 |
ENST00000537950.1
ENST00000665145.1 |
KCNJ8
|
potassium inwardly rectifying channel subfamily J member 8 |
| chr1_-_247458105 | 0.04 |
ENST00000641149.1
ENST00000641527.1 |
OR2B11
|
olfactory receptor family 2 subfamily B member 11 |
| chr11_-_6683282 | 0.04 |
ENST00000532203.1
ENST00000288937.7 |
MRPL17
|
mitochondrial ribosomal protein L17 |
| chr3_-_62373015 | 0.04 |
ENST00000475839.1
|
FEZF2
|
FEZ family zinc finger 2 |
| chr4_-_163613505 | 0.04 |
ENST00000339875.9
|
MARCHF1
|
membrane associated ring-CH-type finger 1 |
| chr11_-_30016945 | 0.04 |
ENST00000328224.7
|
KCNA4
|
potassium voltage-gated channel subfamily A member 4 |
| chr11_-_33722710 | 0.03 |
ENST00000415002.7
ENST00000437761.6 ENST00000445143.6 |
CD59
|
CD59 molecule (CD59 blood group) |
| chr3_+_173398438 | 0.03 |
ENST00000457714.5
|
NLGN1
|
neuroligin 1 |
| chr9_+_74497308 | 0.03 |
ENST00000376896.8
|
RORB
|
RAR related orphan receptor B |
| chrX_+_49879475 | 0.03 |
ENST00000621775.2
|
USP27X
|
ubiquitin specific peptidase 27 X-linked |
| chr10_+_17951906 | 0.03 |
ENST00000377371.3
ENST00000377369.7 |
SLC39A12
|
solute carrier family 39 member 12 |
| chr8_-_27838034 | 0.03 |
ENST00000522944.5
|
PBK
|
PDZ binding kinase |
| chr15_-_78811415 | 0.03 |
ENST00000388820.5
|
ADAMTS7
|
ADAM metallopeptidase with thrombospondin type 1 motif 7 |
| chr3_+_35679614 | 0.03 |
ENST00000474696.5
ENST00000412048.5 ENST00000396482.6 ENST00000432682.5 |
ARPP21
|
cAMP regulated phosphoprotein 21 |
| chr2_-_40430257 | 0.03 |
ENST00000408028.6
ENST00000332839.8 ENST00000406391.2 ENST00000405901.7 |
SLC8A1
|
solute carrier family 8 member A1 |
| chr12_-_11395556 | 0.03 |
ENST00000565533.1
ENST00000389362.6 ENST00000546254.3 |
PRB2
PRB1
|
proline rich protein BstNI subfamily 2 proline rich protein BstNI subfamily 1 |
| chr16_+_10877181 | 0.03 |
ENST00000618207.4
ENST00000618327.4 ENST00000324288.14 ENST00000381835.9 |
CIITA
|
class II major histocompatibility complex transactivator |
| chr7_-_23470469 | 0.03 |
ENST00000258729.8
|
IGF2BP3
|
insulin like growth factor 2 mRNA binding protein 3 |
| chr2_+_165294031 | 0.03 |
ENST00000283256.10
|
SCN2A
|
sodium voltage-gated channel alpha subunit 2 |
| chr10_+_18260715 | 0.02 |
ENST00000615785.4
ENST00000617363.4 ENST00000396576.6 |
CACNB2
|
calcium voltage-gated channel auxiliary subunit beta 2 |
| chr10_+_84194527 | 0.02 |
ENST00000623527.4
|
CDHR1
|
cadherin related family member 1 |
| chr2_-_157444044 | 0.02 |
ENST00000264192.8
|
CYTIP
|
cytohesin 1 interacting protein |
| chr3_+_12351470 | 0.02 |
ENST00000287820.10
|
PPARG
|
peroxisome proliferator activated receptor gamma |
| chr10_-_114144599 | 0.02 |
ENST00000428953.1
|
CCDC186
|
coiled-coil domain containing 186 |
| chr3_+_196744 | 0.02 |
ENST00000256509.7
ENST00000397491.6 |
CHL1
|
cell adhesion molecule L1 like |
| chrX_+_40581035 | 0.02 |
ENST00000447485.6
|
ATP6AP2
|
ATPase H+ transporting accessory protein 2 |
| chr11_-_96343170 | 0.02 |
ENST00000524717.6
|
MAML2
|
mastermind like transcriptional coactivator 2 |
| chr7_+_44044663 | 0.02 |
ENST00000456905.5
ENST00000440166.5 ENST00000452943.5 ENST00000448521.6 ENST00000468694.5 ENST00000494774.5 |
DBNL
|
drebrin like |
| chr8_-_71547626 | 0.01 |
ENST00000647540.1
ENST00000644229.1 |
EYA1
|
EYA transcriptional coactivator and phosphatase 1 |
| chr8_-_25424260 | 0.01 |
ENST00000421054.7
|
GNRH1
|
gonadotropin releasing hormone 1 |
| chr12_+_25052732 | 0.01 |
ENST00000547044.5
|
IRAG2
|
inositol 1,4,5-triphosphate receptor associated 2 |
| chr6_+_167111789 | 0.01 |
ENST00000400926.5
|
CCR6
|
C-C motif chemokine receptor 6 |
| chr9_+_33290493 | 0.01 |
ENST00000379540.8
ENST00000318524.6 |
NFX1
|
nuclear transcription factor, X-box binding 1 |
| chr8_+_49911396 | 0.01 |
ENST00000642720.2
|
SNTG1
|
syntrophin gamma 1 |
| chr12_+_110614027 | 0.01 |
ENST00000550703.6
ENST00000551590.5 |
TCTN1
|
tectonic family member 1 |
| chr1_+_197268222 | 0.01 |
ENST00000367400.8
ENST00000638467.1 ENST00000367399.6 |
CRB1
|
crumbs cell polarity complex component 1 |
| chr1_+_197268204 | 0.01 |
ENST00000535699.5
ENST00000538660.5 |
CRB1
|
crumbs cell polarity complex component 1 |
| chr8_+_49911604 | 0.01 |
ENST00000642164.1
ENST00000644093.1 ENST00000643999.1 ENST00000647073.1 ENST00000646880.1 |
SNTG1
|
syntrophin gamma 1 |
| chr6_-_87589961 | 0.01 |
ENST00000369536.10
|
RARS2
|
arginyl-tRNA synthetase 2, mitochondrial |
| chr6_-_34146080 | 0.01 |
ENST00000538487.7
ENST00000374181.8 |
GRM4
|
glutamate metabotropic receptor 4 |
| chr2_-_206218024 | 0.01 |
ENST00000407325.6
ENST00000612892.4 ENST00000411719.1 |
GPR1
|
G protein-coupled receptor 1 |
| chr13_-_46897021 | 0.00 |
ENST00000542664.4
ENST00000543956.5 |
HTR2A
|
5-hydroxytryptamine receptor 2A |
| chr20_-_59940289 | 0.00 |
ENST00000370996.5
|
PPP1R3D
|
protein phosphatase 1 regulatory subunit 3D |
| chr7_+_44044634 | 0.00 |
ENST00000490734.6
|
DBNL
|
drebrin like |
| chr1_-_92486916 | 0.00 |
ENST00000294702.6
|
GFI1
|
growth factor independent 1 transcriptional repressor |
| chr5_+_140855882 | 0.00 |
ENST00000562220.2
ENST00000307360.6 ENST00000506939.6 |
PCDHA10
|
protocadherin alpha 10 |
| chr7_+_155644635 | 0.00 |
ENST00000401878.8
ENST00000392759.7 |
RBM33
|
RNA binding motif protein 33 |
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:1904117 | response to vasopressin(GO:1904116) cellular response to vasopressin(GO:1904117) |
| 0.1 | 0.4 | GO:0050925 | cerebral cortex tangential migration using cell-cell interactions(GO:0021823) postnatal olfactory bulb interneuron migration(GO:0021827) chemorepulsion involved in postnatal olfactory bulb interneuron migration(GO:0021836) negative regulation of negative chemotaxis(GO:0050925) |
| 0.1 | 0.6 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 0.1 | 0.2 | GO:0032304 | negative regulation of icosanoid secretion(GO:0032304) |
| 0.1 | 0.4 | GO:0003025 | regulation of systemic arterial blood pressure by baroreceptor feedback(GO:0003025) vagus nerve development(GO:0021564) |
| 0.1 | 0.6 | GO:2000535 | regulation of entry of bacterium into host cell(GO:2000535) |
| 0.0 | 0.4 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.1 | GO:0042357 | thiamine diphosphate metabolic process(GO:0042357) |
| 0.0 | 0.1 | GO:0046586 | regulation of calcium-dependent cell-cell adhesion(GO:0046586) |
| 0.0 | 0.1 | GO:0042270 | protection from natural killer cell mediated cytotoxicity(GO:0042270) |
| 0.0 | 0.1 | GO:1902594 | viral penetration into host nucleus(GO:0075732) multi-organism nuclear import(GO:1902594) |
| 0.0 | 0.2 | GO:2000230 | negative regulation of pancreatic stellate cell proliferation(GO:2000230) |
| 0.0 | 0.1 | GO:0045643 | regulation of eosinophil differentiation(GO:0045643) positive regulation of eosinophil differentiation(GO:0045645) |
| 0.0 | 0.1 | GO:1904956 | neural crest cell fate commitment(GO:0014034) regulation of midbrain dopaminergic neuron differentiation(GO:1904956) |
| 0.0 | 0.2 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.0 | 0.1 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.0 | 0.3 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.0 | 0.1 | GO:0030505 | inorganic diphosphate transport(GO:0030505) |
| 0.0 | 0.1 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.0 | 0.2 | GO:0046618 | drug export(GO:0046618) |
| 0.0 | 0.4 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.0 | GO:0044179 | cytolysis by symbiont of host cells(GO:0001897) hemolysis by symbiont of host erythrocytes(GO:0019836) hemolysis in other organism(GO:0044179) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 0.0 | 0.1 | GO:0000961 | negative regulation of mitochondrial RNA catabolic process(GO:0000961) |
| 0.0 | 0.1 | GO:0086073 | bundle of His cell-Purkinje myocyte adhesion involved in cell communication(GO:0086073) |
| 0.0 | 0.1 | GO:1904220 | regulation of serine C-palmitoyltransferase activity(GO:1904220) |
| 0.0 | 0.2 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.0 | 0.1 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.0 | 0.3 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.0 | 0.1 | GO:0046549 | retinal cone cell differentiation(GO:0042670) retinal cone cell development(GO:0046549) |
| 0.0 | 0.1 | GO:0044860 | protein localization to plasma membrane raft(GO:0044860) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:1990723 | cytoplasmic periphery of the nuclear pore complex(GO:1990723) |
| 0.0 | 0.1 | GO:0030895 | apolipoprotein B mRNA editing enzyme complex(GO:0030895) |
| 0.0 | 0.1 | GO:0016939 | kinesin II complex(GO:0016939) |
| 0.0 | 0.1 | GO:0071754 | dimeric IgA immunoglobulin complex(GO:0071750) secretory dimeric IgA immunoglobulin complex(GO:0071752) IgM immunoglobulin complex(GO:0071753) IgM immunoglobulin complex, circulating(GO:0071754) pentameric IgM immunoglobulin complex(GO:0071756) |
| 0.0 | 0.1 | GO:0035061 | interchromatin granule(GO:0035061) |
| 0.0 | 0.4 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.1 | GO:0097454 | Schwann cell microvillus(GO:0097454) |
| 0.0 | 0.2 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.0 | 0.1 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 0.6 | GO:0002114 | interleukin-33 receptor activity(GO:0002114) |
| 0.0 | 0.6 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.4 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.0 | 0.4 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.2 | GO:0015307 | drug:proton antiporter activity(GO:0015307) |
| 0.0 | 0.1 | GO:0005137 | interleukin-5 receptor binding(GO:0005137) |
| 0.0 | 0.4 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.0 | 0.2 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.0 | 0.1 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.0 | 0.3 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.0 | 0.1 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.0 | 0.1 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.0 | 0.1 | GO:0019862 | IgA binding(GO:0019862) |
| 0.0 | 0.2 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.0 | 0.1 | GO:0035529 | NADH pyrophosphatase activity(GO:0035529) |
| 0.0 | 0.1 | GO:0034040 | lipid-transporting ATPase activity(GO:0034040) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 0.4 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 0.3 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.3 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.0 | 0.4 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |